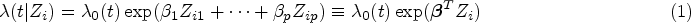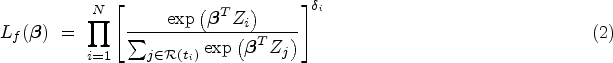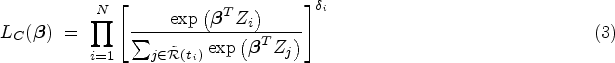Abstract
Analyses of distributed data networks of rare diseases are constrained by legitimate privacy and ethical concerns. Analytical centers (e.g. research institutions) are thus confronted with the challenging task of obtaining data from recruiting sites that are often unable or unwilling to share personal records of participants. For time-to-event data, recently popularized disclosure techniques with privacy guarantees (e.g., Differentially Private Generative Adversarial Networks) are generally computationally expensive or inaccessible to applied researchers. To perform the widely used Cox proportional hazards regression, we propose an easy-to-implement privacy-preserving data analysis technique by pooling (i.e. aggregating) individual records of covariates at recruiting sites under the nested case-control sampling framework before sharing the pooled nested case-control subcohort. We show that the pooled hazard ratio estimators, under the pooled nested case-control subsamples from the contributing sites, are maximum likelihood estimators and provide consistent estimates of the individual level full cohort HRs. Furthermore, a sampling technique for generating pseudo-event times for individual subjects that constitute the pooled nested case-control subsamples is proposed. Our method is demonstrated using extensive simulations and analysis of the National Lung Screening Trial data. The utility of our proposed approach is compared to the gold standard (full cohort) and synthetic data generated using classification and regression trees. The proposed pooling technique performs to near-optimal levels comparable to full cohort analysis or synthetic data; the efficiency improves in rare event settings when more controls are matched on during nested case-control subcohort sampling.
Introduction
Large cohort biomarker studies of rare diseases or expensive-to-recruit studies often require merging datasets collected across multiple study centers or databases. Integrating individual data from contributing centers/nodes is a major challenge due to legitimate privacy and ethical concerns. 1 Analytical centers thus resort to disclosure control techniques such as intermediate statistics release or synthetic data generation to conduct etiologic studies or make predictions about relevant clinical endpoints.2–5 Sampling 6 and noise perturbation 7 are also sometimes used.
In recent years, the strengths of proposed privacy-preserving techniques have improved considerably. Traditional attempts at inducing privacy, for example, k-anonymity, were primarily focused on masking quasi-identifiers using de-identification techniques such as generalization and suppression.8,9 However, with recent advancements in computational power and our presence in multiple social network databases, the potential risk of re-identification is high. 10 More refined techniques like t-closeness promise better privacy guarantees but the strength of their effectiveness is largely dependent on the reliability of our assumptions about the intruder. 11 Techniques motivated from cryptography remain the strongest privacy guarantees; they often require different parties run known learning algorithms on a merged version of the local datasets without revealing individual data. Prediction from the learned model however requires the participation of each unit to implement a private scoring algorithm. Consequently, a major challenge of such techniques is the need for high communication and computational cost. 12
Although not motivated by data privacy concerns, sample aggregation (referred to as pooling henceforth) was initially proposed as a method to circumvent the challenge of testing individual samples in a limited resource setting during World War II. 13 The method allowed practitioners to combine samples from multiple individuals for a single test in order to reduce the cost of resources needed to test for syphilis. The technique has seen some advancements in recent years, especially for identifying infectious agents in low prevalence settings 14 and more recently for testing Covid-19. 15 Alongside these developments, many variations of pooling have been proposed to infer associations between various clinical outcomes of interest and relevant covariates.16–18 These techniques have become particularly appealing for the analysis of high-dimensional data where the strategy is utilized to compensate for limited samples or high biological variation. 19 Pooling is also sometimes used in the literature to mean combining records from various contributing data sites; however, we restrict our use of the term to mean aggregating individual-level covariate.
Of interest in the current article is the application of pooling to preserve patient privacy during the analysis of time-to-event (or survival time) data in a distributed data network. Survival analysis plays a fundamental role in biomedical research, where survival probabilities are routinely computed to inform clinical decisions.20,21 Several approaches have been proposed for protecting patient privacy when dealing with survival time data.22–25 For example, O’Keefe and Rubin 22 introduced a privacy technique based on data suppression (e.g. removal of censored events), smoothing, and data perturbation. Yu et al. 23 also proposed a method based on affine projections of the Cox model. However, these techniques have been challenging to adapt in practice for several reasons: (i) require sustained communication between the analytical center and data node to train the model, for example, distributed regression techniques, (ii) high computational cost to generate synthetic data at the study node, for example, classification and regression trees (CART) synthetic microdata generation, 4 PATE-GAN, 26 Differentially Private Generative Adversarial Network (DPGAN), 27 etc., and (iii) inaccessible to applied researchers without formal statistical training due to intricacies associated with sampling synthetic data from posterior predictive distributions. While pooling has been adapted to preserve patient privacy when analyzing binary outcome data in multi-center studies, 28 the framework has not been extended to survival-time data under privacy restrictions.
The objectives of the current study include (i) to propose an aggregation based, easy-to-implement, study design for time-to-event outcome under disclosure limitation, (ii) to estimate the hazard ratio of the postulated Cox proportional hazards (PH) model using the aggregate data from the proposed pooling design, and (iii) to estimate the full-cohort survival curves based on the subset of patients that were included in the pooling design. The rest of the manuscript is organized as follows. In Section 2, we first introduce the pooled study design for time-to-event outcome based on a nested case-control subset of the full cohort. We then introduce the full cohort (individual) and pooled subcohort Cox partial likelihood functions under nested case-control sampling. We show that the individual and pooled likelihoods are of the same form and inference could be carried out using existing analytic tools designed for individual likelihood functions. A sampling technique for generating the survival curve based on the subjects included in the pooled design is proposed, and compared to the full cohort and synthetic data. Section 3 discusses various performance assessment metrics relevant for our setting. We present simulations and a real-life example in Section 4 and close with a discussion in Section 5.
Methods
Consider a clinical study involving sensitive biomedical data, such as the time-to-death records of HIV/TB co-infected patients, distributed among multiple study centers. Assume that, due to confidentiality agreements and legitimate privacy concerns, the centers are unable to share individual data with analytical centers (see Figure 1 for a schematic representation of the distributed data setting). Our proposed design, based on nested case-control (NCC) subcohort sampling, provides an alternative approach to estimate the hazard ratio (HR) or overall survival curves of time-to-event data while limiting the disclosure of sensitive information from individual-level data. The approach is executed in three stages:
NCC design stage: NCC sampling of participants conditional on the event times, Pooling design stage: aggregation/pooling of sampled subcohorts based on event status prior to data sharing, and Estimation stage: estimation of model parameters from the pooled subcohorts and reconstructing approximate event times for pooled cases and controls that constitute the matched sets to estimate overall survival curves.
Steps (1) and (2) are sequentially conducted at each contributing data nodes before the final step (3) is performed at the central analyses node.
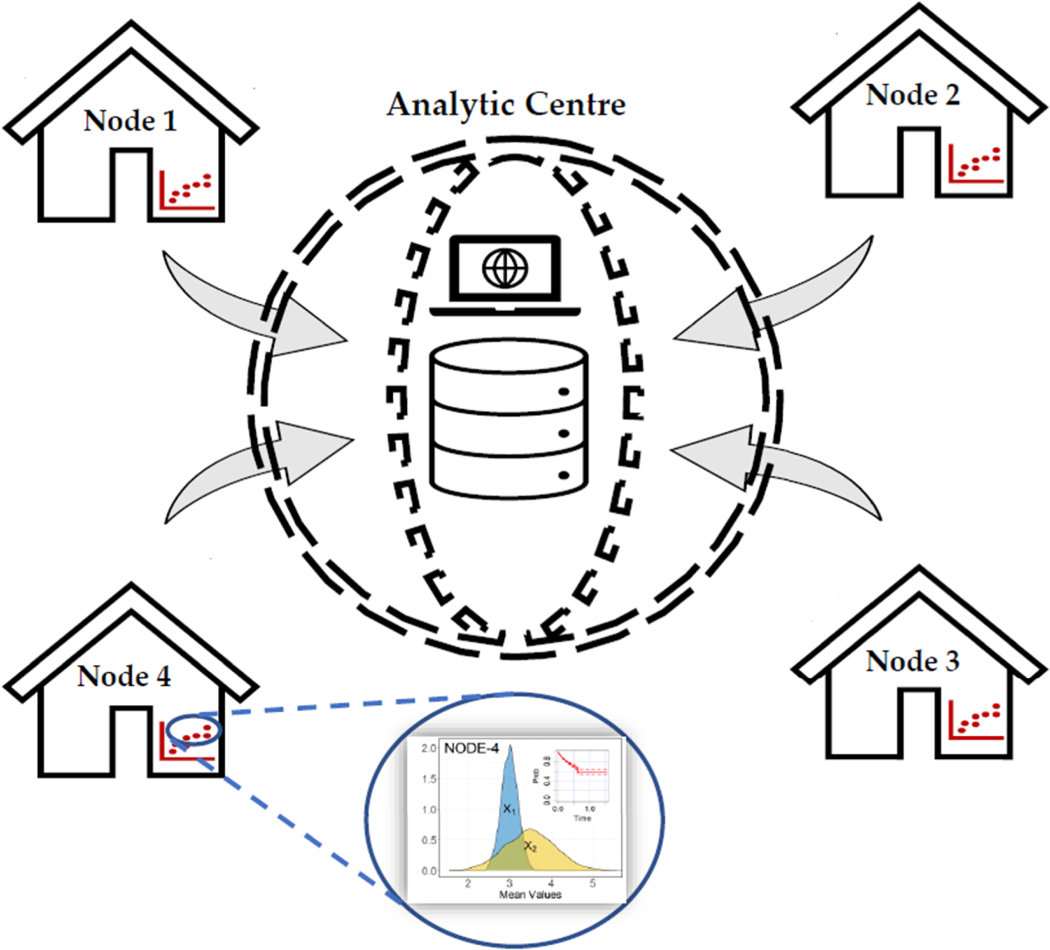
Problem setting—horizontally partitioned time-to-event data between study nodes 1–4. The nodes are only allowed to share aggregate/pooled data with the analytic center. Individual data cannot be shared between nodes or with the center.
Notation and methods
Let
Likelihood for individual-level data
Let the at-risk indicator for subject
NCC subcohort sampling provides an attractive alternative to full cohort analysis, particularly when the analysis is constraint by the amount of available resources 30 or the amount of information that could be released. 31 This idea provides a key motivation for the proposed method where aggregate values of NCC subcohorts are utilized to preserve individual records during Cox regression modeling. NCC sampling is also sometimes referred to as riskset sampling or incidence density sampling. A hypothetical example is presented in Figure 2.
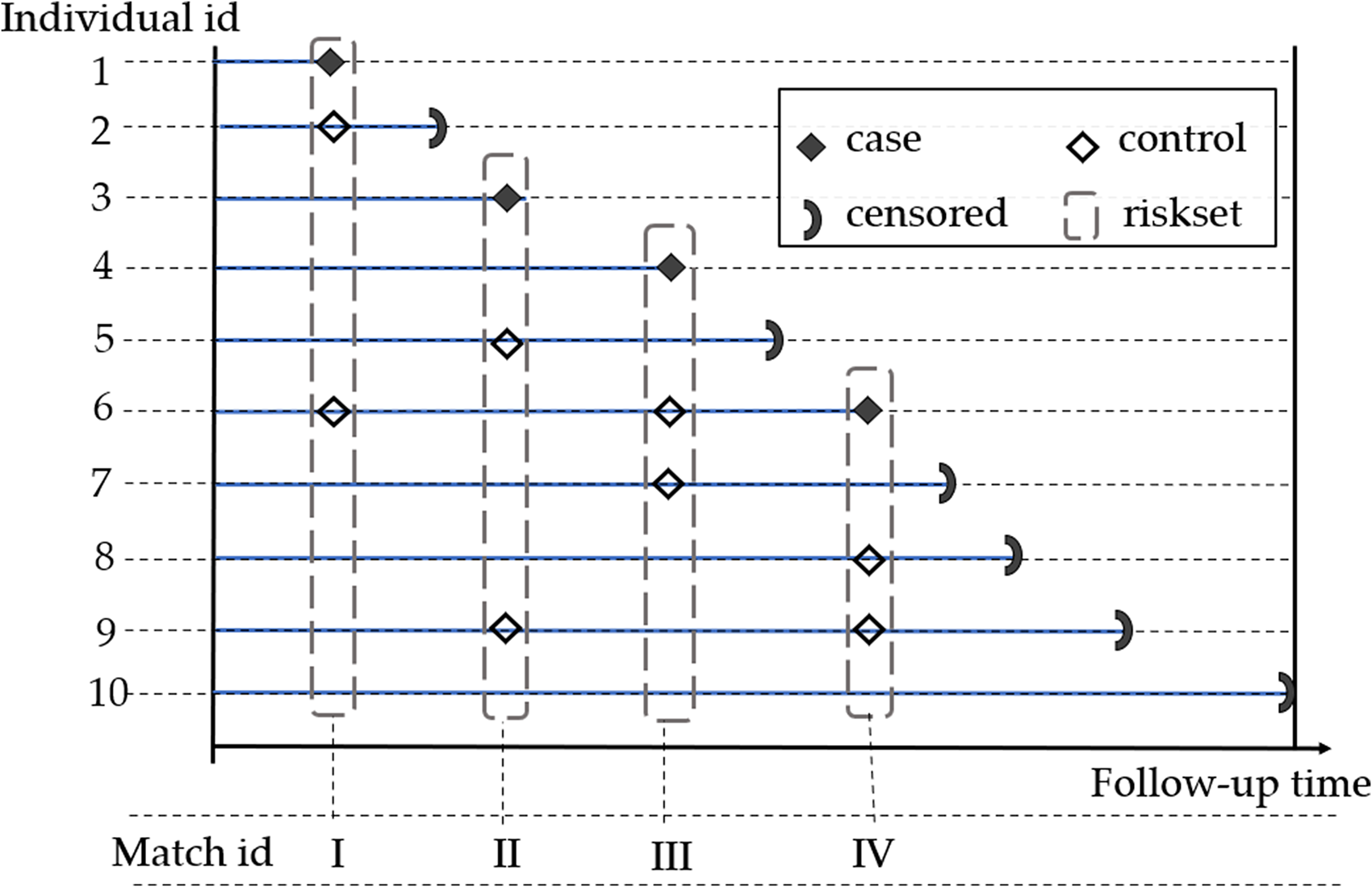
Nested case-control sampling example of two controls per case in a hypothetical cohort of 10 patients. At each event time, two controls are randomly selected in the riskset and matched to the case. Selected controls can become future cases or controls.
In NCC sampling, we embed a case-control study in a cohort such that for all individuals with the outcome of interest (cases), we randomly sample
Aggregating individual-level data takes place at the contributing nodes following NCC sampling in stage 1. Upon reducing the Cox partial likelihood in (2) to the NCC subcohort likelihood given in equation (3), the nodes set out to follow the outcome dependent aggregation scheme described by Saha-Chaudhuri and Weinberg 18 and Saha-Chaudhuri and Juwara. 31
For simplicity, we assume individual records of the time-to-event data at each node are sensitive and comprise of individual records of event times, outcome status, exposure
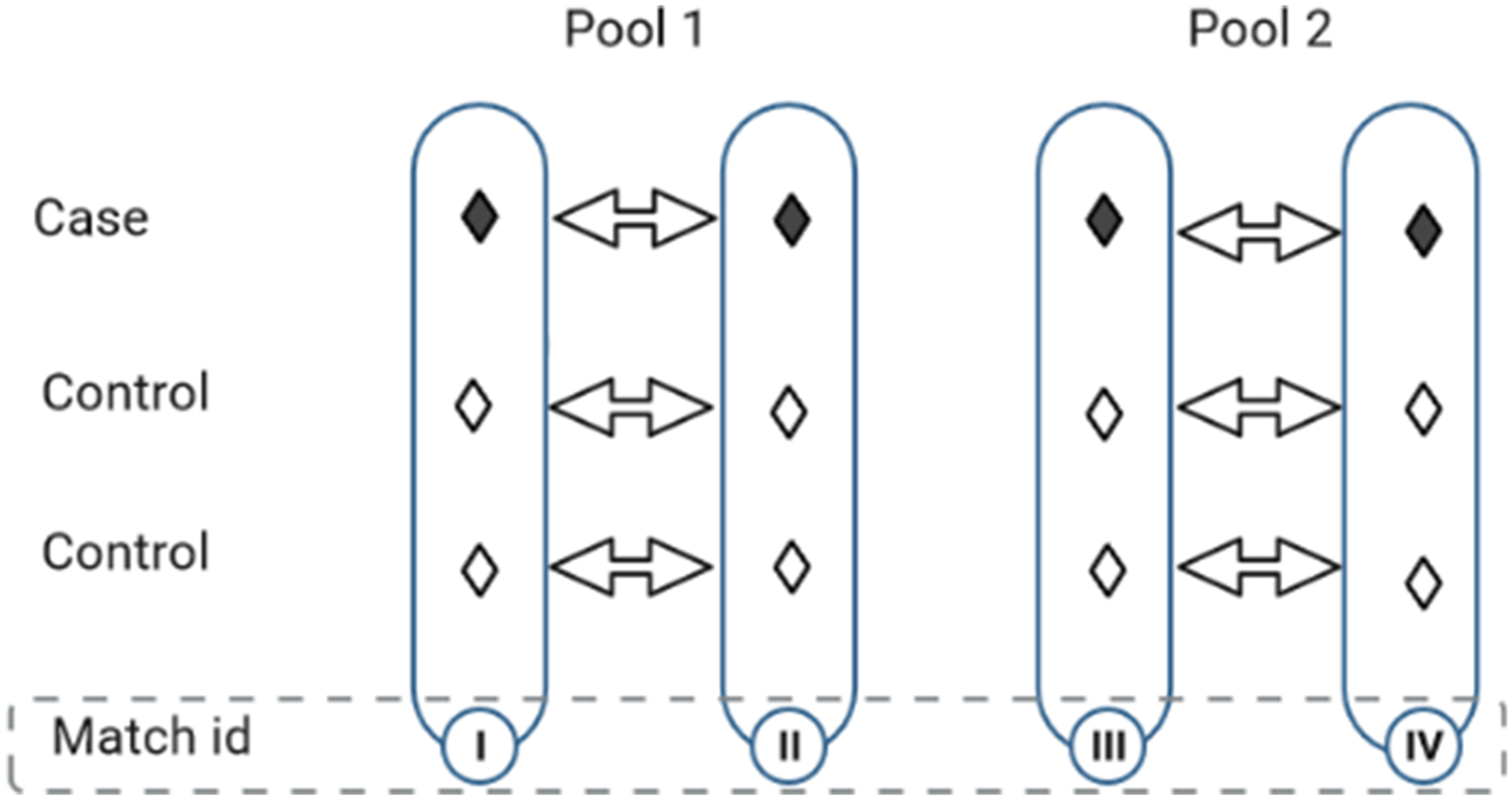
Pooling nested case-control matched sets of participants followed in Figure 2. Each pool is randomly formed from two matched sets by aggregating cases (with cases) and controls (with controls), independently.
A snippet of the node-level pooled subcohort that is anticipated for release at each node is presented in Table 1. In the example provided, the pooled exposure values of pool id 1 are obtained by adding the covariate values of cases in matched set I with cases in matched set II (i.e.
Pooled survival data created at each node under NCC subcohort sampling using the toy data of Figure 2. Information marked
The proposed pooling design can be extended to include pooled versions of additional confounders. Thus, if variables are available based on individual determinations, for example, by questionnaires, one could form set-based versions by summing the values across the individuals in the set. For categorical data, indicator variables of the categories could be used and pooled accordingly. Similarly, transformations of covariates (e.g. polynomial or log) and effect modifiers can be handled, but need more care.
Once the pooled NCC subcohorts have been created at the nodes, the next step is to share the aggregate values (as shown in Table 1) with the analytical center where they are combined into a single data file for analyses. We mainly focus on HR estimation under the postulated Cox model and estimation of overall survival curves.
HR estimation with pooled NCC likelihood
We now describe the likelihood contribution of individuals pooled at a single node. Consider a 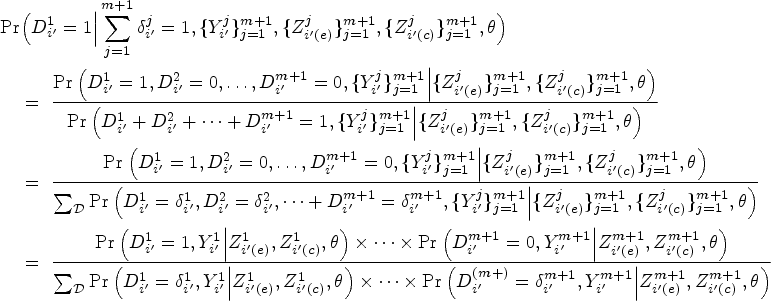
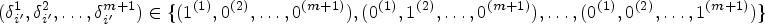
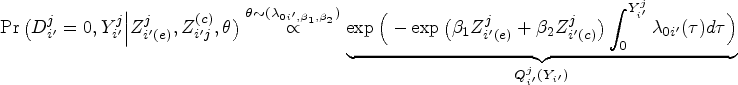
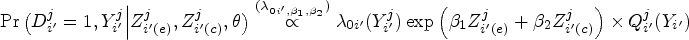
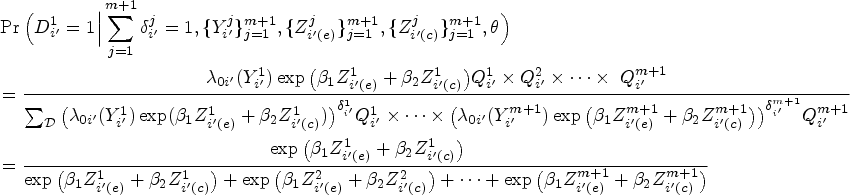
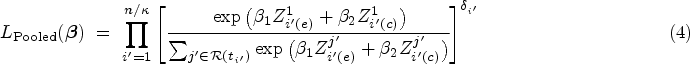
Utilizing the well-established equivalence between likelihood of the NCC subcohort and the likelihood of conditional logistic regression,39,36 inference on the MLEs could be carried out by using readily available packages for conditional logistic regression or stratified Cox regression. 34 The estimated parameters are interpreted as log HRs rather than the traditional log odds ratios derived from conditional logistic likelihood. Moreover, standard inference techniques applicable to conditional logistic likelihood can be employed to estimate the SEs of the pooled subcohort estimators. Of note, the units of analysis for the pooled NCC subcohort are the pools themselves, as opposed to individual measurements.
As mentioned earlier, the AC receives pooled NCC subcohorts (see Table 1) from each contributing node. This includes partial information on the observed event times of the pooled cases, the number of individuals making up each riskset at the node level if it is more than five individuals, the pool event status (1 if cases are pooled and 0 if controls are pooled), and their corresponding aggregate covariate values. While the log HRs associated with the covariates could be estimated using the pooled subcohorts without any need for the matched event times; to estimate the overall survival curves, the individual event times of subjects making up the pools would need to be recovered.
Suppose we want to reconstruct the overall survival curve 
In order to accurately estimate the survival curve based on pooled NCC subcohorts, a procedure for regenerating the observed event/censoring times of the cases and controls making up the pooled matched sets is needed. To perform this reconstruction, the AC has access to the pooled survival times of cases, the size of the riskset at each disclosed event time, and the event status of the pools. Of note, only the minimum event time of the pooled cases from a given pooled matched set is passed on to the AC. The survival time for the rest of the cases making up the pool and all of the controls would need to be re-generated to estimate the survival curves.
Let the pooled event times be ordered such that
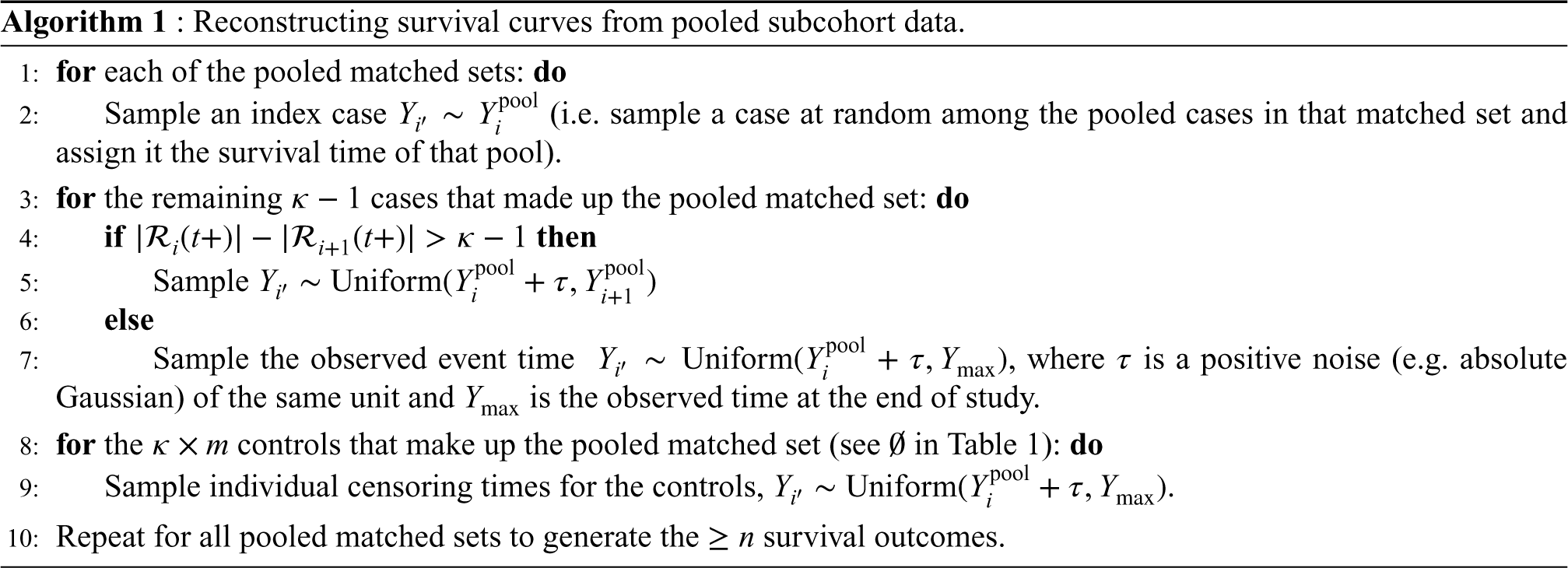

These choices of sampling ensure the ranking of event status (of individuals in the full cohort) are appropriately reconstructed. In other words, the number of individuals that make up each riskset is correctly retrieved. Even though the sampling may not generate precise estimates of observed event times per se, it preserves their ranking and consequently the count of individuals in each riskset. Hence, the estimator in equation (5) could be employed to accurately estimate the survival risk of individuals given a reasonable time window.
Of note, the proposed reconstruction generates excess controls resulting from repeated sampling of controls during NCC sampling. Therefore, to ensure a fair comparison of survival curves generated from different pooled subcohorts, we recommend randomly sampling out the excess controls. For example, the pooled toy example of Figure 3 constitutes 12 data points whereas the original full cohort only included 10 individuals. Hence the reconstruction (of individual survival outcomes) will generate event times for two more controls than what we started with which would need to be randomly sampled out to ensure a fair comparison to full cohort data. We demonstrate the performance of the sampling technique for estimating absolute survival risk after time
In recent years, various synthetic data generation methods (e.g. CART synthesis, DP-GAN, etc.) have gained popularity as advancements over traditional anonymization techniques like suppression, aggregation, and noise perturbation.3,4,26,27 These methods have emerged as effective approaches to facilitate the sharing of sensitive microdata while preserving data integrity. CART synthetic data generation, in particular, has attracted a lot of interest and is employed by various national agencies for data disclosure, for example, the Scottish Longitudinal Study Development and Support Unit (https://sls.lscs.ac.uk/guides-resources/synthetic-data/). We thus use it as a standard technique for generating synthetic data and compare its performance to the proposed method.
The CART algorithm partitions the predictor space into subsets, using unit partitions, in order to obtain subsets with consistent outcomes.40,41 Homogeneity at each split is tracked in CART through the utilization of an impurity function. The best split is determined by thoroughly exploring all variables and split values, ultimately selecting the split that minimizes impurity the most. Examples of such functions include the entropy criterion and the gini index. The optimal size of the CART model is determined by a complexity measure that carefully balances the accuracy of the model and the size of the resulting tree. Initially, the tree is grown to its maximum size and subsequently pruned to refine its structure. We present a tree structure representation of the binary splitting of CART in the Supplemental Appendix.
CART is frequently used for imputing missing categorical variables; however, in the current setting, we are primarily interested in its adoption for synthesizing confidential data. The algorithm is employed to replace individuals at high risk of re-identification with records generated by sampling from the posterior predictive distribution of the outcome fitted with the CART model. In practice, we use the
Performance metrics
The performance of the pooled subcohorts, under the postulated Cox PH regression framework, is assessed based on several metrics. Four main performance metrics are considered in assessing the usefulness of the released data. For the estimation of the log HR, we considered: (1) mean absolute error (bias) of the log HRs
The mean absolute bias estimate for each log HR Similar to standard likelihood theory, the variance The relative efficiency (Reff) of the log HR estimates compares the median model-based variance of the log HR to the empirical variance computed from log HR estimates of the re-sampled datasets.
44
This is expressed as The survival curves generated from the pooled subcohorts and synthetic data are each compared to the survival curve from the full cohort data. This comparison is based on their empirical mean survival probabilities and SEs obtained from multiple simulated datasets.
Applications
We used simulations and real data examples to evaluate the performance of the proposed method. The utility of pooled NCC data, under the proposed framework, was assessed using the performance metrics presented in previous sections. Our analysis considered varying combinations of pooled sizes, case-control matched sets, and censoring proportions. To do this, we compared PH regression models for the full cohort (i.e. individual data), pooled NCC subcohorts, and CART-generated synthetic data.
Simulation study
The simulations were based on 1000 repetitions, each with a sample size 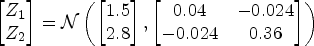
For each constructed data type, we estimated the log HRs associated with the covariates, the model-based SE, mean absolute bias (bias), and relative efficiency (Reff), and compared them to the gold standard estimates of the full cohort. Specifically, the metrics were computed for full cohort, pooled NCC subcohorts of pool sizes 2 and 4, and CART-generated synthetic datasets. The empirical coverage (Cov) was also assessed for each data type and compared to the nominal coverage of 0.95. Additionally, the robustness of the subcohorts constructed to varying parameter values were assessed by alternatively fixing one of
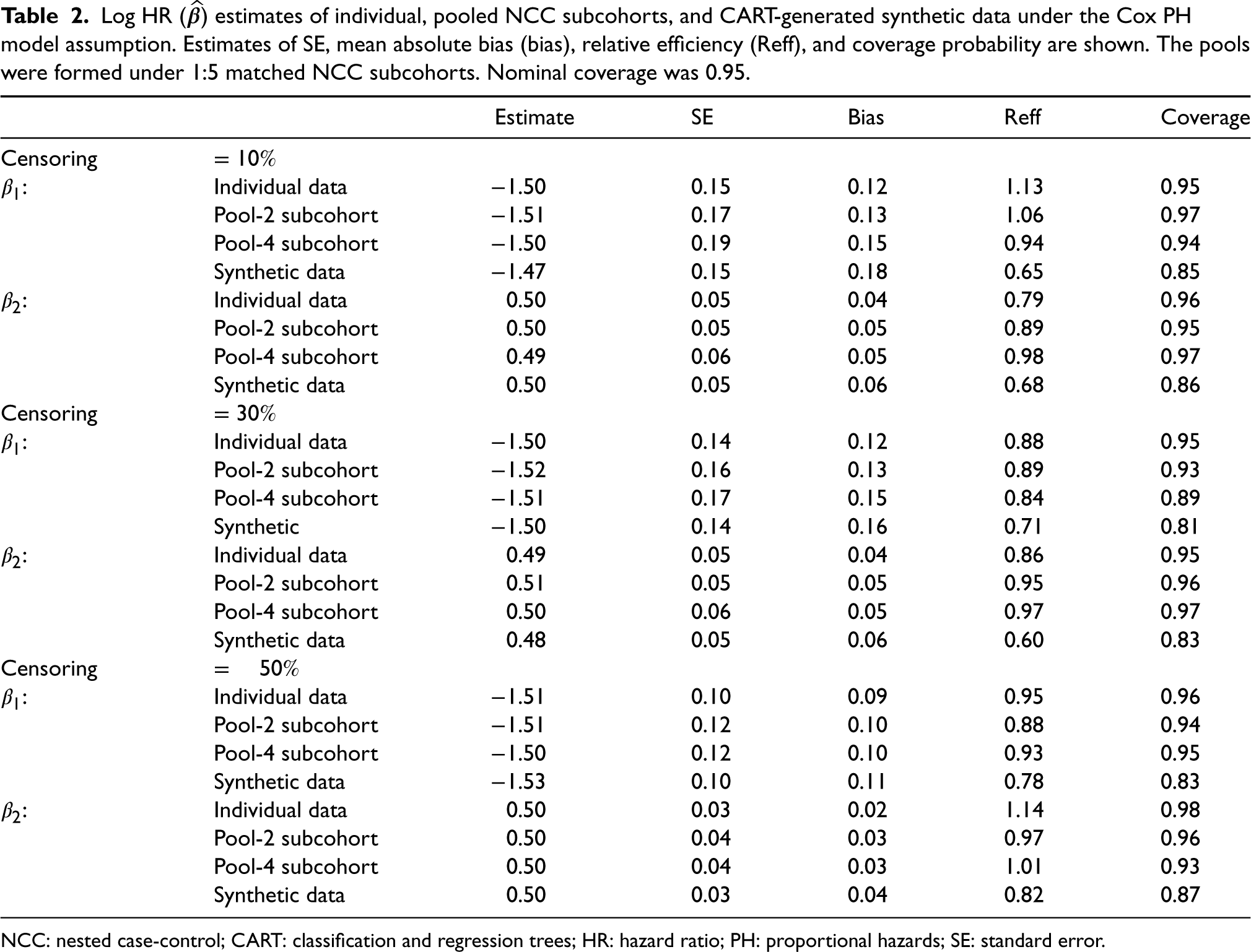
Log HR (
) estimates of individual, pooled NCC subcohorts, and CART-generated synthetic data under the Cox PH model assumption. Estimates of SE, mean absolute bias (bias), relative efficiency (Reff), and coverage probability are shown. The pools were formed under 1:5 matched NCC subcohorts. Nominal coverage was 0.95.
Log HR (

NCC: nested case-control; CART: classification and regression trees; HR: hazard ratio; PH: proportional hazards; SE: standard error.
Table 2 shows the estimates obtained for three different event prevalence rates (10%, 30%, and 50%) with each pooled NCC subcohort generated under five controls per case matching. The log HR estimates obtained from the pooled NCC subcohorts are accurate and comparable to the gold standard. However, the SE and bias estimates were generally worse for pooled subcohorts than individual data. We see an inflation of the SE estimates as more samples are pooled from individual to pool sizes 2 and 4, respectively. The confidence interval coverage of the pooled data are also consistent with the nominal
Power calculations for likelihood ratio tests on
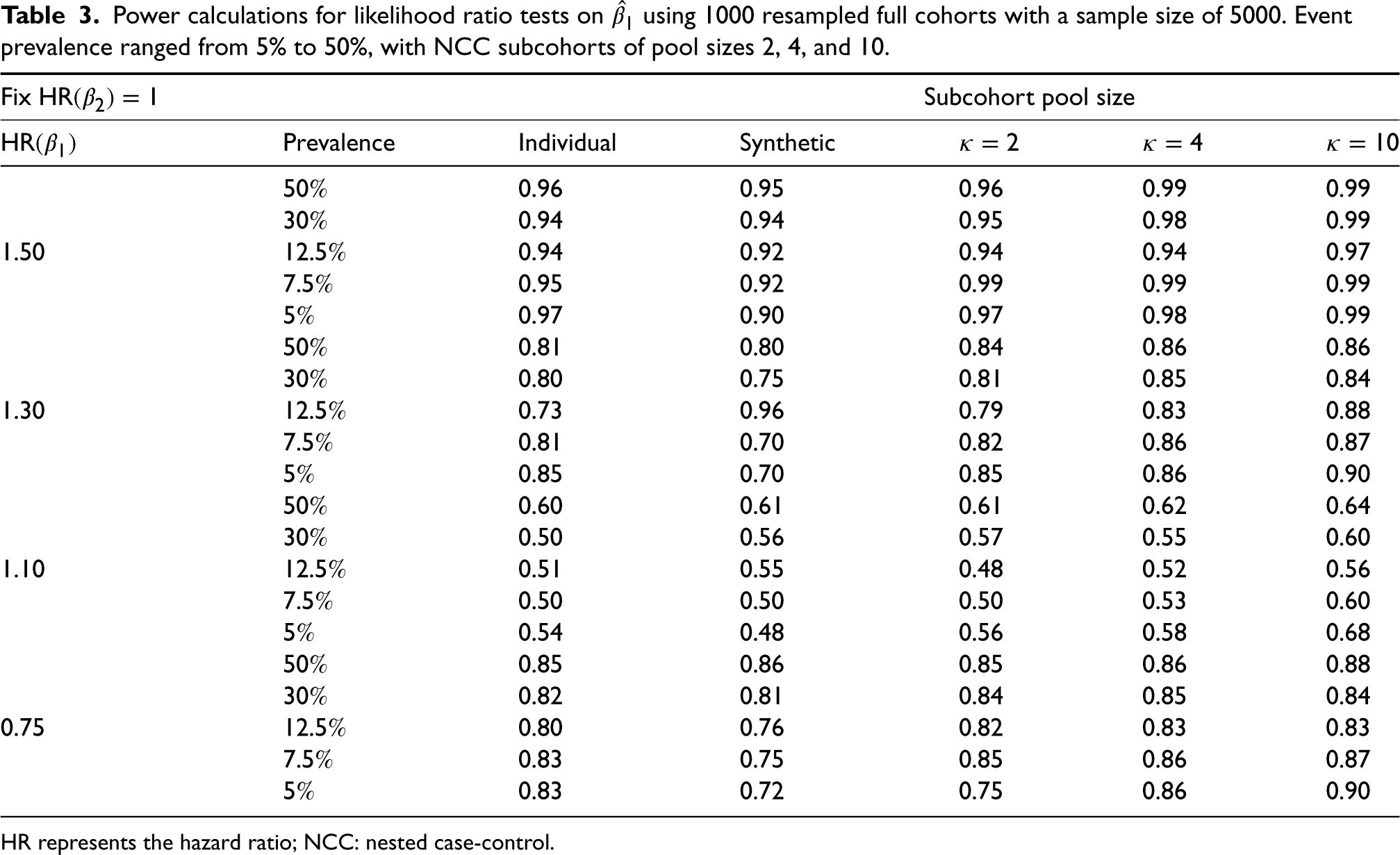
HR represents the hazard ratio; NCC: nested case-control.
Finally, pseudo-observed survival times were constructed for the full cohort data using the pooled NCC subcohorts created in Section 2. Mean survival probability plots were then generated for these reconstructions and compared to the exact curve derived from the original full cohort. Figure 4 shows a representative plot of mean survival probability by follow-up time. Overall, our results indicate that the pooled NCC subcohorts are comparable to the full cohort (considered the gold standard) and synthetic data when considering the mean survival probabilities across 1000 simulated samples. Summaries of the median survival time and the restricted mean survival time (a numeric expression of the area under the KM survival curve) are also presented in Table 4. In summary, the proposed reconstructions of pseudo-event times provide a good approximation for the number of individuals comprising the risksets during follow-up.
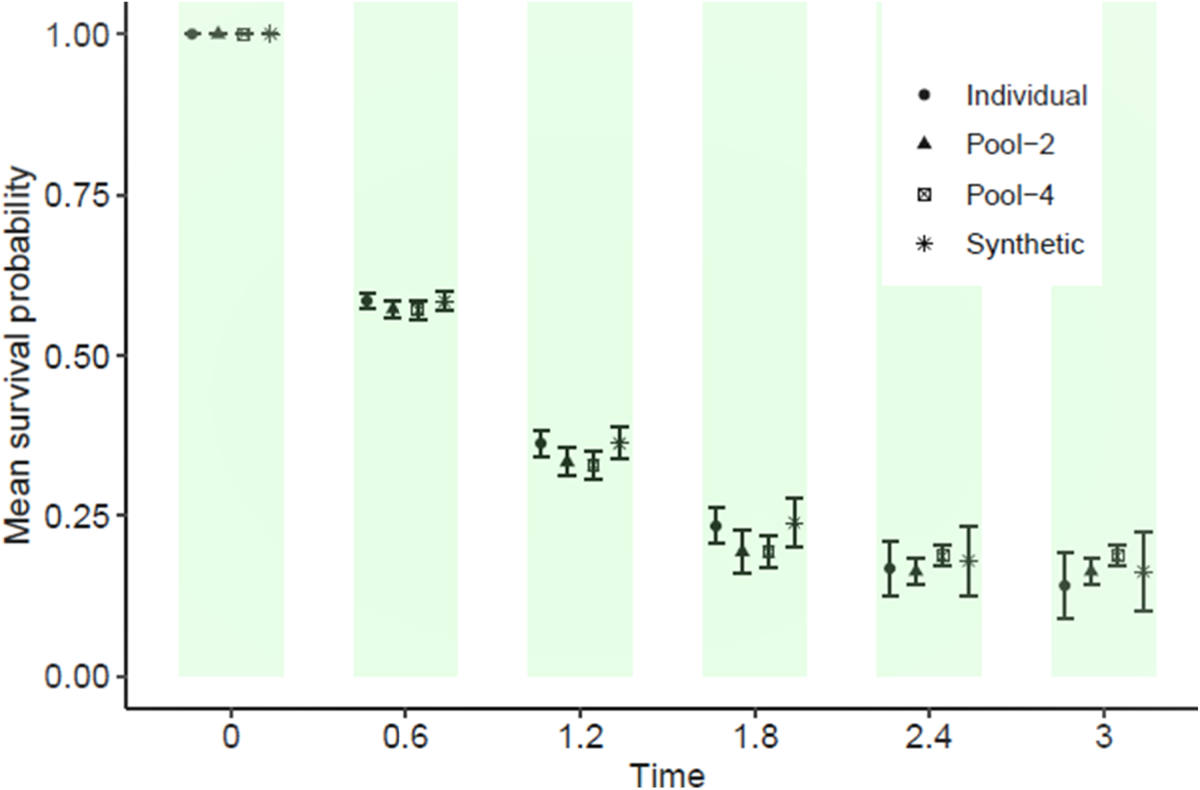
Average estimates of survival probabilities comparing individual, reconstructed pooled subcohorts, and classification and regression trees (CART)-generated synthetic data from 1000 simulated datasets each of size 5000. Mean survival probabilities and error bars are provided.
KM curve summaries of median survival time (
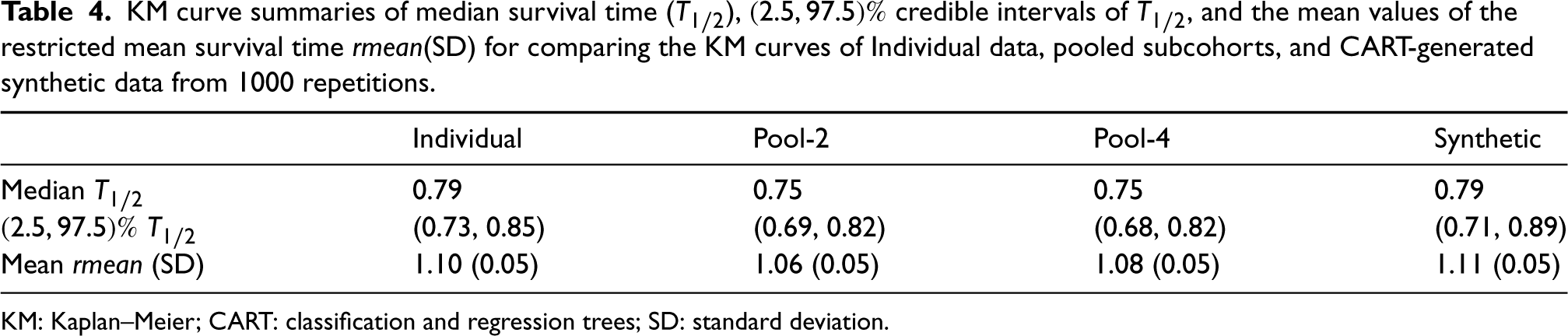
KM: Kaplan–Meier; CART: classification and regression trees; SD: standard deviation.
Baseline characteristics of the lung cancer data (full cohort), pooled NCC subcohorts, and synthetic data.
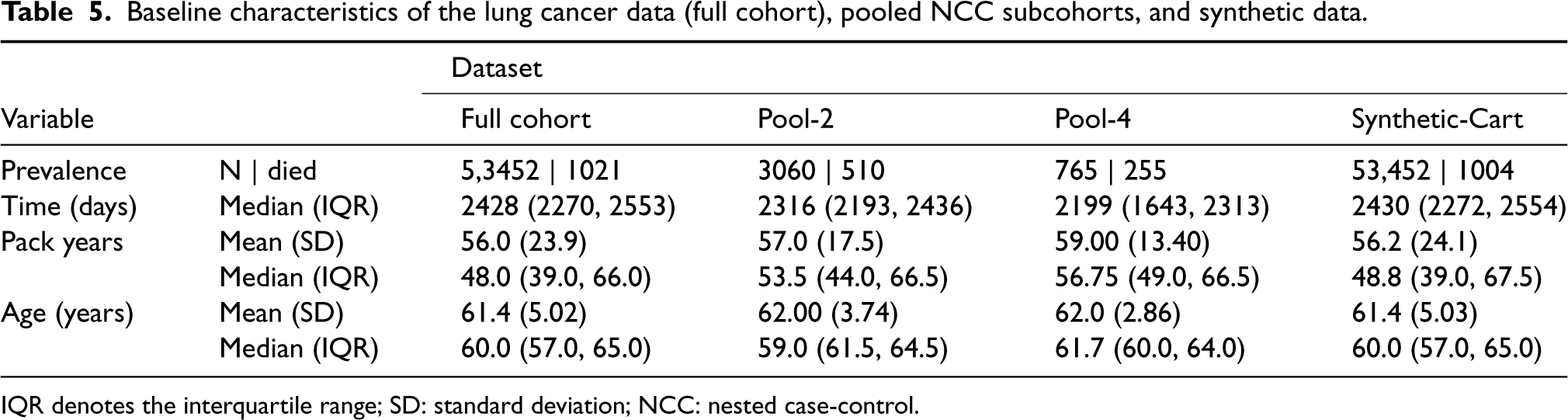
IQR denotes the interquartile range; SD: standard deviation; NCC: nested case-control.
The National Lung Screening Trial (NLST) lung cancer data
45
was used to assess the utility of the proposed method. NLST is a randomized controlled trial designed to determine whether screening for lung cancer with low-dose helical computed tomography reduces mortality in high-risk individuals relative to screening with chest radiography. The study enrolled
Cox PH regression was performed to assess the utility of the datasets in characterizing the association between the outcome and the exposure and confounder. The results presented in Table 6 compare the log HR estimates from six data sources: full cohort data of individual level observations (gold standard), Pool-2 and four subcohorts (each of 1:2 and 1:5 matched sets) that combines node-derived pooled data, and synthetic data generated based on the CART data synthesis.
HR and 95% CI estimates for the associations of pack years and age with lung cancer deaths across the full cohort (individual), pooled NCC subcohorts (1:2 and 1:5 matched sets), and CART synthetic datasets.
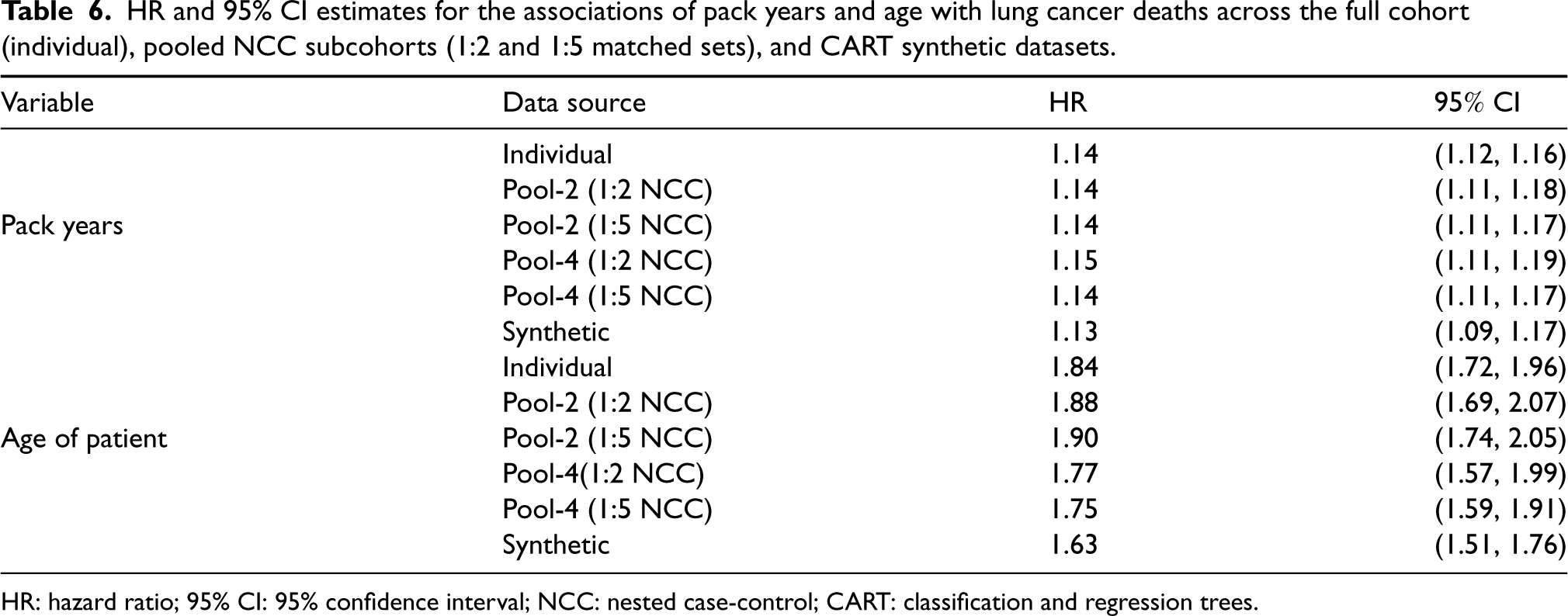
HR and 95% CI estimates for the associations of pack years and age with lung cancer deaths across the full cohort (individual), pooled NCC subcohorts (1:2 and 1:5 matched sets), and CART synthetic datasets.

HR: hazard ratio; 95% CI: 95% confidence interval; NCC: nested case-control; CART: classification and regression trees.
Similar to the simulated data results, the HR estimates obtained for the full cohort NLST data are comparable to estimates obtained from the pooled NCC subcohorts and synthetic data. The SE estimates increase as the pool size is increased from 2 to 4 records per pool. As expected with the NCC sampling design, the precision of the parameter estimates also improve as more samples were matched on (e.g. 1:2 versus 1:5 matched case-control sets). For example, the 95% confidence interval of the HR associated with age tightens from
In this article, we propose a privacy-preserving analysis technique for time-to-event data within the NCC sampling framework. The technique leverages the pooling scheme introduced by Weinberg and others16,18,31 to randomly aggregate individual records across matched sets within the same NCC subcohorts, stratified by the outcome status. Only the aggregated covariate levels of the NCC subcohorts are shared with the analytical site. We employ several data utility metrics to assess the usefulness of the shared aggregate records for Cox PH regression and survival curve estimation under data privacy restrictions. These metrics are applied to the following datasets: full cohort (individual) data, pooled NCC subcohorts with pool sizes of 2 and 4, and CART-generated synthetic data.
The results of our simulations and real data example demonstrate that the HR estimates and SEs obtained through pooled data analysis are comparable to those obtained from individual full cohort records. The estimators derived from the pooled NCC framework are MLEs of the HRs, providing consistent estimates of the individual-level HRs and asymptotically normal results. We also find that matching on more controls during NCC sampling, prior to pooling, leads to a substantial improvement in the efficiency of effect estimates (HRs), as previously reported by Langholz and Goldstein 30 and Kim. 46 Bias and relative efficiency estimates of pooled NCC subcohorts are similar to those of individual full cohort Cox regression. The empirical coverage, computed under pooled NCC subcohorts, closely approximates the 95% nominal coverage in most scenarios. Furthermore, survival times reconstructed based on the proposed sampling method are adequate for making meaningful clinical predictions of survival risk using the KM estimator.
The proposed method offers several advantages over competing techniques. For example, matching cases to controls before aggregating the matched sets based on event status is computationally inexpensive and readily accessible via standard statistical software. The communication cost for transferring pooled data across the network is also fairly low, as pooling reduces the overall sample size. The cost efficiency improves as the pool size increases. Moreover, the technique is intuitively simple and accessible to researchers in other fields. In comparison, methods for synthetic data generation are often quite expensive, especially for a large quantity of attributes (e.g. three or more attributes), and might require special and difficult-to-program algorithms. Another noteworthy advantage is that NCC sampling minimizes both selection and recall biases, reducing the risk of re-identification before matched set aggregation. Inclusion of binary effect modifiers and additional confounders is also easily attainable without major modifications to the underlying likelihood. 18 For example, outcome pooling followed by stratifying by an effect modifier could improve efficiency of estimation. Sometimes, such further stratification is recommended to ensure a fair comparison of measurements. For instance, when the exposures of interest are biomarker measurements from frozen biological material, the cases and controls should be matched based on factors such as storage time or condition. However, the current study recommends random pooling of covariates based on only the outcome status, as the privacy gain in inferential or re-identification risk outweighs the efficiency gain when the patients are further stratified.
Inferential and re-identification disclosure risk are fundamental concerns in assessing privacy. The former is still an active area of research and as such assessing the associated disclosure risk of individuals in the released microdata remains a challenging task.47–49 In principle, an intruder (e.g. the analyst or data steward) could infer or learn information about a participant even if exact records were not disclosed. Inferential disclosure risk has been reported to be a major concern in traditional techniques such as suppression of quasi-identifiers or noise perturbation, partly because disclosed records often retain local data structures potentially aiding inference. Pooled NCC subcohorts, in comparison, smooth out underlying local perturbations via summation rendering the risk of inferential disclosure extremely small. 50
While a comparison of the risk of patient re-identification disclosure of individual records is not covered in the current manuscript, we believe the risk is low in NCC subcohort data for several reasons: 51 (1) under the generous assumption that all attributes in the database are quasi-identifiers, pooled NCC subcohort data is non-overlapping with individual records of external data sources because aggregates do not retain underlying structures of individual elements, (2) any unique combination of pooled covariates records cannot be an identifying class of the individual components due to random pooling, and (3) any pattern of population overlap between the sensitive individual record and an external source is destroyed via pooling.50,52 Furthermore, the distribution of any single aggregate attribute is not an accurate indicator of re-identification disclosure risk, or in the case of a categorical variable the results may be falsely reassuring. In the instances the combination of aggregate attributes leads to a unique combination, individuals remain protected as the aggregates are non-overlapping with external databases of individual records.
Our method has some limitations. First, data aggregation decreases statistical power, potentially reducing the utility of pooled NCC subcohorts compared to individual full cohort data. Consequently, meaningful data patterns could be distorted or completely destroyed via aggregation. Second, selecting a suitable pool size requires the analyst to strike a balance between privacy and utility. While a pool size of 4 has been shown to offer good privacy protection and data utility, even with small to moderate samples,36,28 it may be helpful for individual data nodes to assess various pool sizes before sharing the pooled data. For instance, each node could perform a pool-level assessment and share data when the estimates from the largest pool size are comparable to the site’s individual-level estimate. Third, another limitation of publishing pooled NCC subcohort data is the lack of a provable privacy guarantee for the randomized summation mechanism used to generate pools. In recent years, synthetic data generation methods with differential privacy guarantees 53 have gain popularity. Various classical and modern (deep generative) approaches have since been developed to generate differentially private synthetic datasets. In the future, the development of a pooling mechanism with a mathematically provable privacy guarantee warrants exploration. Lastly, the current method cannot be implemented in heterogeneous data settings where the data nodes harbor different distributions 54 or with vertically partitioned databases where the attributes are split among the data nodes. In the former case, our approach assumes that the datasets stored at the nodes follow the same distributions and that site-specific subcohorts can be combined without taking between-node differences into account. 55 A potential modification to account for these differences might be to introduce a weighted likelihood function for evaluating the pooled subcohorts. The latter case remains an active area of research.
Despite these limitations, the trade-off between disclosure risk and utility of pooled disclosed data makes the technique appealing. Pooling individual records at contributing data sites result in minimal utility loss whilst preserving the confidentiality of patients. The additional efficiency gain when more controls are matched-on makes the technique particularly desirable, especially in rare disease settings or in emerging infectious disease studies where the outcome prevalence is still low. In conclusion, the proposed pooling technique of survival time data under NCC subcohort sampling preserves patient privacy while ensuring consistent estimation of effects, suitable standard errors, and accurate survival curves comparable to individual full cohort data.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802231215804 - Supplemental material for Privacy-preserving analysis of time-to-event data under nested case-control sampling
Supplemental material, sj-pdf-1-smm-10.1177_09622802231215804 for Privacy-preserving analysis of time-to-event data under nested case-control sampling by Lamin Juwara, Yi Archer Yang, Ana M Velly and Paramita Saha-Chaudhuri in Statistical Methods in Medical Research
Footnotes
Acknowledgements
The authors are grateful to the editor, associate editor, and two referees for their insightful comments and suggestions, which significantly improved the quality of the paper. We thank the funding agencies Mitacs Accelerate Fellowship, Natural Sciences and Engineering Research Council of Canada (RGPIN-2017-06100), and Fonds de la recherche en santé du Québec (Salary Award for PS-C) for supporting our research.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Data availability
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
Supplemental material
Supplemental materials for this article are available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.