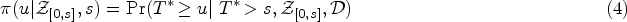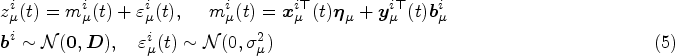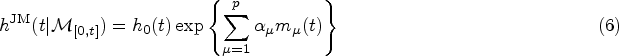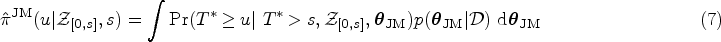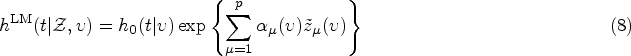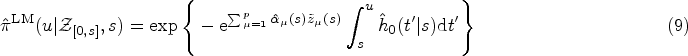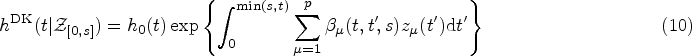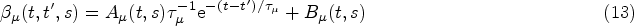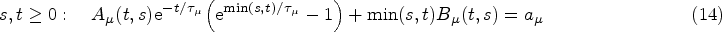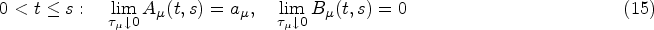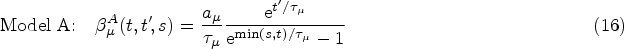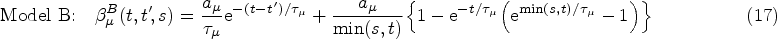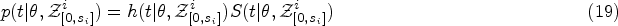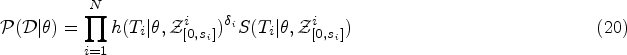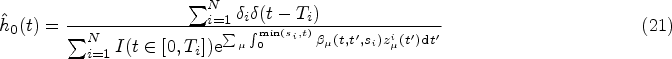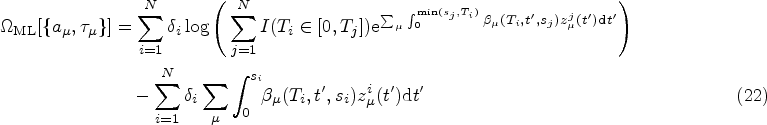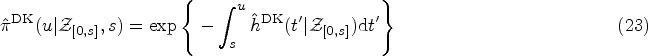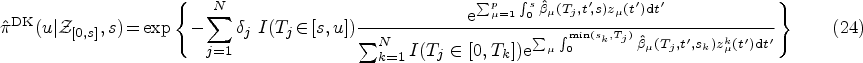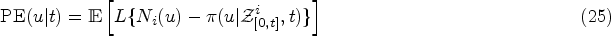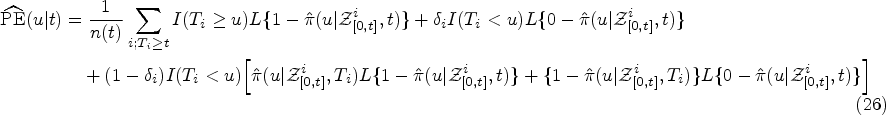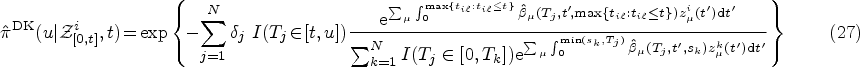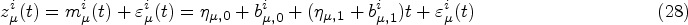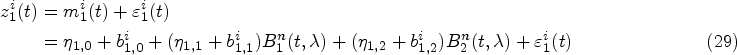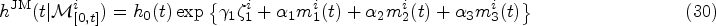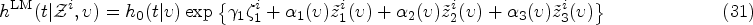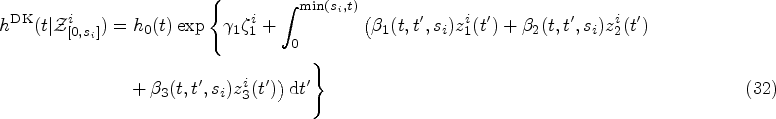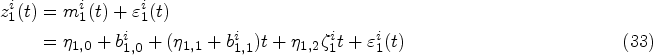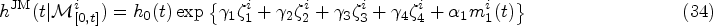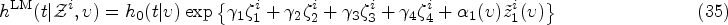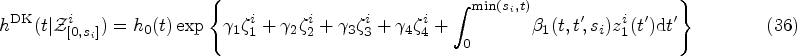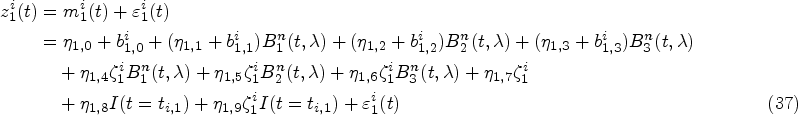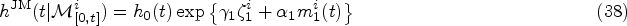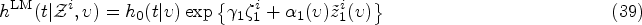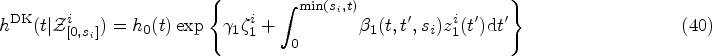Abstract
Predicting patient survival probabilities based on observed covariates is an important assessment in clinical practice. These patient-specific covariates are often measured over multiple follow-up appointments. It is then of interest to predict survival based on the history of these longitudinal measurements, and to update predictions as more observations become available. The standard approaches to these so-called ‘dynamic prediction’ assessments are joint models and landmark analysis. Joint models involve high-dimensional parameterizations, and their computational complexity often prohibits including multiple longitudinal covariates. Landmark analysis is simpler, but discards a proportion of the available data at each ‘landmark time’. In this work, we propose a ‘delayed kernel’ approach to dynamic prediction that sits somewhere in between the two standard methods in terms of complexity. By conditioning hazard rates directly on the covariate measurements over the observation time frame, we define a model that takes into account the full history of covariate measurements but is more practical and parsimonious than joint modelling. Time-dependent association kernels describe the impact of covariate changes at earlier times on the patient’s hazard rate at later times. Under the constraints that our model (a) reduces to the standard Cox model for time-independent covariates, and (b) contains the instantaneous Cox model as a special case, we derive two natural kernel parameterizations. Upon application to three clinical data sets, we find that the predictive accuracy of the delayed kernel approach is comparable to that of the two existing standard methods.
Keywords
Introduction
Survival analysis is a well-established field of medical statistics that involves modelling the probability of survival until some specified irreversible event such as death or the onset of disease. Of particular clinical interest is the prediction of patient-specific survival based on a set of observed biomarkers or ‘covariates’. 1 Such predictions aid clinicians in making treatment and testing decisions and provide personalized information for patients about their health. 2
Cox’s proportional hazards (PH) model
3
remains the most widely used model in survival analysis.4,5 In this context, survival is assumed to depend on a set of covariates,
Survival prediction in the standard Cox model is based on the survival function
In reality, covariates are often measured repeatedly over time. This means that multiple observations of time-dependent covariates
However, in practice, one does not have access to the full covariate trajectories
Recently, there has been much interest in so-called “dynamic prediction”.9,10 These methods aim to make survival predictions based on the longitudinal history of biomarker data and update these predictions as more data becomes available. Such analysis is clinically valuable as it allows patients and clinicians to review disease progression over time and update the prognosis at each follow-up visit. 11 Currently, there are two main approaches to dynamic prediction; joint modelling and landmarking. Landmarking was an early approach to the problem, 12 whereby a standard Cox model is fitted to patients in the original data set who are still at risk at the time point of interest, using their most recent covariate measurements. More recently, joint modelling (JM) has become an established method.13–16 Here one models the time-dependent covariate trajectory using a parameterized longitudinal model, and this complete trajectory is then inserted into an instantaneous Cox-type survival model. A joint likelihood of the longitudinal and survival sub-models is constructed, and the model parameters are estimated via maximum likelihood (ML) or Bayesian inference.
Both methods have limitations. In particular, joint models are demanding both conceptually and computationally. Correctly modelling the longitudinal trajectories can be difficult when patient measurements exhibit varied non-linear behaviour 11 and mis-specification of this trajectory has been found to lead to bias. 17 Furthermore, the number of model parameters increases rapidly with the inclusion of multiple longitudinal markers. This means that many software packages cannot handle more than one longitudinal covariate,18–20 and those that can quickly become computationally intensive.21,22 For these reasons, the landmarking model is often seen as the only practical option. 11 However, the relative simplicity of the landmarking approach comes with its own drawbacks. By using only the ‘at risk’ data set to make predictions at a certain time (discarding patients who had an event before the landmark time), landmarking makes use of only a subset of the available data. In standard landmarking approaches, the history of the covariate values are not taken into account directly, and a new model must be fitted every time one wishes to update the predictions.
In this work, we present a new approach to dynamic prediction that conceptually and in terms of computational complexity lies somewhere in between the JM and landmarking methods. Rather than modelling the covariate trajectory at future times, as in the JM approach, we model the probability of survival conditioned directly on the observed covariates measured from the baseline time up to a subject-specific final observation time. Unlike the landmark approach, a single model is fitted to all of the available data, using the full history of the covariate values. We do, however, maintain well-established and desirable features of the Cox model, so that our model contains the instantaneous Cox model as a special case, and reduces automatically to the standard Cox model for covariates that are observed to be fixed over time. Within these constraints, we define time-dependent parametric association kernels,
The remainder of this article is set out as follows. In Section 2, we introduce the motivating data sets. In Section 3, we then provide details of the dynamic prediction models. We begin by describing the longitudinal and time-to-event data, and briefly outline the standard methods: JM (Section 3.2) and landmarking (Section 3.3). In Section 3.4, we introduce the delayed kernel (DK) approach. We start by defining the hazard rate conditioned on the observed data and then develop two natural parameterizations for the association kernels that meet our requirements. We outline the ML method for parameter estimation for these models and show how the DK approach can be used to make dynamic predictions. In Section 3.5, we describe a simple simulation study to aid the interpretation of our model parameters. Via application to the real data sets, in Section 4, we compare the performance of the delayed kernel approach to the standard methods using an established measure of predictive accuracy. Finally, we discuss and summarize our results in Section 5.
In our work, we will assess the predictive capabilities of the different models for dynamic prediction using three clinical data sets that contain both longitudinal covariate measurements and time-to-event data. All three data sets are publicly available in the
Primary biliary cirrhosis (PBC)
The first motivating data set is from a study conducted by the Mayo Clinic from 1974 to 1984 on patients with primary biliary cirrhosis (PBC), a progressive chronic liver disease.
27
We will refer to this as the PBC data. The study involved
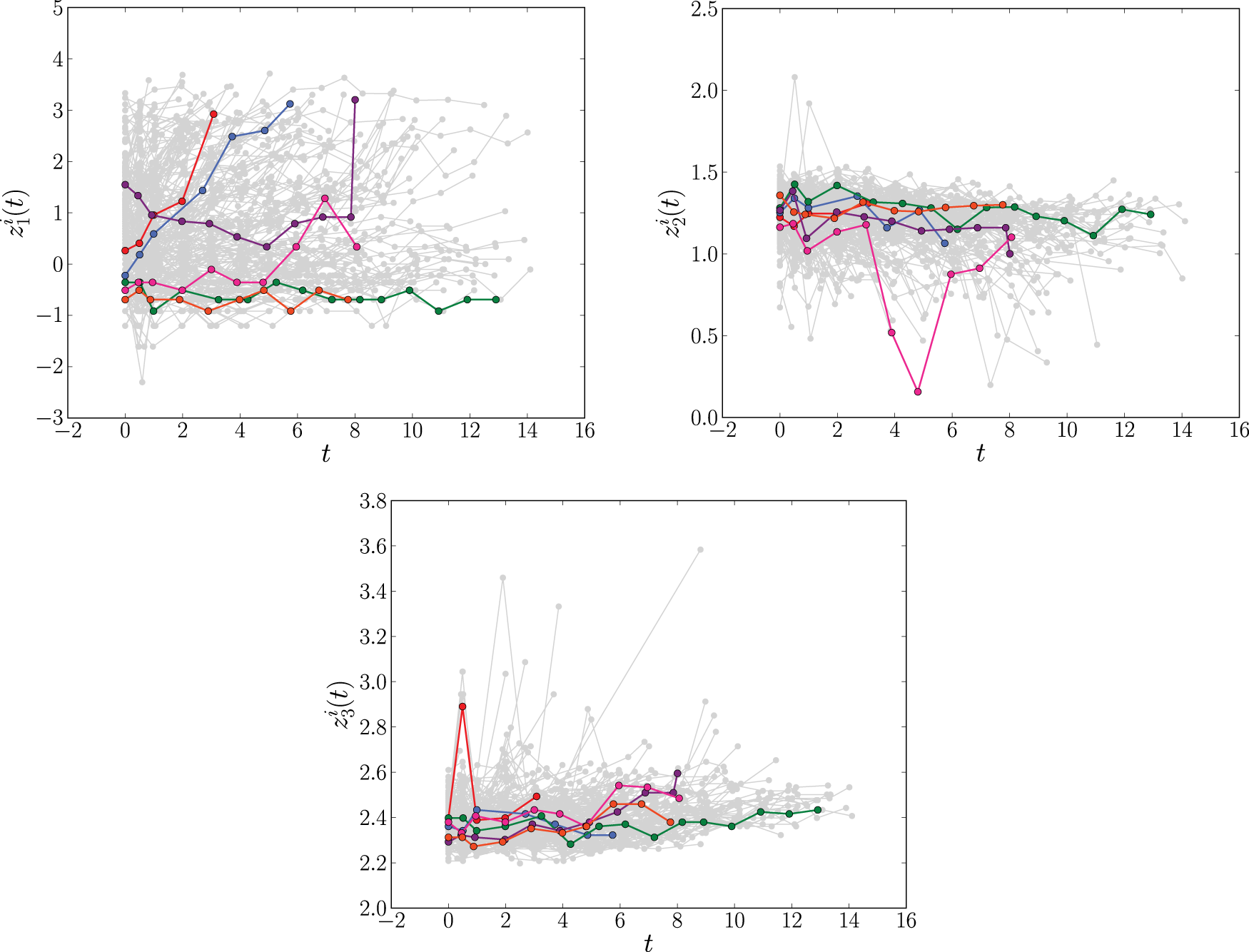
The longitudinal profiles of the time-dependent covariates log serum bilirubin (
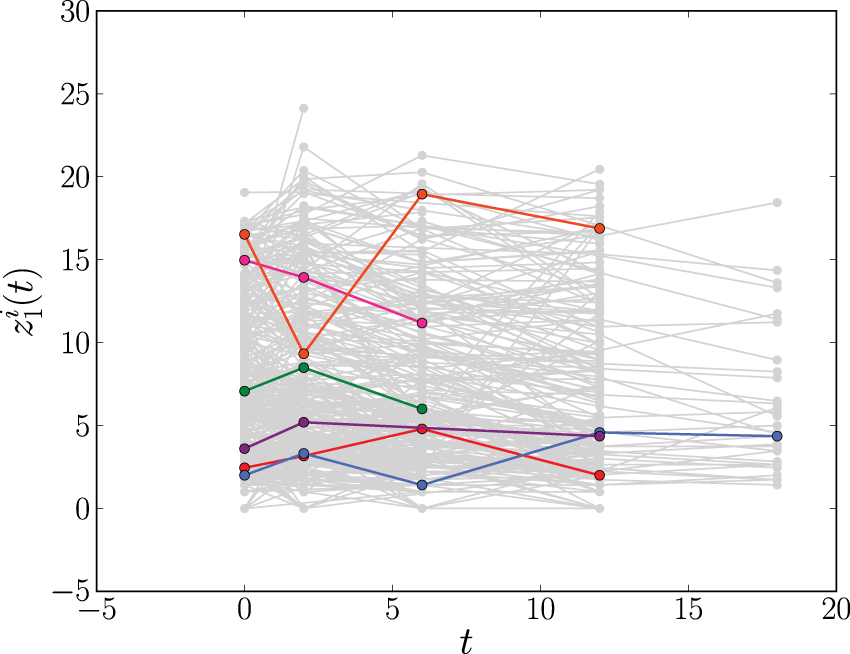
The longitudinal profiles of CD4 count (
The second data set involves
Liver cirrhosis
The third data set is from a trial conducted between 1962 and 1974, involving
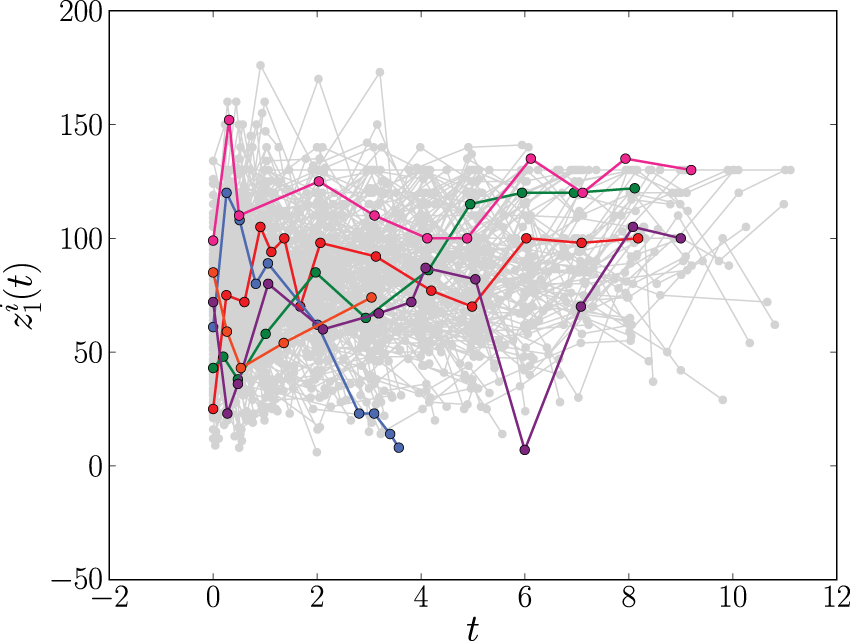
The longitudinal profiles of the prothrombin index (
Setup and notation
In this work, we consider longitudinal survival data of the form
We are interested in predicting survival probabilities for some new subject with longitudinal measurements
In JM one specifies two model components: a longitudinal model for the trajectory of the time-dependent covariates, and a survival model which relates to the covariate trajectory via shared parameters. In the
Longitudinal modelling component
Mixed-effects models are typically specified for the longitudinal covariate trajectories, where it is assumed that the observed value
The hazard at time
It has also been proposed to introduce a weight function to capture cumulative effects, writing
Finally, in the JM framework we use, the quantity
Joint models have undergone much development over recent years, with various extensions making the approach flexible in a range of different scenarios. However, the JM approach requires the ability to correctly specify both the longitudinal and survival model. This can involve modelling assumptions which are not always easy to verify. Indeed, simulations have demonstrated that the JM approach is biased under mis-specification of the longitudinal model. 17 In addition, as more longitudinal outcomes are included, the dimensionality of the random effects increases, and fitting the joint model becomes computationally intensive. Depending on the longitudinal model specified, it can be difficult to include more than three or four longitudinal covariates.11,21 This is amplified when cumulative or weighted cumulative effects are used in the survival model (as numerical integration of the longitudinal model is required). As a result, there are cases where joint models are not a viable option and, instead, one must rely on approaches such as landmarking. 11
Landmarking
Description of the landmarking procedure
The landmarking approach to dynamic prediction is based on the standard Cox model.1,12,33 Upon denoting with
To estimate the quantity
Landmarking is computationally and conceptually much simpler than the JM approach. For data sets with multiple longitudinal covariates, disparate non-linear covariate trajectories or categorical time-dependent covariates, landmarking is often the preferred approach.
11
However, it also has limitations. For example, the standard model focuses only on the most recent value observed before time
Delayed kernel (DK) approach
We now introduce our DK approach to dynamic prediction. It aims to overcome some of the limitations of the standard JM and landmarking methods. Unlike landmarking, the DK approach aims to incorporate the entire data set, including the full history of covariate values while, at the same time remaining conceptually and computationally simpler than joint models.
General setup
The starting point for the DK approach is an expression for the hazard rate that resembles that of weighted cumulative exposure models,23,24
The assumptions we make in writing expression (10) for the hazard function at time
In our view, there is no direct link between our present model (DK) and existing models such as JM, landmarking, or the more recent ‘landmarking 2.0’ (see Putter and van Houwelingen
40
for details of this model); the latter follow different strategies. In JM one models the dynamics of the covariates in order to predict (probabilistically) their values at times beyond
In an early WCE model from Abrahamowicz et al.,
41
the past covariate values are summarized in a time-dependent cumulative dose, which at time
In their simplest form, all existing models build on the assumption that the instantaneous Cox model would hold if all covariate values at all times were known in full. They differ in how, from this starting point, the non-availability of covariate information in the time interval
The relation between the three approaches is further summarized in the schematic diagram in Figure 4.
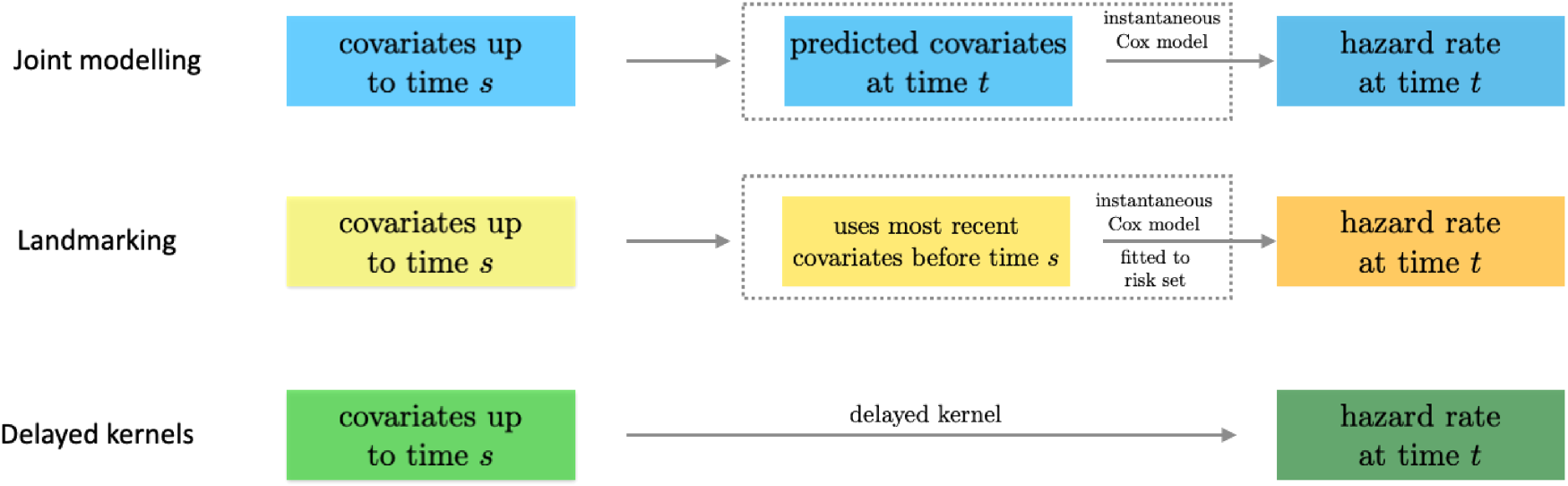
Relation of the delayed kernel approach to joint modelling and landmarking. All three approaches use covariates up to time
The kernel Exponential decay of covariate impact. We assume that the impact of each covariate Equivalence with standard Cox model for stationary covariates. Our second requirement is that expression (10) reduces to the standard Cox model in equation (1) in the case of a constant covariate, that is, when Equivalence with instantaneous Cox model for short impact time scales. Finally, for
From (i), it follows that our kernel
Finally, from the remaining family of models (those that satisfy equations (14) and (15)), we choose the two simplest members. These are defined by demanding that either
Equations (16) and (17) only hold for
We note that we condition on knowledge of the covariates observed over a specific time interval
As in the standard Cox model, we use ML inference to determine the most plausible values of the model parameters based on the observed data. For simplicity, in this section, we will mostly omit the superscript DK from the hazard function. We write
A full derivation of the ML equations for models of the form in equation (10) is provided in Section S2.1 of the Supplemental Material. Here we present only the results. The ML estimator of the baseline hazard function is the direct analogue of the Breslow estimator,
39
Once we have obtained the ML estimates of the model parameters
Using the ML estimates
Any uncertainty in the prediction (23) reflects uncertainty in the inferred model parameters. It is hence quantified by the variance of the covariate-conditioned survival function, calculated with respect to the posterior parameter distribution (given the data). The parameter distribution is approximated by a Gaussian distribution (for sufficiently large sample sizes), centered at the ML point, and with a covariance matrix obtained via the Fisher information.
So far, we have defined the DK models conditional on covariate trajectories
We choose the simple “last observation carried forward” (LOCF) procedure, commonly used in instantaneous Cox models and landmarking approaches. That is,
Simulation study
To understand better the DK model and how to interpret the associated parameters, we perform a simple simulation study. Generating data according to the DK model is non-trivial since event times are dependent on the period over which the covariates are observed. Instead, we generate data according to a simple joint model (with longitudinal measurements generated from a linear random slope and random intercept model) and compare parameter estimates between the models. We do this for a joint model with an instantaneous association function in the survival sub-model (scenario 1) and for a joint model with a cumulative association (scenario 2). In each scenario we fit both DK models, a landmarking model and two joint models (one with an instantaneous survival model and one with a cumulative survival model). We repeat the simulation 50 times and inspect the average behavior of the different models.
Details of the methodology and results of the simulation are provided in Section S3 of the Supplemental Material. We briefly describe the main findings here. First, we find that the fixed associated parameter
Finally, we find that even when events are generated from a survival sub-model with an instantaneous association, the DK models estimate a non-zero decay parameter,
In a couple of iterations of the simulation, the DK models estimated values of
Application to clinical data
Methods
Cross-validation
For each of the data sets in Section 2, we assess the predictive accuracy of the different dynamic prediction models using 10-fold cross-validation. The data is split randomly into 10 equally sized groups. At each iteration, one group is assigned as the test data set and the remaining groups make up the training data set. Each model is fitted to the training data and the resulting model is used to make survival predictions about individuals in the test data set. Predictive accuracy is assessed by comparing these predictions to the true event times in the test data (see Section 4.1.3). The procedure was repeated 10 times with each group being assigned as the test data.
Fitting models
We implemented the DK models using our own
Measuring predictive accuracy
Following Rizopoulos et al.,
2
we quantify the predictive accuracy of the different models using the expected error of predicting future events. Dynamic prediction is concerned with predicting the survival of individuals to a given time
Again following Rizopoulos et al.,
2
in this article, we use the overall prediction error
To compare the predictive accuracy of JM, landmarking and the DK approach we insert into equation (26) the respective estimators
The term
To perform the prediction error calculation for the DK models, we use our own
First, we compare the predictive accuracy of the three methods by specifying a fixed base time
Fixed prediction window
In our second test, we vary the base time
Based on previous analysis of the PBC and Liver data,16,32 we choose three prediction windows:
Based on the event time distribution of the AIDS data, we choose prediction windows
PBC data set
We fit each model to the PBC training data set using
The PBC data set contains event-time information for two events, death and liver transplant. The most appropriate way of analysing this data is to use a competing risks model. However, for simplicity, we here treat the transplant event as a censoring event. Another simple way to analyse this data is to treat the two events as a single composite event. We provide the results of the latter analysis in Section S5 of the Supplemental Material. The two analyses are found to give similar results.
Models
For the joint model, we first fit a simple multivariate linear mixed model to each of the three time-dependent covariates
Figure 1 suggests that the covariate trajectories in the PBC data may be non-linear for some individuals. Hence, for extra flexibility, we also fit a second joint model that includes natural cubic splines in both the fixed and random effects parts of the model. Following Rizopoulos,
25
the log serum bilirubin (
For both the linear and spline longitudinal models, the hazard function of the survival sub-model in the JM framework is
For the DK approach, we specify the hazard function as
Figure 5 shows plots of the overall prediction error
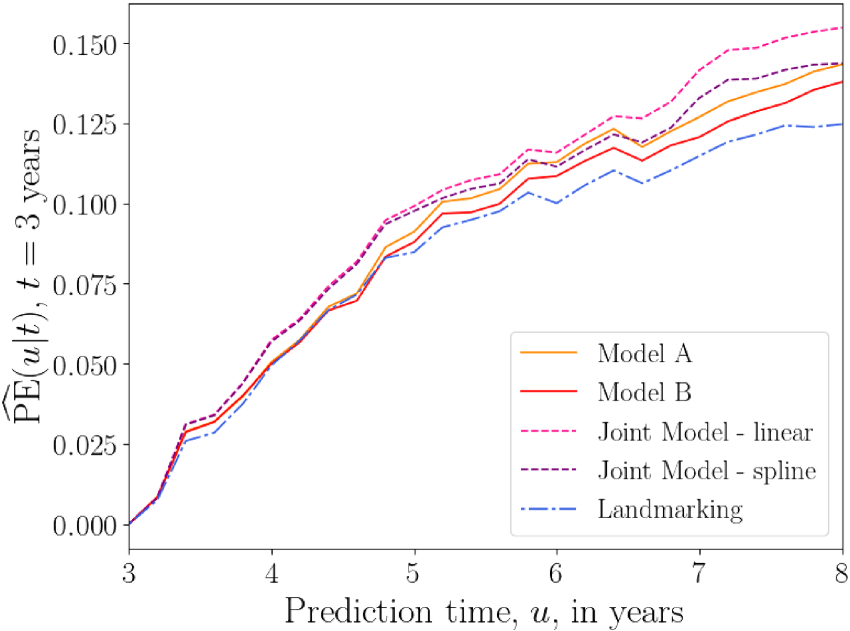
Overall prediction error
Plots of the average prediction error
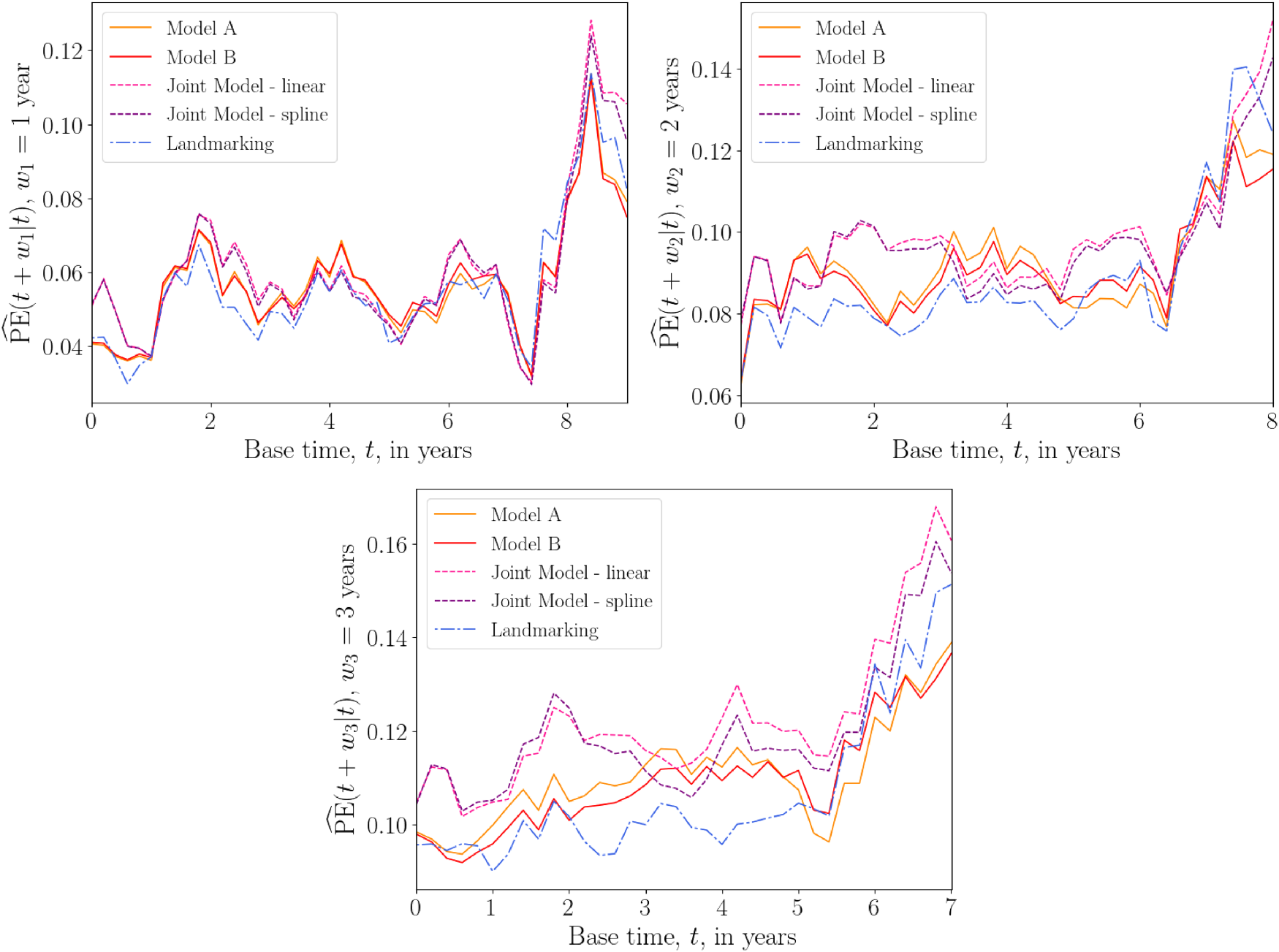
Overall prediction error
The results of the above tests suggest that, for the PBC data, the DK approach performs as well as, and sometimes better than, existing methods. Care should be taken when interpreting these results, as we have not used a competing risks model in our analysis. This data set does, however, serve as an illustration that, without sacrificing accuracy, the DK model can serve as a simpler alternative to JM when considering multiple longitudinal covariates.
In the AIDS data set, we focus on a single longitudinal covariate, the CD4 count
Models
The JM framework allows us to model the dependence of CD4 count on the patients drug group. Following Rizopoulos,
16
we fit the linear mixed model
The plots of
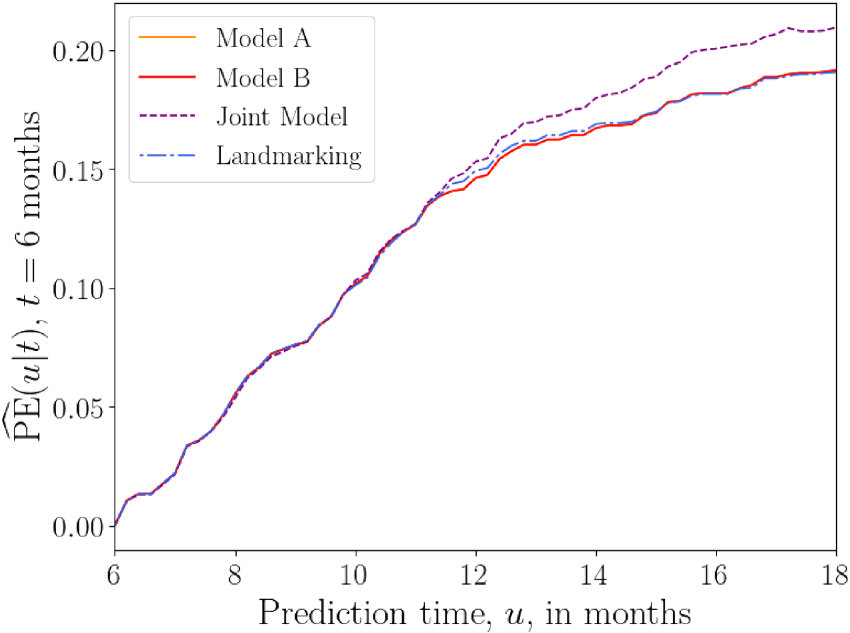
Overall prediction error
Figure 8 shows plots of
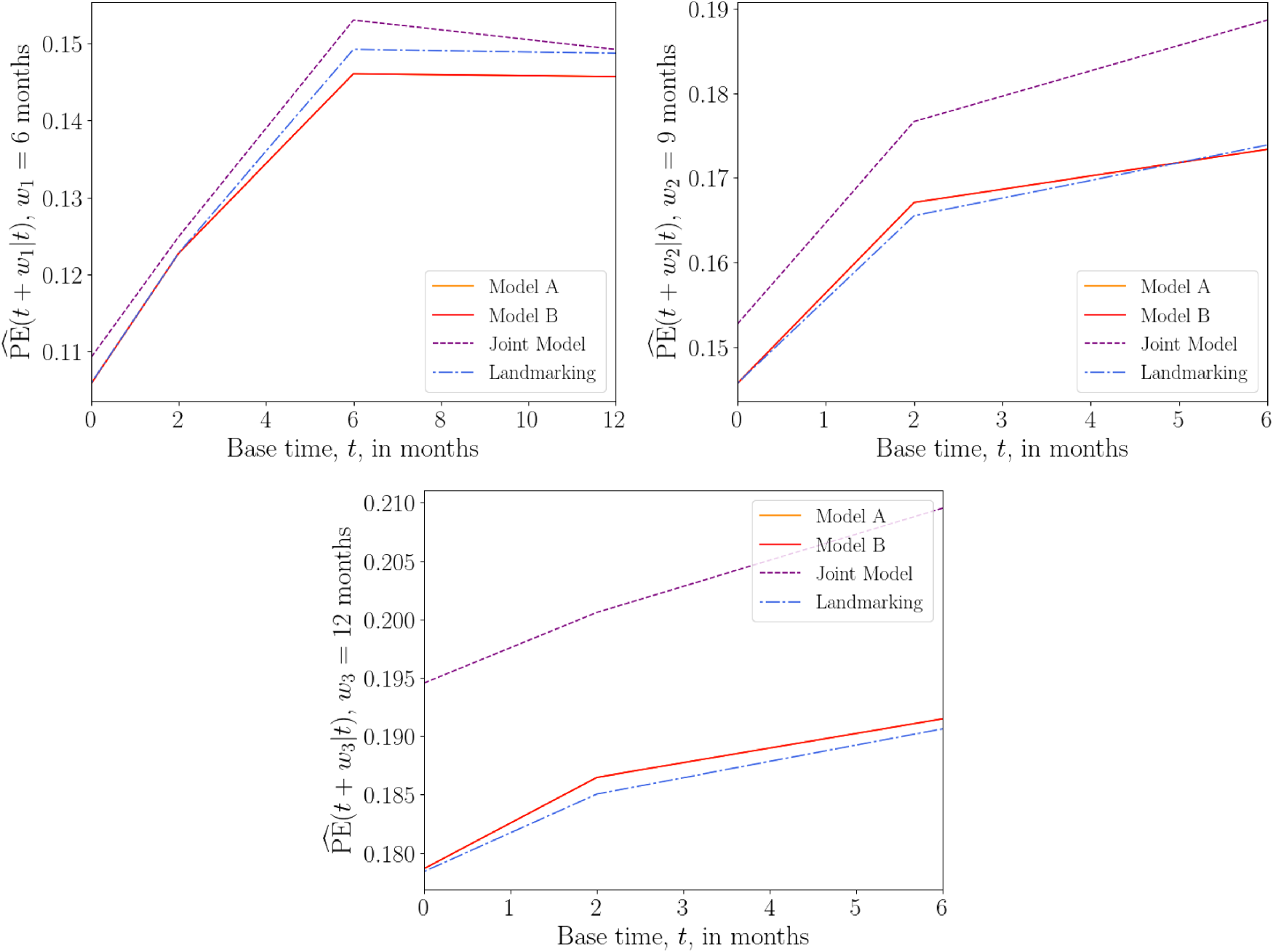
Overall prediction error
The above results suggest that, for the AIDS data set, the joint model has the worst predictive accuracy overall, while the landmarking and DK models perform similarly.
For the liver data set, we model prothrombin index as our one longitudinal covariate
Models
Following Rizopoulos,
16
we define a flexible longitudinal model for the subject-specific prothrombin trajectories, using natural cubic splines with different average profiles for each drug group. Rizopoulos
16
also suggests to include a separate indicator variable of the baseline measurement, to capture sudden changes in the prothrombin index in the early part of follow-up. The longitudinal model then takes the form
The hazard functions for the joint model and the landmark model (with landmark time
Figure 9 shows the prediction error
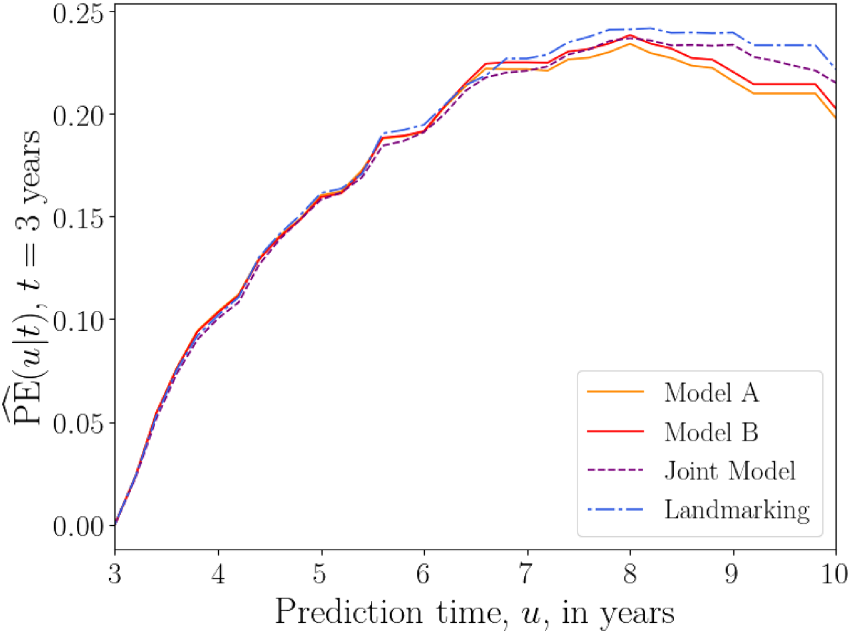
Overall prediction error
Plots of average
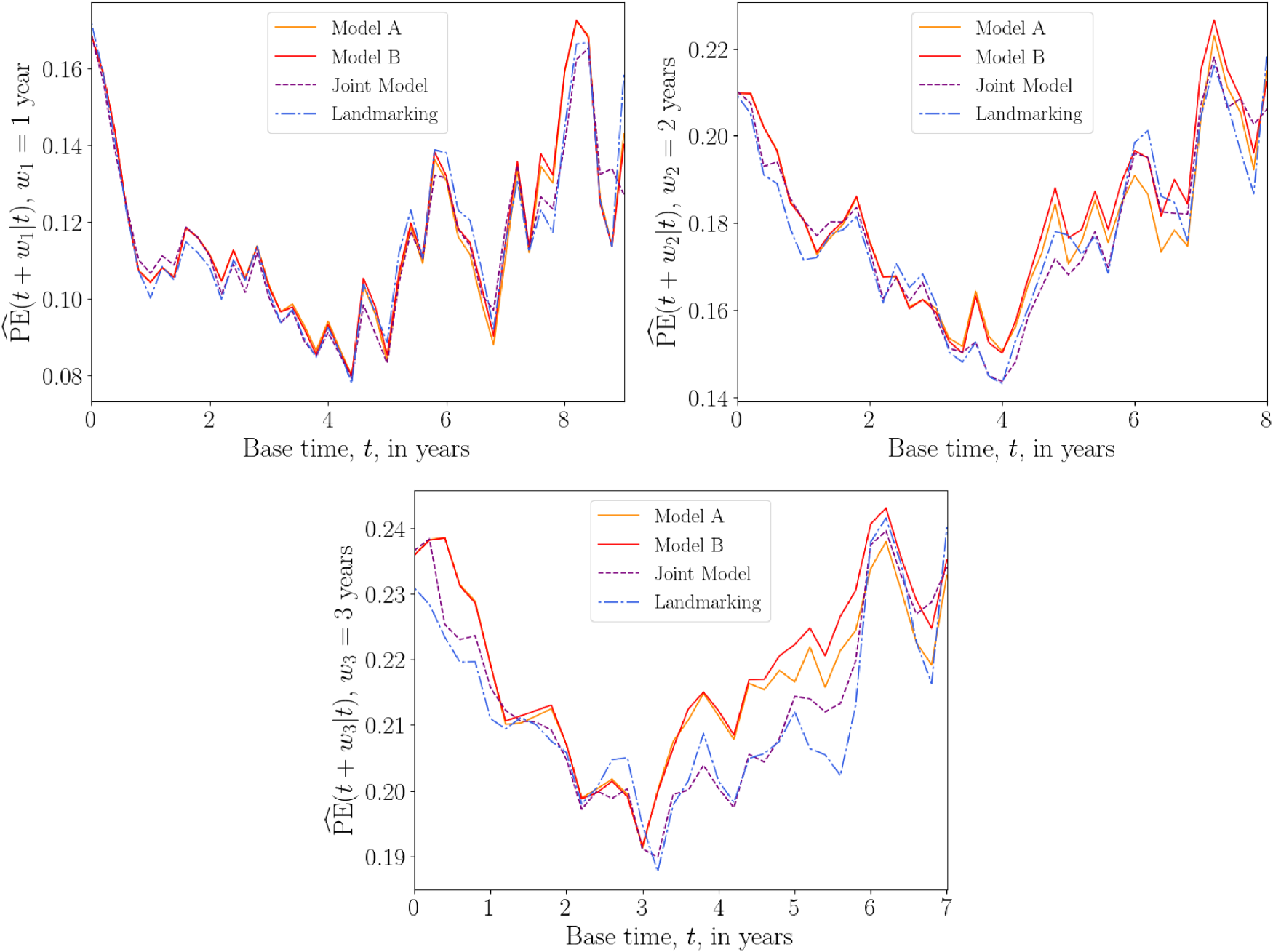
Overall prediction error
For the liver data set, the above results suggest that the DK models have a predictive accuracy that is comparable to those of standard methods.
In this work, we propose a DK approach to dynamic prediction in survival analysis. In terms of complexity, our method comes somewhere between the two standard approaches, JM and landmarking. It is more parsimonious than JM, as it does not model the longitudinal covariate trajectory, and it makes no assumptions about the baseline hazard function. This makes the method more practical for data with multiple time-dependent covariates. The DK approach conditions only on the observed covariates and, unlike JM, makes no assumptions about covariate values in the future. This makes it more suitable for covariates that cannot easily be predicted, such as categorical ones. Compared to landmarking, the DK approach makes use of more of the available data. In landmarking, a new model is fitted at each landmark time, discarding individuals in the data set who have experienced the event before this landmark time. Additionally, standard landmarking only uses covariate measurements that are most recent before the landmark time. In contrast, the DK approach fits a single model that incorporates information from all individuals in the data set, using the full history of their covariate measurements. We note that in extensions of the landmarking approach one could, in principle, fit a landmarking model using multiple covariate measurements, for example, by replacing the most recent measurement with the mean value of measurements up to that time.
The DK approach relies on parameterization of the association kernels
In tests on medical data, we found that the DK approach performs similarly to the two standard approaches in terms of predictive accuracy. Depending on the data set, base time and prediction window, each method (JM, landmarking, or delayed kernels) had at some point the highest or the lowest prediction error; none appeared to be consistently superior or inferior across the scenarios we tested. An open question that should be the focus of future work is how to establish which longitudinal survival model will perform best for a given scenario. In simulations with data generated from joint models, the DK models had larger prediction errors than both joint models and landmarking. Here, landmarking even outperformed correctly specified joint models in some cases. Our results, therefore, suggest that the straightforward landmarking approach might still be a good choice in practice, while joint models suffer from mis-specification.
Although the DK models do not consistently outperform existing methods yet, we still see value in expressing a problem in a new formulation. We believe that our approach is less ‘ad-hoc’ than landmarking, and we expect that, in the future, suitable delayed kernels can be defined that reproduce the landmarking method. This would then significantly enrich the landmarking idea since many different kernels can be developed, and tailored to characteristics of the data sets at hand. In this spirit, we were careful to include standard models as special cases of our own.
There is scope to further develop the DK approach. For example, we used a naïve interpolation procedure (LOCF) but could try smoother interpolation methods such as Gaussian convolutions.45,46 We could also take a Bayesian inference approach, using non-informative prior distributions or incorporating existing knowledge into informative ones. While JM takes into account measurement errors, we have not attempted to do this for the DK model. Cox models were indeed found to be biased in the presence of such errors.17,49,50 Hence, future work could involve building measurement error effects into the DK approach. One could also build on methods from WCE models 37 and use a wider class of association kernels, for example, those estimated via spline functions.
In this work, we compared against standard landmarking models, though extensions to these models exist.2,33,51 Similarly, in our analysis of real data sets we only considered joint models with instantaneous dependence on the ‘true’ covariate trajectory
In summary, we have developed a ‘delayed kernel’ approach to dynamic prediction that overcomes some limitations of existing methods. By conditioning the hazard rate on observed covariates over a given time frame, it offers a simpler alternative to joint models without disregarding portions of longitudinal covariate data, as is the case with landmarking methods. Using three different clinical data sets we have demonstrated that DKs can have a predictive accuracy comparable to that of established methods. Therefore, we believe that the DK method is a promising addition to the toolbox of dynamic prediction methods.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802241275382 - Supplemental material for Delayed kernels for longitudinal survival analysis and dynamic prediction
Supplemental material, sj-pdf-1-smm-10.1177_09622802241275382 for Delayed kernels for longitudinal survival analysis and dynamic prediction by Annabel Louisa Davies, Anthony CC Coolen and Tobias Galla in Statistical Methods in Medical Research
Footnotes
Acknowledgements
ACCC would like to thank Marianne Jonker for valuable discussions.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: AD acknowledges funding by the Engineering and Physical Sciences Research Council (EPSRC UK), grant EP/R513131/1. TG is grateful for partial financial support by the Maria de Maeztu Program for Units of Excellence in R&D (MDM-2017-0711).
Data availability
![]() . The data sets analysed are available publicly via the
. The data sets analysed are available publicly via the
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.