Abstract
Cullin4A (CUL4A) is implicated in many cellular events including cell survival and growth. However, the specific function and underlying mechanisms of CUL4A in cancer invasion have not yet been elucidated. In this work, we were focused on investigating the role of CUL4A in human osteosarcoma (OS). The expression level of CUL4A was evaluated by immunohistochemical (IHC) assay in human OS tissues. Lentivirus-mediated CUL4A shRNA (Lv-shCUL4A) constructed by us was transfected into OS cells for assessing its effects on cell proliferation and invasive potential, respectively detected by MTT and Transwell assays. It was demonstrated that the expression of CUL4A protein was markedly increased in OS tissues compared with the adjacent non-cancerous tissues (ANCT) (57.8% vs. 25.6%,
Introduction
Osteosarcoma (OS) is the most common primary bone tumor in adolescents and young adults. 1 In spite of the advances in therapeutic strategies including surgery, chemotherapy, radiotherapy, and biological treatment, the overall survival of OS patients remains not optimistic. 2 Therefore, it is indispensable to identify predictive biomarkers and develop novel treatment strategies for the treatment of OS.
Cullin4A (CUL4A) encodes a core subunit of ubiquitin ligase that targets proteins for ubiquitin-mediated p27 degradation and is involved in regulation of cell cycle exit and differentiation during erythropoiesis. 3 CUL4A is essential for hematopoietic stem cell engraftment, self-renewal, and survival, and its deficiency causes cell apoptosis, proliferation defects, and genome instability.4–6 Some studies have shown that Cul4A is overexpressed in mesothelioma cells, 7 breast cancer, 8 and multiple myeloma, 9 indicating an oncogenic role in the pathogenesis of these cancers.
Functionally, CUL4A promotes pituitary cell proliferation, migration, and invasion, leads to pituitary tumor development and progression, 10 but represses apoptosis and DNA damage through regulation of the apoptosis-related proteins (Bcl-2, caspase-3, p53, and p73) and cell cycle regulator (p27).10–12 In addition, CUL4A induces tumorigenesis of skin cancer by inhibition of DNA repair and accelerating S phase entry, 13 and results in inactivation of RASSF1A and promoting cell cycle progression, 14 suggesting that CUL4A may play a critical role in regulation of some biological processes. 15
Up to now, few studies report the link between CUL4A expression and OS. To confirm the function and molecular mechanisms of CUL4A in OS cells, using a tissue microarray procedure, we examined the expression of CUL4A in OS tissues by IHC assay, and constructed Lv-shCUL4A vector for investigating its effects on biological behaviors of OS cells. We hypothesized that knockdown of CUL4A might suppress growth and invasion of GC cells.
Materials and methods
Materials
The human OS MG-63 cell line used in the experiments was from Institute of Biochemistry and Cell Biology (Shanghai, PR China). Lentivirus mediated Lv-shCUL4A vector and negative control (NC) vector were from Genechem (Shanghai, PR China). The primer of p27 was synthesized by ABI (Framingham, MA, USA). The tissue microarray of human OS was purchased from the branch of Biomax (Xi’ an, PR China). All antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA).
Drugs and reagents
Dulbecco’s Modified Eagle medium (DMEM) and fetal bovine serum (FBS) were from Thermo Fisher Scientific Inc (Waltham, MA, USA); TRIzol Reagent and Lipofectamine 2000 were from Invitrogen (Carlsbad, CA, USA); M-MLV Reverse Transcriptase was from Promega (Madison, WI, USA); SYBR Green Master Mixture was from Takara (Otsu, Japan). ECL-PLUS/Kit was from GE Healthcare (Piscataway, NJ, USA).
Clinical samples and data
The tissue microarray was prepared for immunohistochemical (IHC) test. Human OS tissues and the ANCT were collected in a total of 45 consecutive cases of OS in our hospital from October 2008 to February 2014. This study was approved by Medical Ethics Committee of Shanghai Jiao Tong University and written informed consent was obtained from the patients before sample collection. Two pathologists checked these samples.
IHC staining
Anti-CUL4A antibody was used for IHC test of the expression of CUL4A protein in this tissue microarray. IHC examination was performed on 3 mm thick sections. For anti-CUL4A IHC, unmasking was performed with 10 mM sodium citrate buffer, pH 6.0, at 90°C for 30 min. Sections were incubated in 0.03% hydrogen peroxide for 10 min at room temperature, to remove endogenous peroxidase activity, and then in blocking serum for 30 min at room temperature. Anti-CUL4A antibody was used at a dilution of 1:200. The antibody was incubated overnight at 4°C. Sections were then washed three times for 5 min in PBS. Finally, staining was developed using diaminobenzidine substrate, and sections were counterstained with hematoxylin. Normal serum or PBS was used to replace anti-CUL4A antibody in negative control.
Quantification of protein expression
The expression of CUL4A was semi-quantitatively assessed as the total immunostaining scores, calculated as the product of a proportion score and an intensity score. The proportion and intensity of the staining was estimated by two observers. The proportion score indicated the fraction of positive staining cells (score 0, <5%; score 1, 5–10%; score 2, 10–50%; score 3, 50–75%; score 4, >75%), and the intensity score represented the staining intensity (score 0, no staining; score 1, weak positive; score 2, moderate positive; score 3, strong positive). Finally, a total expression score was given in the range of 0–12. Based on this analysis, CUL4A was considered as negative expression if the score <2, and positive expression if the score was ⩾2.
Cell culture and transfection
OS cells were cultured in DMEM medium supplemented with 10% heat-inactivated FBS, 100 U/mL of penicillin and 100 μg/mL of streptomycin. They were all placed in a humidified atmosphere containing 5% CO2 at 37°C. OS cells were replated at 5×105 cells/well in 24-well plates with serum-free growth medium. Once OS cells arrived at 50% confluence, they were transfected with Lv-shCUL4A and cultured at 37°C and 5% CO2 for 4 h. Then supernatant was discarded and serum containing growth medium was added. Positive stable transfectants were expanded for further study.
Quantitative real-time PCR
To quantitatively determine the mRNA expression of CUL4A, real-time PCR was performed. Total RNA of each clone was extracted with TRIzol according to the manufacturer’s protocol. Reverse-transcription was carried out using M-MLV and cDNA amplification was carried out using SYBR Green Master Mix kit according to the manufacturer’s protocol. CUL4A gene was amplified using specific oligonucleotide primer and human GAPDH gene was used as an endogenous control. Three separate experiments were performed for each clone.
Western blot assay
OS cells were harvested and extracted using lysis buffer. Cell extracts were boiled for 5 min in loading buffer and then equal amount of cell extracts were separated on 15% SDS-PAGE gels. Separated protein bands were transferred into polyvinylidene fluoride (PVDF) membranes and the membranes were blocked in 5% skim milk powder. The primary antibodies against p27, MMP-2, and p53 were diluted according to the instructions of antibodies and incubated overnight at 4°C. The relative protein level was normalized to β-actin concentration. Three separate experiments were performed for each clone.
Cell proliferation assay
OS cells transfected with Lv-shCUL4A were incubated in 96-well-plates at a density of 1×105 cells per well with DEME medium supplemented with 10% FBS. Cells were pretreated with 20 μL MTT dye and then incubated with 150 μL of DMSO. The color reaction was measured at 570 nm with enzyme immunoassay analyzer.
Transwell invasion assay
Transwell filters were coated with matrigel on the upper surface of a polycarbonic membrane. Harvested cells in serum-free DMEM were added into the upper compartment of the chamber. A total of 200 µL conditioned medium derived from NIH3T3 cells was used as a source of chemoattractant, and was placed in the bottom compartment of the chamber. After 24 h incubation at 37°C with 5% CO2, the medium was removed from the upper chamber. The non-invaded cells on the upper side of the chamber were scraped off with a cotton swab. The cells that had migrated from the matrigel into the pores of the inserted filter were fixed with 100% methanol, stained with Hematoxylin, and mounted and dried at 80°C for 30 min. Each assay was repeated three times.
Cell apoptosis analysis
To detect cell apoptosis, OS cells were trypsinized, washed with cold PBS, and resuspended in binding buffer according to the instruction of the apoptosis kit. FITC-AnnexinV and PI were added to the fixed cells for 20 min in darkness at room temperature. Then, Annexin V binding buffer was added to the mixture before the fluorescence was measured on FAC sort flow cytometer. The cell apoptosis was analyzed using the Cell Quest software (Becton Dickinson, East Rutherford, NJ, USA). Three separate experiments were performed for each clone.
Statistical analysis
Statistically significant difference was analyzed by SPSS version 20.0. The difference in each group was considered as significance by Kruskal-Wallis H test and ANOVA analysis of variance.
Results
The expression of CUL4A in human OS
The expression of CUL4A in human OS tissues was examined by IHC staining. As shown in Figure 1, the staining of CUL4A in OS tissues was found with strong reactivity rate compared to that in the ANCT, indicating the increased expression of CUL4A in human OS tissues (

The expression of CUL4A in human OS (×200). IHC staining was used to detect the expression of CUL4A in human OS, indicating increased expression of CUL4A in OS tissues compared to that in the ANCT.
The expression of CUL4A in human OS.

ANCT, adjacent non-cancer tissue; OS, osteosarcoma.
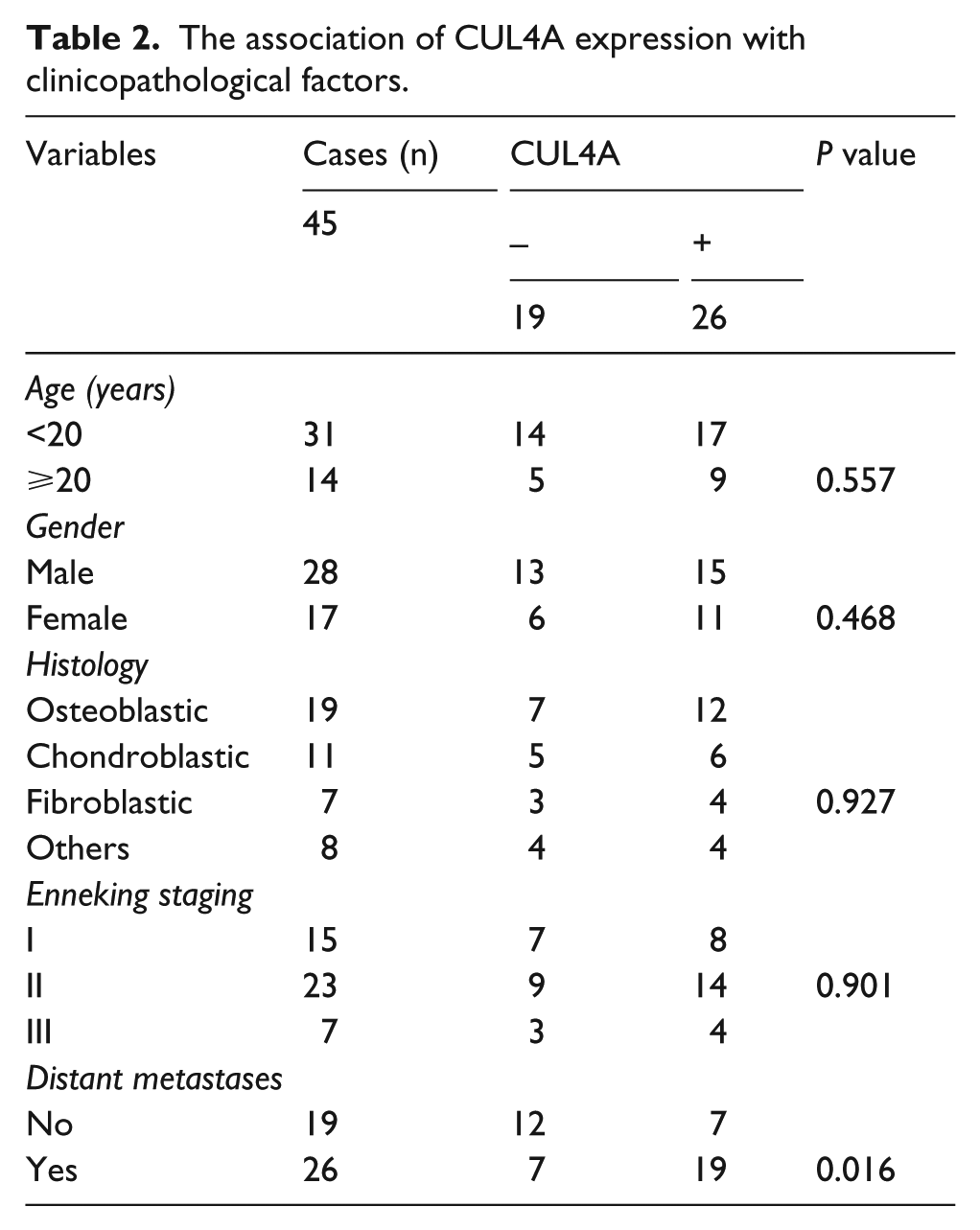
Furthermore, the association between CUL4A expression and the clinicopathologic characteristics of OS patients was analyzed. As shown in Table 2, increased expression of CUL4A was positively associated with the distant metastases in OS patients (
The association of CUL4A expression with clinicopathological factors.
The effect of CUL4A knockdown on cell proliferation
After OS cell line (MG-63) was transfected with Lv-shCUL4A for 24 h, MTT assay was carried out to determine cell proliferative activities. It was found that knockdown of CUL4A significantly reduced cell proliferative activities in a time-dependent manner in comparison with the NC and CON groups (Figure 2a, **

The effect of CUL4A silencing on cell proliferation. (a) MTT assay was used to evaluate cell proliferative activities, showing that silencing of CUL4A gene significantly decreased cell proliferation in a time-dependent manner in comparison with the NC and CON groups (**
The effect of CUL4A knockdown on cell invasion
After OS cells were transfected with Lv-shCUL4A for 24 h, Transwell assay was performed to assess cell invasive capability. It was found that knockdown of CUL4A markedly decreased cell invasive potential in comparison with the NC and CON groups (Figure 3a, b, **

The effect of CUL4A silencing on cell invasion (×200). (a, b) Transwell assay was performed to determine cell invasive capability, revealing that knockdown of CUL4A markedly suppressed cell invasive potential in comparison with the NC and CON groups (**
The effect of CUL4A knockdown on cell apoptosis
After OS cells were transfected with Lv-shCUL4A for 24 h, flow cytometry analysis was performed to determine cell apoptosis. It was found that knockdown of CUL4A significantly induced cell apoptotic indexes in comparison with the NC and CON groups (Figure 4a, b, **

The effect of CUL4A silencing on cell apoptosis. (a, b) Flow cytometry analysis was used to determine cell apoptosis, indicating knockdown of CUL4A significantly augmented cell apoptotic indexes in comparison with the NC and CON groups (**
Discussion
CUL4A has been verified to be overexpressed in breast cancers, hepatocellular carcinomas, squamous cell carcinomas, adrenocortical carcinomas, childhood medulloblastomas, and malignant pleural mesotheliomas. 13 Overexpression of CUL4A is involved in proliferation and cell cycle progression of breast cancer 16 and associates with the invasive potential and poor outcome in ovarian carcinoma. 17 But, few studies show that high expression of CUL4A predicts improved progression-free survival and overall survival of breast cancer. 18 In the present study, we further evaluated the expression of CUL4A in OS tissues and found that upregulation of CUL4A, mainly localized in the nucleus, was positively correlated with distant metastases in OS patients but was negatively associated with p27 expression, 10 suggesting that nuclear accumulation of CUL4A might promote the tumorigenesis of OS.
Other studies have indicated that CUL4A contributes to cell growth, epithelial-mesenchymal transition, and metastasis of breast cancer through regulation of ZEB1 expression and antitumor immune response,19,20 enhances tumor growth and sensitizes lung cancer cells to erlotinib via regulation of EGFR 21 and determines the response to thalidomide treatment in prostate cancer, 22 involving in the multi-drug resistance in cancer. 23 Inhibition of CUL4A expression can augment DNA damage response and fight against the progression of skin cancer. 24 To clarify the function of CUL4A in cancer, using the loss-of-function experiment, we found that knockdown of CUL4A suppressed the growth and invasion and induced cell apoptosis in OS cells. These studies suggest that CUL4A may be the potential therapeutic target for the treatment of OS.
Bcl-2, initially discovered at the t14,18 chromosomal breakpoint in follicular lymphomas, inhibits cell apoptosis, and plays an important role in cell development and maturation of GC. 25 Expression of p27 is found to be implicated in cell proliferation and differentiation of GC and serves as a prognostic marker in GC patients. 26 The expression of MMP-2 exerts an important feature of tumor cell escape from the primary neoplasm and entry into the secondary body site and determines the invasive capacity of a cancer cell. 27 Mechanistically, CUL4A modulates the expression of p53, p27, and Bcl-2 in pituitary tumor cells.10,11 Similarly, we found that knockdown of CUL4A downregulated the expression of MMP-2 and upregulated the expression of p27 and p53 in OS cells, suggesting that CUL4A might be involved in the development of OS cells via regulation of the expression of p27, p53, and MMP-2.
In conclusion, our present findings indicate that high expression of CUL4A is positively correlated with tumor metastases in OS patients, and silencing of CUL4A gene suppresses the growth and invasion and indices cell apoptosis in OS cells, suggesting that CUL4A may serve as a potential therapeutic target for the treatment of GC.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
