Abstract
This study demonstrates the application of open-source deep learning models from the computer vision domain to streamline tasks in near-infrared (NIR) hyperspectral imaging (HSI) data processing. Specifically, it demonstrates a challenging case of dry matter prediction in mango fruits under conditions typical for fruit importers and exporters, who often need to assess the quality parameters of fruit packed in boxes. NIR HSI offers a non-destructive alternative to traditional hot air oven drying for dry matter determination. However, processing HSI images of fruit boxes presents challenges due to objects touching, overlapping, or being partially hidden, which complicates traditional HSI analysis. Modern artificial intelligence (AI) approaches can simplify such HSI data analysis, and integrating AI with chemometric modeling represents a promising future direction for NIR HSI data processing.
Introduction
Mango quality can be assessed using either destructive or non-destructive methods. Destructive methods, such as cutting or probing, render the fruit unsellable. In contrast, non-destructive techniques preserve fruit integrity, are faster, require less labor, and maintain commercial value. Due to the drawbacks of destructive testing, there is growing interest in predictive models that use non-destructive data to estimate quality attributes traditionally measured destructively. This is typically achieved by training machine learning models on combined datasets, allowing predictions based solely on non-destructive inputs during deployment.
Hyperspectral imaging (HSI) is a promising non-contact, non-destructive method for rapid quality assessment of multiple fruits in a single scan, making it suitable for conveyor-based sorting systems. Near-infrared spectroscopy, particularly in the 400–1000 nm range, is effective for predicting dry matter in mangoes. A key challenge in HSI is analyzing multiple, variably oriented samples within a single image. Once samples are segmented and their mean spectra extracted, conventional chemometric methods can be applied. While most studies use controlled imaging conditions, practical applications like sorting lines or warehouse inspections require robust methods for handling random orientations and batch imaging.
This study presents a realistic HSI application, imaging mangoes in crates with variable orientations. Challenges include segmenting individual fruits from the background and neighboring objects, defining optimal regions of interest, and addressing reduced pixel density per fruit. Artificial intelligence (AI) models can effectively segment mangoes from complex backgrounds, although such approaches are underutilized in the NIR domain. This work aims to introduce AI-based segmentation techniques to NIR practitioners, highlighting their potential to simplify HSI data analysis. Integrating AI with chemometric modeling is a promising direction for future NIR HSI research.
Materials and methods
Data collection was conducted in December 2023, using 99 Kent and 391 Palmer mangoes sourced from Brazil. Upon arrival in Wageningen, fruits were stored at 8°C, 15°C, or 22°C to induce varying ripeness stages, then equilibrated at 20°C and 60% relative humidity one day prior to measurement. HSI was performed using an all-in-one VNIR system, 1 which automated image acquisition and calibration (black and white reference). Each hyperspectral image consisted of 224 bands spanning 399–1002 nm, with a spatial resolution of 870 × 1024 pixels. Individual mangoes were labeled with unique handwritten IDs for traceability. Dry matter content was determined by weighing each fruit before and after oven drying at 60°C for 60 h.
Each HSI file contained one or two carton crates, with up to 19 mangoes in total. For model building, a mask needed to be created for each individual mango. The open-source AI model ‘Segment-anything’ from Meta was used for this purpose. 2 Since this segmentation model could not take full HSI data as input, a copy of spectral band 125 from each HSI file was converted to a grayscale .jpg file and saved. This spectral band was chosen for its high contrast and low noise. To use the converted images in the model, they were loaded and then converted from grayscale to a BGR format, which the Segment-anything model accepted. The OpenCV library was used for image saving, loading, and conversion. 3 To minimize computing time, the ‘ViT-B SAM model’ was utilized.
After generating all masks, they were filtered to remove those not covering a mango. They were filtered based on overlays, maximum height and width, and minimum size. To filter out overlapping masks, a threshold of 90% overlap between two masks was set. If the threshold was exceeded, the smallest mask was discarded. Based on the masks, the mean spectra of the fruit were extracted and used for PLS regression analysis. In this study, the model was trained on the Palmer mango cultivar and tested on the Kent cultivar to demonstrate independent model testing.
Results
Segmentation using ‘Segment-Anything’ worked adequately, as a mask was created for every mango. Most masks were also correctly filtered, as shown with an example in Figure 1. Figure 2 shows an example input and output after filtering. From this output, one incorrect mask was still present in the top-left corner, which was filtered away manually. Although this manual step could be automated, there were only a limited number of incorrect masks, so it was more time-efficient to perform such filtering manually. For inline implementation, strategies such as the use of spectral information could be employed to filter such masks.
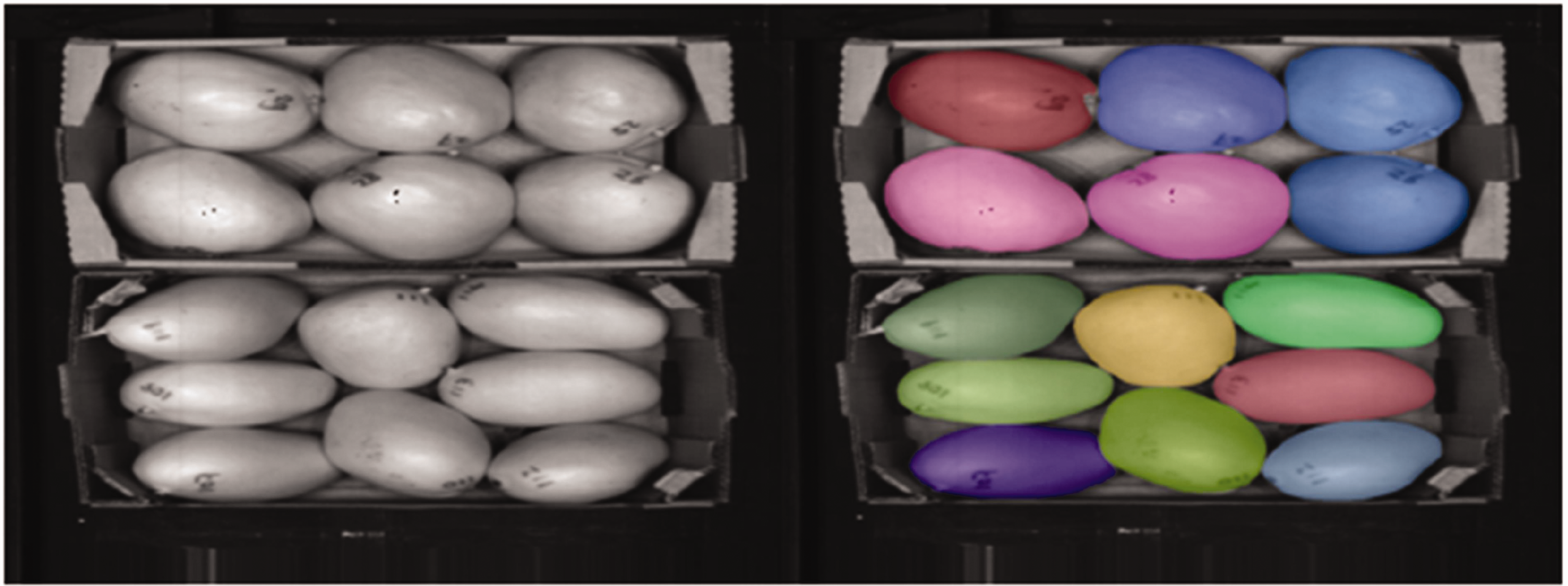
On the left, a visualization of a false colour hyperspectral image. This image was used as an input on the Segment-Anything model. To the right, the masks that are generated and all correctly filtered as shown.
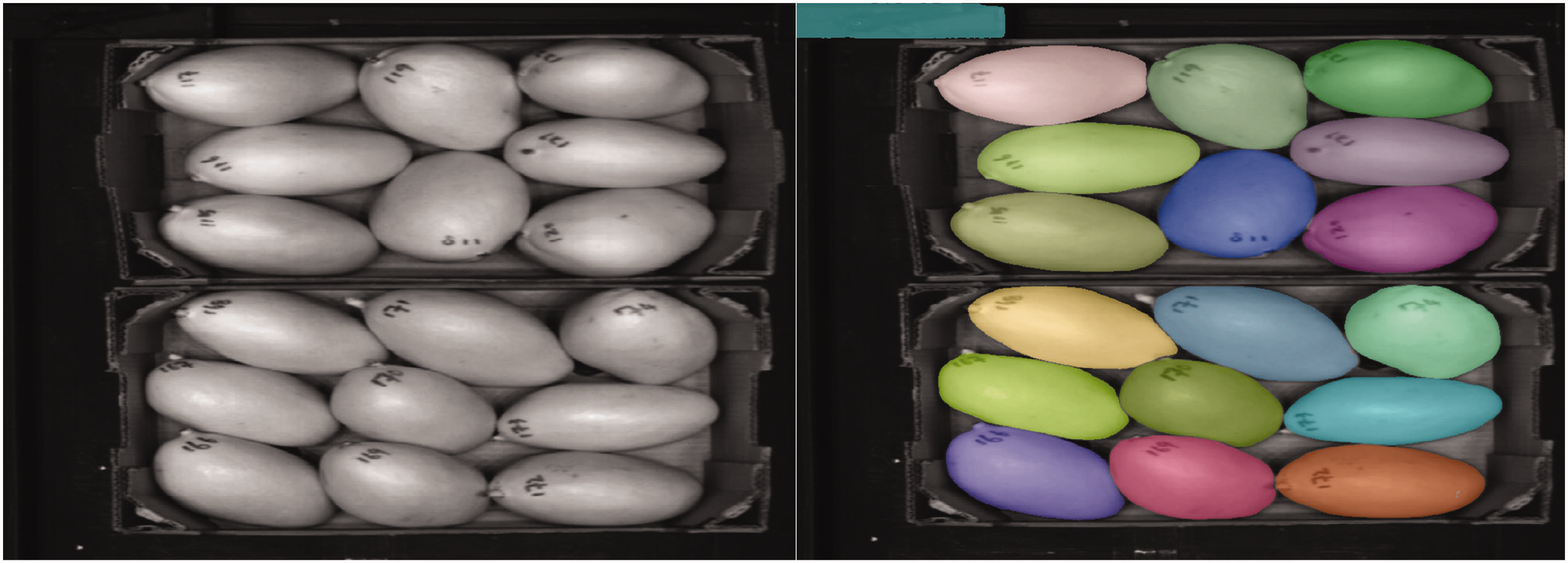

On the left, a visualization of a false colour hyperspectral image. This image was used as an input on the Segment-Anything model. To the right, the masks that are generated and filtered are shown, with one incorrect mask in the top left corner.
In the literature, different threshold-based segmentation methods have been presented for fruit segmentation; for example, the normalized difference vegetation index (NDVI) is also commonly used for this segmentation step. 1 However, the mangoes and the carton crates have similar reflectance on red and near-infrared wavelengths, which are used in NDVI calculation. Therefore, NDVI was not suitable for segmenting these hyperspectral images. Furthermore, many mangoes touch each other, which makes the implementation of a simple threshold a challenging task in this case. Once an individual mango is identified, the mean spectra of individual fruit can be extracted. Finally, the PLS model was calibrated on the Palmer cultivar and tested on the Kent cultivar, as demonstrated in Figure 3.
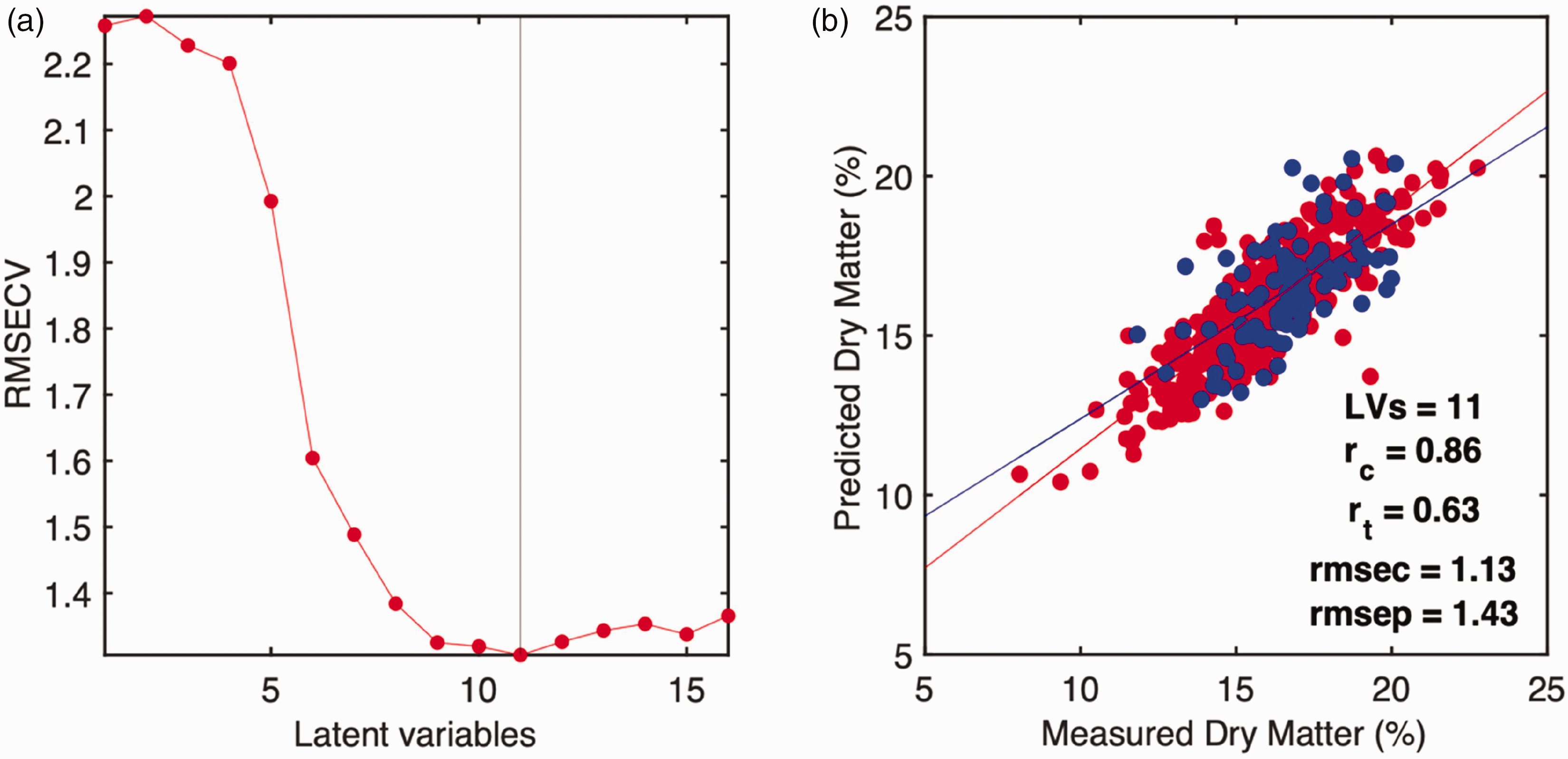
Cross-validation error plots and prediction plots for calibration of brix and dry matter. (a) Cross-validation curve for brix, and (b) prediction plot for dry matter.
Conclusions
The study aimed to demonstrate the potential of open AI models, such as the “Segment-Anything model,” to simplify some steps of HSI data handling, including object detection, segmentation, and spectra estimation. Thanks to the advanced AI algorithm, the segmentation of mangoes was achieved with minimal model adjustment. The extracted data were then used to build a predictive model for dry matter content in mangoes. This study is unique in that it was performed on minimally prepared samples, which is often the case when dealing with fruit import and export quality control. The study involved imaging mangoes directly in the boxes as supplied by the importers. This presented the challenge that mangoes were closely packed to each other, and AI appears to have handled the problem very well. Notably, no transfer learning or adjustments were performed on the AI model. The performance of the model in this research was based solely on two mango cultivars, Palmer and Kent. It would be interesting to see how this AI model and method can be generalized to different mango cultivars. The data processing in this study was done in batches, and optimizing processing time was not a top priority. For a real-world industrial application, real-time processing would be required. For model deployment based on the workflow of this study, it would be possible to convert it to a real-time prediction model and operate fully unsupervised. However, optimizing the Segment-Anything model and using a high-performance computer would be required to achieve real-time processing. AI can play a major role in simplifying HSI data analysis.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
