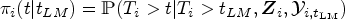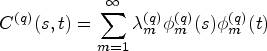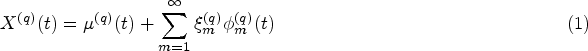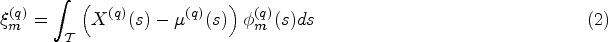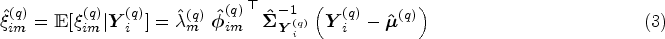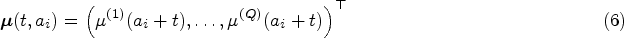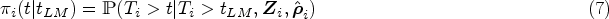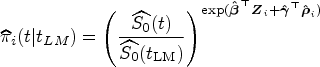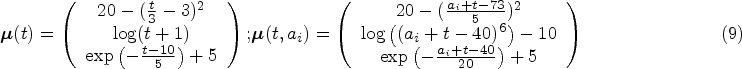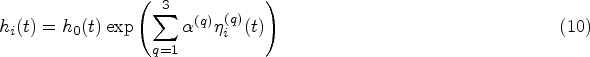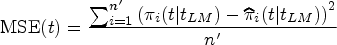Abstract
Dynamically predicting patient survival probabilities using longitudinal measurements has become of great importance with routine data collection becoming more common. Many existing models utilize a multi-step landmarking approach for this problem, mostly due to its ease of use and versatility but unfortunately most fail to do so appropriately. In this article we make use of multivariate functional principal component analysis to summarize the available longitudinal information, and employ a Cox proportional hazards model for prediction. Additionally, we consider a centred functional principal component analysis procedure in an attempt to remove the natural variation incurred by the difference in age of the considered subjects. We formalize the difference between a ‘relaxed’ landmarking approach where only validation data is landmarked and a ‘strict’ landmarking approach where both the training and validation data are landmarked. We show that a relaxed landmarking approach fails to effectively use the information contained in the longitudinal outcomes, thereby producing substantially worse prediction accuracy than a strict landmarking approach.
Introduction
Routine collection of a wide array of repeatedly measured patient health outcomes is becoming more and more common, providing vital information about the current status of a patient’s health. Dynamically predicting future (adverse) events for individuals using their currently available information has therefore rapidly become of great interest. One of the main questions for this prediction problem is how to properly extract and use the information contained in the repeated measurements, especially if many variables are available for each subject.
Many different models already exist allowing for the prediction of survival probabilities from longitudinal data. Two commonly used methods are joint modelling (JM) and landmarking. JM of longitudinal outcomes and time-to-event data has risen in popularity as it can be used to simultaneously model the progression of longitudinal data and the survival outcome, allowing information between the two to be shared. A summary of recent developments and issues in JM approaches can be found in Hickey et al. 1 There are a few downsides to JM: a (linear) model must be specified for the longitudinal outcomes, misspecifying the random effect structure in JM can lead to biased estimates and modelling many longitudinal outcomes quickly becomes computationally expensive. To deal with the last problem, Mauff et al. 2 proposed a corrected two-stage approach to cut down computation time significantly, but to the best of the authors’ knowledge it is still not feasible to incorporate more than approximately ten longitudinal covariates in a JM model. The landmarking approach 3 uses a Cox model with a landmarked data set containing only the values of the longitudinal variables until a so called landmark time. As an example, Van Houwelingen et al. 4 have used this model to predict five-year failure-free survival after bone marrow transplantation. Nicolaie et al. 5 further developed this model by incorporating competing risks and proposing a smoothed estimate over a collection of multiple landmarked data sets. As values of longitudinal covariates are not always known at the landmark time, an appropriate model (such as a mixed model) can be used to extrapolate these values in an approach that Ferrer et al. 6 call ‘two-stage’ landmarking. The greatest advantage of landmarking methods is that they are very simple to implement: researchers can simply fit existing models on adjusted versions of the available data.
The traditional landmarking approaches described above utilize only the value of a longitudinal variable at the landmark time, ignoring information contained in the variable progression. A different approach can be taken where the longitudinal covariates and time-to-event data are modelled separately. This requires variable progression to first be described described using an appropriate model. For dynamic prediction this then entails a Landmark longitudinal and survival data. Describe longitudinal trajectories using an appropriate model and extract summaries. Supply above summaries to a survival model and use for future predictions.
In step
An assumption often made when analysing longitudinal data is that subjects are comparable at their entry time into the study. This assumption is implicitly present in FPCA models, where the mean progression of longitudinal variables is assumed to be the same for all subjects. Especially in an observational study this might not be the case, as participants will differ significantly in age at baseline. This gives rise to the hypothesis that there should be an effect of age on the longitudinal trajectories. As an example, brain mass has been shown to increase and decrease throughout the human lifespan. 14 Seeing as age can be included in the baseline predictors, we would therefore like to eliminate the natural variation in the longitudinal trajectories caused by the age disparity between subjects before performing further analyses. We propose an age-based centred (ABC) mFPCA procedure, where the mean value of the longitudinal outcomes is assumed to depend on the age of the single subject. A comparable idea called generalized landmarking analysis has recently been explored by Yao et al. 15
The main focus of this article will be to examine whether we can improve the predictions of the MFPCCox model proposed by Li et al. 8 by using a strict landmarking approach and eliminating the natural variation in the longitudinal variables by using an ABC mFPCA procedure.
This paper is structured as follows. In Section 2 we introduce the notation and discuss the proposed methods. Section 2.6 provides a brief overview of the considered methods. We evaluate the proposed models in Section 3 by means of a simulation study. In Section 4 we apply all different methods on an observational study on Alzheimer’s disease (AD). The article ends with a discussion in Section 5.
Methods
In this section, we describe a three-step approach to dynamically predict survival probabilities.
Notation
Consider a study with patients
At the first patient visit,
Let
In (dynamic) prediction modelling, the data is usually separated into a training and test data. The training data is used to build a prediction model and the performance of the model is then evaluated on the test data. A common approach is to use (repeated)
Step 1: Landmarking
The first step in our method will be to landmark the data, removing information that can skew predictions for future patients.
Strict landmarking
We consider a ‘strict landmarking’ approach for dynamic prediction as proposed by Van Houwelingen.
3
We landmark both the training and the testing data set by removing all patients with an observed event before a landmark time
We only train our model on the longitudinal trajectories of the remaining patients until landmark time. In other words, we truncate the data
Relaxed landmarking
A popular approach is to use what we call ‘relaxed landmarking’. In this landmarking approach, the training model uses the complete information of all individuals in the training set. For the prediction set, only the observations up until landmark time are used. This means that more information is used to build the model than is available at the time of prediction. This can lead to a biased model, as patients that have already experienced an event at the landmark time are not likely to follow the same longitudinal patterns as patients without an event at the landmark time. The relaxed landmarking approach has already been used before on the Alzheimer Disease Neuroimaging Initiative (ADNI) data set considered in Section 4.1.8,9,16
Step 2: Functional principal component analysis
For each
uFPCA
For each
Consider the values of the
Instead of decomposing each of the
Happ and Greven
19
show that there is a one-to-one correspondence between the univariate and multivariate decomposition(s). Using this correspondence it is possible to obtain the multivariate scores and eigenfunctions from
By fitting the model on the training data we obtain eigenfunctions and eigenvalues, as well as patient specific scores. To determine scores for new patients in the test data, we can follow the same procedure, starting with equation (3). Note that except for
In equations (1) and (4) both the mean functions and eigenfunctions are modelled on the time-on-study scale. However, for most covariates of interest modelling their dynamic evolution as a function of time-on-study may be unrealistic, as one would expect the value of the longitudinal covariates to depend on the age of the patient, rather than on the time since their entry into the study. For example, brain mass is unlikely to depend on the time spent in a study but has been shown to depend on the age of the subject.
14
For this reason, we consider an alternative modelling approach where the mean of the longitudinal processes
To obtain an estimate
When using relaxed landmarking, the uFPCA models are built on the complete trajectories of patients in the training set. This means that there is usually plenty of information to train the model, even when there are a lot of missing observations. When strict landmarking is applied, the training models are only built on the trajectories until landmark time. Besides this, patients with an event time before the landmark time are not considered, meaning that the prediction model is trained on a much smaller set of data. Combined with missingness in the data, this can lead to difficulties in the estimation of the mean functions
Another issue arises for data with a large proportion of missing data. The scores for both training and test data are calculated using PACE (see equation (3)). For each patient this requires the calculation of
Step 3: Dynamic prediction of survival
In step 2 we estimated patient specific scores
When the number of longitudinal covariates is large, the number of scores
Model validation
We evaluate the accuracy of the predicted survival probabilities
We employ repeated cross-validation to obtain unbiased estimates of predictive performance (as measured by tdAUC and BS) by repeatedly splitting all available data randomly into
Summary
There are three choices to be made in the methods discussed above:
Strict/Relaxed Landmarking. Standard/ABC mFPCA. Standard/Regularized Cox regression.
The methods are also graphically summarized in Figure 1. The model with relaxed landmarking, the standard mFPCA procedure and standard Cox regression is the MFPCCox framework proposed by Li and Luo.
8
We will consider MFPCCox as a reference method and evaluate the effect of the considered additions by comparing the performance between methods in a simulation study as well as on a real data set.
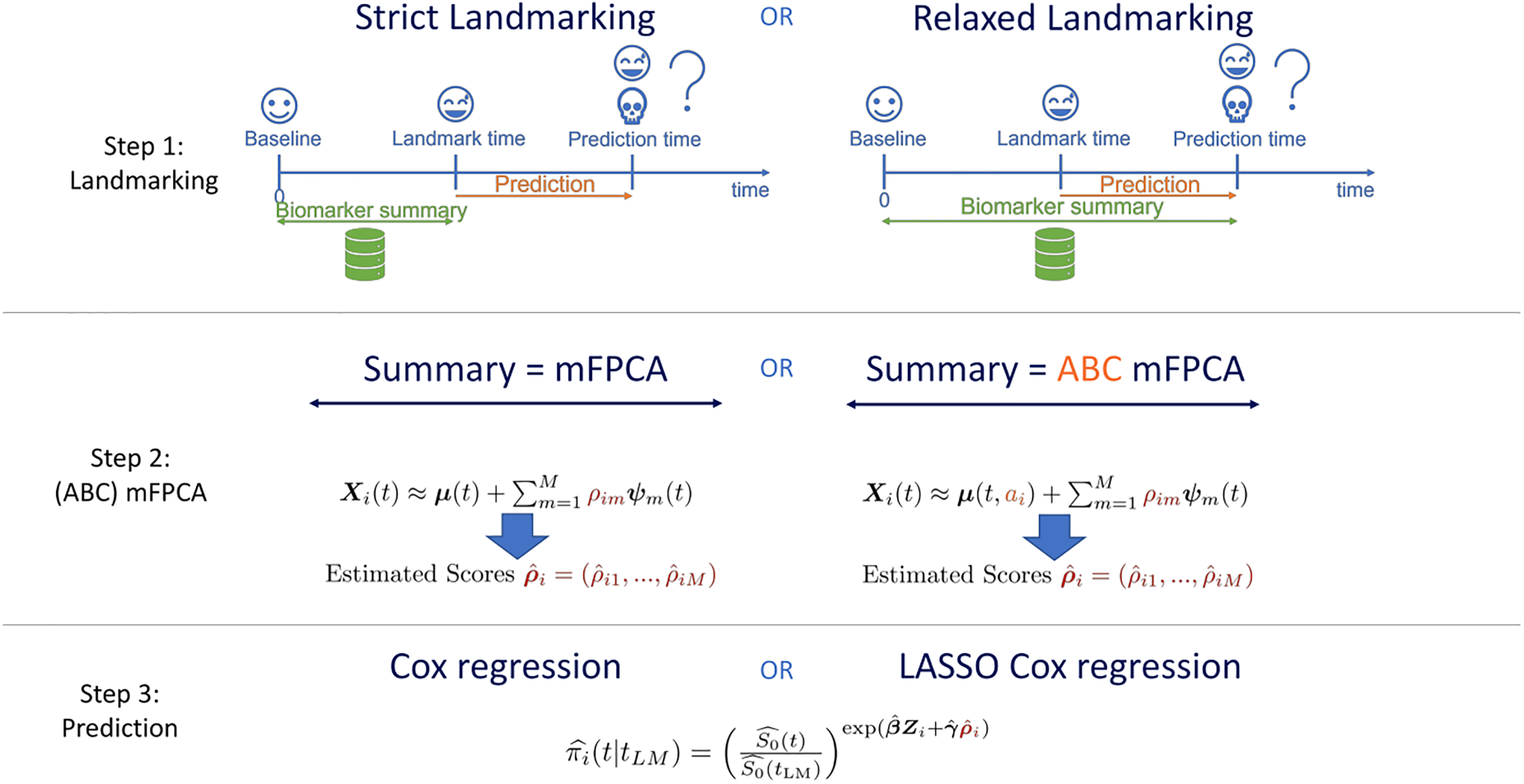
Graphical summary of the methods proposed in Section 2. See also Section 2.6.
In this section, we perform a simulation study to evaluate the impact on prediction accuracy of the following choices in
Data generation
We consider a study where
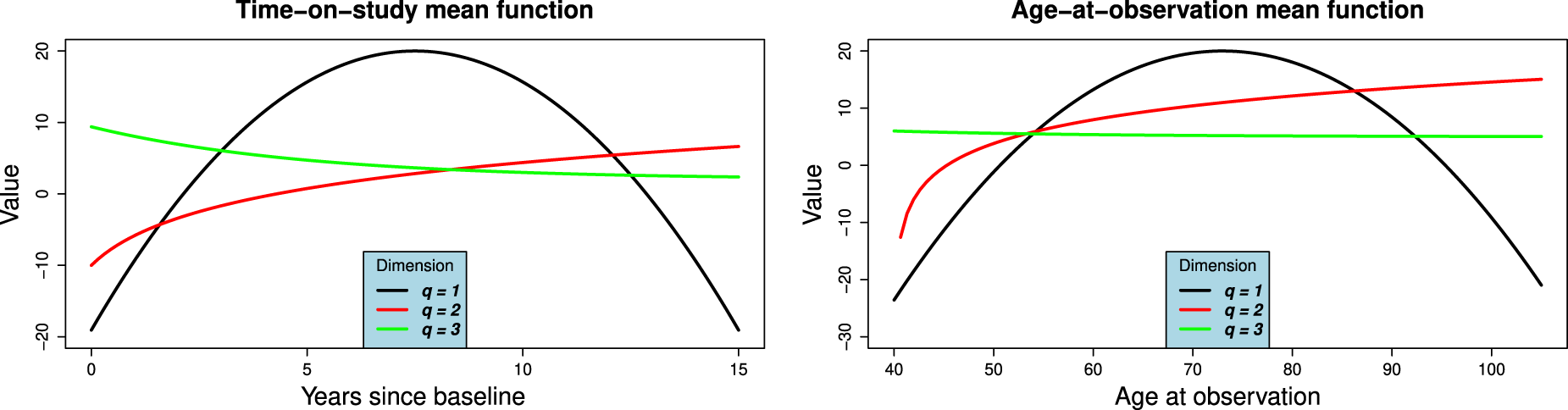
(a) Time-on-study and (b) Age-at-observation mean functions for simulation study. The functions correspond to equation (9).
Survival times are generated by applying the inverse transformation method
32
as follows. Consider the subject-specific hazard function:
Time-on-study data and light censoring Time-on-study data and median censoring Time-on-study data and heavy censoring Age-at-observation data and light censoring Age-at-observation data and median censoring Age-at-observation data and heavy censoring
Note that the hazard used to simulate survival outcomes (equation (10)) is not the same as the one used by the models (equation (8)). If we were to simulate survival times from the hazard in equation (8) we would favour relaxed landmarking approaches as the scores that influence the survival times can only accurately be determined from the full follow-up duration in that case. We therefore take a different approach, where the hazard depends on the current value of the longitudinal trajectories for each patient. With this procedure neither the strict nor relaxed landmarked model is correctly specified. We believe however that this simulation procedure is more realistic, as it seems more likely that the progression of biomarkers is predictive for the occurrence of an event as opposed to some underlying latent factor that influences the biomarkers trajectories. An important consideration in this simulation study is that the age of a subject does not influence their survival directly, but only through how well the true scores can be recovered from the generated data after substracting the estimated mean function.
Having simulated longitudinal covariates (both on the time-on-study and age-at-observation scale) and survival times in each scenario, we now apply the models discussed in Section 2. We do not consider regularization in the simulation study, as we are working in a low-dimensional setting and the true scores are uncorrelated. We choose
The main goals of our simulation study are to evaluate the effect on prediction accuracy of three components in the proposed methods. First of all, we are interested in what happens when the data-generating mechanism has been misspecified in the model: What happens to ABC models in a time-on-study scenario and what happens when not performing ABC in an age-at-observation scenario? For this reason, we generate data in both mechanisms. Secondly, we want to explore whether strict landmarking can outperform relaxed landmarking or vice versa? Which one yields the best predictions and does this depend on the landmark time? To examine this, we consider three different landmark times at three, six and nine years after the start of the study. With the employed survival generation mechanism, most people will still be at risk at the first landmark time, approximately half will have had an event at six years and most people will have failed by the last landmark time. Finally, we are curious to see how censoring patterns affect the predictive performance of the models and whether it influences the previous two points. We therefore consider the three different censoring patterns (light/median/heavy).
To keep the results clear, we only show the results for median censoring (scenarios
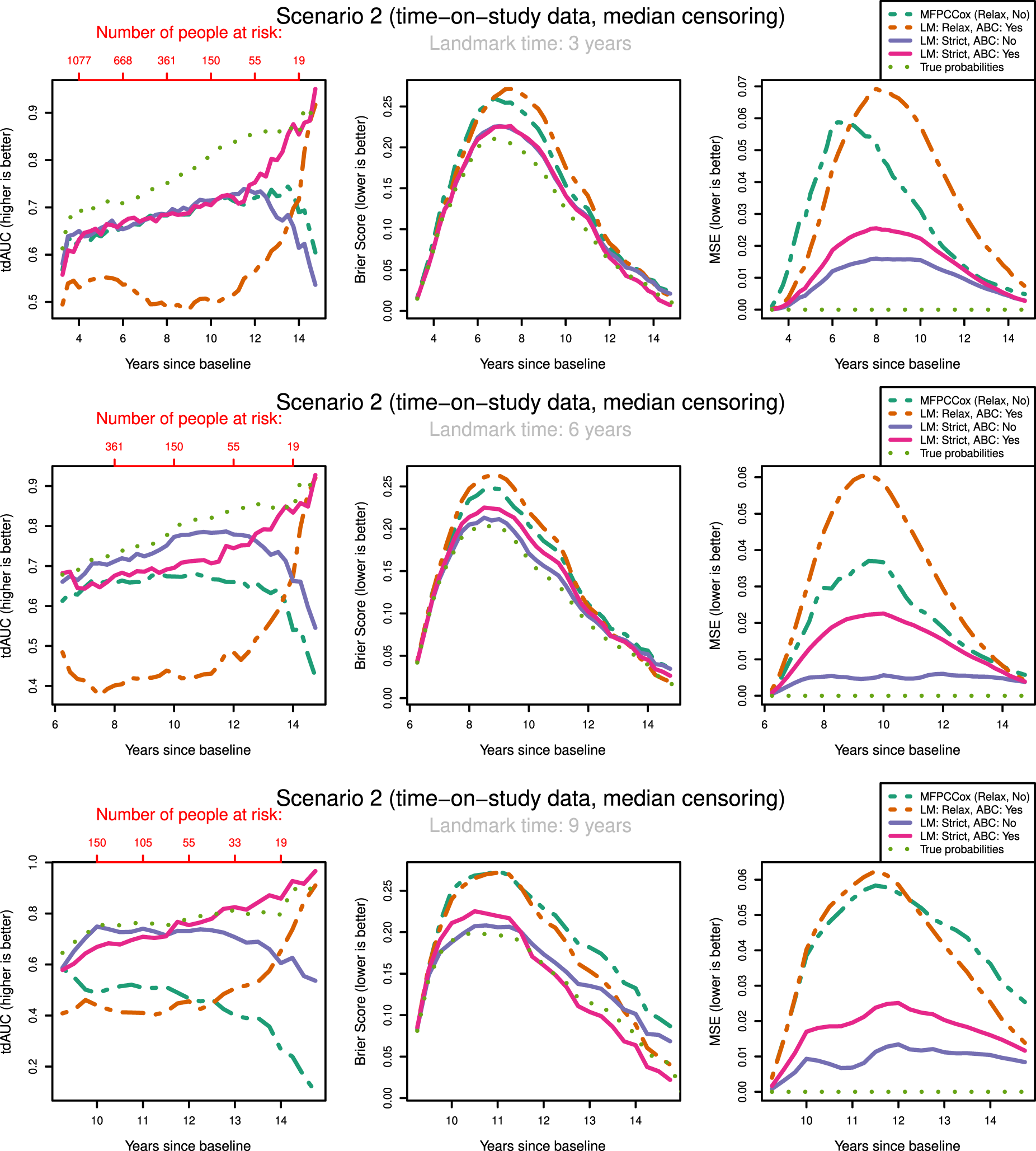
tdAUC, Brier Score and MSE in the second scenario (time-on-study data, median censoring) for the considered methods over landmark times at
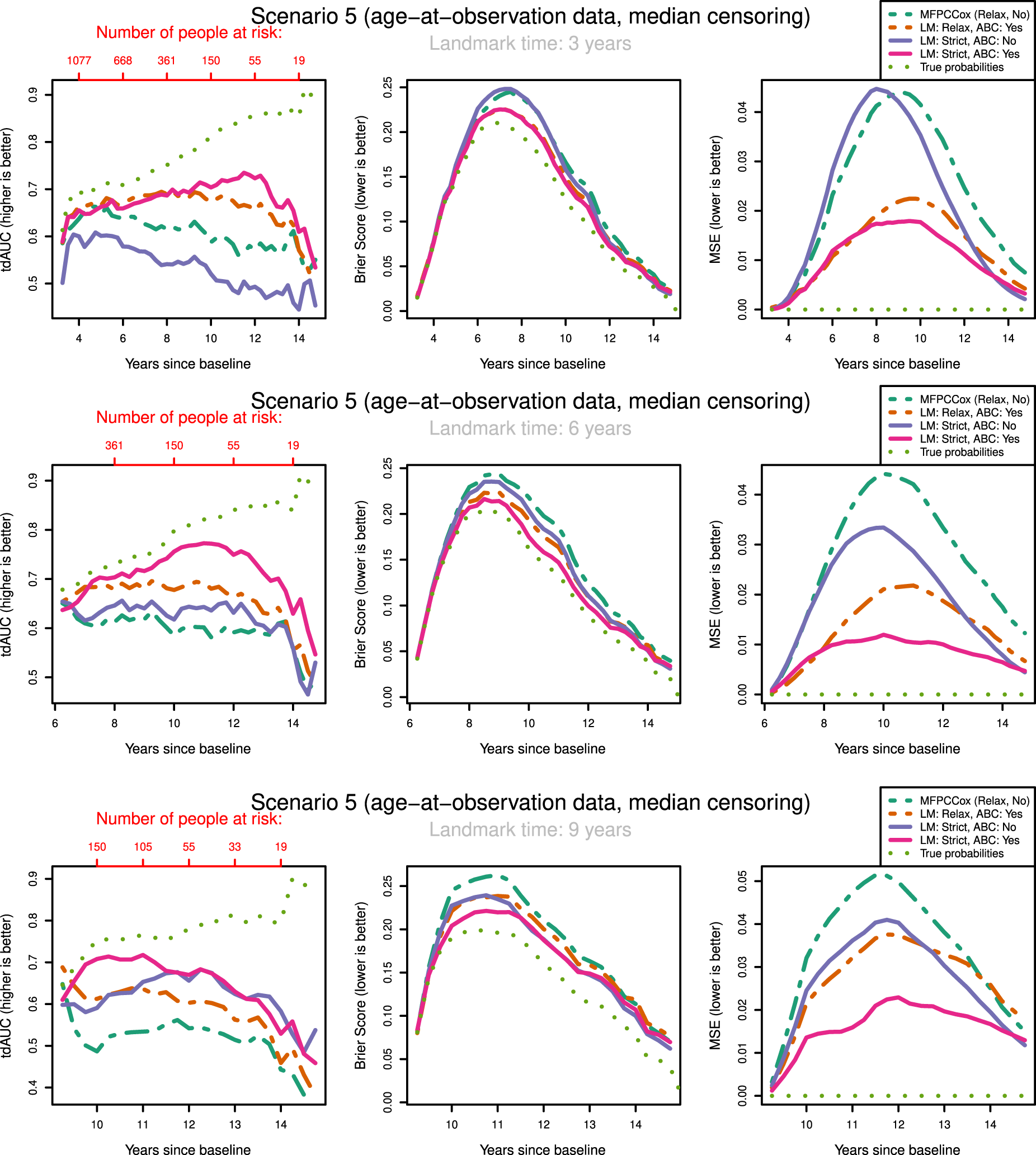
tdAUC, Brier Score and MSE in the fifth scenario (age-at-observation data, median censoring) for the considered methods over landmark times at
The most notable result is that in both scenarios and over all landmark times the strictly landmarked correctly specified model performs best in MSE. Let us compare the MSE of the ABC methods between strict and relaxed landmarking in Scenario
We might expect that strict landmarking methods will perform worse at later landmark times as they are using less data to build the models, seeing as most subjects will have had an event at later times and therefore be removed from the training set. This reverse is the case, with relaxed landmarking performing worse as the landmark time increases. This is likely due to relaxed methods using a very unsuitable training set: using data from people who have already had an event and using future observations to predict past ones.
Surprisingly, we find that Strict ABC landmarking can perform extremely well in the time-on-study scenarios when considering tdAUC and BS. At some prediction time points, the predictive performance of this model can even exceed that of the perfect prediction model, see for example Figure 3(c). Additionally, the Relaxed ABC method can also perform very well at later time points. Although looking at the Brier score we would conclude that both these methods are performing better than the correctly specified strict landmarking, we can see that in MSE they are performing significantly worse. Upon examining the intermediary results of these models, we found that the misspecified relaxed/strict models were estimating scores with extremely high variances which were not found to have predictive power in the resulting Cox model. Further inspecting the predicted probabilities of these models, we found that these were indeed very suitable to discriminate between subjects at later time points, but recovered the true probabilities very poorly. Note that this only ‘improves’ their predictive capabilities when very few people are at risk. When looking only at tdAUC and BS measures however, this can lead us to believe that they are performing very well. This can be a problem for evaluating the prediction of the models on real-life data.
The question arises whether correctly specifying the landmarking approach or the mean function is most important. We can see in Figure 3 that strict landmarking approaches perform best (in MSE), with correctly specified ABC performing best. The relaxed methods display significantly worse MSE over all landmark times and all censoring patterns (see also Supplemental Figures
In our discussions above it seems that there are multiple biases working against each other, making it difficult to pinpoint what exactly causes one method to perform better or worse than the other. In general, we can conclude that relaxed landmarking introduces an estimation bias into the mFPCA procedure due to using future observations and an unsuitable training set. We find this bias to be largest when the training sets between the methods differ most, which in our simulation study is at later landmark times as then most subjects will already have experienced an event. On the other hand, there is also the bias of misspecifying the data generation mechanism. This can result in worse estimates for the survival probabilities, but has less of an impact in the time-on-study scenarios.
Let us examine how censoring influences the predictive potential of the models. In both the time-on-study and age-at-observation scenarios, the degree of censoring does not influence the performance between strict landmarking methods in MSE. For relaxed landmarking models, it is unclear what the influence of censoring is on the performance. For light and median censoring correctly specified relaxed landmarking performs better than incorrectly specified relaxed landmarking, especially at earlier landmark times. At later landmark times, both have comparable performance in MSE. In heavy censoring scenarios, incorrectly specified relaxed landmarking can perform better than correctly specified relaxed landmarking in MSE (see Supplemental Figure 2). As this is also pronounced in the BS, we expect to notice this in real-life applications as well. Censoring degrees seem to only influence the correctly specified relaxed landmarking methods, whereas incorrectly specified never show good predictive potential and are therefore not influenced much by the censoring rates.
Overall scenarios and all landmark times, the results in BS and tdAUC align very well: whenever a method performs better/worse in tdAUC it also does so in BS. Unfortunately, the patterns we see in MSE are completely not visible in either tdAUC or BS. This is alarming as it means that in a real-life application it will be impossible to conclude whether a method is really recovering the true survival probabilities well and therefore whether it is appropriate to use for dynamic prediction purposes.
In conclusion, misspecifying the data generation mechanism can lead to very unstable estimates in the models. This can lead to poor predictive power, but sometimes the reverse can be visible in Brier or tdAUC scores. It is crucial to perform strict landmarking to obtain good predictive performance. Using a relaxed approach might seem like a good idea due to the increase in available information to build the model, but results in biased estimates and poor predictions. The degree of censoring only has an influence on relaxed landmarking models, and only at heavy degrees of censoring around
In this section, we compare the performance of all methods previously discussed to predict time to dementia (DM) in a real-life study on AD which is a degenerative brain disease and is the most common form of DM, 33 causing patients to experience progressively worsening cognitive capabilities. A large population of healthy and afflicted individuals is currently being followed by researchers in the ongoing ADNI study (https://adni.loni.usc.edu/). Multiple genetic and longitudinal biomarkers are being collected over the duration of the study, such as structural MRI/PET scans and questionnaires assessing the neurocognitive abilities of participants. There are currently no disease-modifying treatments available to reverse the damage caused by AD, but treatments to slow the progression of the disease are available. 34 Seeing as many countries have an aging population, estimating the remaining time in which an individual will be disease free is becoming more and more important.
ADNI data description
ADNI is an ongoing observational study initiated in October
Our goal is to predict time to DM for individuals that enter the study as MCI or CN. Therefore we restrict our attention to participants without DM at baseline. As a result, we exclude
Description of the ADNI data set.
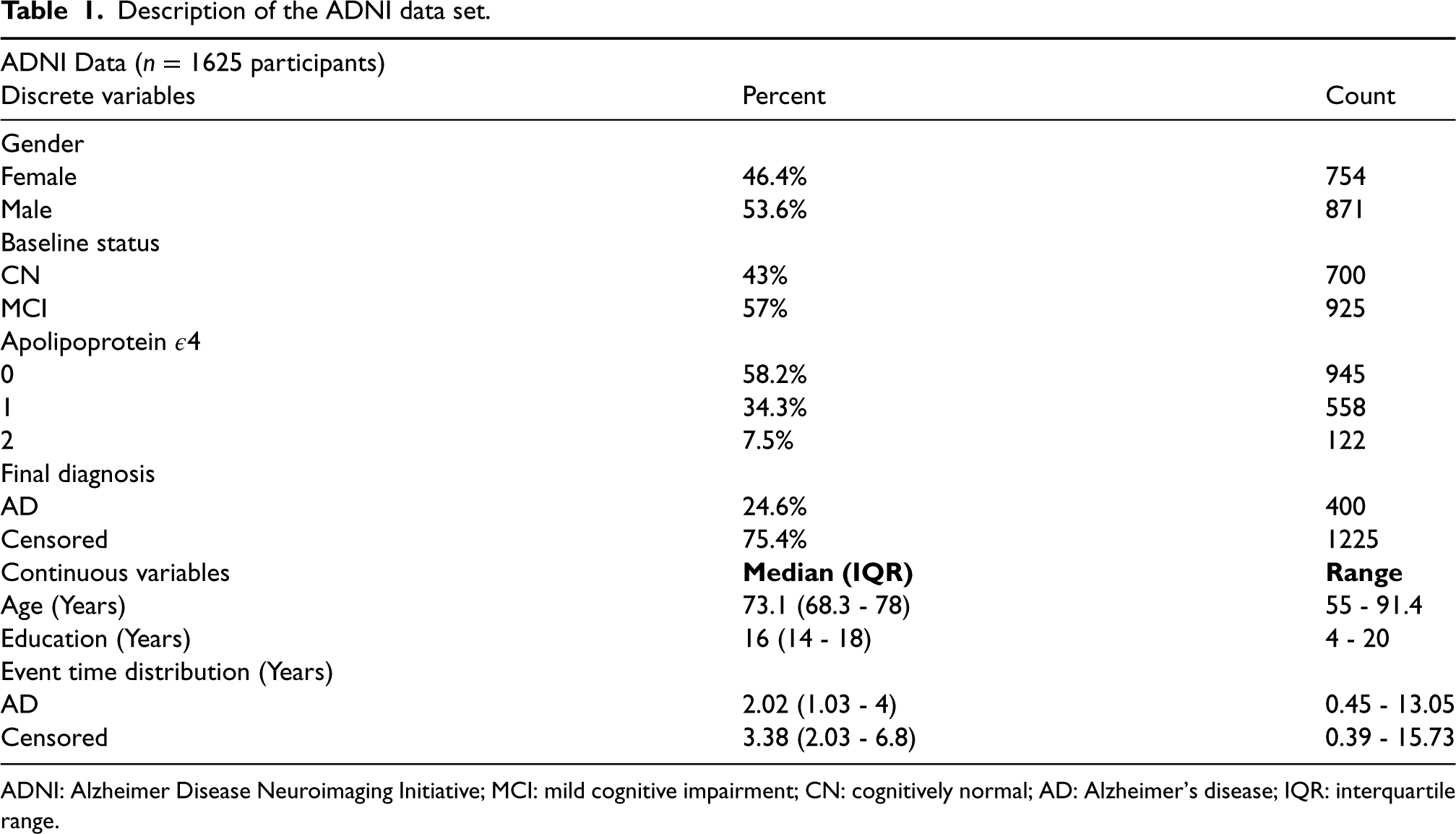
Description of the ADNI data set.

ADNI: Alzheimer Disease Neuroimaging Initiative; MCI: mild cognitive impairment; CN: cognitively normal; AD: Alzheimer’s disease; IQR: interquartile range.
Study participants were followed up over half-year intervals for at most
As discussed in Section 2.3.4 the strictly landmarked model is built on less data than the relaxed model. Due to the low density of observations at early time points in some variables, we cannot fit the strictly landmarked model on all
The results are shown in Figure 5. We can see a clear distinction in performance between strict landmarking and relaxed landmarking methods, with strict landmarking performing better on both tdAUC and BS. The overall best-performing method seems to be the LASSO regularized strictly landmarked MFPCCox (lightgreen line). MFPCCox has the worst predictive performance over all landmark times, with any considered addition improving on this. Over all landmark times, we can see that methods performing better in tdAUC also perform better in BS and vice versa. We also observed this phenomenon in our simulation study in Section 3. Additionally, the best-performing relaxed landmarking method changes from regularized MFPCCox at early landmark times to age-based centered MFPCCox at later landmark times. The age-based centered and uncentered strictly landmarked methods have near identical performance, at all times outperforming the best relaxed landmarking method.
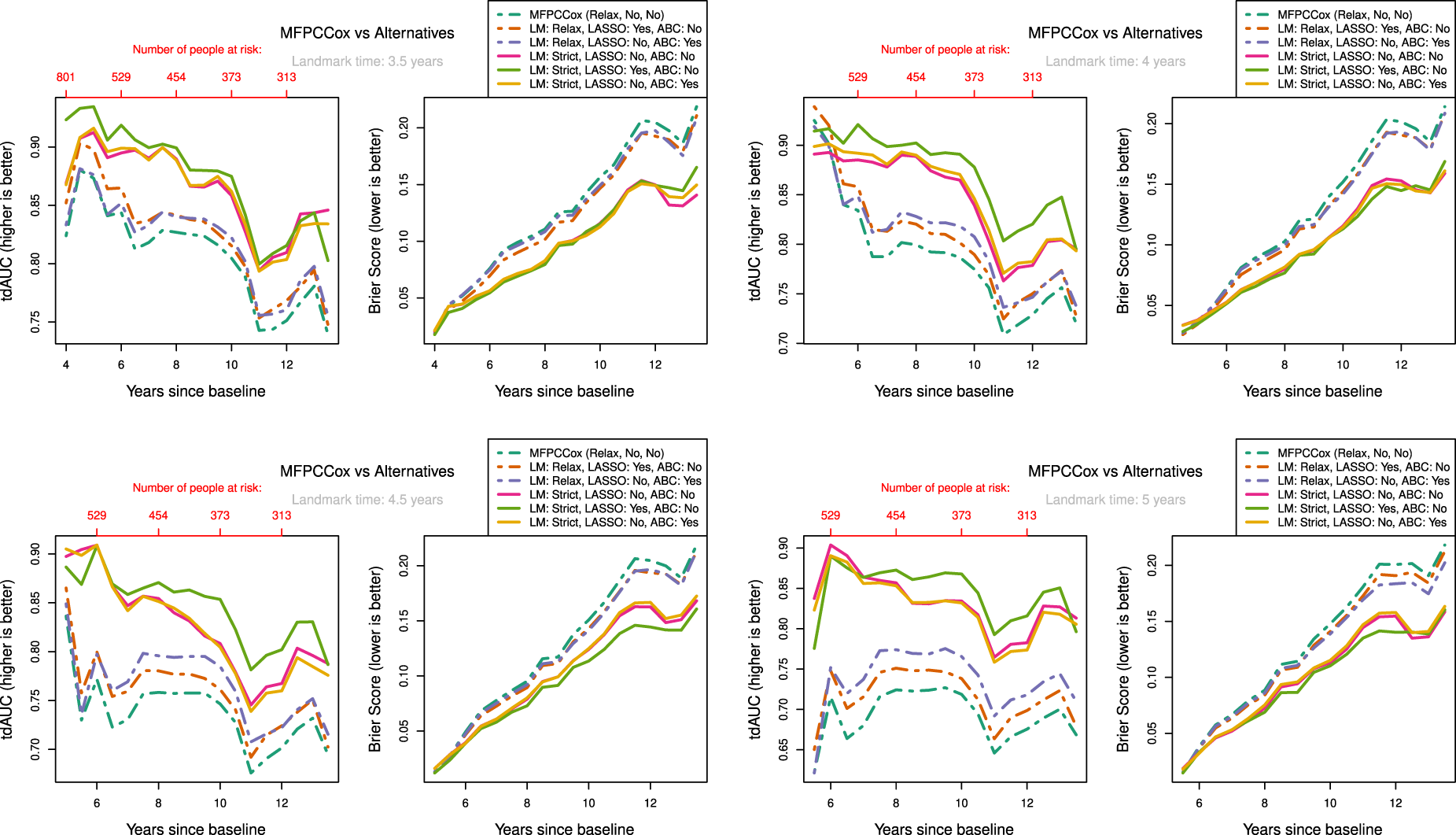
Measure of performance for LM, LASSO regularization (LASSO) and ABC methods at different landmark times on ADNI data. Validation scores were determined by using
Interestingly, for strictly landmarked models age-based centering does not seem to improve prediction accuracy at all, while for relaxed landmarked models age-based centering improves prediction as the landmark time becomes larger (see Figure 5(c) & (d)). A possible reason for this is that with strict landmarking the mean on the age time scale
A notable effect of using relaxed landmarked methods is the sharp drop in prediction accuracy right after the landmark time when compared to strictly landmarked methods. This happens because the relaxed model fit on the training data is not necessarily representative for participants who are event-free at landmark time and is one of the main reasons to consider strict landmarking. Finally, LASSO regularization seems to improve prediction accuracy for both strictly and relaxed landmarked methods, although not by much.
Remember that age is included as baseline predictor in all considered methods. Examining the Cox prediction models, we find that when not using ABC age is not found to be a significant predictor of DM. This is the case for both strict and relaxed landmarking methods. On the other hand, when using ABC age is found to be a significant predictor. As an example, in one of the evaluation folds we found p-values for age of
Let us compare the results of the application with those of the simulation study in Section 3. In the ADNI data around
In this article we introduced the concept of relaxed and strict landmarking. We developed an age-based centred mFPCA procedure, which can be used to remove the variation due to the difference in age at baseline of subjects participating in a study. Even though our methodology focused on age as centring variable, the procedure can be extended to any time-dependent variable. Finally, we used a regularized Cox model to dynamically predict time to DM.
Results based on the simulation study and on the real data application show that improperly landmarking can lead to biased results in dynamic prediction models, thereby strongly decreasing predictive accuracy. It is therefore important to use strict landmarking approaches so that accurate predictions can be made. In practice, it might not always be possible or desirable to use strict landmarking. For the ADNI data, we did not consider landmark times before
In the analysis of the ADNI data set, there is a big improvement in prediction accuracy when using age-based centring methods with relaxed landmarking. The results from our simulation study suggest that although using an age-based centred procedure might be appropriate for the ADNI data, the largest gain in predictive potential can be gained by using a strict landmarking procedure. In other words, the landmarking bias outweighs the bias incurred by incorrectly specifying the underlying data generation mechanism. A limitation is that we have considered only two options: Age-based centring for all variables or for none. Exploring all possible combinations of variables to use for age-based centring was not feasible; with proper medical knowledge it might be possible to consider only few covariates for age-based centring. It is therefore possible that an improvement in prediction accuracy can be achieved when considering only a relevant subset of covariates. This is especially emphasized by the result from the simulation study which showed that using a time-on-study model for age-at-observation data resulted in very poor predictive accuracy.
In the ADNI study, all subjects did not follow the study plan, either by not showing up to the planned follow-ups or at the planned dates. As a consequence the resulting longitudinal information is sparse and the observed time grid is not regular. Even though FPCA methods do not strictly require a regular grid, it becomes computationally infeasible to estimate the mean and covariance functions on an irregular grid when observations are very sparse. We therefore assumed the grid to be regular by considering the follow-up to have happened at the closest planned assessment time. This can have a great effect on the estimation procedures, affecting the resulting scores and eigenfunctions such that they do not represent subject progressions appropriately. On the other hand, mixed modelling approaches such as
JM approaches are often employed to model survival and longitudinal outcomes jointly. Li et al. 8 have performed a simulation study to compare the performance of MFPCCox with that of a JM approach. They found that when the parametric form of the joint model was misspecified, MFPCCox achieved comparable predictive performance while more accurately recovering the underlying longitudinal trajectories. As our proposed methodology has been shown to improve on MFPCCox, we expect to outperform JM approaches even further. Li et al. 8 also discuss the computational burden of JM approaches, noting that the JM approaches are not computationally feasible with more than six longitudinal variables. We can therefore not fit a joint model on the ADNI data considered in Section 4.
A different approach for the multivariate principal component analysis decomposition (mFACEs), using tensor product B-splines to estimate the covariance function directly was proposed in Li et al. 16 The authors show in a simulation study in a sparse setting that mFACEs captures cross-correlation between functions and recovers the eigenfunctions and eigenvalues better than mFPCA, as well as better predicting the longitudinal biomarkers in the ADNI data. As mFACEs uses a multivariate version of PACE (3) for score estimation, this could mitigate the problem of missing data discussed in Section 2.3.4, possibly allowing for earlier predictions with strict landmarking. Additionally, age-based centring can also be incorporated into the mFACEs procedure. Lin et al. 9 showed that mFPCA performed slightly better in survival prediction than mFACEs when using relaxed landmarking with a Cox model, both in a simulation study as well as on the ADNI data. They also concluded that RSFs outperformed Cox models for survival prediction. These results should be re-evaluated with a strict landmarking approach, as the bias incurred by relaxed landmarking might favour RSF over Cox models.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802231224631 - Supplemental material for Dynamic prediction of survival using multivariate functional principal component analysis: A strict landmarking approach
Supplemental material, sj-pdf-1-smm-10.1177_09622802231224631 for Dynamic prediction of survival using multivariate functional principal component analysis: A strict landmarking approach by Daniel Gomon, Hein Putter, Marta Fiocco and Mirko Signorelli in Statistical Methods in Medical Research
Footnotes
Acknowledgement
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The data used in the application section were obtained from the ADNI project. The project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012).
Footnote
The ![]() .
.
Data Availability Statement
Data used in the application section were obtained from the ADNI database (http://adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this paper. A complete listing of ADNI investigators can be found at: ![]() .
.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.