Abstract
Circ-ITCH, a novel circRNA, was generated from several exons of itchy E3 ubiquitin protein ligase (ITCH). Recently, circ-ITCH has been demonstrated to be involved in cancer development. However, there have been few investigations on the specific role of circ-ITCH in glioma. In this study, we performed quantitative real-time polymerase chain reaction analysis and identified that circ-ITCH was significantly downregulated in glioma tissues and cell lines. The function assays showed that upregulation of circ-ITCH inhibited glioma cell proliferation and invasion
Introduction
Glioma, one of the most common brain tumors, accounts for about 80% of all primary tumors in the central nervous system 1,2 . This disease usually shows various malignant features such as intense cell proliferation, diffuse infiltration, and aggressive progression 3,4 . These features result in a poor prognosis for most glioma patients 5 . Despite the great advancement in therapeutic approaches including chemotherapy and radiotherapy, the median survival time of glioma patients is short (12 to 15 months) and the 5-year survival rate is even less than 5% 6 –8 . Therefore, it is imperative to identify specific glioma-related biomarkers to fight against gliomas and improve the clinical management of glioma patients.
Circular RNAs (circRNAs) are a novel class of endogenous noncoding RNAs 9 . They are formed by backsplicing and feature a covalently closed loop 10 . CircRNAs play a critical role in regulating multiple biological processes such as cellular growth, survival, and differentiation 11 –13 . Recently, a growing number of circRNAs have been reported to be aberrantly expressed in cancer, suggesting a close association of circRNAs with tumorigenesis 14,15 . Thus, circRNAs have become the focus of research on cancer progression because the exploration of their functions will provide a great benefit to early diagnosis and prognosis prediction. Circ-ITCH, a novel circRNA, was generated from several exons of itchy E3 ubiquitin protein ligase (ITCH) and was first identified by Memczak et al 16 . In a variety of studies, circ-ITCH has been demonstrated to be involved in cancer development. For example, Guo et al. found that circ-ITCH was lowly expressed in hepatocellular cancer tissues and could be considered a prognostic indicator for hepatocellular cancer 17 . Furthermore, Yang et al. reported that circ-ITCH was downregulated in bladder cancer tissues and cell lines and enforced expression of circ-ITCH inhibited the aggressive biological behaviors of bladder cancer 18 . However, there have been few investigations on the specific role of circ-ITCH in glioma.
In this study, we found that circ-ITCH was downregulated in glioma tissues and cell lines. Upregulation of circ-ITCH suppressed glioma cell proliferation and invasion
Materials and Methods
Patients and Tissue Samples
Human glioma tissues and adjacent normal brain tissues were collected from 48 glioma patients who received surgery at Nanjing First Hospital, Nanjing Medical University (Nanjing, China). No patients underwent chemotherapy or radiotherapy before surgery. The study was approved by the Ethics Committee of Nanjing First Hospital. All tissue samples were collected with written informed consent from each patient and then frozen in liquid nitrogen and stored at −80 °C.
Cell Lines and Cell Culture
Human glioma cell lines (U251, U87, SHG44, and A172) and normal glial cell line HEB were purchased from the Chinese Academy of Sciences (Shanghai, China) and cultured in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, Rockville, MD, USA) supplemented with 10% fetal bovine serum (FBS; Gibco), 100 U/ml penicillin, and 100 µg/ml streptomycin. All cell lines were incubated at 37 °C in a humidified atmosphere with 5% carbon dioxide.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
Total RNA was isolated from tissues or cells using the Trizol reagent (Invitrogen, Carlsbad, CA, USA) and reversely transcribed into complementary DNA (cDNA) using the PrimeScript RT reagent kit (Takara, Tokyo, Japan). RT-qPCR was performed using the SYBR Green PCR kit (Takara) on an ABI PRISM 7500 System (Applied Biosystems, Foster City, CA, USA). The following primers were used: circ-ITCH, 5′-CCTTGAGCAAGAAGACTATGCCAAT-3′ (forward) and 5′-CCGCATTCTGTGGTAAGCAATCA-3′ (reverse); SASH1, 5′-CGGGAAAGCGTCAAGTCGGA-3′ (forward) and 5′-ATCTCCTTTCTTGAGCTTGAG-3′ (reverse); GAPDH, 5′-GTCAACGGATTTGGTCTGTATT-3′ (forward) and 5′-AGTCTTCTGGGTGGCAGTGAT-3′ (reverse); miR-106a-5p, 5′-GATGCTCAAAAAGTGCTTACAGTGCA-3′ (forward) and 5′-TATGGTTGTTCTGCTCTCTGTCTC-3′ (reverse); U6, 5′-GCTTCGGCAGCACATATACTAAAAT-3′ (forward) and 5′-CGCTTCACGAATTTGCGTGTCAT-3′ (reverse). The relative expression was analyzed using the 2−ΔΔCt method.
Western Blot Analysis
Total protein was extracted from tissues or cells with radioimmunoprecipitation assay lysis buffer (Beyotime, Shanghai, China), separated by 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis, and then transferred onto polyvinylidene difluoride membranes (Millipore, Billerica, MA, USA). After blocking in 5% skim milk, the membranes were incubated overnight at 4 °C with primary antibodies against SASH1 and β-tubulin. Subsequently, the membranes were washed 3 times and incubated with appropriate secondary antibodies at room temperature. All antibodies were obtained from Cell Signaling Technology (Danvers, MA, USA). The protein bands were visualized with an enhanced chemiluminescence system (Thermo Fisher Scientific, Waltham, MA, USA) and analyzed using the Quantity One system (Bio-Rad, Hercules, CA, USA).
Cell Transfection
Circ-ITCH overexpression plasmid pcDNA3.1-circ-ITCH and pcDNA3.1 vector were purchased from GenePharma (Shanghai, China). Short hairpin RNA (shRNA) targeting SASH1 (shSASH1), miR-106a-5p mimics, and corresponding negative controls were purchased from GeneCopoeia (Guangzhou, China). Cell transfection was performed using Lipofectamine 2000 (Invitrogen).
CCK-8 Assay
Cell proliferation was detected using the cell counting kit-8 (CCK-8) assay. Cells were seeded in a 96-well plate at a density of 2 × 103 cells/well and cultured for different time periods. Then, CCK-8 reagents (Dojindo, Tokyo, Japan) were added to each well, and cells were further incubated for 4 h. The absorbance was measured at 450 nm using a microplate reader.
Transwell Assay
Transwell chambers (8 mm pores) were used to measure cell invasion. 1 × 105 cells in serum-free DMEM medium were added to the Matrigel-coated upper chamber and DMEM medium containing 10% FBS was added to the lower chamber. After incubation for 24 h, cells invading the lower side of the insert were fixed and stained with 0.1% crystal violet. The number of invading cells from 5 random fields was counted under a microscope.
Luciferase Reporter Assay
The potential binding sites in the circ-ITCH and SASH1 3′-untranslated region (UTR) were predicted by bioinformatics analysis. Sequences containing wild-type (Wt) or mutant-type (Mut) binding sites were constructed into the pmirGLO vector (Promega, Madison, WI, USA). For the luciferase reporter assay, the reporter vectors and miRNAs were co-transfected into cells. After 48 h, luciferase activity was measured using the dual-luciferase reporter assay system (Promega).
In Vivo Xenograft Tumor Assay
The
Statistical Analysis
Statistical analysis was performed using SPSS 17.0 software. Data were presented as means ± SD. The differences between groups were evaluated using Student’s
Results
Circ-ITCH Is Downregulated in Glioma Tissues and Cell Lines
In an attempt to explore the role of circ-ITCH in glioma, we first analyzed its expression in 48 pairs of glioma tissues and adjacent normal tissues by RT-qPCR. The results indicated that circ-ITCH had a lower expression in glioma tissues than in adjacent normal tissues (Fig. 1A). We further analyzed circ-ITCH expression in glioma cell lines. As shown in Fig. 1B, circ-ITCH expression was downregulated in U251, U87, SHG44, and A172 cell lines in comparison with the normal glial cell line HEB.
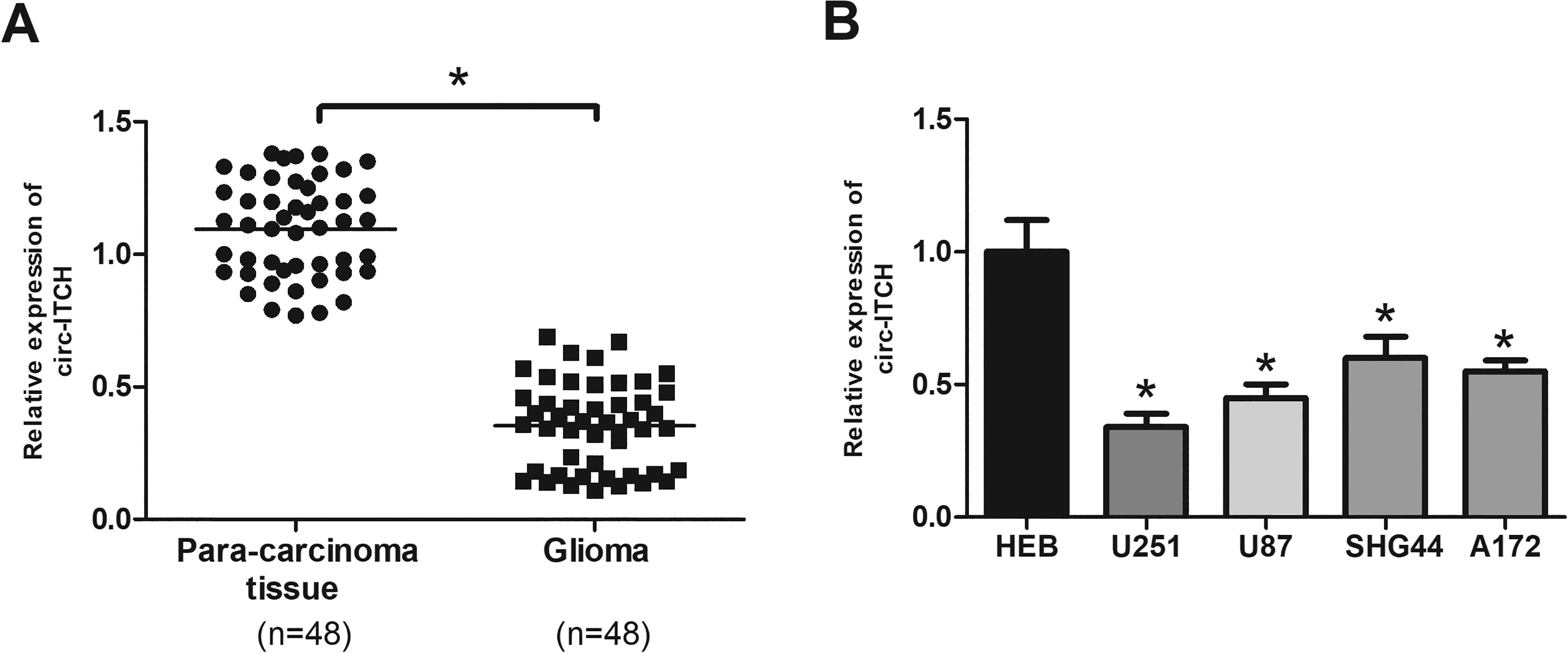
Circ-ITCH is downregulated in glioma tissues and cell lines. (A) Circ-ITCH expression in 48 paired glioma tissues and adjacent normal tissues was measured by RT-qPCR. (B) Circ-ITCH expression in glioma cell lines (U251, U87, SHG44, and A172) and normal glial cell line HEB. *
Upregulation of Circ-ITCH Inhibits the Proliferation of Glioma Cells
To investigate the effect of circ-ITCH on glioma, U251 and U87 cells were transfected with the circ-ITCH expression vector to upregulate the expression of circ-ITCH in cells (Fig. 2A and B). Then, we evaluated cell proliferation. The CCK-8 assay showed that overexpression of circ-ITCH evidently suppressed U251 cell proliferation in comparison with the control group (Fig. 2C). Consistently, the proliferative ability of U87 cells was also significantly inhibited by circ-ITCH upregulation (Fig. 2D).
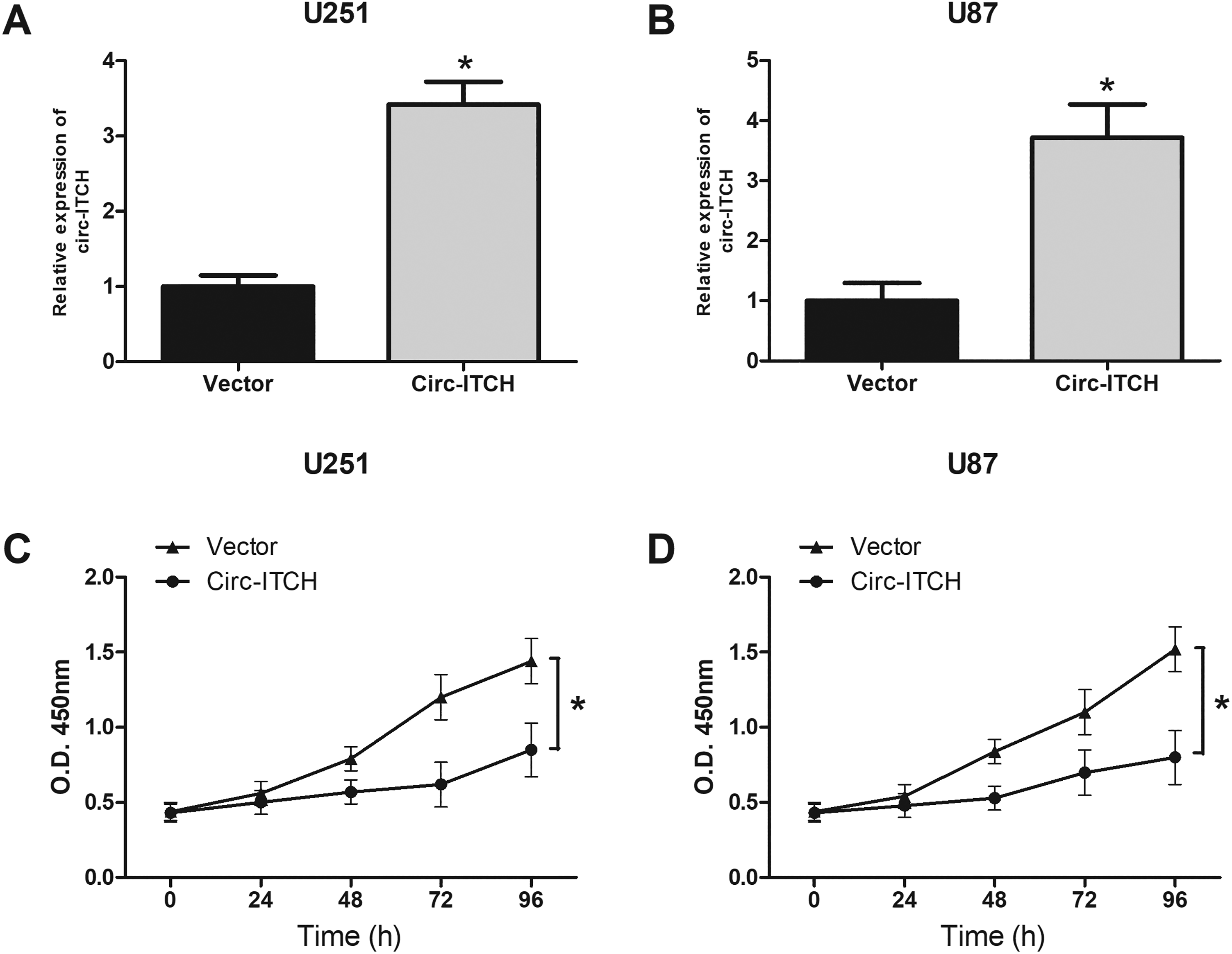
Upregulation of circ-ITCH inhibits the proliferation of glioma cells. Quantitative real-time polymerase chain reaction analysis of circ-ITCH expression in U251 (A) and U87 (B) cells after transfection. The cell counting kit-8 assay was performed to evaluate U251 (C) and U87 (D) cell proliferation. *
Upregulation of Circ-ITCH Inhibits the Invasion of Glioma Cells
Then we assessed the effect of circ-ITCH upregulation on glioma cell invasion. The transwell assay indicated that the number of invading U251 cells was markedly reduced by circ-ITCH upregulation (Fig. 3A and B). We obtained similar results in U87 cells (Fig. 3A, B).
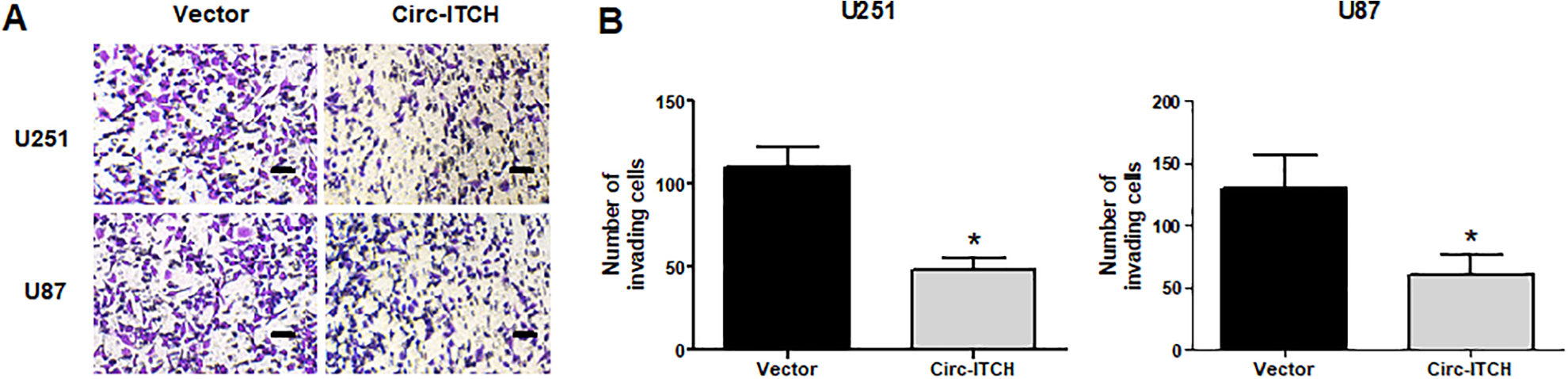
Upregulation of circ-ITCH inhibits the invasion of glioma cells. (A, B) The transwell assay was conducted to assess U251 and U87 cell invasion. *
Circ-ITCH Serves as a Sponge of MiR-106a-5p
Previous studies reported that circRNAs could interact with miRNAs and participate in pathological processes 19,20 . To explore the regulatory mechanism of circ-ITCH in glioma, we analyzed the potential targets of circ-ITCH using bioinformatics tools and identified miR-106a-5p as the most possible target of circ-ITCH (Fig. 4A). We performed the luciferase reporter assay to confirm the direct interaction between miR-106a-5p and circ-ITCH. The results showed that the relative luciferase activity of circ-ITCH-Wt was obviously decreased by miR-106a-5p mimics in U251 and U87 cells (Fig. 4B, C). In addition, upregulation of circ-ITCH obviously reduced miR-106a-5p expression in U251 and U87 cells (Fig. 4D). We also found that miR-106a-5p expression was significantly increased in glioma tissues in comparison with the normal tissues (Fig. 4E). Furthermore, Pearson’s correlation analysis showed that miR-106a-5p expression was inversely correlated with the circ-ITCH level in glioma tissues (Fig. 4F).
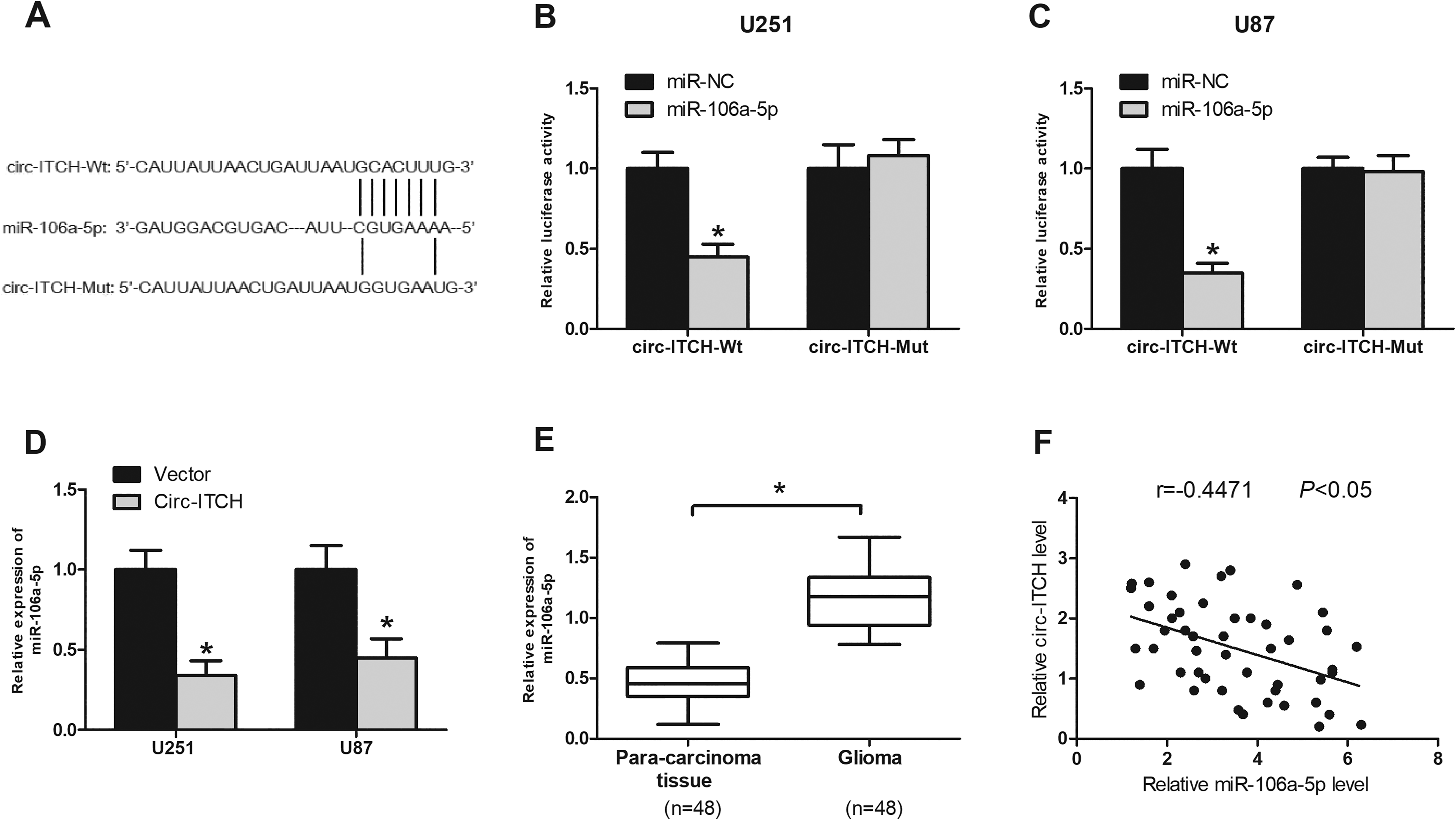
Circ-ITCH serves as a sponge of miR-106a-5p. (A) The predicted binding site between circ-ITCH and miR-106a-5p. The luciferase assay of U251 (B) and U87 (C) cells co-transfected with indicated miRNAs and luciferase reporter containing circ-ITCH-Wt or circ-ITCH-Mut. (D) MiR-106a-5p expression was decreased in U251 and U87 cells transfected with circ-ITCH. (E) Relative expression of miR-106a-5p was determined by quantitative real-time polymerase chain reaction analysis in glioma tissues. (F) The correlation between miR-106a-5p and circ-ITCH expression in glioma tissues. *
MiR-106a-5p Mimics Reverse the Effect of Circ-ITCH on Proliferation and Invasion of Glioma Cells
To better understand the link between circ-ITCH and miR-106a-5p in glioma progression, we co-transfected U251 and U87 cells with miR-106a-5p mimic and circ-ITCH overexpression vector, followed by determination of cell proliferation and invasion. The CCK-8 assay showed that upregulation of miR-106a-5p by transfection with miR-106a-5p mimics significantly abrogated the inhibitory effect of circ-ITCH on U251 and U87 cell proliferation (Fig. 5A, B). The transwell assay obtained consistent results (Fig. 5C, D).
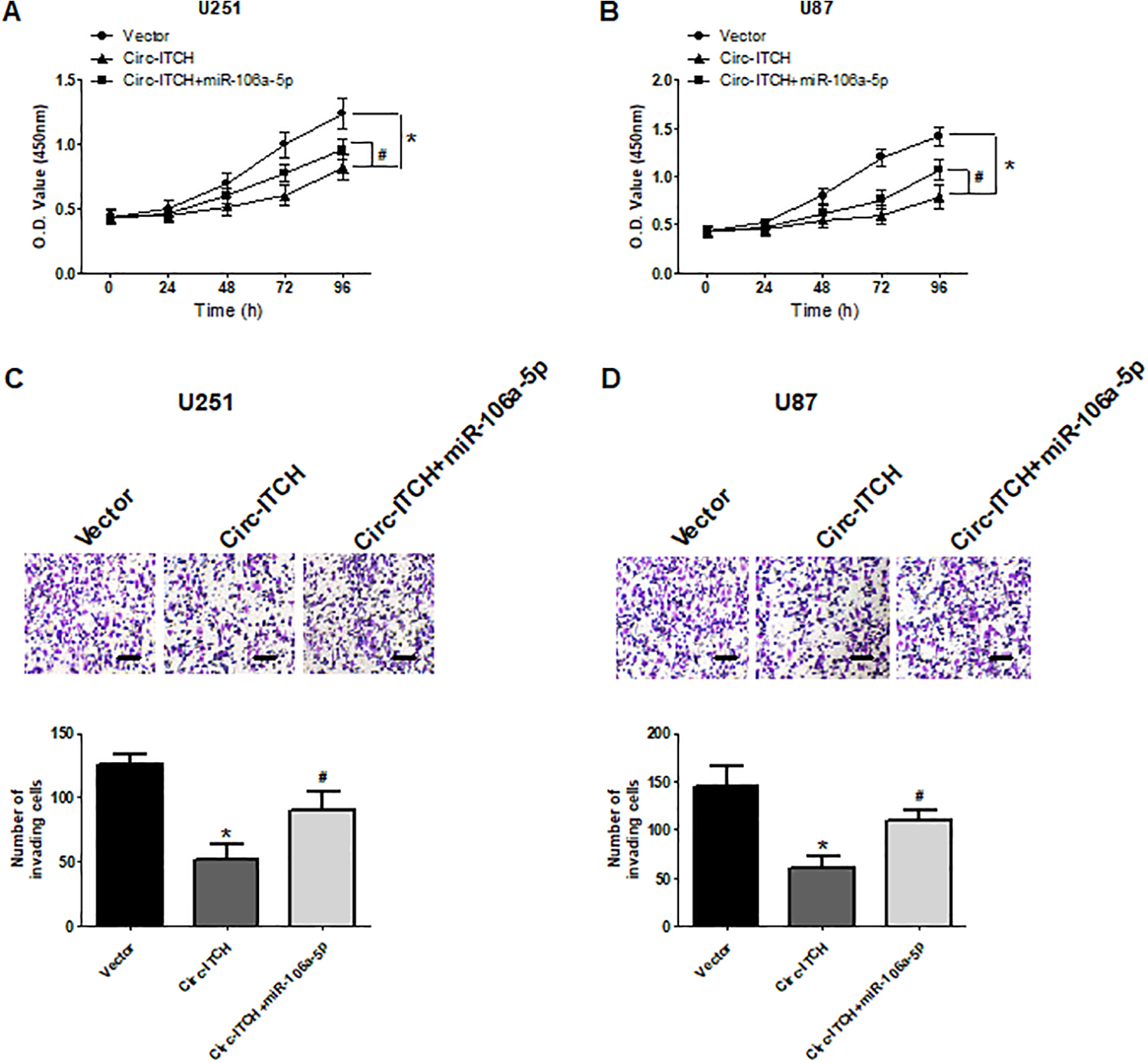
MiR-106a-5p mimics reverse the effect of circ-ITCH on proliferation and invasion of glioma cells. U251 and U87 cells were co-transfected with miR-106a-5p mimic and circ-ITCH expression vector. The cell counting kit-8 and transwell assays were conducted to determine U251 and U87 cell proliferation (A, B) and invasion (C, D), respectively. *
SASH1 is a Direct Target of MiR-106a-5p
To further explore the underlying mechanism by which miR-106a-5p exerted its effects on glioma cells, we used bioinformatics tools and identified SASH1 as a potential downstream target of miR-106a-5p (Fig. 6A). The luciferase reporter assay showed that miR-106a-5p mimics significantly reduced the luciferase activity of SASH1-Wt in U251 and U87 cells (Fig. 6B, C). In addition, the western blot analysis showed that miR-106a-5p mimics markedly decreased the protein levels of SASH1 in U251 and U87 cells (Fig. 6D, E). We also investigated SASH1 expression in glioma tissues. The results showed that SASH1 had a lower expression in glioma tissues than in the normal tissues (Fig. 6F).
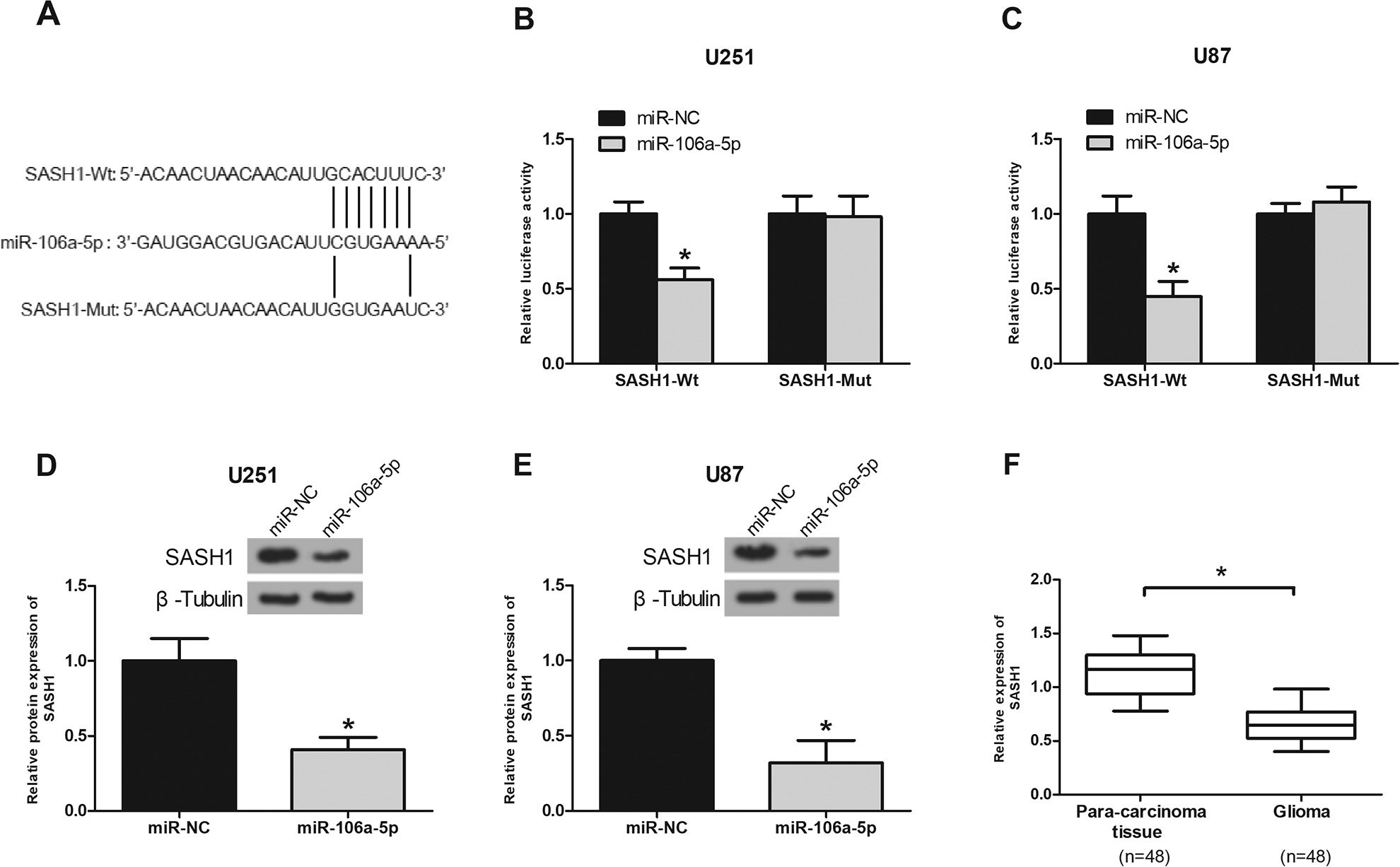
SASH1 is a direct target of miR-106a-5p. (A) The predicted binding site between SASH1 and miR-106a-5p. The luciferase assay of U251 (B) and U87 (C) co-transfected with indicated miRNAs and luciferase reporter containing SASH1-Wt or SASH1-Mut. SASH1 expression was detected in U251 (D) and U87 (E) cells using the western blot analysis after transfection with miR-106a-5p mimics. (F) SASH1 expression was measure in glioma tissues using the quantitative real-time polymerase chain reaction analysis. *
Upregulation of Circ-ITCH Inhibits Glioma Progression via Regulating the MiR-106a-5p/SASH1 Axis
To explore whether circ-ITCH regulated SASH1 expression via mediating miR-106a-5p, we investigated the effects of circ-ITCH on SASH1 expression. The results showed that circ-ITCH upregulation obviously increased the protein levels of SASH1 in U251 and U87 cells, and this effect was abolished by miR-106a-5p mimics (Fig. 7A, B). These data suggested that circ-ITCH promoted SASH1 expression by sponging miR-106a-5p. Furthermore, we confirmed the significance of SASH1 regulation by circ-ITCH in glioma cell proliferation and invasion. The results showed that the proliferative and invasive abilities of U251 and U87 cells were decreased by circ-ITCH upregulation, and these effects were reversed by SASH1 downregulation (Fig. 7C–F).
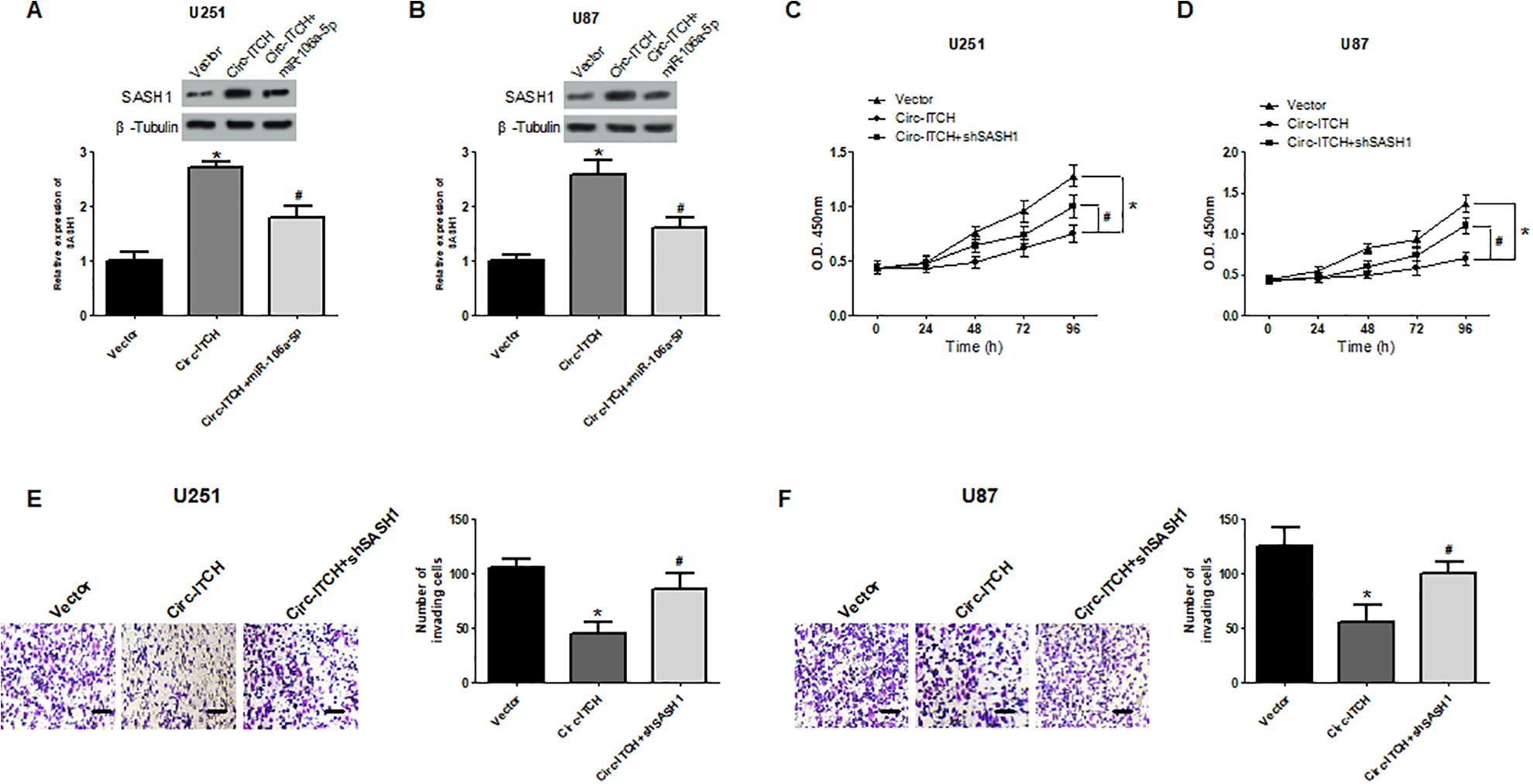
Upregulation of circ-ITCH inhibits glioma progression via regulating the miR-106a-5p/SASH1 axis. The protein levels of SASH1 in U251 (A) and U87 (B) cells were measured using the western blot analysis after co-transfection with miR-106a-5p mimic and circ-ITCH expression vector. The cell counting kit-8 and transwell assays were performed to assess U251 and U87 cell proliferation (C, D) and invasion (E, F) respectively, after different treatments. *
Upregulation of Circ-ITCH Inhibits Glioma Cell Growth In Vivo
To determine the
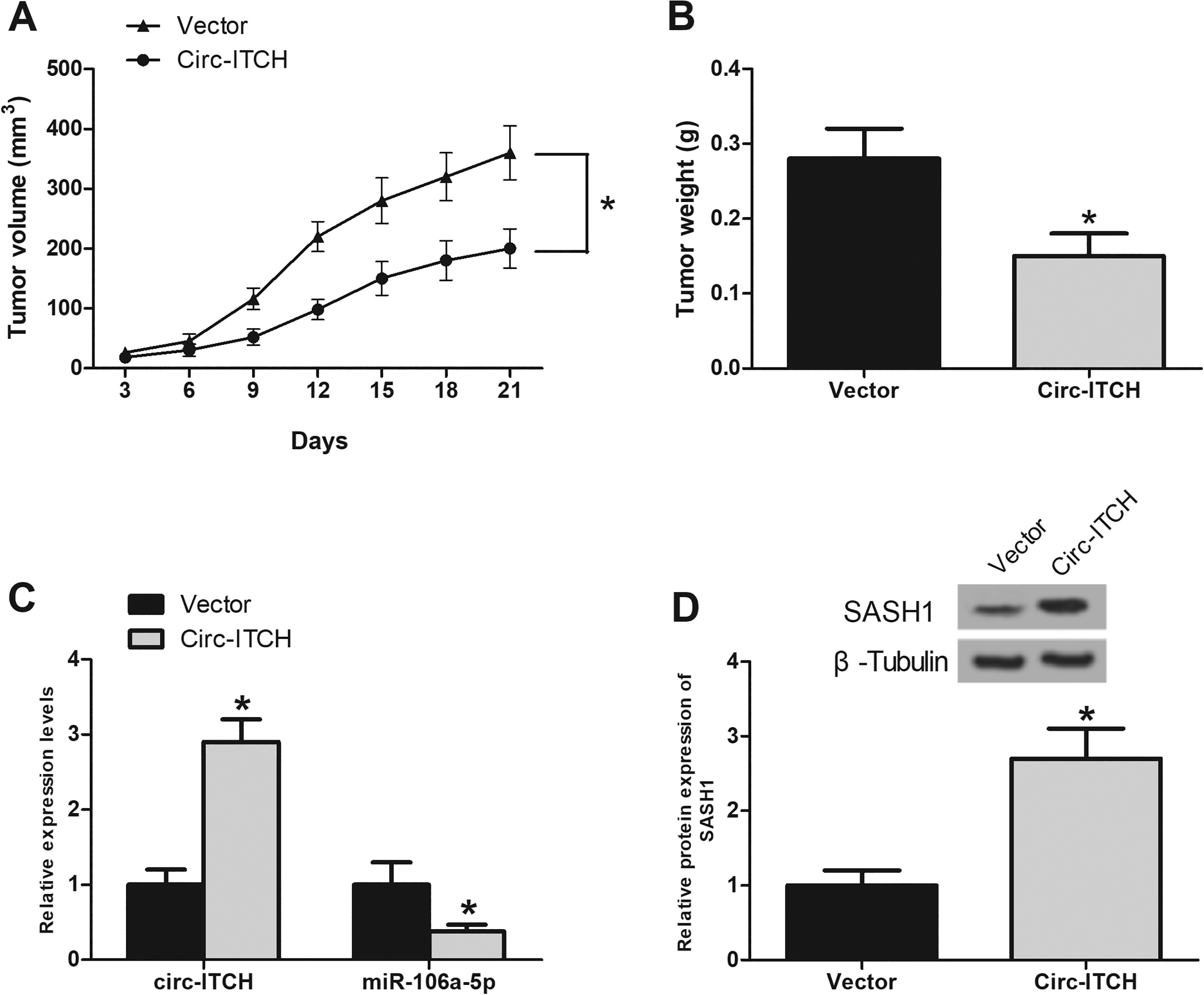
Upregulation of circ-ITCH inhibits glioma cell growth
Discussion
Previous studies have demonstrated that noncoding RNAs are important participants in the regulation of cellular functions and physiological development
21
. CircRNAs belong to a novel family of noncoding RNAs and have been reported to play a significant role in cancer progression
9,22,23
. For example, Wang et al. showed that circ-ITCH was lowly expressed in thyroid cancer cells, and its overexpression inhibited cell proliferation and invasion
In the present study, we found that circ-ITCH was downregulated in glioma tissues and cell lines. Function assays showed that upregulation of circ-ITCH inhibited glioma cell proliferation and invasion. Moreover, our xenograft tumor assay demonstrated that circ-ITCH overexpression suppressed glioma cell growth
CircRNAs have been reported to regulate diverse physiological and pathological processes by serving as sponges of miRNAs 19,20 . For example, Liang et al. showed that circ-ABCB10 acted as the sponge of miR-1271 to promote breast cancer progression 26 . Han et al. found the regulatory role of the circ-BANP/miR-503/LARP1 pathway in lung cancer 27 . In this study, we showed that circ-ITCH could directly bind to miR-106a-5p. In diverse types of cancers, miR-106a-5p is generally documented as a crucial tumor suppressor. For example, He et al. showed that miR-106a-5p could suppress the proliferation, migration, and invasion of osteosarcoma cells by targeting HMGA2 28 . Conversely, Li et al. demonstrated that miR-106a-5p facilitated glioblastoma cell proliferation and invasion by activating the Wnt/β-catenin pathway 29 . Thus, miR-106a-5p plays a significant role in tumor development. In the present study, RT-qPCR analysis showed that miR-106a-5p expression was markedly reduced in glioma cells after circ-ITCH upregulation. We also found that miR-106a-5p expression was obviously increased in glioma tissues. Moreover, we found SASH1 acting as a downstream target of miR-106a-5p. SASH1, belonging to the family of signal adapter proteins, is necessary for interaction between proteins and formation of signaling complexes 30,31 . Recent studies have indicated that SASH1 functions as a tumor suppressor in a variety of cancers. For example, Chen et al. reported that forced expression of SASH1 suppressed cervical cancer cell proliferation and invasion via regulating the FAK pathway 32 . Zong et al. showed that SASH1 overexpression inhibited TGF-β1-induced EMT in gastric cancer cells 33 . In this study, we showed that SASH1 had a lower expression in glioma tissues than in normal tissues. Besides, SASH1 expression was significantly increased in glioma cells after circ-ITCH upregulation, and this effect was abolished by miR-106a-5p mimics. In addition, rescue experiments showed that the proliferative and invasive abilities decreased by circ-ITCH upregulation could be reversed by SASH1 downregulation in glioma cells. These findings suggested that circ-ITCH might affect glioma progression by regulating the miR-106a-5p/SASH1 pathway.
In conclusion, we identified that circ-ITCH inhibited glioma cell proliferation and invasion through sponging miR-106a-5p and positively regulating SASH1 expression. Our study provided a new molecular target for glioma treatment.
Footnotes
Author Contributions
WC and MW were responsible for study design. STC contributed to data analysis. YZ and ZL conducted experiments. LSL wrote the manuscript. All authors revised and approved the manuscript.
Ethical Approval
Ethical approval was obtained for all experimental procedures by the Ethics Committee of Nanjing Medical University (Nanjing, China).
Statement of Human and Animal Rights
All procedures involving human subjects and animals in this study were approved by the Ethics Committee of Nanjing Medical University.
Statement of Informed Consent
All participants in this study have provided written informed consent.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
