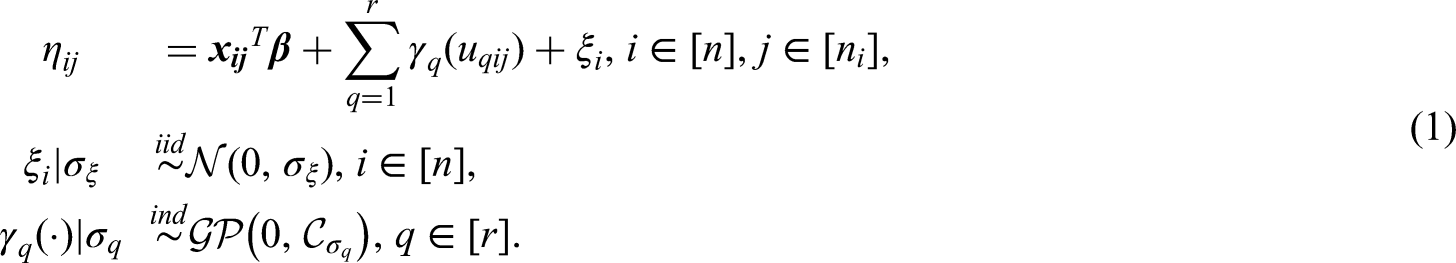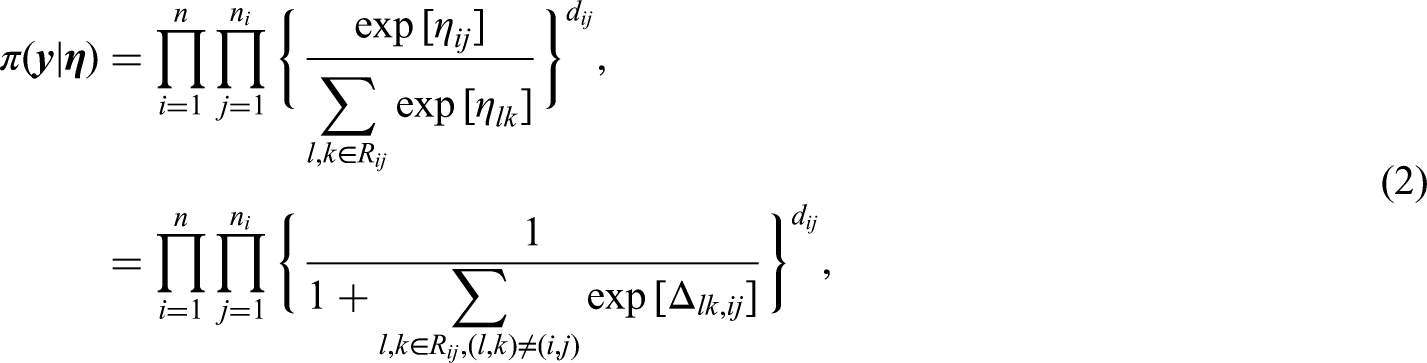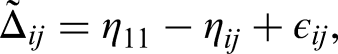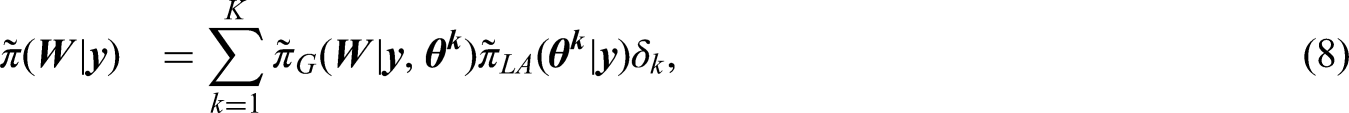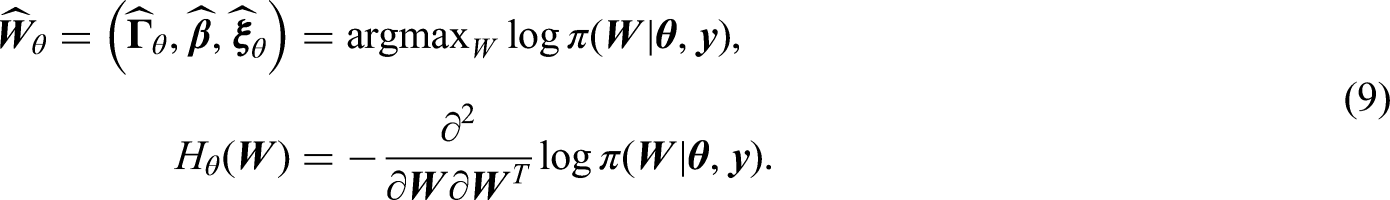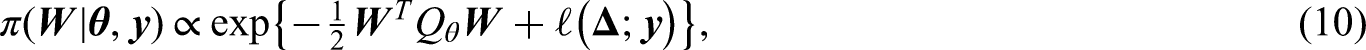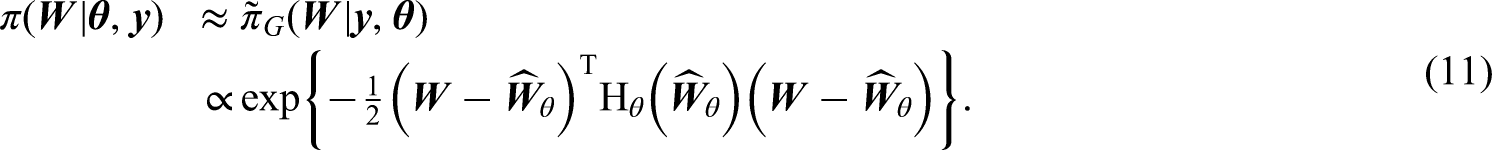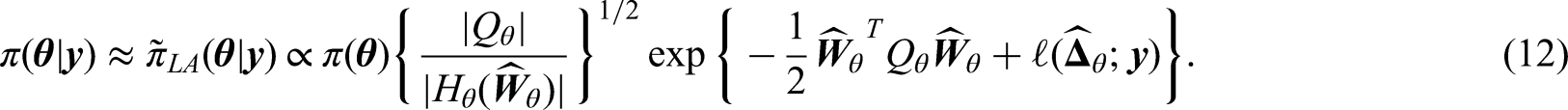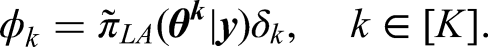Abstract
We propose a flexible and scalable approximate Bayesian inference methodology for the Cox Proportional Hazards model with partial likelihood. The model we consider includes nonlinear covariate effects and correlated survival times. The proposed method is based on nested approximations and adaptive quadrature, and the computational burden of working with the log-partial likelihood is mitigated through automatic differentiation and Laplace approximation. We provide two simulation studies to show the accuracy of the proposed approach, compared with the existing methods. We demonstrate the practical utility of our method and its computational advantages over Markov Chain Monte Carlo methods through the analysis of Kidney infection times, which are paired, and the analysis of Leukemia survival times with a semi-parametric covariate effect and spatial variation.
Keywords
Introduction
For problems involving time-to-event data, the combination of Cox proportional hazard (Cox PH) models and inference via partial likelihood has been the dominant methodology following its development by Cox. 1 The Cox PH model assumes that any two subjects’ event hazards are proportional as a function of time, with the ratio depending on unknown covariate effects which are inferred from the observed data. Event times may be correlated within the sample, for example when the response is time to kidney failure for the left and right kidneys from the same subject. Inference that is conducted via partial likelihood does not require assumptions to be made about the form of the baseline hazard. Further, the use of Bayesian inference with the Cox PH model is desirable as this (a) allows the use of substantive prior information on the hazard ratios and (b) provides uncertainty quantification for all parameters of interest in the presence of complex models for the hazard, which would be difficult to achieve otherwise. However, existing methods for approximate Bayesian inference based on integrated nested Laplace approximations (INLA) 2 cannot be applied to the Cox PH model with partial likelihood because the Hessian matrix of the log partial-likelihood is fully dense while INLA requires this matrix to be diagonal.
Alternative methods of making Bayesian inference for this kind of survival model have been considered in the literature. Dykstra and Laud 3 considered a fully non-parametric approach for Bayesian survival analysis, where the entire hazard function is modeled with an extended gamma process prior and the posterior distribution is derived to be another extended gamma process. Kim and Kim 4 considered Bayesian analysis on Cox PH model on partial likelihood and on full likelihood with an extended gamma process prior for the baseline hazard, and carried out inferences using Markov Chain Monte Carlo (MCMC). Martino et al. 5 considered application of the INLA methodology to the Cox PH model, using the full likelihood with baseline hazard modeled semi-parametrically. Kalbfleisch 6 derived the partial likelihood to be the limiting posterior when baseline hazards are modeled with non-informative priors, and Sinha et al. 7 later extended the result to allow the inclusion of grouped survival data, and implemented partial likelihood-based Bayesian inference with a Gibbs sampling algorithm. Henschel et al. 8 proposed a Bayesian inference method using MCMC on the full likelihood, with baseline hazard function modeled either as piecewise constant or as linear combination of B-splines, and they accommodated the inclusion of different types of frailties in their method. Hennerfeind et al. 9 developed a general geo-additive Cox PH model that allows the inclusion of components such as non-linear covariate effect, spatial effect and group level frailties, with inference carried out using MCMC on full likelihood and baseline hazards modeled using P-splines. Kneib 10 generalized the method of Hennerfeind et al. 9 to accommodate left truncation, left censoring, and interval censoring of the survival times. Most of the existing methods for Bayesian inference of Cox PH model have been utilizing MCMC method to obtain the posterior, and are based on the full likelihood with an explicit form to model the unknown baseline hazard.
Stringer et al. 11 developed an approximate Bayesian inference methodology for case-crossover model, which can be viewed as a special case of Cox PH model, by applying the approximation strategy of INLA to a log-partial likelihood with a non-diagonal Hessian matrix. Their methodology includes nonlinear covariate effects and yields full posterior uncertainty for the corresponding smoothness parameters, an improvement over existing frequentist methods. The partial likelihood they considered corresponds to one of the simplest special case[s] of the general Cox PH model, and the Hessian matrix of their log-partial likelihood is block-diagonal and sparse. In contrast, the Hessian matrix of log-partial likelihood of Cox PH model is generally fully dense, leading to increased computational burden when compared to the model considered by Stringer et al. 11 Further, they use a manual integration strategy which requires the user to supply their own quadrature points, which requires specialist knowledge to do properly. This limits the practical utility of their method. In order to make approximate Bayesian inferences for the Cox PH model with partial likelihood, an alternative computational strategy is needed.
In this paper, we develop an approximate Bayesian inference method for Cox PH models with partial likelihood, that allows linear and nonlinear covariate effects, spatial effects and frailties for modeling correlations between survival times. The proposed inference method utilizes the Laplace approximation-based strategy2,11 in a novel way that accommodates the use of partial likelihood. Through two simulation studies, we illustrate the circumstances under which the proposed method yields improved results compared to existing methods based on full likelihood, and demonstrate the accuracy of the posterior approximation and its computational advantages compared to partial likelihood method fit with MCMC. We then further demonstrate the practical utility of the proposed method through the analysis of two classical datasets with correlated survival times, nonlinear covariate effects and spatial variations.
The remainder of this paper is organized as follows. In section 2 we describe the Cox PH model and the method of semi-parametric smoothing that will be used in the inference of the nonlinear covariate effect in this paper. In section 3, we describe our proposed methodology and the introduced improvements to solve the computational challenges presented by the complicated partial likelihood. In section 4 we illustrate advantages of the proposed methodology in two simulation studies and through the analysis of the Kidney catheter data analyzed by McGilchrist and Aisbett 12 and Leukemia survival data analyzed by Martino et al. 5 We conclude in section 5 with a discussion.
Model
A general Cox PH model
Suppose we observe
Define
Stringer et al. 11 considered a general linear predictor that accommodates both linear fixed and nonlinear semi-parametric covariate effects, but the type of likelihood they considered is one of the simplest special cases of the general partial likelihood of Cox PH model, and does not allow the estimation of group-level correlation of survival times.
To accommodate nonlinear covariate effects and correlated survival times, we define an additive predictor
The nonlinear covariate effects
To infer the infinite-dimensional parameters
We put a sum-to-zero constraint such that
Finally, define the variance parameter vector
The proposed method can also accommodate the inference of spatial variations, by replacing the covariance function of
Partial likelihood
Our inference is carried out via a partial likelihood function. Define the risk set
The partial likelihood (2) can be written in the following form:
Further define
In existing Laplace approximations for posterior distributions,2,5,11 the latent parameters
Such posterior approximation methods have the advantage that, when the likelihood can be factored out in the form of (4), the resulting log-likelihood Hessian matrix is diagonal and hence efficient to be computed and stored.
2
Alternatively, if the likelihood is in the form of (5), the Hessian matrix is still sparse even it is no longer diagonal.
11
However, if one considers applying such approximate Bayesian inference on Cox PH model with partial likelihood, the resulting Hessian matrix will be completely dense and with the number of elements growing quadratically with sample size
Approximate Bayesian inference
To deal with the problem of dense Hessian matrix, we proposed a new way to utilize the Laplace approximation, by defining the latent parameter vector to only include the parameters of interest,
When nonlinear semi-parametric covariate effect is included in the model,
Define
For the posterior of variance parameter (7), we follow the procedure of Stringer et al.
11
to approximate it with its corresponding Laplace approximation
For any fixed
To sample from
Computing the approximations (8) requires choosing a quadrature rule consisting of nodes
To mitigate the computational challenges associated with applying a manual quadrature rule for (8), we implement Adaptive Gauss-Hermite Quadrature (AGHQ). This technique has been motivated as a useful tool for Bayesian inference
26
and work has been done to show that it is very accurate when using only a very small number of quadrature points,28,27 for example attaining
Computing the AGHQ rule requires computation of the mode of the Laplace approximation:
In this section, we present two simulation studies and two data analysis examples. All the codes are available in the online supplementary materials.
Simulation studies
We will provide two simulation studies to demonstrate the accuracy of our proposed method and under which situations the accuracy is improved over the existing full likelihood method INLA. Also, we will show that the proposed method provides posterior approximations that are comparable to results of MCMC, with much shorter runtime.
Simulation with sparse frailties
In the first simulation study, we considered the Bayesian inference problem for models with sparse frailties. In other words, survival times were correlated within groups while the number of observations in each group is small. We randomly generated
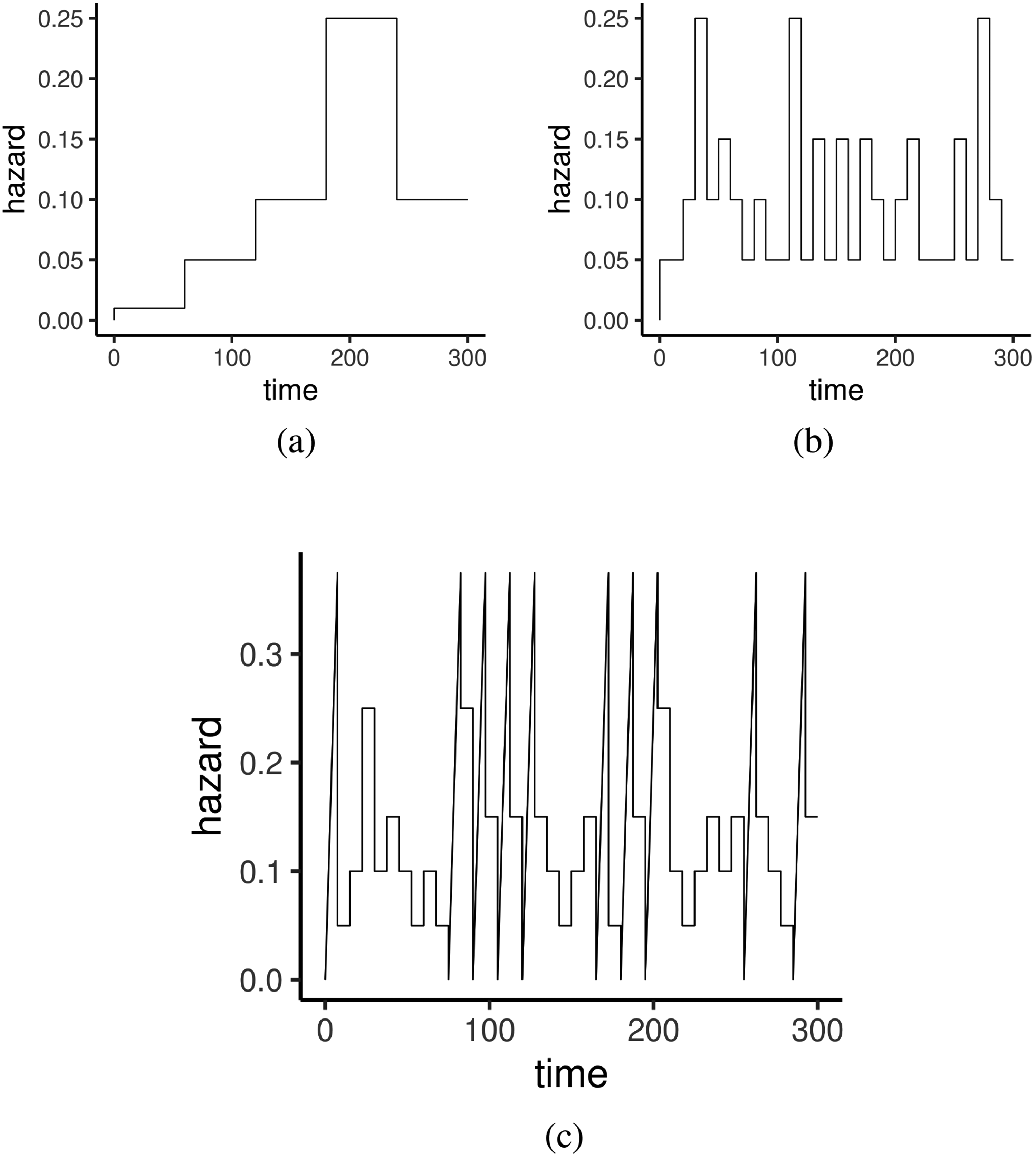
True Baseline Hazards in the two examples in section 4.1. (a) Simple stepwise baseline. (b) Oscillating stepwise baseline. (c) Complicated baseline.
The fixed effect
The comparison metrics when
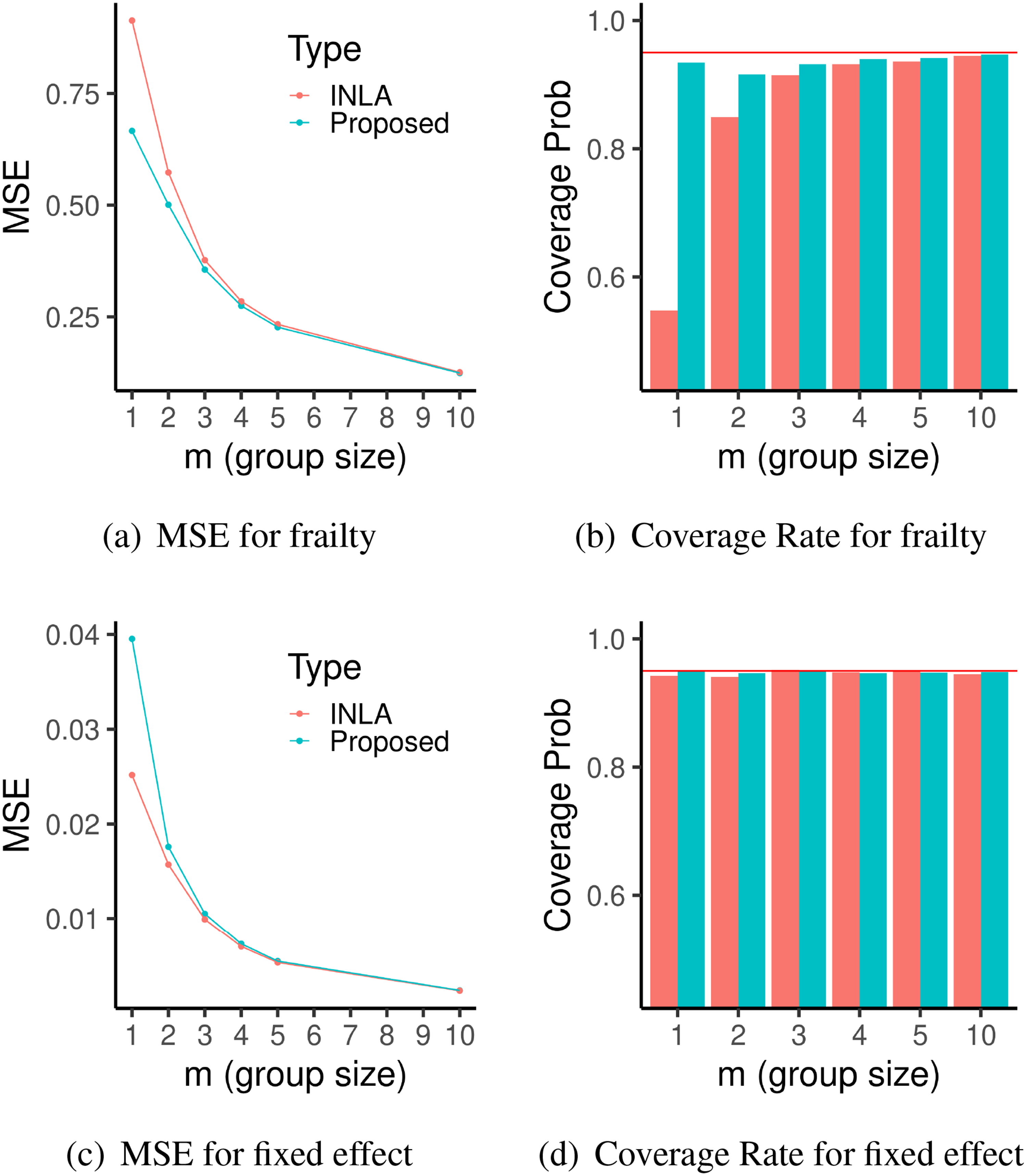
Results for the first simulation with sparse frailty in section 4.1. Left: Plots of MSE of frailties (a) and fixed effect (c) from 5000 independent replications with different
For completeness, we implemented the same simulation setting for
We then study the performance of the two inference methods when the group sizes
To compare the performance at different levels of censoring rates, we use the setting of
The simulation result above seems to be related to the type of full likelihood INLA utilized in its inference.
5
In Cox,
32
Cox pointed out one problem associated with the use of the type of full likelihood considered by Martino et al.,
5
that is the large number of parameters introduced in order to model the unknown baseline hazard function. In this simulation study, the sample size is only
To demonstrate the accuracy of our proposed approximation and the computational advantage compared to existing method, we also fitted the same partial likelihood model using MCMC method, through STAN’s No U-turn Sampler (NUTS),
33
using two replications where
The difference between posterior distributions yielded by the MCMC method and the approximate posterior obtained by the proposed method is quantified using Kolmogorv–Smirov (KS) statistic, which denotes the maximal absolute difference between the two cumulative posterior distributions, with a larger value indicating less similarity between two distributions. The KS statistics have been computed for both
KS statistic for each parameter in the first simulation study with sparse frailty in the “Examples section”, to compare the proposed approach with MCMC.
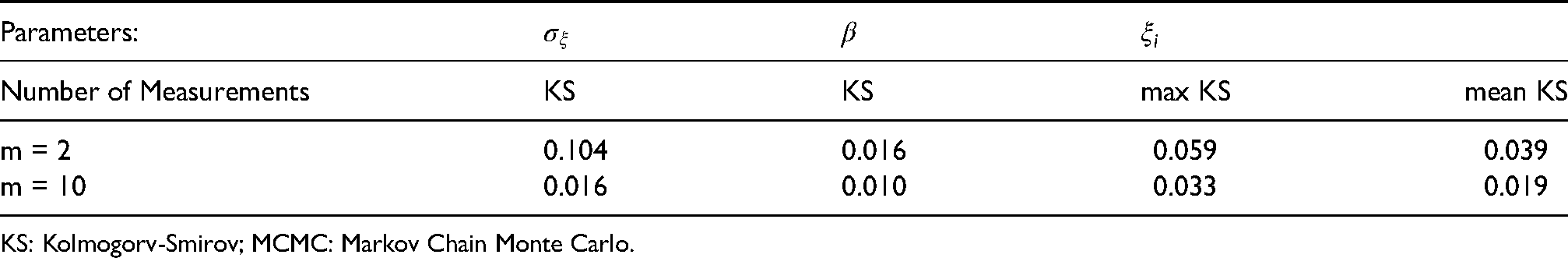
KS: Kolmogorv-Smirov; MCMC: Markov Chain Monte Carlo.
To compare the accuracy of our method with INLA when the smoothness assumption for baseline hazard function is violated, we performed our second simulation study. We generated
To infer the unknown risk function
The comparison metrics under the three settings of baseline hazard are shown in Figure (3). Figure (3)(a) shows boxplots of the mean squared errors provided by the two methods for
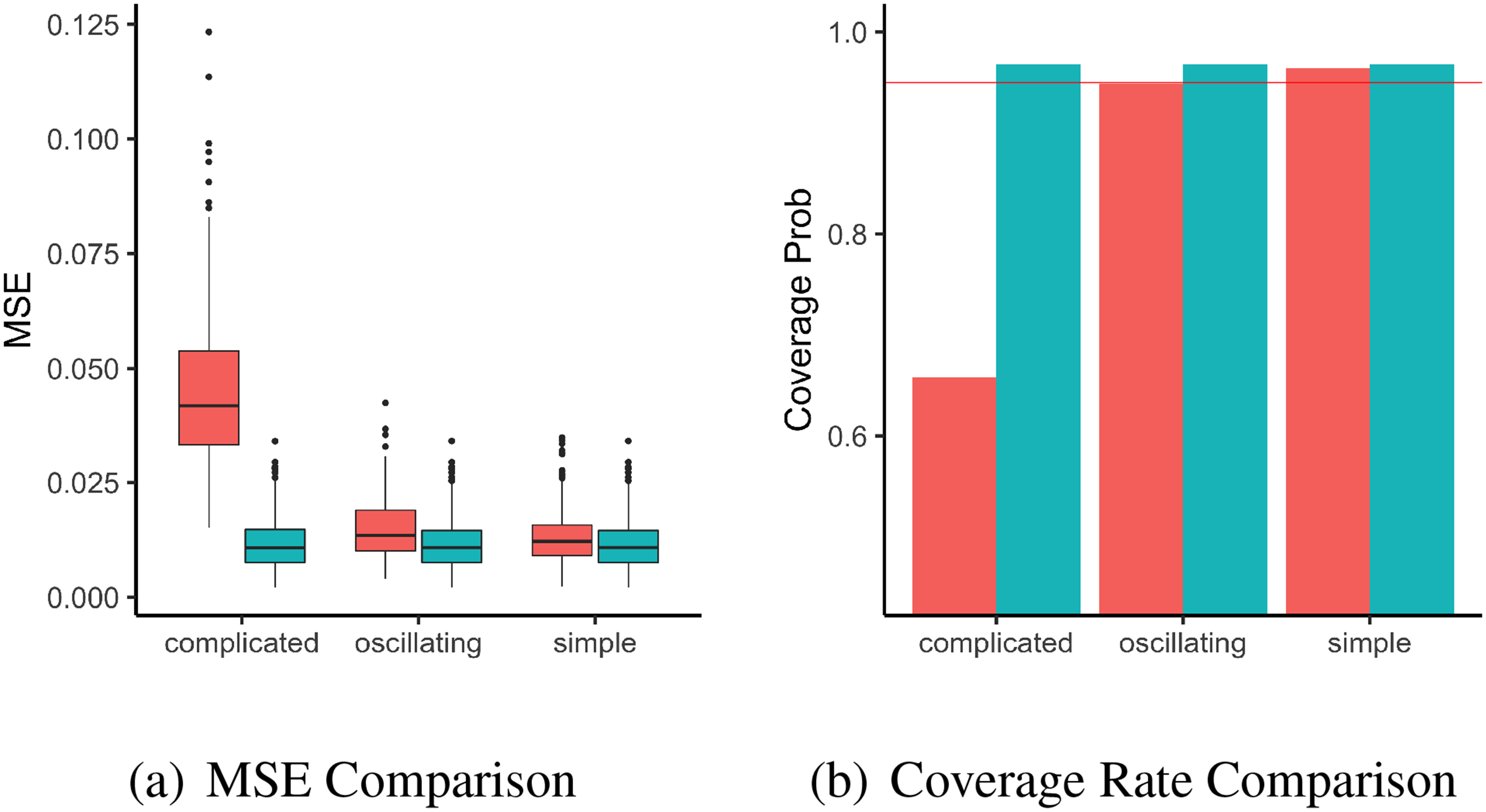
Results for the second simulation with non-smooth baseline in the section 4.1.2. (a): Plots of MSE from 1000 replications with different baseline settings, using INLA (red) and the proposed method (blue). (b): Bar-plot of Coverage Rate of 95% posterior credible intervals from 1000 replications with different baseline settings, using INLA (red) and the proposed method (blue). The red horizontal line is the nominal rate of 95%. INLA: integrated nested laplace approximation; MSE: mean square error.
While the performance of the proposed method is not affected by the choice of true baseline hazard function, the performance of INLA is sensitive to the true baseline hazard function, as shown in the corresponding boxplots and barplots. This is not unexpected as the full-likelihood used in INLA’s inference implicitly requires that the baseline hazard is smooth enough to be approximated well by its first-order random walk, which will not hold under setting such as Figure (1)(c) where the baseline hazard is varying rapidly as time changes. On the other hand, the inference of our proposed method relies on the partial likelihood, which makes no assumption on the form of the baseline hazard, and hence unaffected by the wiggliness of the baseline hazard in this study.
Again, to assess the accuracy of the proposed posterior approximation, we fitted the same partial likelihood model using MCMC using NUTS with four chains each with 10,000 iterations and 8000 warmups, for an arbitrarily chosen replication. The KS statistic for variance parameter
Therneau et al.
35
analyzed a Kidney Catheter dataset using their proposed penalized partial likelihood method. The Kidney Catheter dataset contains 76 times to infection at the point of insertion of a catheter, for
We first analyzed this dataset on partial likelihood using the proposed method, with
Table (2) shows the results of our procedure compared to the results obtained using INLA and MCMC. Based on the table, our procedure gave different posterior means and reported larger posterior standard deviations compared to INLA, especially for the effects of different disease types. These results are also summarized in Figure (4), where the posteriors obtained from the proposed method are shown to be much closer to MCMC than posteriors obtained from INLA. As we have shown in the section 4.1, when sparse frailties exist, Bayesian inference on partial likelihood tends to be more stable than on full likelihood.
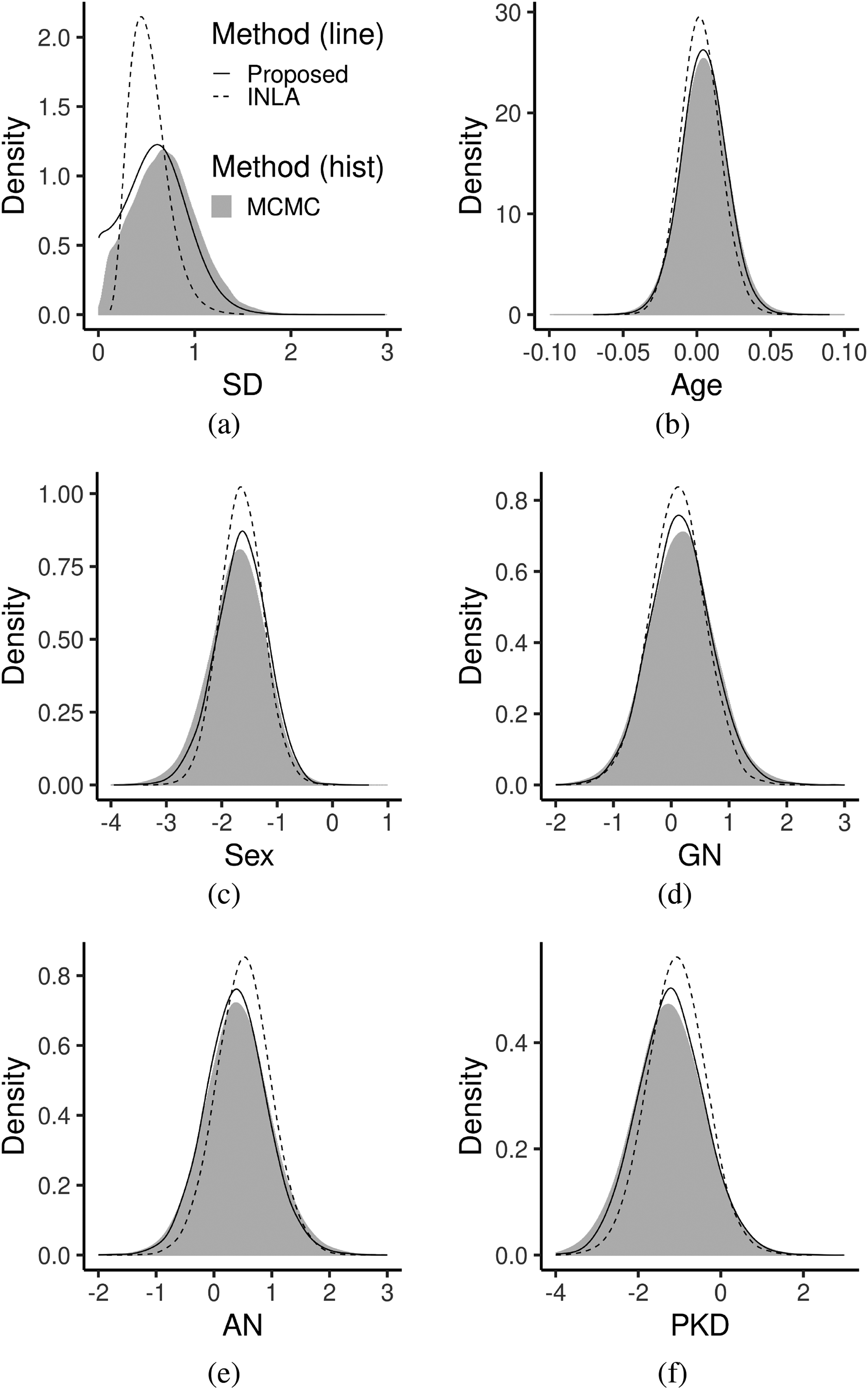
Posteriors of parameters for the kidney data in the section 4.2. Posterior distribution was obtained using the proposed method(solid line), using MCMC(gray histogram) and using INLA (dashed). (a) Posteriors of σξ. (b) Posteriors of βAge. (c) Posteriors of βSex. (d) Posteriors of βGN. (e) Posteriors of βAN. (f) Posteriors of βPKD. MCMC: Markov Chain Monte Carlo; INLA: integrated nested Laplace approximation.
Estimated means and standard deviations of linear effects by proposed method, INLA and MCMC for the kidney data in the section 4.2.
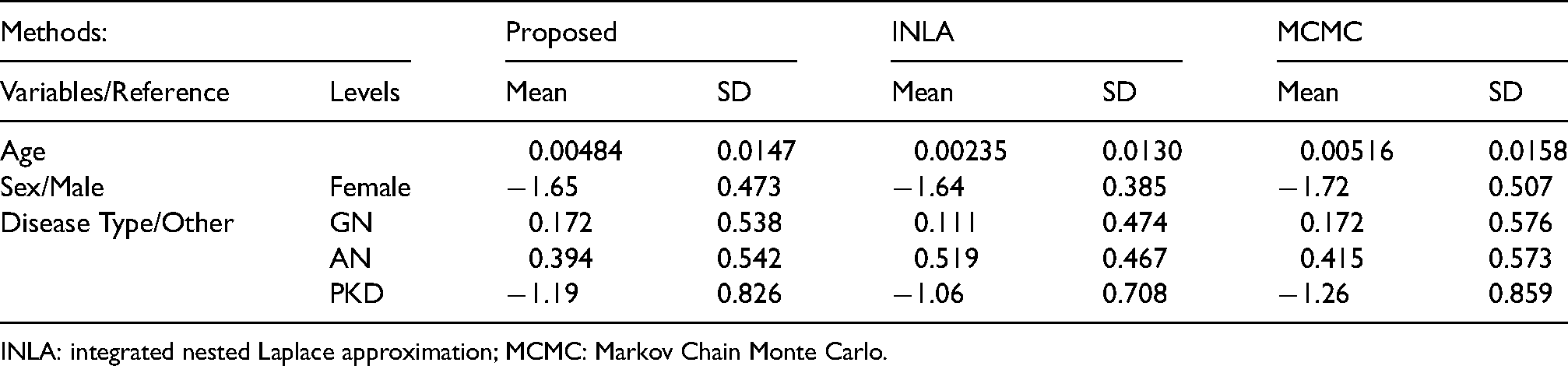
INLA: integrated nested Laplace approximation; MCMC: Markov Chain Monte Carlo.
Again, to quantify the difference between the proposed posterior approximation and the posterior yielded by MCMC, we computed the KS statistics for the five fixed covariate effects, the
The KS statistics show that the proposed method give similar posteriors to MCMC method, for most of the parameters. As shown by the KS statistics, the posterior for the variance parameter
In this example, we implemented our proposed procedure to fit a Cox PH model to the Leukemia data set analyzed by Martino et al.
5
as well as previously by Lindgren et al.
37
; Henderson et al.
38
The dataset contains information from
Based on the model comparison result from Kneib and Fahrmeir,
39
we follow the same analysis in Martino et al.
5
to model the effects of age, sex and white blood cell count linearly, and model the effect of the deprivation index (tpi) nonlinearly with the semi-parametric model described in the section 2 using 50 equally spaced knots. Prior distributions
Figure 5(a) shows the posterior mean and posterior 95 % credible interval of the exponentiated covariate effect of tpi using the proposed approach. Figure 5(b) shows the posterior and prior for the spatial correlation parameter
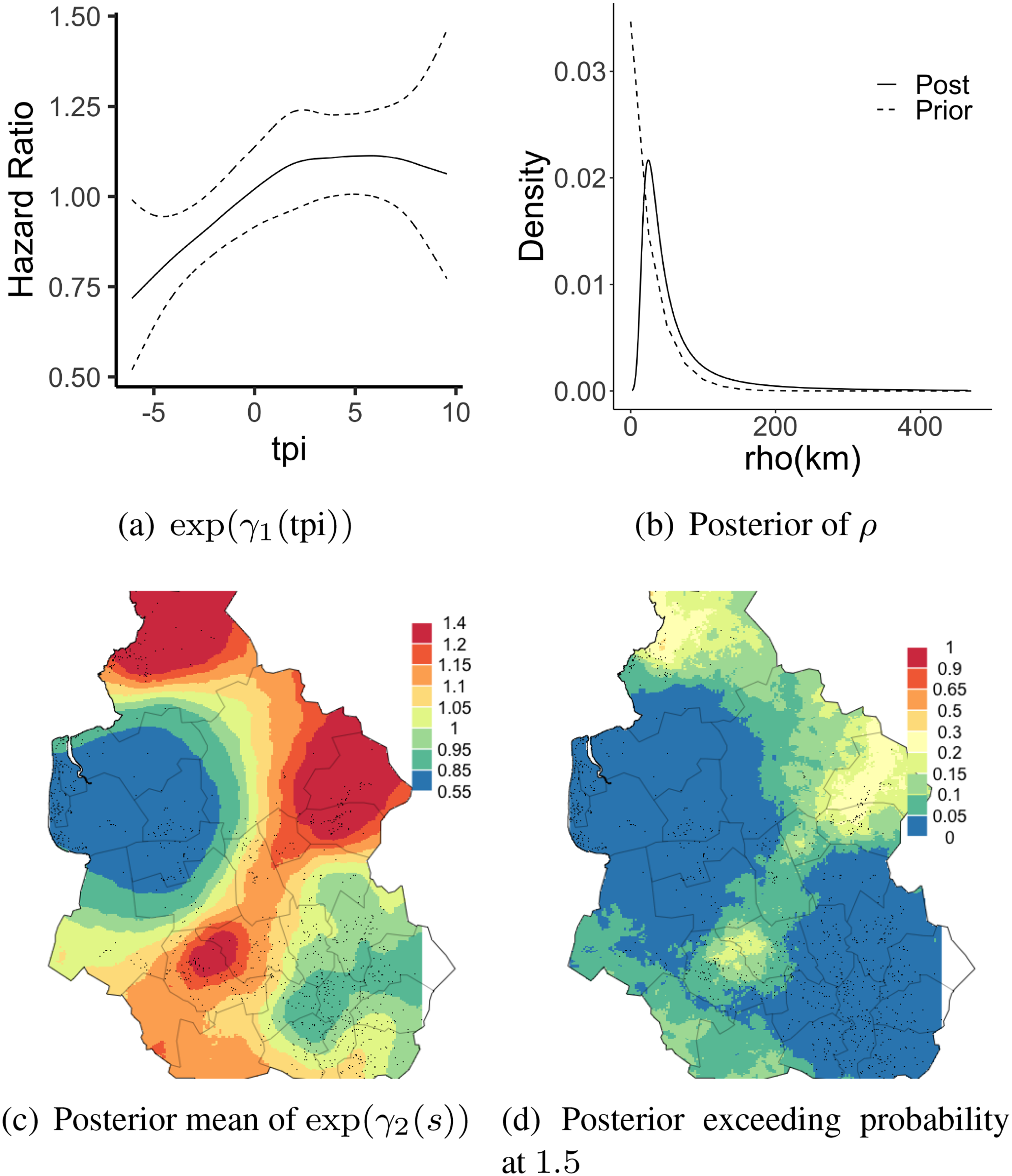
Results for the leukemia data in the section 4.3, (a): Posterior mean of (exponentiated) tpi effect (solid) and its 95% credible interval (dashed). (b) Posterior (solid) and prior (dashed) for the spatial
There are two major differences between our analysis and the analysis in Martino et al.
5
First, our analysis is carried out on partial likelihood using the proposed method, but the analysis in Martino et al.
5
is carried out on full likelihood and hence requires assumption on the form of true baseline hazard function. Second, the proposed approach is able to achieve higher resolution estimate for the spatial variation than the approaches taken by Martino et al.
5
because we include the full continuously sampled locations
To contrast the proposed inference method with the method of Stringer et al.,
11
we compared the runtimes of Cholesky factorization as well as the memory requirements for storage, of
The methodology we proposed in this paper provides a flexible way to carry out Bayesian inference for Cox PH models with partial likelihood, that accommodates the inference for nonlinear covariate effects, spatial variations and correlated survival times. The use of partial likelihood does not require any assumption on the baseline hazard function, which is an advantage over existing approaches for Bayesian inference in this model. We have demonstrated the accuracy and the computational efficiency of our new approach through simulation studies and analysis of two classical datasets in survival analysis. Our proposed method is an appealing option to adopt for the analysis of time-to-event data, when the inference of baseline hazard is not of primary interest.
As shown in our first simulation example, when frailties are sparse, Laplace approximation-based methods tend to yield less accurate approximation for
The framework of this proposed methodology can be extended to fit more complex models, by modifying the covariance structure of the covariate with nonlinear semi-parametric effect or the covariance structure of the spatial variations. Because we accommodate the dense Hessian matrix of the log-likelihood, our approach could be extended to approximate Bayesian inference for other models with a dense Hessian matrix. We leave such extensions to future work. A R package that implements our proposed method called abcoxp is available at https://github.com/AgueroZZ/abcoxpGitHub.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802221134172 - Supplemental material for Bayesian inference for Cox proportional hazard models with partial likelihoods, nonlinear covariate effects and correlated observations
Supplemental material, sj-pdf-1-smm-10.1177_09622802221134172 for Bayesian inference for Cox proportional hazard models with partial likelihoods, nonlinear covariate effects and correlated observations by Ziang Zhang, Alex Stringer, Patrick Brown and Jamie Stafford in Statistical Methods in Medical Research
Footnotes
Data availability statement
The simulated data of the two examples in the “Simulation studies section” can be found at ![]() , together with the scripts to replicate the results in the paper. Data for example 4.2 were obtained from R package “INLA” and are freely available. Data for example 4.3 were obtained from R package “survival” and are freely available.
, together with the scripts to replicate the results in the paper. Data for example 4.2 were obtained from R package “INLA” and are freely available. Data for example 4.3 were obtained from R package “survival” and are freely available.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.