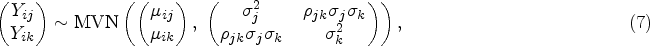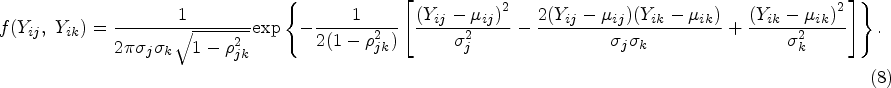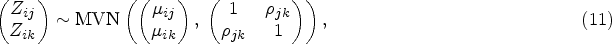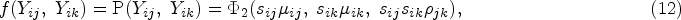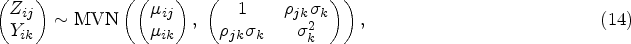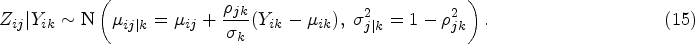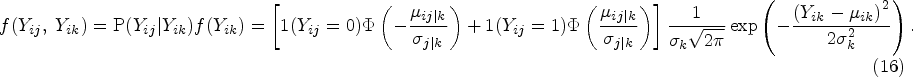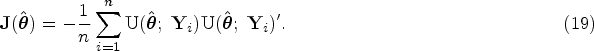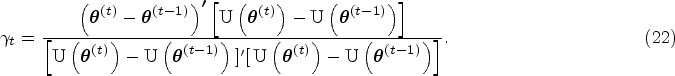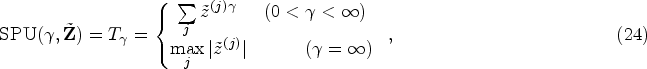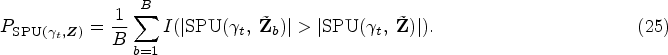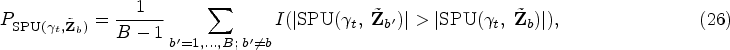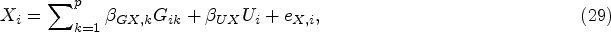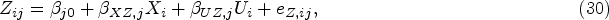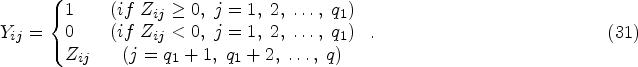Abstract
In clinical research, it is important to study whether certain clinical factors or exposures have causal effects on clinical and patient-reported outcomes such as toxicities, quality of life, and self-reported symptoms, which can help improve patient care. Usually, such outcomes are recorded as multiple variables with different distributions. Mendelian randomization (MR) is a commonly used technique for causal inference with the help of genetic instrumental variables to deal with observed and unobserved confounders. Nevertheless, the current methodology of MR for multiple outcomes only focuses on one outcome at a time, meaning that it does not consider the correlation structure of multiple outcomes, which may lead to a loss of statistical power. In situations with multiple outcomes of interest, especially when there are mixed correlated outcomes with different distributions, it is much more desirable to jointly analyze them with a multivariate approach. Some multivariate methods have been proposed to model mixed outcomes; however, they do not incorporate instrumental variables and cannot handle unmeasured confounders. To overcome the above challenges, we propose a two-stage multivariate Mendelian randomization method (MRMO) that can perform multivariate analysis of mixed outcomes using genetic instrumental variables. We demonstrate that our proposed MRMO algorithm can gain power over the existing univariate MR method through simulation studies and a clinical application on a randomized Phase III clinical trial study on colorectal cancer patients.
Keywords
Introduction
With the increasing collection of data on various clinical and patient-reported outcomes (e.g. toxicities and quality of life measures) in clinical and observational studies, there is growing interest in detecting the causal relationship between an exposure variable (X) and multiple outcomes of interest (Y).1–5 For example, some baseline clinical characteristics may affect a patient's risk of experiencing certain toxicities after receiving treatment. Finding and studying such causal effects may help clinicians predict the risk of adverse events and make better treatment decisions.6,7 This can be illustrated through the analysis of data from the CO.20 trial conducted by the Canadian Cancer Trials Group, which was a phase III randomized, placebo-controlled study aiming to examine the effect of the addition of brivanib (BRI) to cetuximab on colorectal cancer patients.8–10 The original analysis of the study showed that compared to cetuximab plus placebo treatment, cetuximab plus BRI was associated with increased toxicities and did not significantly improve the overall survival of patients. 8 It is of interest to investigate whether certain baseline variables have any effect on the selected toxicities that a patient may experience when undergoing the cetuximab plus placebo treatment, with the mixed-type measurements for these toxicities.
There are two major challenges, however, in these analyses. One challenge is that, although patients may be randomized by treatment, they may not be balanced by the baseline variable X of interest, and therefore, directly modelling the relationship between X and Y without considering the confounders, whether measured or unmeasured, may lead to misleading results. 11 The other challenge is that sometimes the multiple outcomes of interest may be correlated and mixed with different types of distributions (e.g. binary and continuous). 12 Studying each outcome separately is a commonly used straightforward approach but does not use the correlation information between outcomes, while modelling mixed outcomes together may utilize more information but require more complex models.13,14
From a clinical standpoint, there exists a major analytical gap in our ability to simultaneously study the influence of a causal effect on multiple outcomes, particularly in the context of toxicities. In the areas of rheumatology, immunology/transplantation, and oncology, it is common for patients and physicians to accept multiple moderate to severe toxicities in their treatments, as these treatments are considered life-saving. Most of the drugs used in these settings exhibit widely variable toxicities from person to person. Understanding the etiological factors that lead to significant toxicities in some patients but not others is fundamental to precision medicine. Innovative methods for analyzing individual etiological factors on multi-dimensional toxicities, particularly those that are graded differently (e.g. ordinal symptoms vs continuous laboratory-based toxicities) are greatly needed.
Facing the challenge of confounders, people have been applying the Mendelian randomization (MR) approach, which uses genetic variants as instrumental variables. 15 Under certain instrumental variable assumptions, MR methods can efficiently estimate and make inferences on the causal effect of the exposure on a single outcome. Different MR methods have been proposed to address a variety of problems, such as how to handle invalid instruments and how to incorporate multiple exposures.16–21 Nevertheless, we are not aware of any existing MR techniques that allow researchers to jointly model mixed outcome variables. When multiple outcomes need to be examined, people usually conduct univariate analysis, which analyzes each outcome separately. This may not be ideal for testing the overall hypothesis, whether the exposure has any effects on any outcomes, since the correlation structure between different outcomes is not considered, and Bonferroni correction is usually required, which is known to be conservative. 22
To address the above issues, it is desirable to develop a multivariate MR approach that can handle mixed outcomes. Different methods have been proposed to directly model mixed outcomes jointly,12,23–26 while it is computationally challenging to implement most of them into the MR framework, especially when the full likelihood is considered. Bai et al. 27 proposed using a composite-likelihood method with the Newton–Rapson algorithm to alleviate the computational issue. 28 All these existing methods only model the relationship between X and Y without using instrumental variables, which means they cannot handle unmeasured confounders.
We have developed a two-stage multivariate MR method for mixed outcomes, denoted by MRMO that combines the two-stage MR approach with the composite-likelihood-based mixed response model to assess the causal effects of the exposure on different outcomes. For instance, MRMO can be used to test whether a baseline variable (e.g. baseline magnesium) has a causal effect on potential disease outcomes such as rash and nausea, as its MR component can handle the effects of potential confounders such as age, gender, drinking status, etc. We propose using the gradient descent algorithm when jointly modeling mixed outcomes, which has been shown to have a number of advantages over the commonly used Newton–Rapson algorithm.29–31 In terms of testing the overall hypothesis, it is natural to consider the Wald test for multivariate models. Besides that, we propose conducting the aSPU (adaptive sum of powered score) test for multiple outcomes,32,33 which can combine a class of powered sum tests to gain power in certain scenarios. Through extensive simulations and an application on a randomized Phase III clinical trial study (CO.20) on colorectal cancer patients,8–10 we demonstrate that compared to the commonly used univariate MR analysis, our proposed multivariate approach is able to improve the power of testing the overall hypothesis while controlling type I errors, which can help clinicians find potential associations and generate new hypotheses.
This manuscript is structured as follows. In section 2, we introduce the concept of univariate MR in the presence of multiple outcomes and then present our multivariate MR approach to model mixed outcomes jointly. We also propose different algorithms and tests for our multivariate analysis. In section 3, we show the settings and results of our simulation studies, as well as an application to the CO.20 study data to compare the performance of different methods. In section 4, we present discussions summarizing the advantages and limitations of our new approach and propose some potential future directions.
Methods
Univariate model for Mendelian randomization with multiple outcomes
In this section, we illustrate the basic framework of traditional MR analysis, depicted by Figure 1(A). Suppose we are interested in studying the causal relationship between an exposure (X) and a single outcome (Y1). Directly modelling Y1 versus X may give us very problematic results, since there may be confounders (U) that are unmeasured or not included in the model. MR analysis incorporates genetic variants G, typically SNPs (single-nucleotide polymorphisms), to serve as instrumental variables (IVs) for causal inference. Using IVs can help us avoid the confounding problem if certain IV assumptions are met (i.e. IVs should be associated with X; IVs should not affect or be affected by U; and IVs should not affect Y1 through any pathways other than X). For example, in a colorectal cancer study,8–10 clinicians may be interested in examining whether an exposure variable like body mass index (BMI) has a causal effect on a disease outcome like quality of life. With the causal estimand defined as the average change in the quality of life for a unit change in BMI, which can be denoted as
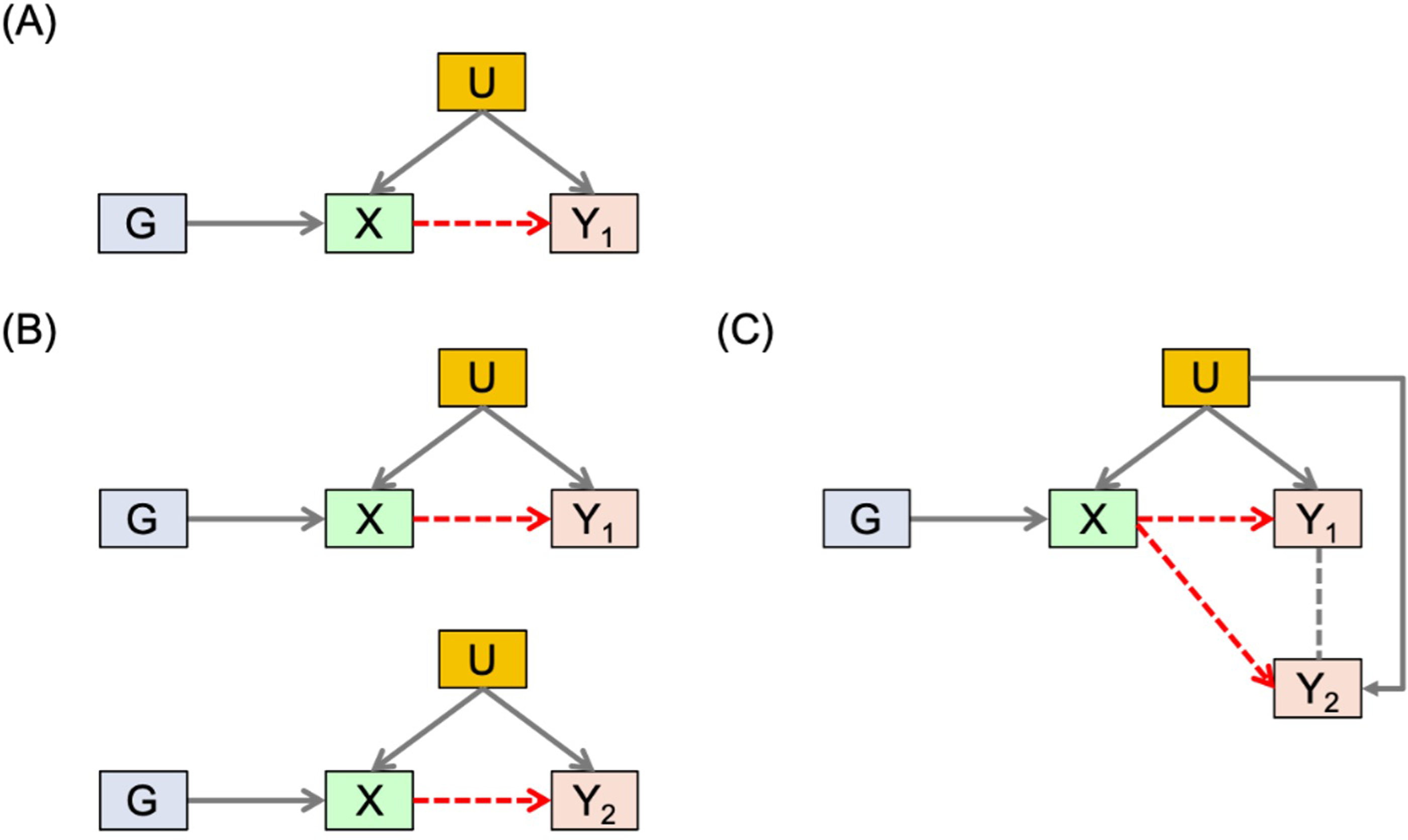
Basic framework. (A) MR analysis with a single outcome. (B) MR with multiple outcomes and univariate analysis. (C) MR with multiple outcomes and multivariate analysis.
When there are multiple outcomes of interest (e.g. different types of toxicities such as rash, nausea, and vomiting in cancer trials), people normally conduct several univariate analyses separately, which applies MR to one outcome at a time, as depicted in Figure 1(B) for two outcomes Y1 and Y2. However, analyzing each outcome separately means ignoring the correlation information from the multiple outcomes, which may lead to loss of power, especially when we want to test the overall null hypothesis: the exposure does not affect any of the outcomes. Meanwhile, multivariate analysis, illustrated in Figure 1(C), models and analyzes multiple outcomes in a single model simultaneously. It may give us a better framework for testing the null hypothesis while accounting for the correlation structure of the different outcomes.
Before presenting our multivariate approach, we would like to give a brief overview of the two-stage univariate MR analyses using individual level data in a two-sample scenario, where we have two different datasets with different subjects for the exposure and outcomes. Suppose we have an exposure variable X, p genetic IVs and q outcomes, and the first sample set contains
The first-stage regression model for the univariate MR is
In this section, we propose our two-stage multivariate MR method for mixed outcomes, which is expected to be more efficient than the univariate method when testing the overall null hypothesis (the exposure does not affect any of the outcomes). The first-stage model is the same as what we have described in the univariate two-stage MR method, of which the main purpose is to model the association between the genetic information and the exposure for the second sample. In the second stage, instead of modeling each outcome separately, we propose to model different outcomes jointly using a multivariate mixed response model.26,27 The relationship between each outcome and the fitted exposure can be modelled using a generalized linear model
To model all the outcomes simultaneously while incorporating their correlation structure, applying a conventional likelihood-based approach is possible but computationally very challenging. Using the pairwise composite likelihood approach as described in
27
may be a better option. For each pair of outcomes
If a pair of outcomes
If one outcome j is binary, and another outcome k is continuous, we assume that
To estimate Choose the initial values of the parameters, denoted by Apply the gradient descent method to update the parameter estimates. For iteration Calculate the new score function using the updated parameters. Repeat the previous two steps until convergence. By default, the algorithm is stopped when
Note that the choice of step size
In this section, we discuss more on the different choices for testing the overall hypothesis
For the multivariate analysis, once we have
Meanwhile, we propose to apply the aSPU test as an alternative method, which can combine a class of tests and may gain power when testing
In this section, we present the general procedure for applying our new approach to study the causal relationship between an exposure and multiple outcomes, as shown in Figure 2. We would like to point out that the second step is crucial for MR analysis since including SNPs that violate the IV assumptions may lead to problematic results. Nevertheless, for this manuscript, we emphasize the fifth step, which mainly compares the multivariate approach with the univariate approach for modelling the relationship between the exposure and outcomes. We would also like to note that when conducting two-sample MR, it is more common to apply various MR techniques that use summary statistics (e.g. MR-Egger). We choose to focus on the two-stage MR method that is more comparable to our new multivariate approach, which requires individual-level data. Another reason why we would like to focus on the two-stage methods is that two-stage regression allows correlated instrumental variables in the first-stage model, whereas for summary-based MR methods, the chosen instrumental variables are required to be independent. In situations where there are not many available instrumental variables that are independent, it may be better to choose a looser correlation cutoff and apply the two-stage approach. We will discuss more on the strengths and drawbacks of our approach in the Discussion section. Besides, it is worth mentioning that the two-stage methods can also be applied to the one-sample situation, where we have a single dataset containing information on G, X and Y. However, we would like to focus on the two-sample scenario, since the literature has shown that only using one sample is less preferred, as it is more likely to result in biased estimates and inflated type I errors.15,35 Some simulation results and discussion for the one-sample scenario are provided in Appendix A of the Supplementary materials.
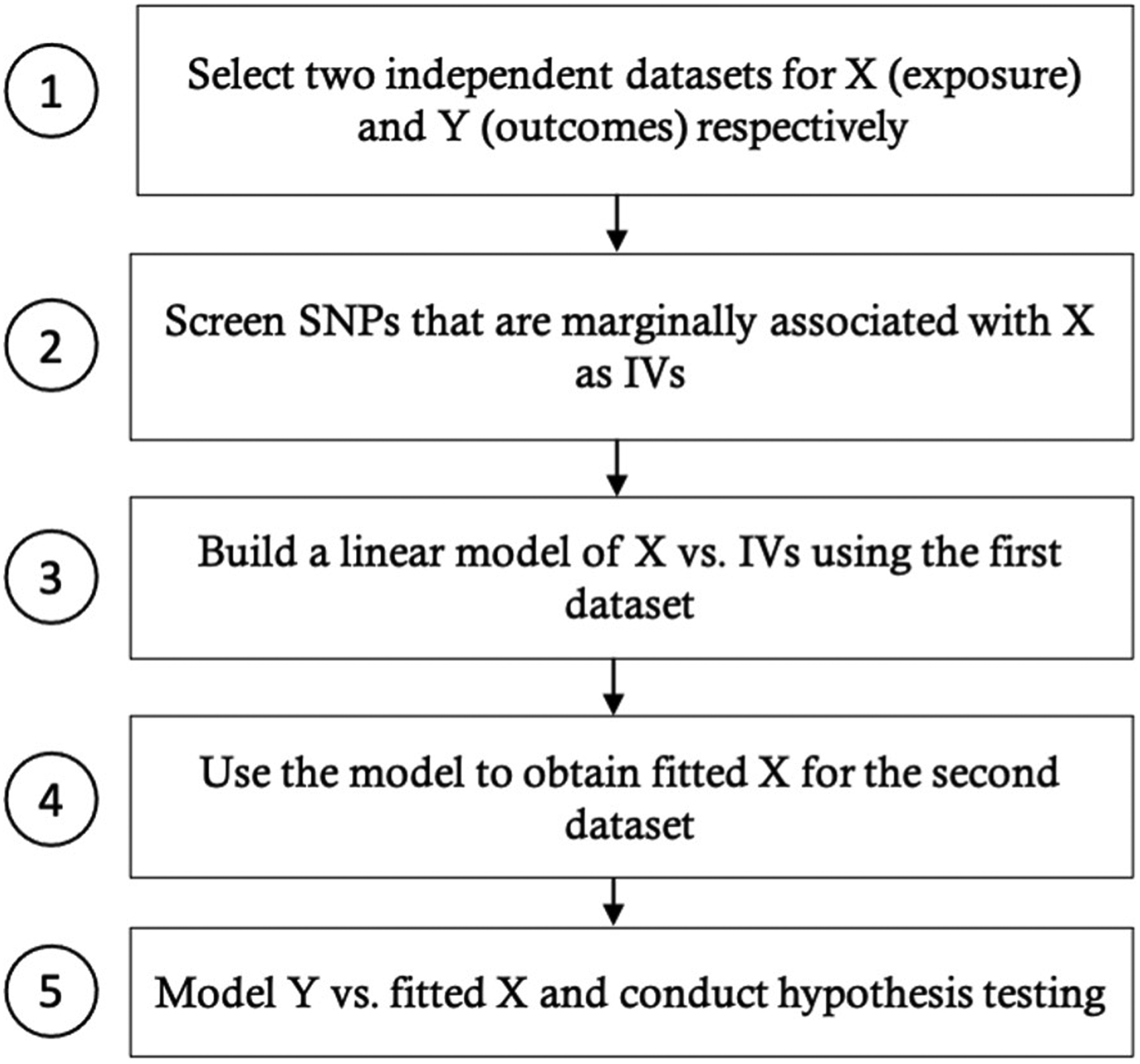
General procedure for applying MR to multiple outcomes using two samples with individual-level data.
Simulation studies
We conduct simulation studies to compare the performance of our new approach and the univariate two-stage MR method. Among the common variants that show some marginal association (p < 1e-4) with baseline magnesium (MG) based on the CO.20 genotype data,8–10 we randomly select 19 of them with weak or no correlations (pairwise correlations between −0.1 and 0.1). All of these SNPs have minor allele frequencies (MAFs) greater than 0.05, and 16 of them have MAFs greater than 0.1. In the simulations, we assume that these SNPs are the true association SNPs and simulate the genotypes for two independent samples, each with a sample size of 559, using multivariate binomial distributions and maintaining the MAF of each SNP. Since the correlations among the selected SNPs are very weak, for convenience, we simulate the SNPs as independent unless otherwise stated. Our experience shows that using weakly correlated SNPs or independent SNPs does not really affect the results. Suppose there are
We simulate U, X and Y using the following models:
Following Slob and Burgess,
46
we generate
At first, we consider a mixture of two binary and two continuous outcomes. We set
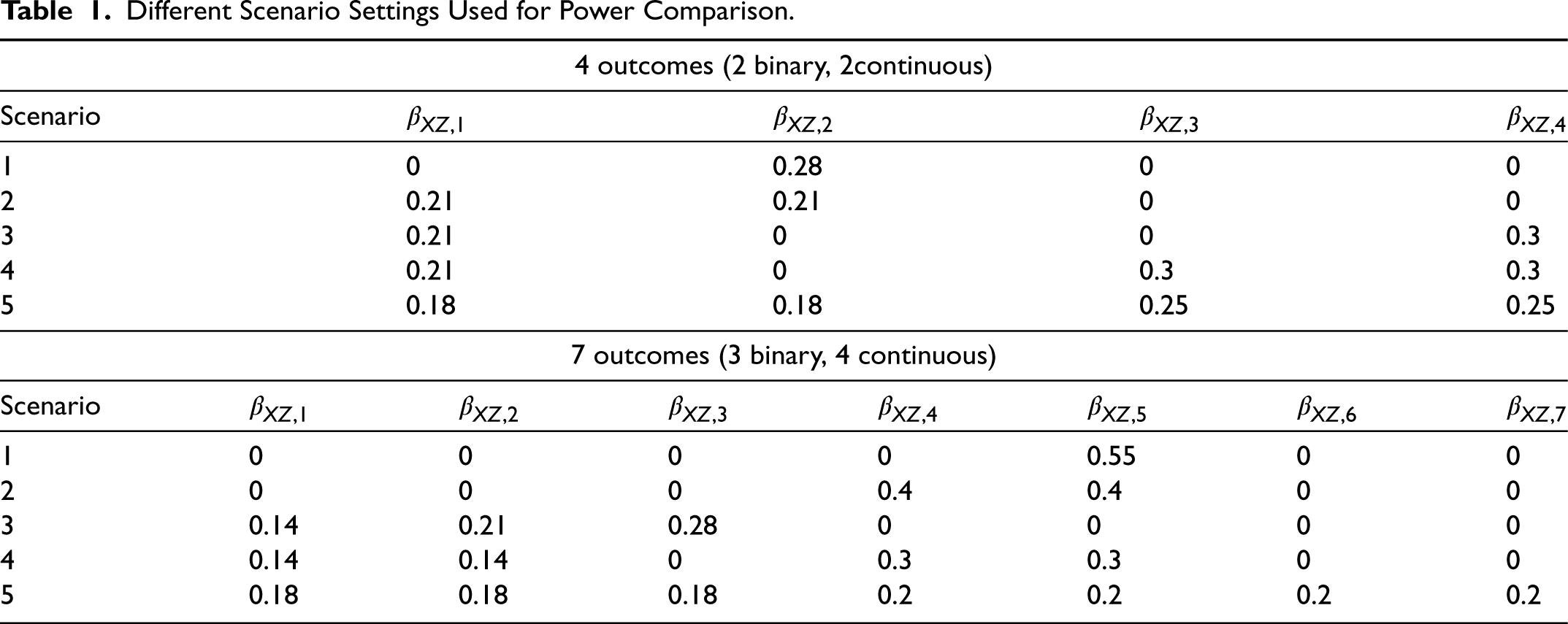
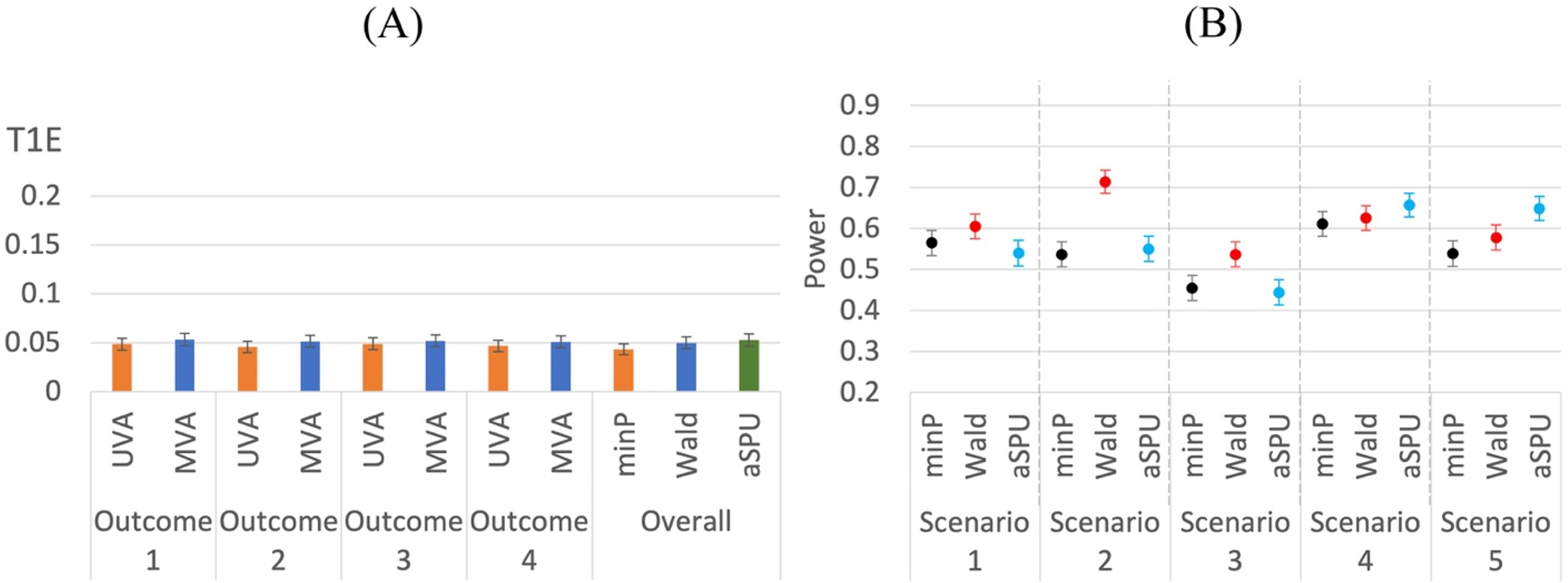
As Figure 3 shows, both univariate and multivariate analyses are able to control type I errors when testing the overall hypothesis or each single outcome. In terms of power, we examine various scenarios, for which the different parameter settings are included in Table 1. Though we mainly focus on our proposed gradient descent algorithm for this manuscript, our experience shows that the results of gradient descent and Newton's method are very consistent under most circumstances, as shown in Figure 4, for example.
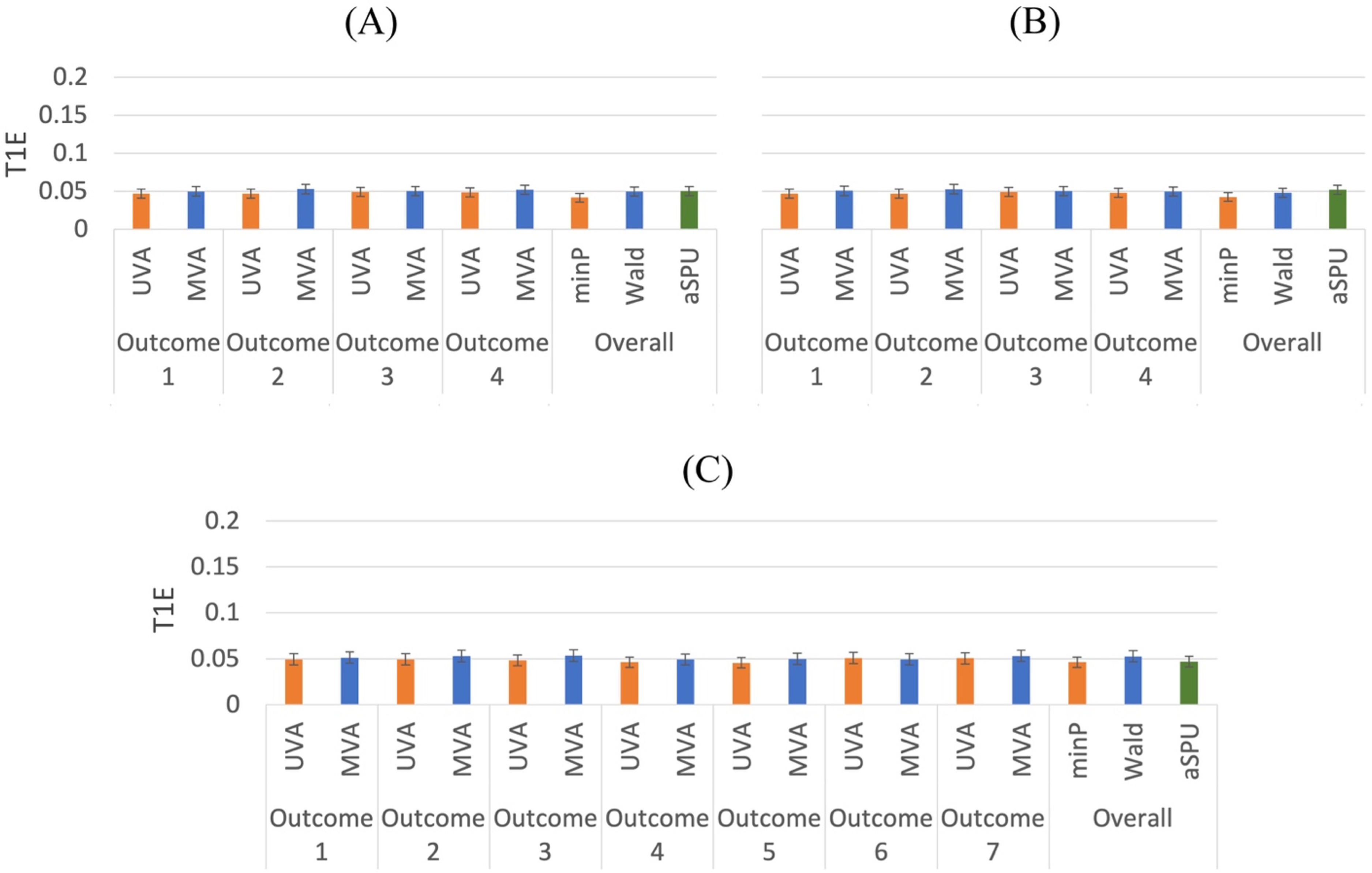
Type I error comparison of univariate and multivariate methods for mixed outcomes. 5000 replications. (A) 2 binary and 2 continuous outcomes.
Different Scenario Settings Used for Power Comparison.
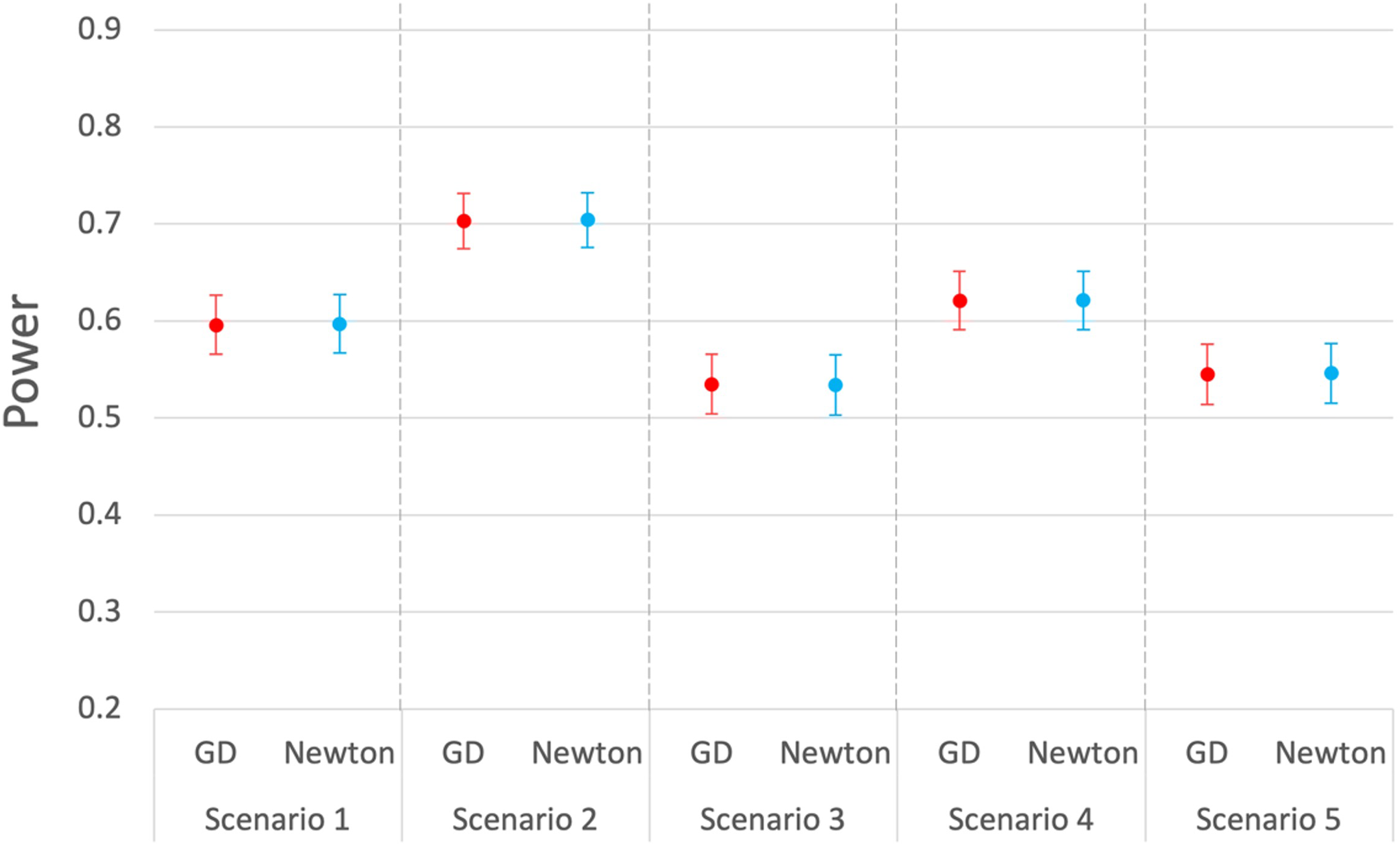
Power comparison of the Wald tests for mixed outcomes (2 binary, 2 continuous) using gradient descent and Newton's method. 1000 replications.
Comparing the power performances of univariate and multivariate analysis with regard to the overall test (minP for univariate analysis; Wald, aSPU for multivariate analysis), we find that the Wald test usually has higher power than the minP test, as shown in Figure 5. The aSPU test has the highest power when the proportion of outcomes affected by the exposure is relatively high, though it may be disadvantaged when there is only one outcome affected by the exposure.Our finding is consistent with the literature about the aSPU test.
33
The minP test is close to the SPU(
Note that for the above scenarios, we assume that the IVs are independent, which is usually the case for Mendelian randomization studies using summary statistics. However, for the two-stage methods, we can demonstrate that IVs with moderate correlations can also be used. We explore another situation with 4 mixed outcomes (2 binary, 2 continuous) and 10 correlated IVs. We select Power comparison of univariate and multivariate methods for mixed outcomes. 1000 replications. (A) 2 binary and 2 continuous outcomes. T1E and power comparison of univariate and multivariate methods for mixed outcomes using 10 correlated IVs. 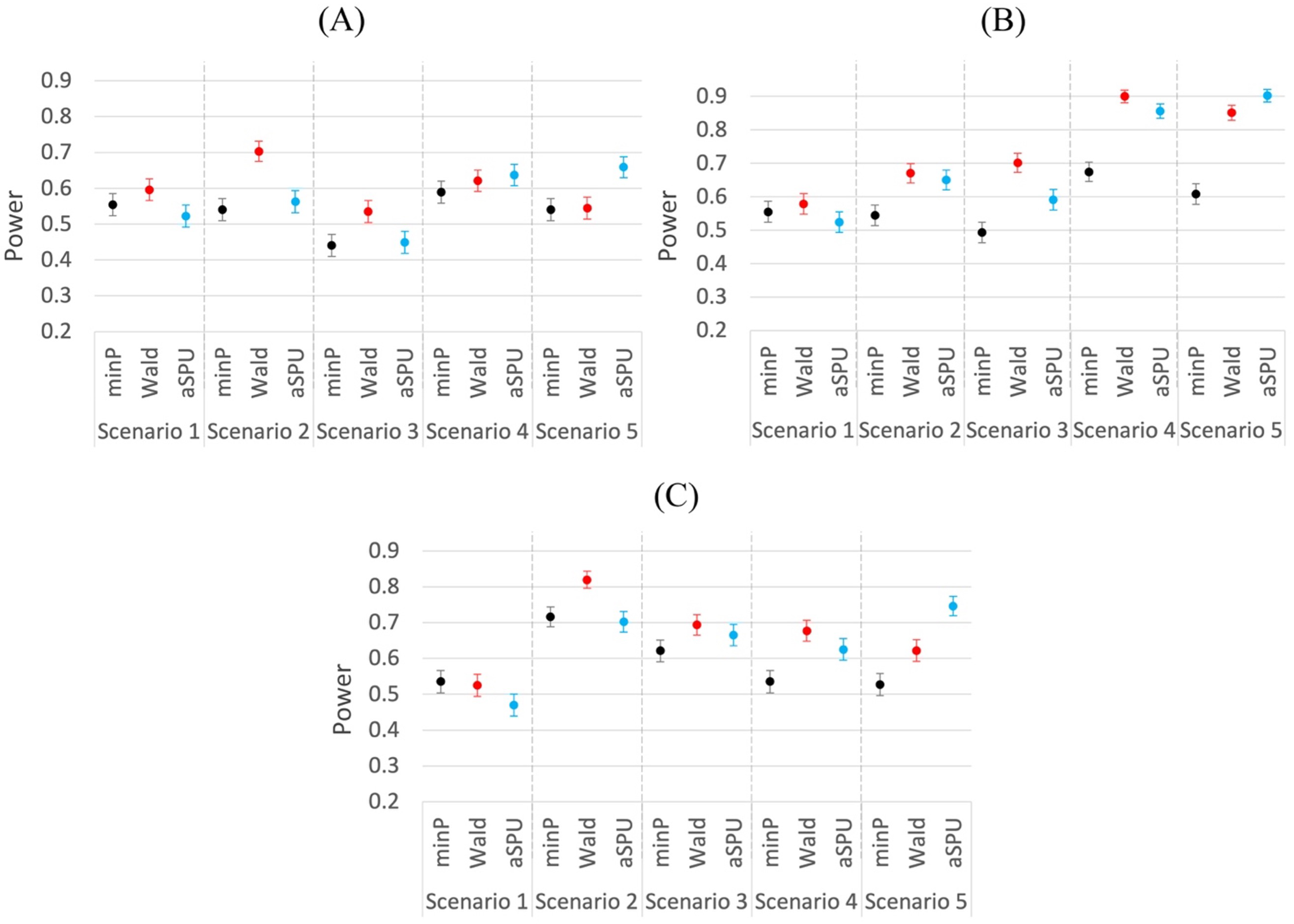
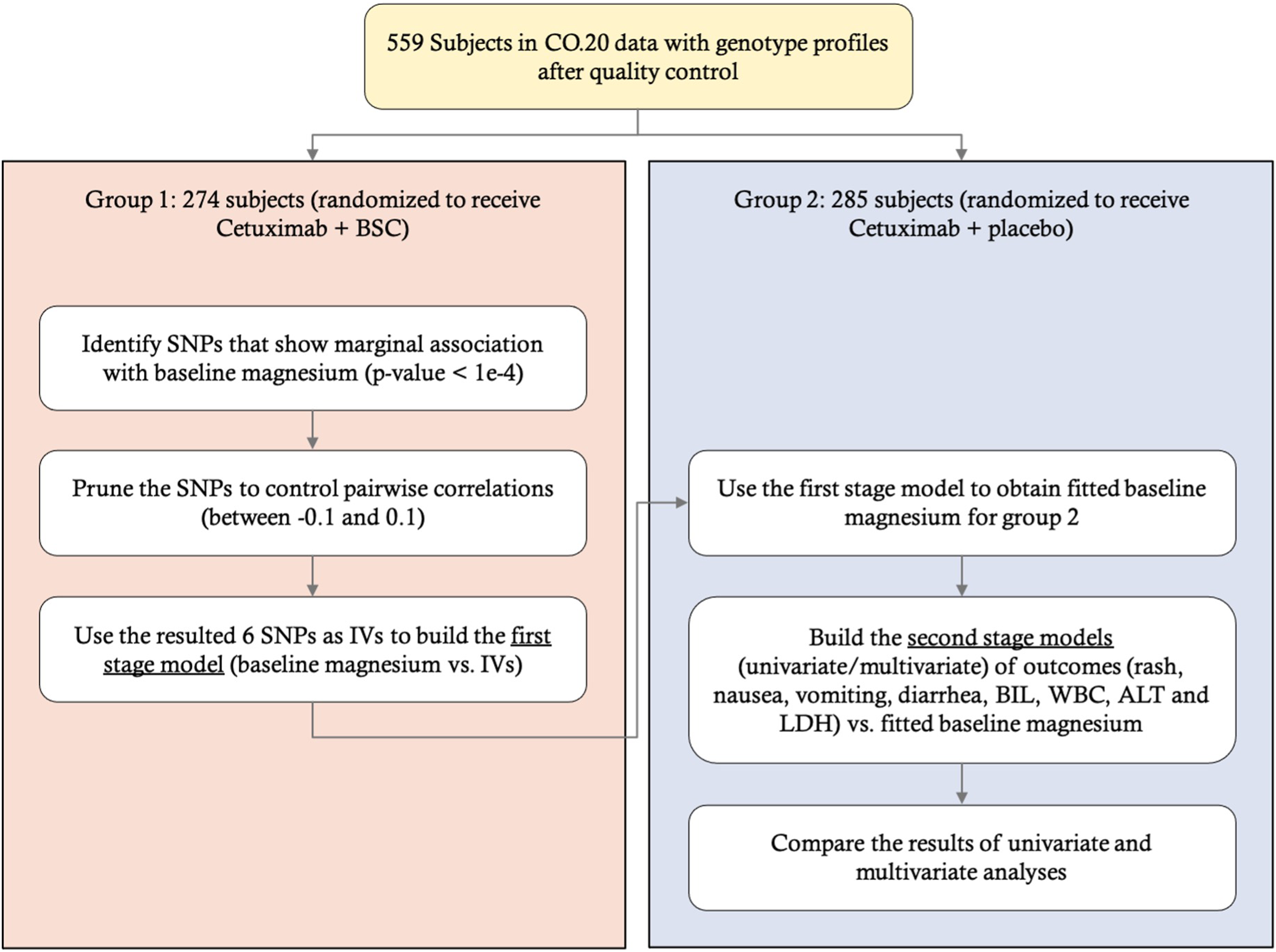
Workflow for the CO.20 study application.
To further demonstrate the difference between univariate and multivariate analyses in practice, we look at the data from the CO.20 trial as mentioned in the introduction, which is randomly assigned into two treatment groups (cetuximab + BRI; cetuximab + placebo), which, for convenience, we denote as Group 1 and Group 2, respectively. We choose baseline magnesium level (scaled by 10) as our exposure of interest X, since it is associated with certain genetic variants, and we are interested in examining whether this variable affects any of the following outcome variables: 3 binary toxicity variables (rash, nausea, and vomiting) and 4 continuous lab variables (bilirubin [BIL], white blood count [WBC], alanine aminotransferase [ALT], and lactate dehydrogenase [LDH]). Each binary toxicity variable is coded as 0 or 1 (has not experienced or has experienced a toxicity event within 8 weeks), and each lab variable is defined as the worst (maximum) value within 8 weeks after allocation. We take log-transformation on BIL, ALT, and LDH and remove 3 subjects with outlying values, defined as more than 4 standard deviations away from the sample mean value. The sample correlation matrix of the 7 outcomes is illustrated in Appendix B of the Supplementary materials, which shows that most outcomes are weakly or moderately correlated.
Using the data from Group 1, we obtain the marginal associations of SNPs with X (adjusted for age and gender) and select SNPs with marginal p-values smaller than 1E-4 and MAFs greater than 0.05. After pruning the SNPs to control pairwise correlations to be within −0.1 and 0.1, we end up with 6 SNPs as our IVs and build a first-stage model of X vs. IVs. Next, we use the first-stage model to obtain fitted X for Group 2, after which we conduct the second-stage analysis, applying the univariate and multivariate modeling approaches to explore whether X has any effects on any of the mixed outcomes. Figure 7 shows a summary of our workflow.
The p-values for testing baseline magnesium's effect on each of the outcomes based on univariate analysis are shown in Table 2. We also include the results of univariate analysis without MR, where each outcome is regressed on observed X instead of fitted X. According to the table, based on univariate MR analysis, BIL and ALT show some significance with p-values less than 0.05. Meanwhile, possibly as a result of unhandled confounders, the p-values based on univariate analysis without MR are usually quite different. This may partially demonstrate the effect and importance of considering instrumental variables for causal inference.
P-Values for Testing Baseline Magnesium's Effect on Each of the Outcomes Based on Two-Stage Univariate MR Analysis and Univariate Analysis Without MR. Exposure: Baseline MG.
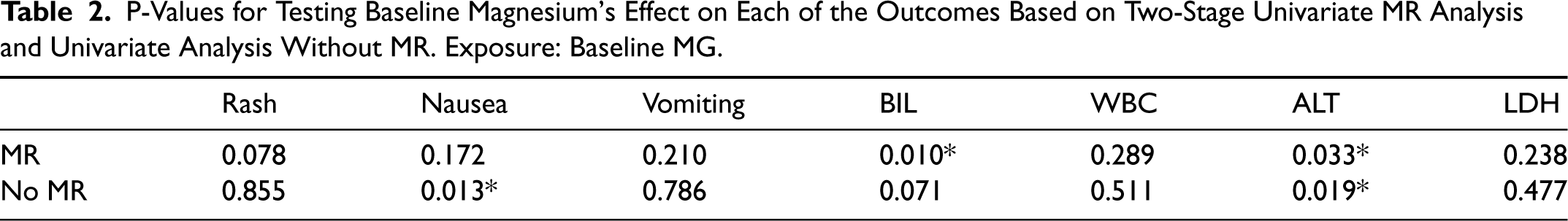
P-Values for Testing Baseline Magnesium's Effect on Each of the Outcomes Based on Two-Stage Univariate MR Analysis and Univariate Analysis Without MR. Exposure: Baseline MG.

Table 3 summarizes the results of univariate and multivariate analyses. For estimating and testing a single effect, univariate and multivariate analyses provide consistent results, though they are not exactly the same. However, the overall hypothesis test results are very different. The p-value of the minP test is not significant, but the p-values of the Wald and aSPU tests are. This agrees with our simulation results, demonstrating the potential power advantage of multivariate analysis.
Comparison Between Univariate and Multivariate Methods for Mixed Outcomes (Rash, Nausea, Vomiting, BIL, WBC, ALT, and LDH). Exposure: Baseline MG.
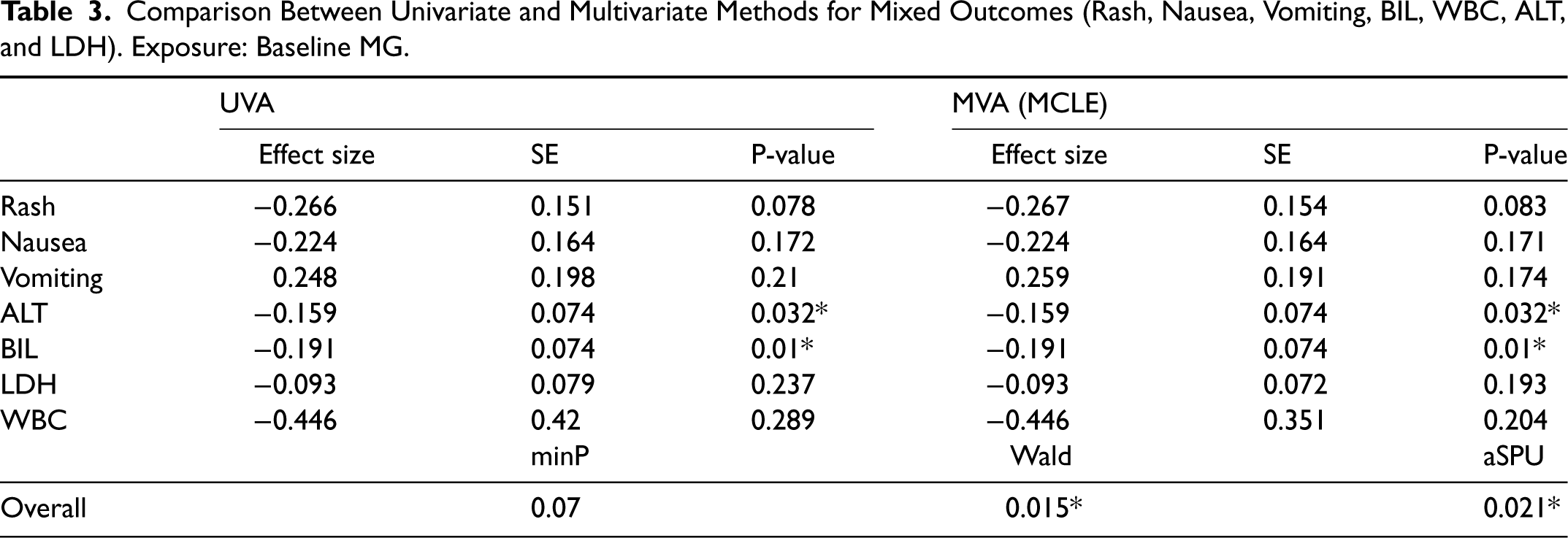
Thus, the proposed MRMO model is shown to identify the statistically significant associations between genetic predisposition of having lower magnesium levels to hepatotoxicities (elevated ALT levels and elevated bilirubin levels) in cetuximab-treated patients.
We have presented a novel approach to conduct two-stage Mendelian randomization with multivariate analysis on mixed outcomes. As shown in our simulations, our innovative approach can increase the power of the overall test over the univariate approach in most scenarios, especially when more than one outcome is affected by the exposure. The two different overall tests based on multivariate analysis, Wald and aSPU tests, have different performances in different scenarios. The aSPU test shows better power when the proportion of the outcomes affected by the exposure is higher, while the Wald test performs the best when the exposure has causal effects on a small to medium proportion of the outcomes. We have also noticed that carrying out the tests using our proposed gradient descent algorithm tends to yield very similar results to those based on the Newton–Rapson method, which was used by Bai et al. 27 Nevertheless, given the possible drawbacks of the Newton–Rapson method in some other scenarios,29–31 we recommend using the gradient descent approach, which may take more iterations to converge but can avoid some potential computational issues.
After applying the univariate and multivariate methods to the CO.20 data, we have found that the parameter estimations for single outcomes were close. However, the minP test for testing the overall hypothesis did not give a significant p-result, whereas both the Wald test and the aSPU test in our new method showed significant results. The increased statistical power offered by multivariate analyses was therefore able to identify a previous latent relationship between predisposition to hypomagnesemia and hepatotoxicity from cetuximab-treated chemo-refractory colorectal cancer patients. This is relevant clinically because hypomagnesemia has been associated with improved outcomes in cetuximab-treated patients. Because we identified that predisposition to low magnesium may lead to increased liver toxicities, new avenues of research have been opened, where predisposition to low magnesium levels may lead either to increased levels of cetuximab (pharmacokinetic association) or increased efficacy of cetuximab on metastatic cancer (pharmacodynamic association). Confounding this association is the knowledge that the elevated bilirubin may be due either to drug therapy or bulky liver metastasis. Because we also found a separate relationship between predisposition to hypomagnesemia and elevated ALT, this result may suggest but does not prove that the relationship may be related more to drug toxicities than liver metastases, because ALT abnormalities are more commonly seen with drug toxicity than with liver metastases.
We would like to point out that our chosen screening criteria for the instrumental variables were relatively loose due to the limited sample size of the CO.20 data, which may raise concerns about the possible violation of instrumental variable assumptions. In order to alleviate this potential problem, one approach we can consider is to combine different datasets to get a larger sample size. This may require modifying our model framework to take the different structures of different datasets and potential correlations into account, which is a future direction worth exploring. Another possible approach is to use the publicly available genome-wide association study (GWAS) results, which are usually based on sufficiently large samples, to select instrumental variables. However, it will not be feasible when there are no available GWAS results on the exposure of interest.
In the future, we may explore the possibility of developing multivariate methods that only require summary statistics, as well as methods that can handle certain invalid instrumental variables. Besides, we have demonstrated that our current approach can handle a mixture of binary and continuous outcomes. It can be directly applied to situations where all outcomes are binary, or all outcomes are continuous as well. We may also look at other types of outcomes (e.g. survival outcomes) and develop new methods that incorporate multivariate Mendelian randomization.
Software
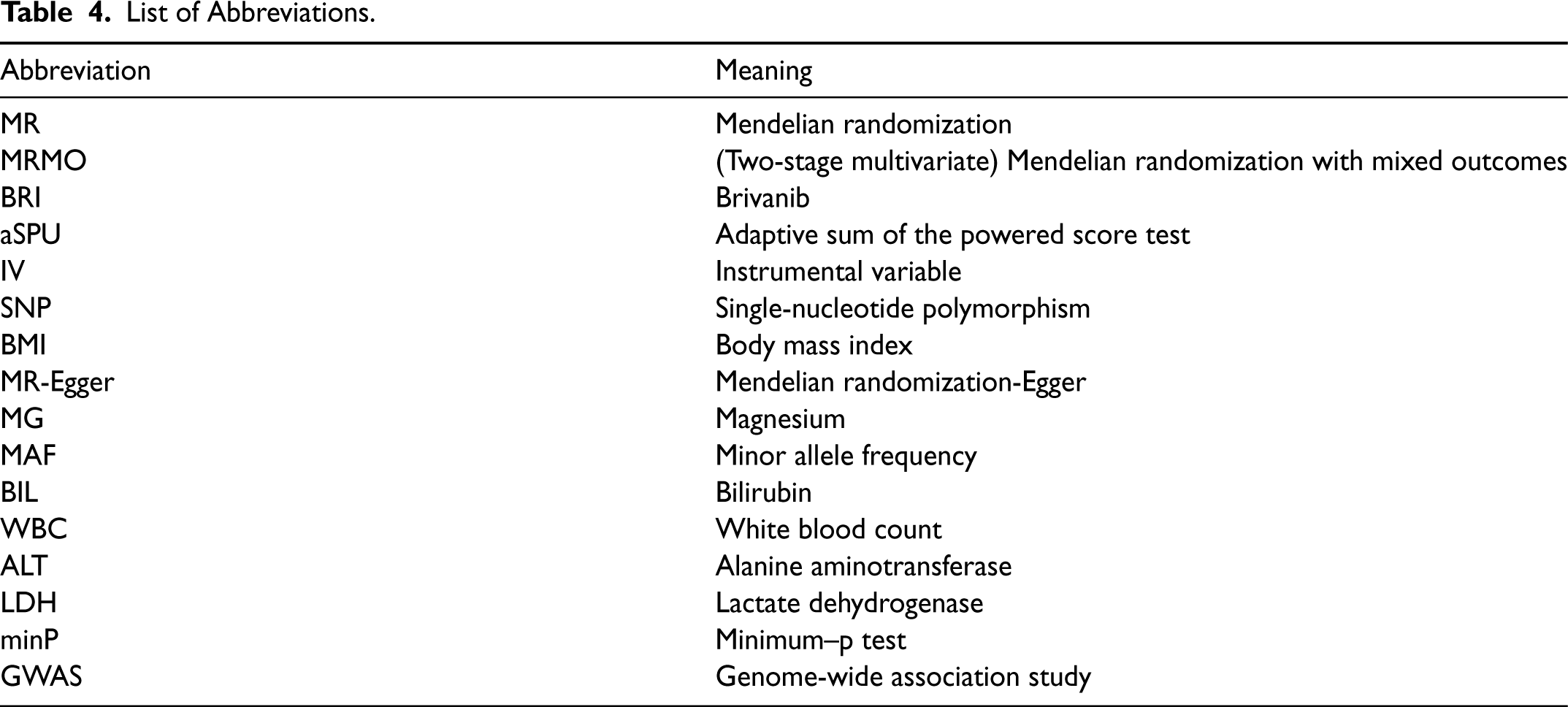
List of Abbreviations.
Supplemental Material
sj-docx-1-smm-10.1177_09622802231181220 - Supplemental material for Two-stage multivariate Mendelian randomization on multiple outcomes with mixed distributions
Supplemental material, sj-docx-1-smm-10.1177_09622802231181220 for Two-stage multivariate Mendelian randomization on multiple outcomes with mixed distributions by Yangqing Deng, Dongsheng Tu, Chris J O'Callaghan, Geoffrey Liu and Wei Xu in Statistical Methods in Medical Research
Footnotes
Acknowledgments
The authors would like to acknowledge the clinical contributions of Lillian Siu.
Declaration of conflicting interests
The author(s) declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Lusi Wong Fund, Princess Margaret Cancer Foundation, Alan Brown Chair in Molecular Genomics (to GL). WX was funded by the Natural Sciences and Engineering Research Council of Canada (NSERC Grant RGPIN-2017-06672).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.