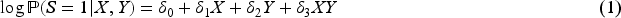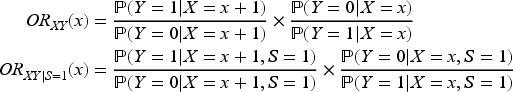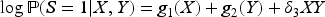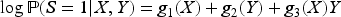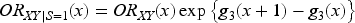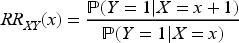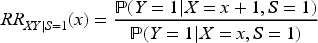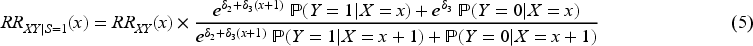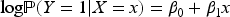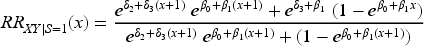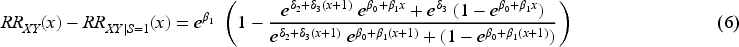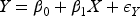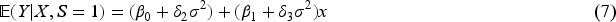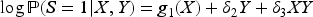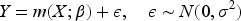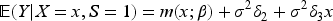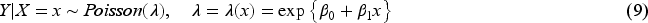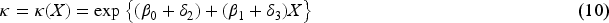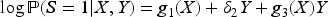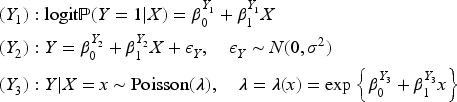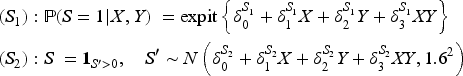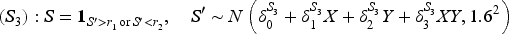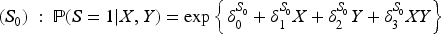Abstract
Collider bias occurs when conditioning on a common effect (collider) of two variables
Keywords
Introduction
Collider bias is a common concern in epidemiological studies. When exploring the association between an exposure
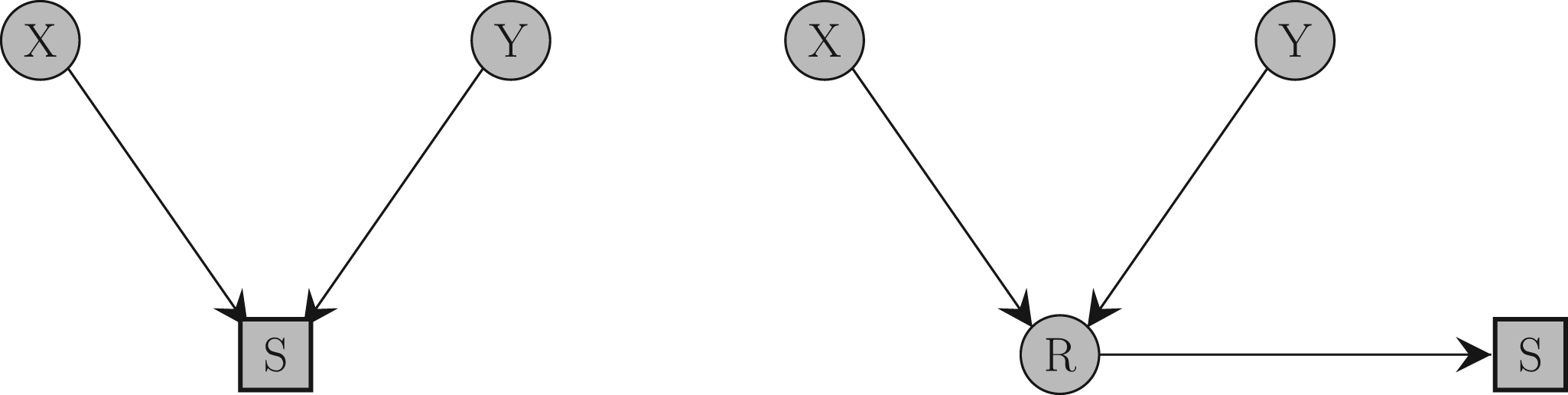
Causal diagram illustrating how collider bias occurs. Two variables
In this article, we focus on collider bias induced when an analysis is restricted to a single level of a binary collider variable
It has long been recognized in the literature that this type of collider bias relates to interactions between the exposure and the outcome on the log-additive scale in their effects on the collider.6,7 Consider the following log-additive model for the collider:
In this article, we expand the literature by considering a wider range of outcome variables, including count outcomes (Poisson regression) and continuous outcomes (linear regression) in addition to binary ones. We also place no conditions on the form of the exposure variable. We show that collider bias will not affect the exposure–outcome association when the exposure and outcome do not interact in their effects on the collider, that is, when
These results have important implications for methods attempting to adjust for collider bias, such as inverse probability weighting (IPW) (Seaman and White
18
). Implementing IPW requires the specification of a model for the collider
The rest of this article is organized as follows. In Section 2, we review the relevant literature. We then consider regression models for a binary outcome
Statement of the problem
As in Section 1, suppose that the objective is to investigate the association between an exposure
As in the previous section, we let the collider
We study collider bias separately for binary, continuous and count outcome variables. For binary outcome variables, we examine collider bias in odds ratios, logistic regression coefficients and risk ratios. For continuous outcomes, we quantify collider bias in linear regression coefficients, and for count outcome variables, we investigate collider bias in the log rate ratio parameters of a Poisson regression model. We work under the assumption that the collider
The relationship between collider bias and exposure–outcome interactions
Binary outcome—Collider bias on the odds ratio scale
Consider first the case of a binary outcome, which has received the most attention in the literature (e.g. Bartlett et al.
8
and Jiang and Ding
15
). Some of these papers also restricted the exposure to be binary; here, we do not place any assumptions on the type of the exposure variable. Our main assumption is that the exposure and outcome affect the collider on the log-additive scale, as in (1). Let
In Supplemental Section 2.1 (see also Jiang and Ding
15
), we prove that
Finally, let the variable
We now explore the magnitude of collider bias on the risk ratio scale. Once again, we assume that the collider
In Supplemental Section 2.2, we explore collider bias on the risk ratio scale under a logistic regression model for the outcome and obtain results similar to those reported here.
We now turn our attention to continuous outcome variables, and assume that the outcome is distributed according to the linear regression model
As with binary outcomes, the above derivation allows for a non-linear exposure-collider effect,
Finally, we consider the case of a count outcome variable distributed according to the Poisson regression model
In Supplemental Section 2.4, we show that if expression (9) holds, then
As with binary and continuous outcome variables, a few extensions of this result are possible, including to analyses with a non-linear exposure-collider association: if expression (9) holds and
Study design
So far we have assumed that the collider
Our analysis here is asymptotic in nature: the aim is to obtain asymptotic results about the relationship between the limiting value of the MLE of
We considered nine data generating mechanisms, obtained by combining three different outcome models and three different models for the selection indicator
The parameters of the three outcome models and the three data-generating models for
There were two parameters to be varied in our numerical asymptotic study: the strength of the exposure–outcome interaction
For each of the nine data generating mechanisms and each set of parameter values, we generated a single dataset of size
Data were generated and models were fitted using Magnitude of collider bias induced in the exposure–outcome regression coefficient by restricting the analysis to selected (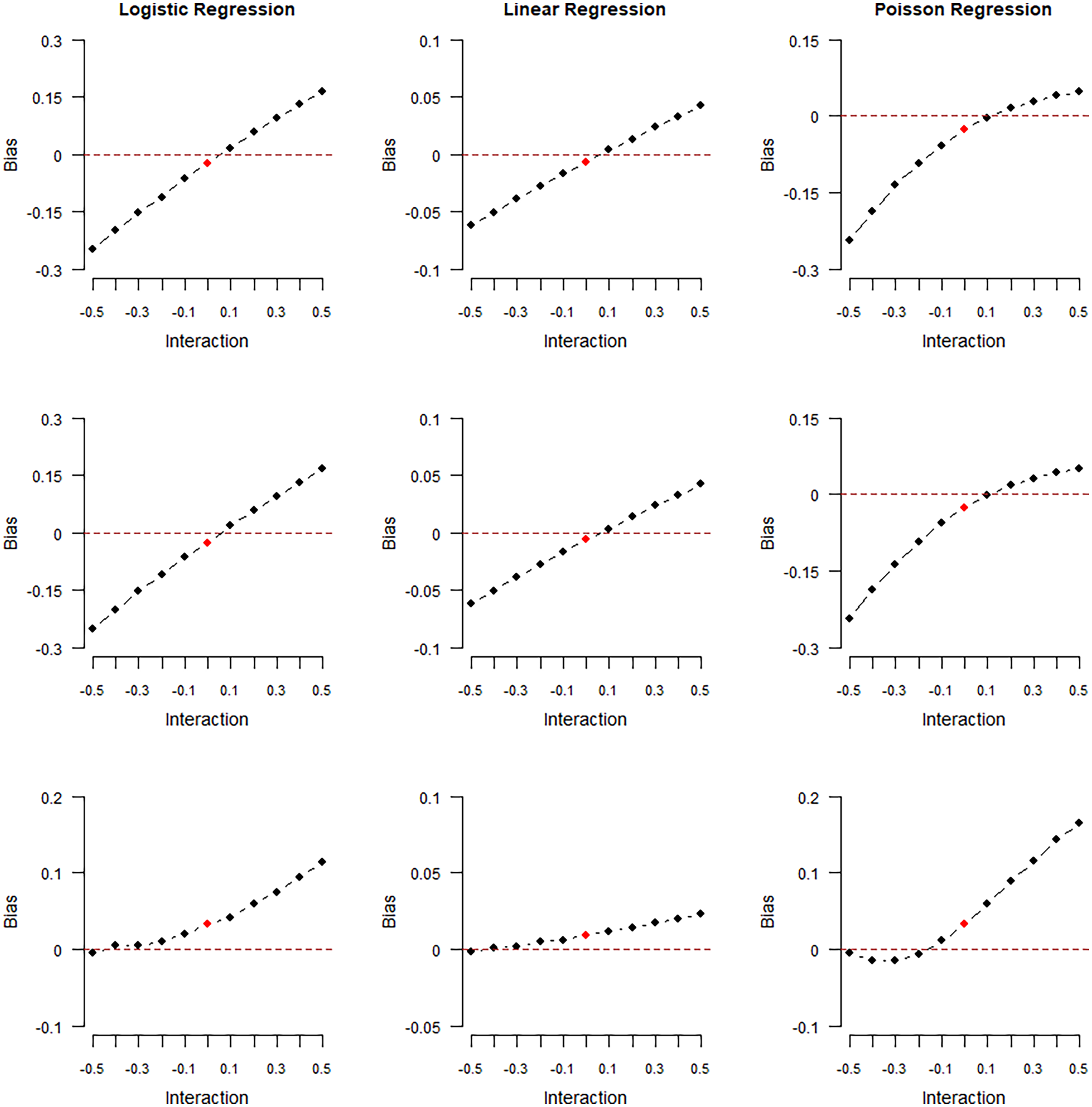
The results of our first numerical experiment are shown in Figures 2 and 3 and reported in the tables in Supplemental Section 3. In the plots of Figure 2, the collider
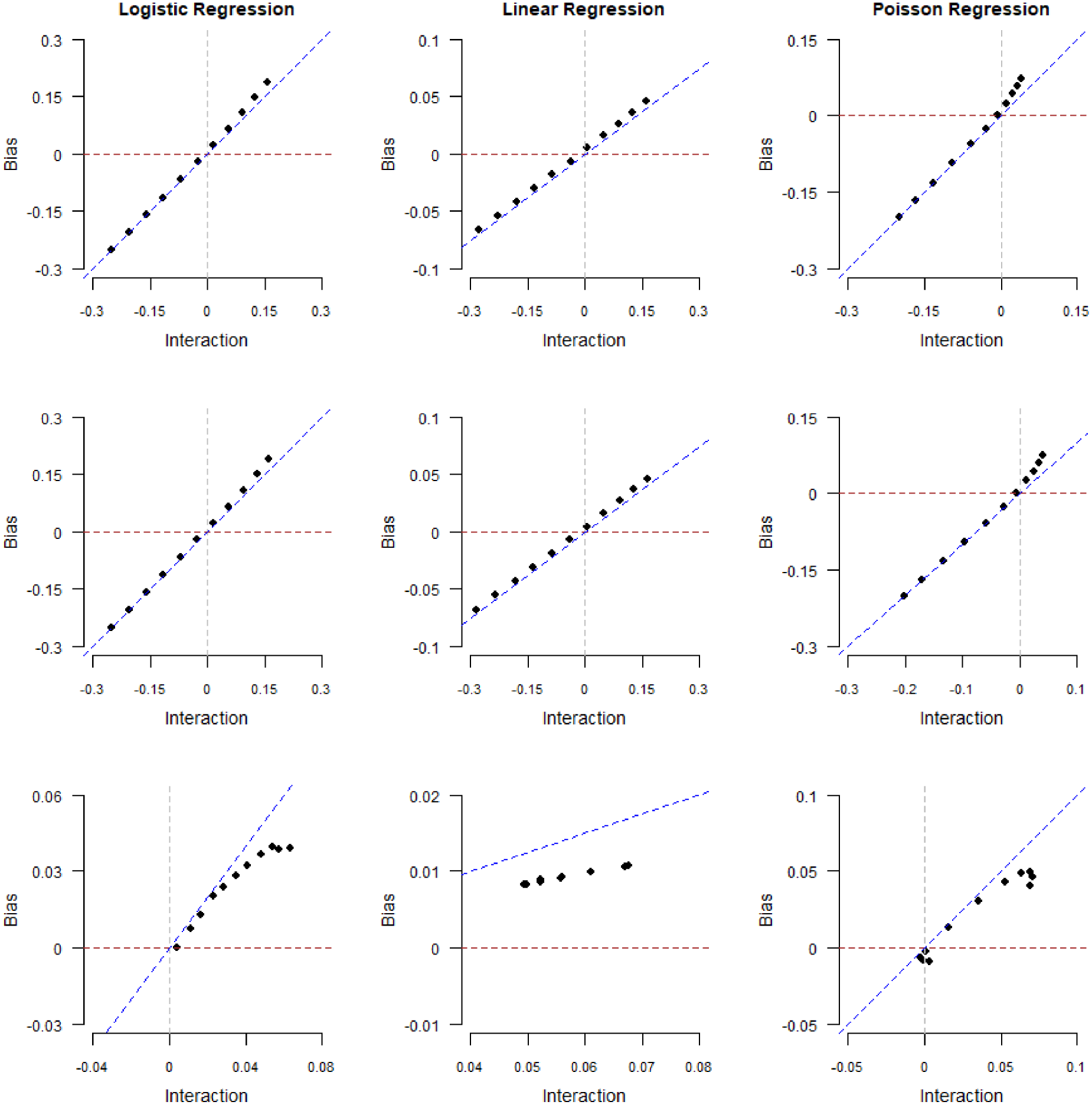
Magnitude of collider bias induced in the exposure–outcome regression coefficient by restricting the analysis to selected (
For some of the models considered here, it is possible to derive analytic expressions for collider bias using arguments similar to those in the previous section. As an example, in Supplemental Section 2.5, we obtain an expression for the collider bias in the exposure–outcome regression coefficient of a logistic regression model when the collider
In Figure 3, we plot collider bias against the limiting values
Note that the linear pattern of collider bias presented in Figure 3 only occurs when the exposure
Results from our second numerical experiment are shown in Figures 4 and 5. In Figure 4, we plot the magnitude of collider bias induced in exposure–outcome regression coefficients against the true value of the exposure–outcome interaction in models (
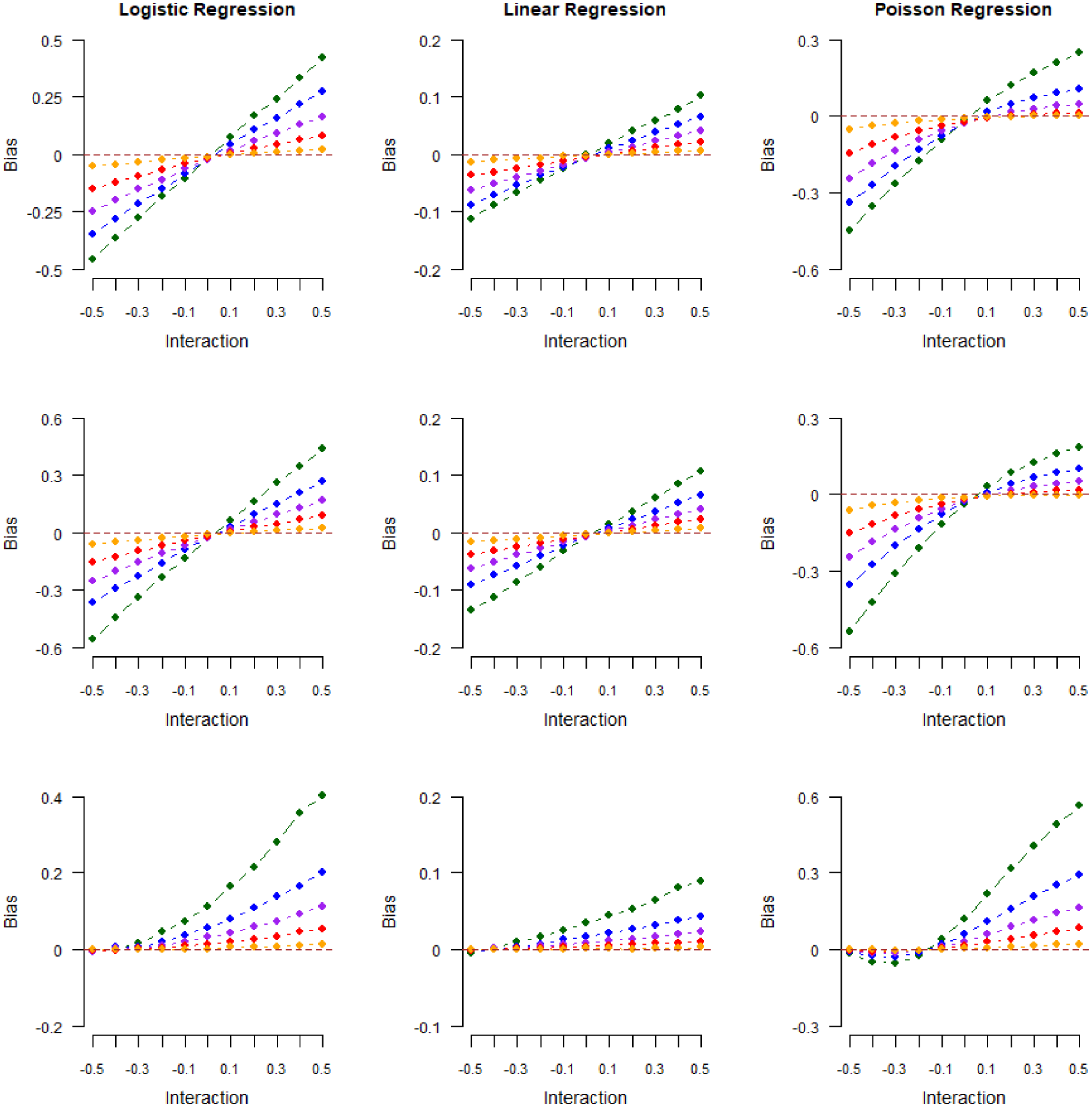
Magnitude of collider bias induced in the exposure–outcome regression coefficient by restricting the analysis to selected (
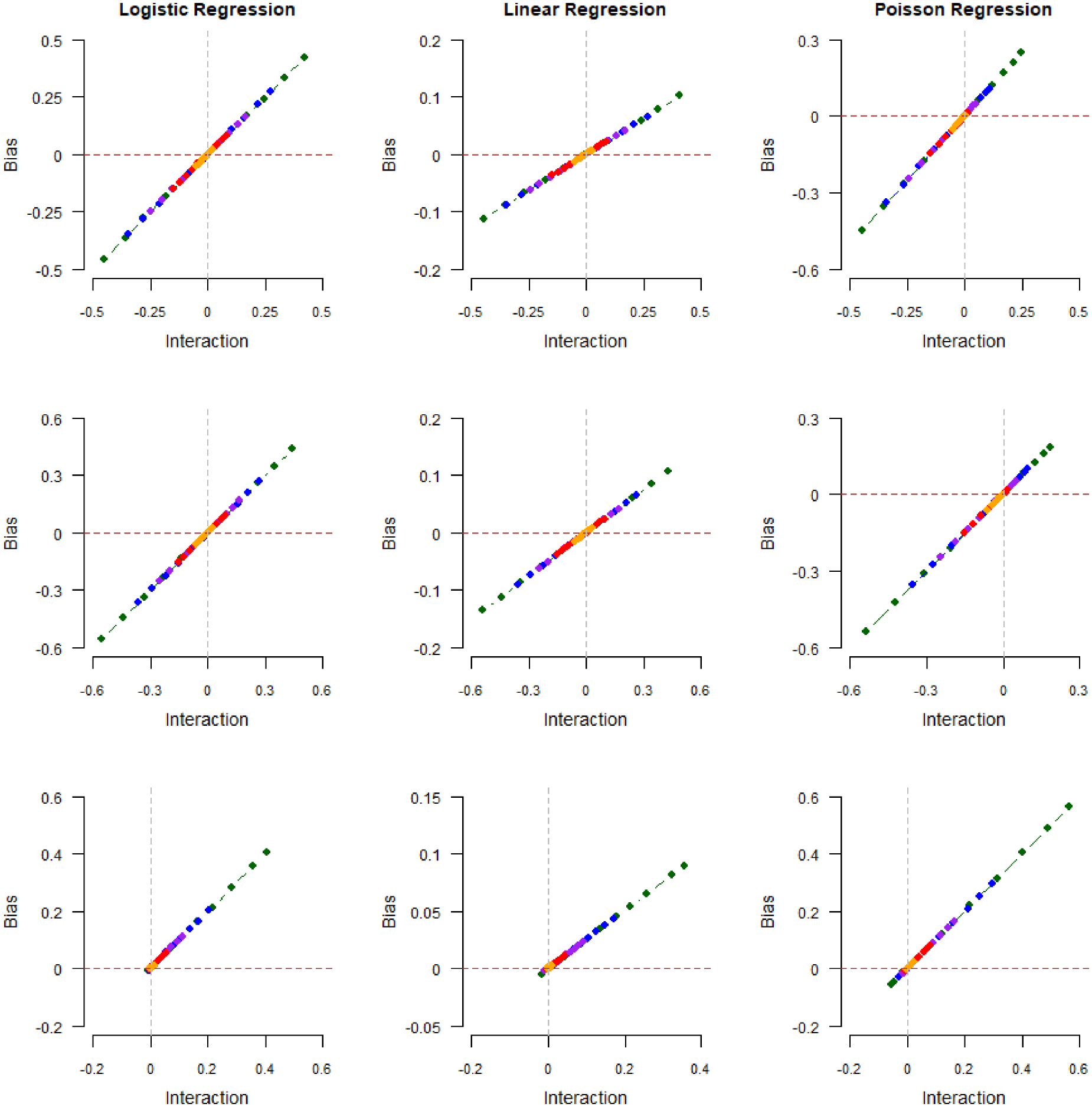
Magnitude of collider bias induced in the exposure–outcome regression coefficient by restricting the analysis to selected (
In summary, our results suggest that there exist scenarios in which the linear relationship between the magnitude of collider bias and the strength of exposure–outcome interaction on the log-additive scale may hold true even if the collider
Data and methods
We also conducted a real-data analysis using data from the Avon Longitudinal Study of Parents and Children (ALSPAC, Boyd et al.
26
and Fraser et al.
27
). ALSPAC is a longitudinal population-based study that recruited pregnant women residing in Avon, UK, with expected delivery dates between 1 April 1991 and 31 December 1992. The study included
The aim of our analysis is to demonstrate, using real data, the relation between collider bias and exposure–outcome interactions in a log-additive model for selection. To do so, we investigated associations of six maternal traits with sex of offspring at birth (hereafter referred to as offspring sex). These maternal traits included age at delivery, highest educational qualification held, pre-pregnancy body mass index (BMI), depression status, pre-pregnancy smoking, and gestational age. Since offspring sex is determined randomly at conception and unaffected by environmental exposures, one would expect its associations with maternal traits to be null in the absence of bias (perhaps with the exception of gestational age, Divon et al.
28
). We obtained maternal trait and offspring sex data for all ALSPAC families; these data were treated as the “complete sample” for the purposes of our analysis. We also obtained participation data for two follow-up stages of ALSPAC: the “Teen Focus 4” (TF4) clinic visit (age 17+) and the “It’s All About You” (CCU) questionnaire (age 20). These two subsamples were considered as the “selected samples” for our analysis. Selection into the two subsamples differed by offspring sex: in TF4, participation rates were
For each of the six maternal traits, we fitted a logistic regression model with offspring sex as the outcome and the maternal trait as exposure. The models were fitted both in the complete ALSPAC sample and in the two subsamples. We fitted a separate model for each maternal trait to mimic the previous parts of our manuscript, where we only considered one exposure variable. In real-data applications, it may be preferable to conduct a single joint analysis instead, with all six maternal traits included as explanatory variables. Such an analysis is presented in Supplemental Section 6.
We computed regression coefficient estimates from the models regressing offspring sex on each maternal trait, fitted either to the full ALSPAC sample or to the TF4/CCU subsamples. The difference between estimates in the TF4/CCU subsamples and in the full ALSPAC sample was taken as a measure of collider bias for each trait. We then fitted log-additive models for TF4/CCU participation, each time using offspring sex and one of the maternal traits as covariates, and compared the interaction estimates in these models to the magnitude of collider bias. In addition, we fitted logistic regression models for TF4 and CCU participation using offspring sex and one of the six maternal traits as explanatory variables but no interaction terms. Logistic regression is often used to assess which variables associate with study participation, or to adjust for collider bias using inverse probability weighting. We investigated whether parameter estimates from the logistic models could be used to quantify the magnitude of collider bias for each maternal trait.
Note that our application here is conducted for illustrative purposes, to demonstrate the connection between interactions and collider bias in a real dataset. In reality, with access only to the TF4/CCU samples, we would not be able to fit the log-additive model for selection, while with access to the complete ALSPAC sample, there would be no need to restrict the analysis to the TF4/CCU subsamples.
For our analyses, we excluded pregnancies that resulted in miscarriage or early termination (
Results
Estimated associations between each maternal trait and offspring sex from the respective logistic models are reported in Table 1. We report parameter estimates, standard errors and p-values of association between each trait and offspring sex, obtained either from all ALSPAC participants or only from TF4/CCU attendants.
Associations of maternal traits with offspring sex in ALSPAC, obtained by fitting six separate logistic regression models for offspring sex, each with one maternal trait as the exposure.
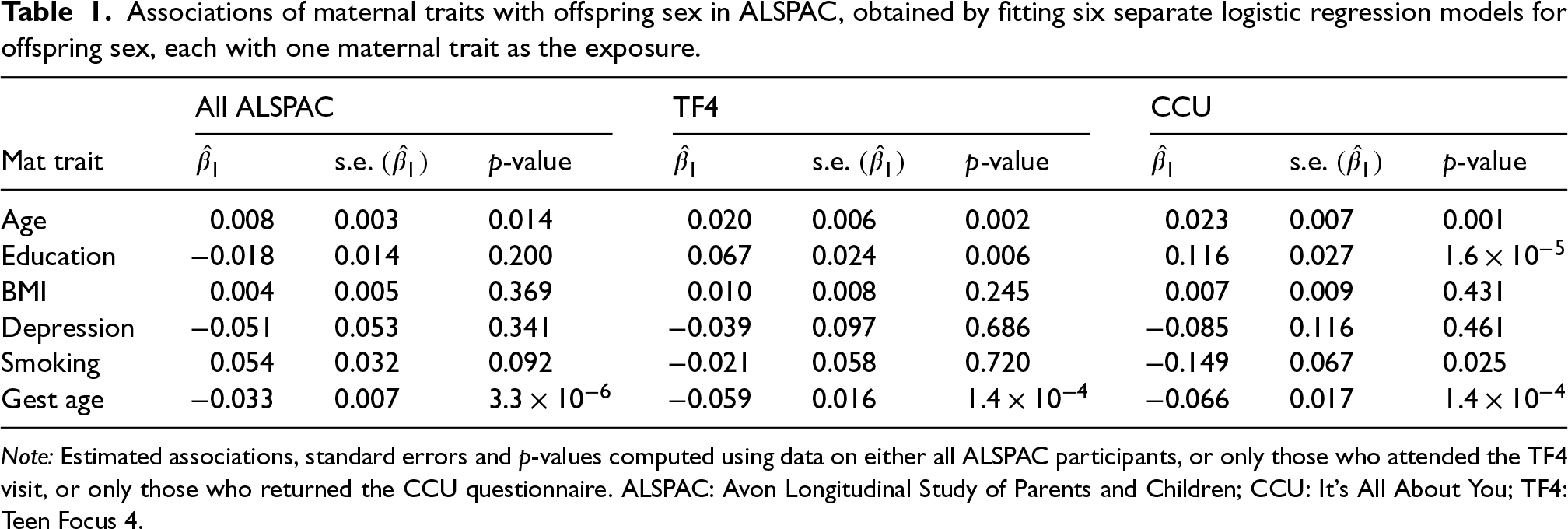
Associations of maternal traits with offspring sex in ALSPAC, obtained by fitting six separate logistic regression models for offspring sex, each with one maternal trait as the exposure.

Note: Estimated associations, standard errors and p-values computed using data on either all ALSPAC participants, or only those who attended the TF4 visit, or only those who returned the CCU questionnaire. ALSPAC: Avon Longitudinal Study of Parents and Children; CCU: It's All About You; TF4: Teen Focus 4.
Mother’s age at delivery and gestational age were associated with offspring sex in all three samples. The observational association between gestational age and offspring sex has previously been noted in the literature 28 and could be due to reverse causation, while the association with mother’s age in ALSPAC was fairly weak and could be due to the missing maternal data. Mother’s education was not associated with offspring sex in the full ALSPAC sample but was seen to associate with offspring sex in both TF4 and CCU. Maternal smoking was associated with offspring sex in the CCU sample but not in the TF4 sample or in the full ALSPAC sample, while BMI and depression before pregnancy exhibited no association with offspring sex in any of the three regression analyses. These results suggest collider bias may be affecting the association of maternal education and smoking with offspring sex. This is not unreasonable, as smoking and education are often associated with participation in scientific studies, and at the same time, participation rates in TF4 and CCU differed between males and females, as mentioned earlier.
Table 2 contains the results of fitting log-additive models for TF4 and CCU participation. Again, six different models were fitted, one for each maternal trait. All models also included offspring sex and an interaction between the maternal trait and offspring sex. We report parameter estimates, standard errors and
Parameter estimates, standard errors and p-values for a log-additive model of TF4 or CCU participation in terms of offspring sex, maternal traits, and interactions between offspring sex and maternal traits. The observed bias in
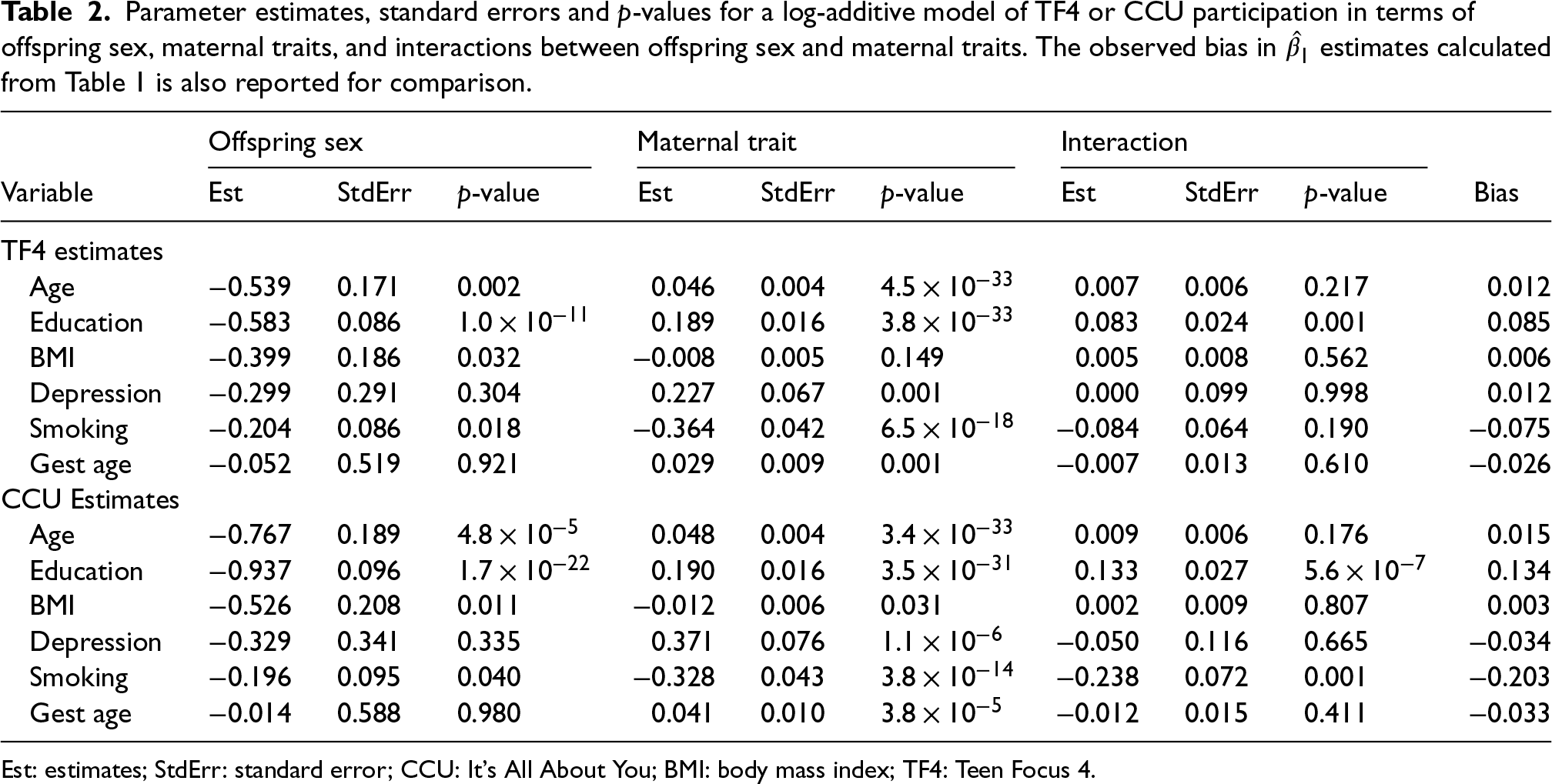
Est: estimates; StdErr: standard error; CCU: It's All About You; BMI: body mass index; TF4: Teen Focus 4.
All maternal traits were associated with CCU participation, and all maternal traits apart from BMI were associated with TF4 participation in their respective models. However, evidence of an interaction between the maternal traits and offspring sex was observed only for maternal education (in both samples) and smoking (in the CCU sample). This was in line with our previous analysis, in which maternal education associated with offspring sex among CCU or TF4 participants, and maternal smoking did so among CCU participants. The regression coefficient for the education-offspring sex interaction was estimated to be positive in both log-additive models; this would suggest positive bias. Indeed the TF4 and CCU regression coefficients in Table 1 were both larger than the regression coefficients in the all-ALSPAC analysis (i.e. positive bias). On the other hand, the smoking-offspring sex interaction in the CCU sample was negative, suggesting negative bias, which was indeed the case based on Table 1. Finally, the interaction parameter estimates were a good approximation of the magnitude of bias caused by restricting to the TF4 or CCU subsamples: for all six traits and for both subsamples, a
The results of fitting the logistic regression models without interactions are given in Table 3. Again, we report parameter estimates, standard errors and p-values for offspring sex and each maternal variable, as well as the bias observed in ALSPAC. These results suggest strong associations between all six maternal traits and participation in both samples, with the exception of BMI in the TF4 sample. In addition, participation is also associated with offspring sex, as expected. By fitting the logistic models, an applied researcher could be led to believe that collider bias will occur when studying the associations of maternal traits with offspring sex in TF4/CCU participants. However, as our analysis in Table 1 indicates, bias is present only for maternal education and smoking, and not for the other four maternal traits considered here. This confirms that using a log-additive model with interactions is more informative about collider bias than the commonly used logistic model without interactions.
Parameter estimates, standard errors and p-values for a logistic regression of TF4 and CCU participation on offspring sex and maternal traits.
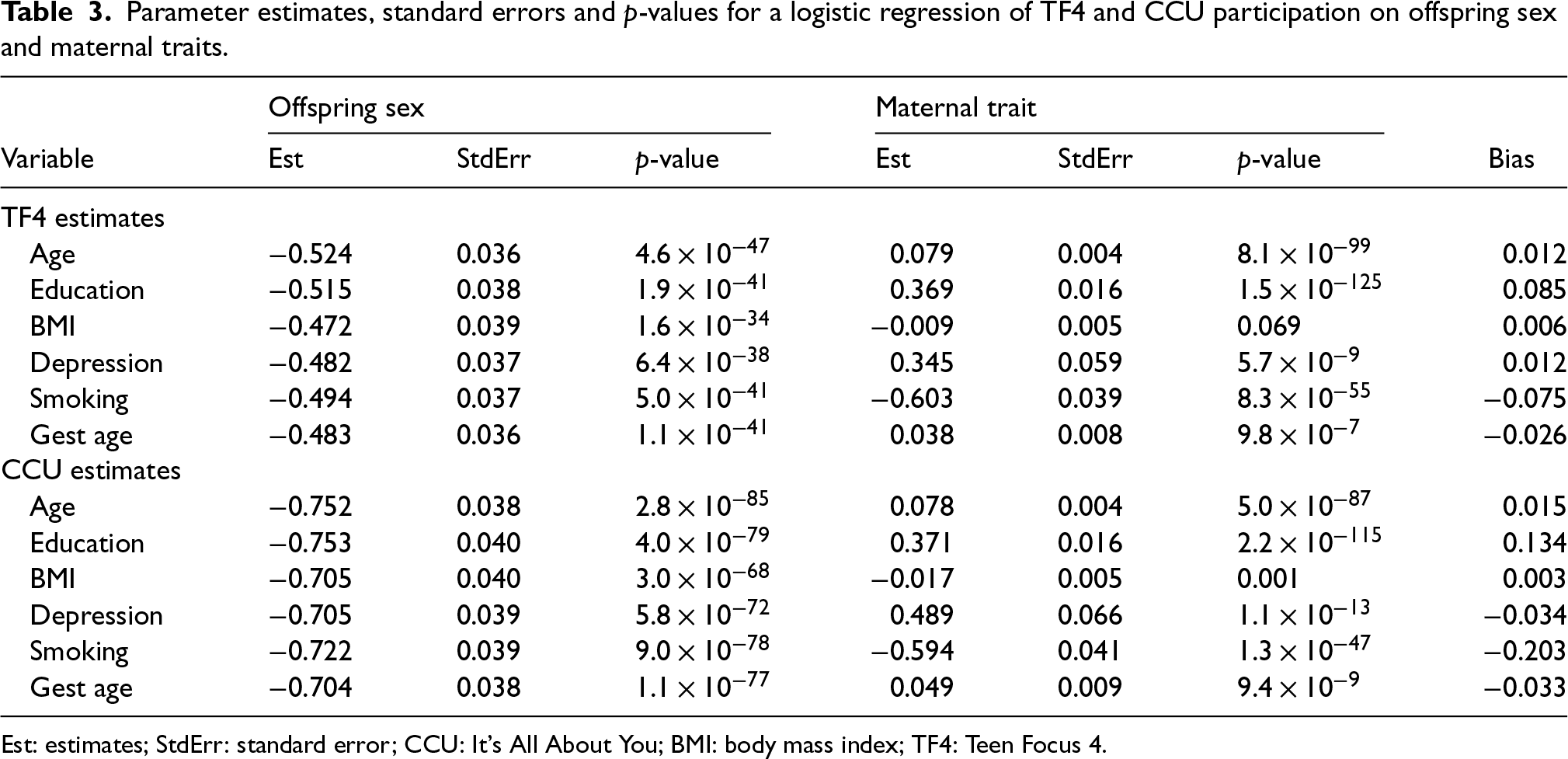
Est: estimates; StdErr: standard error; CCU: It's All About You; BMI: body mass index; TF4: Teen Focus 4.
We have shown that, in three commonly used regression models, the magnitude of collider bias induced in the exposure–outcome association is a linear function of the strength of interaction between the exposure and outcome in a log-additive model for the collider. We have proved these results analytically when the collider is truly distributed according to the log-additive model (1), and explored them via a numerical asymptotic study and a real-data application in cases when the collider does not follow the log-additive model. In Supplemental Tables 1 and 2, we provide a detailed list of all our results, including the distributional assumptions considered for the exposure, outcome, and collider.
Our results can be useful in several ways. As mentioned earlier, modeling selection into a study is an important task for methods that attempt to adjust for collider bias, such as IPW. A key assumption of the IPW method is that the statistical model used to derive weights is correctly specified. If that model is misspecified, the weighted analysis may be biased. There are two aspects to correctly specifying the weighting model: the selection of covariates to be included and the correct specification of the model’s functional form. Advice on the selection of covariates can be found elsewhere; briefly, the variables included in the weighting model should be such that the collider becomes conditionally independent of the exposure and outcome given those covariates.18,30,29 Regarding the model’s functional form, it has become common in the literature to implement IPW using logistic regression without interactions as a weighting model; this choice is often made for convenience, and in some applications there is little reason to believe that the logistic model is correctly specified. In fact, a logistic model without interactions offers relatively little flexibility to capture complex relationships between the collider and other variables, and more flexible statistical models should be preferred. This advice has been given in the literature (e.g. Seaman and White
18
) and several authors have investigated the use of flexible statistical and machine learning approaches for the IPW weighting model.21,20,23,19,22 Although some applied researchers have utilized these approaches, the use of the logistic model without interactions is still common. Our work contributes to this debate by illustrating the relationship between collider bias and interactions in the collider model, and so emphasizing the limitations of the simple logistic model. From that perspective, if
The fact that collider bias only depends on a single parameter in the simple models considered here can also be useful for sensitivity analyses. In some applications, subject-specific knowledge may allow researchers to assess the strength of exposure–outcome interactions, and hence assess whether collider bias is likely to affect their analyses. In addition, simulation studies are sometimes conducted as a form of sensitivity analysis to explore the impact of collider bias in applications. These simulation studies typically work by varying the associations of the exposure, outcome and other relevant variables with study participation and exploring how much collider bias this induces in analysis results. Our work suggests that it is the interactions (on the log-additive scale) that dictate the magnitude of this bias, and therefore that these interactions should also be varied in addition to (or instead of) the exposure-selection and outcome-selection associations.
In addition, simulation studies are used as a tool for assessing the finite-sample performance of novel methods. Our results may therefore be useful to researchers working on methods to detect or adjust for collider bias. For example, it may be desired to design a simulation where complete-case analysis exhibits a specific degree of bias, and compare that with the performance of a newly developed method. This can be done using a log-additive model with an interaction, and specifying the value of the interaction parameter accordingly.
In some applications, additional information about the causal structure of the association between the exposure, outcome and collider may be available. For example, the effect of
Some extensions of our work are possible. Here, we have focused on three simple statistical models for the exposure–outcome association, namely linear, logistic and Poisson regression. It would be interesting to explore whether similar results hold, for example, in survival analysis models. Another potential extension could be to instrumental variable analyses, which are known to suffer from collider bias.31,32
We hope our findings will prove useful to methodologists investigating collider bias, as well as to applied researchers attempting to adjust for the bias in their analyses.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802241306860 - Supplemental material for Relationship between collider bias and interactions on the log-additive scale
Supplemental material, sj-pdf-1-smm-10.1177_09622802241306860 for Relationship between collider bias and interactions on the log-additive scale by Apostolos Gkatzionis, Shaun R Seaman, Rachael A Hughes and Kate Tilling in Statistical Methods in Medical Research
Footnotes
Acknowledgements
AG and KT received funding for this project by the UK Medical Research Council and the University of Bristol (MRC-IEU core funding, MC UU 00032/02). SRS was funded by UKRI (Unit programme numbers MC UU 00002/10 and MC UU 00040/05) and was supported by the National Institute for Health Research (NIHR) Cambridge Biomedical Research Centre (BRC-1215-20014). RAH is supported by a Sir Henry Dale Fellowship that is jointly funded by the Wellcome Trust and the Royal Society (grant 215408/Z/19/Z). The views expressed in this manuscript are those of the authors and not necessarily those of PHE, the NHS, the NIHR or the Department of Health and Social Care. For the purpose of open access, the authors have applied a Creative Commons Attribution (CC BY) licence to any Author Accepted Manuscript version arising.
Access to ALSPAC data was obtained as part of application B4189. The UK Medical Research Council and Wellcome (Grant ref: 217065/Z/19/Z) and the University of Bristol provide core support for ALSPAC. A comprehensive list of grants funding the ALSPAC study is available on the ALSPAC website (![]() ). GWAS data was generated by Sample Logistics and Genotyping Facilities at Wellcome Sanger Institute and LabCorp (Laboratory Corporation of America) using support from 23andMe. We are extremely grateful to all the families who took part in this study, the midwives for their help in recruiting them, and the whole ALSPAC team, which includes interviewers, computer and laboratory technicians, clerical workers, research scientists, volunteers, managers, receptionists and nurses.
). GWAS data was generated by Sample Logistics and Genotyping Facilities at Wellcome Sanger Institute and LabCorp (Laboratory Corporation of America) using support from 23andMe. We are extremely grateful to all the families who took part in this study, the midwives for their help in recruiting them, and the whole ALSPAC team, which includes interviewers, computer and laboratory technicians, clerical workers, research scientists, volunteers, managers, receptionists and nurses.
Data availability statement
The ![]() .
.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.