Abstract
An increasing number of studies have investigated the effects of Cd on human health. Cd-induced dermatotoxicity is an important field of research, but numerous studies have focused on the effects of Cd on the human skin. Moreover, most studies have been performed using HaCaT cells but not primary keratinocytes. In this study, we provide the results describing the cytotoxic effects of Cd exposure on primary human epidermal keratinocytes obtained from different donors. The subtoxic concentration of cadmium chloride was determined via MTT assay, and transcriptomic analysis of the cells exposed to this concentration (25 µM) was performed. As in HaCaT cells, Cd exposure resulted in increased ROS levels, cell cycle arrest, and induction of apoptosis. In addition, we report that exposure to Cd affects zinc and copper homeostasis, induces metallothionein expression, and activates various signaling pathways, including Nrf2, NF-kB, TRAIL, and PI3K. Cd induces the secretion of various cytokines (IL-1, IL-6, IL-10, and PGE2) and upregulates the expression of several cytokeratins, such as KRT6B, KRT6C, KRT16, and KRT17. The results provide a better understanding of the mechanisms of cadmium-induced cytotoxicity and its effect on human epidermal skin cells.
Introduction
Cadmium is a heavy transition metal with a broad range of industrial applications, leading to its release into the environment and severe ecological damage.1,2 Cd does not have any biological functions and is highly toxic to all ecosystems.3–5 Cd contamination leads to dramatic consequences for human health, including neurodegenerative diseases, 6 several types of cancer,1,7 and reproductive dysfunction.8,9 Cd is poorly excreted from the organism and has a half-life of 10–35 years. 10 Dermal exposure to heavy metals has received less attention than oral/inhalation exposure. However, in specific areas such as mine tailings or industrial enterprises, the risk of dermal exposure may be significant.11,12 Notably, Wang et al. demonstrated that, in urban soil samples, Cd exhibited the highest dermal bioaccessibility relative to other heavy metals. 13
In contact with the skin, cadmium may lead to epidermal barrier disruption accompanied by apoptosis induction and genotoxic stress. 14 On the rat model, it was shown that cadmium exposure induced various skin pathologies including hyperkeratosis, acanthosis, ex-pression of pro-inflammatory cytokines and mitotic index increase.15,16 Consequently, these changes may trigger psoriasis, atopic dermatitis and other pathologies. Previously, elevated level of Cd was observed in the skin of psoriatic patients with industrial exposure. 17 Similarly, Liaw et al. showed a correlation between plasma Cd levels and psoriasis severity. 18 Another clinical study revealed a significant difference in blood Cd levels between patients with psoriasis and the control group. 19 These studies provide evidence supporting the significant role of Cd toxicity in the emergence of skin pathologies.
Few studies have focused on Cd cytotoxicity in keratinocytes, and most of these studies used HaCaT keratinocytes as a model cell line. In particular, Cd induces cell cycle arrest, causes genotoxic stress, and increases reactive oxygen species (ROS) formation in HaCaT keratinocytes.20–22 HaCaT cells are a reliable alternative to normal human epidermal keratinocytes (NHEK) and are widely used in cytotoxicity studies.23–26 However, these cells bear two gain-of-function (GOF) mutations in TP53. The novel properties of mutant p53 are realized through two mechanisms. First, mutant p53 is capable of dimerizing with other transcription factors, preventing their binding to DNA. Second, mutant p53 can activate a much larger number of targets through direct interaction with other transcription factors (NF-Y, E2F1, NF-KB, and VDR). 27 These cells are unable to complete differentiation, have altered growth and proliferation rates, and exhibit impaired regulation of apoptosis.27–29 Thus, despite considered normal, HaCaT significantly differ from NHEK, which may limit the application of HaCaT cells as a model cell line for toxicological studies. Given the findings of the comparative proteomic analysis of HaCaT and NHEK, it is recommended to utilize primary cells as a model system, if feasible. 30 However, interpreting experimental results obtained from NHEK can be challenging due to donor-to-donor variability, the limited lifespan of the cultured cells, and differences between passages. 29
Transcriptomic analysis is a powerful tool to investigate gene activity under various conditions, including the impact of toxic substances.31,32 RNA-Seq, a modern RNA sequencing technology, allows for precise identification and measurement of gene activity, including those that are weakly expressed.33,34
The purpose of this study was to evaluate the mechanisms of Cd toxicity in NHEK. A comprehensive transcriptomic analysis was conducted to detect differentially expressed genes between control and Cd-treated NHEK. Donor-to-donor variability was levelled using cells from three independent donors.
Materials and methods
Chemicals and reagents
Cadmium chloride was purchased from Sigma-Aldrich (USA). Keratinocyte serum free medium (KSFM), GlutaMAXTM and penicillin/streptomycin solution were purchased from Gibco (USA). Dulbecco's phosphate-buffered saline (DPBS) and 0.25% trypsin-EDTA solution were from Capricorn Scientific (Germany). Annexin V-FITC/propidium iodide (PI) apoptosis it and the Cycle Assay Kit were from Elabscience (USA). MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide) reagent was from Dia-M (Russia). RNA isolation kit was from QIAGEN (USA). Dimethyl sulfoxide (DMSO) was purchased from Helicon (Russia). IL-1, IL-6 and IL-8 Enzyme-linked immunosorbent assay (ELISA) kits were from Vector-Best (Russia). Prostaglandin E2 (PGE2) ELISA was from Cayman Chemical (USA).
Cell culture and treatment
NHEK were purchased from Perspectiva Research-and-Production Company (Russia). NHEK were cultured in full KSFM (1x) supplemented with 1% GlutaMAXTM and penicillin/streptomycin (100 UI/mL and 100 μg/mL). The cells were cultured in 25 or 75 cm 2 tissue treated flasks (Corning, USA) in a humidified atmosphere containing 5% CO 2 at 37°C. NHEK were passaged upon reaching the confluence of approximately 50%. The medium was replaced with fresh every other day. For all experiments, the cells were harvested by trypsinization, seeded in the appropriate culture plates and grown at 37°C under humidified atmosphere containing 5% CO 2 . For all experimental procedures pooled NHEK obtained from 3 donors were used in order to level donor-to donor variability. Cadmium chloride was dissolved to the desired concentration in regular culture medium. The cell morphology was observed by a Zeiss Primovert microscope (ZEISS, Germany).
MTT assay
For MTT assay the cells were seeded in 96-well tissue-treated plates (Corning, USA) at density of 5.0 × 103 cells/well 24 h prior to cadmium treatment. The day after seeding the culture medium was replaced with medium containing cadmium. The cells were incubated for 24 h as described at 4. 2. After exposition, the culture medium was replaced with fresh containing MTT (1 µg/mL) following a 3 h incubation under culture conditions. Then, the medium was discarded and the cells were dissolved in DMSO. The absorbance was measured via iMark microplate reader (Bio-Rad, USA) at 590 nm.
Annexin V-FITC/PI apoptosis assay and cell cycle analysis
Cell cycle analysis and annexin V-FITC/PI kits were employed to detect apoptosis and cell cycle distribution. The cells were seeded in 60 mm Petri dishes at a density of 5.0 ×105 cells. On the following day, the medium was replaced with containing cadmium or with fresh medium in control groups. After 24 h of exposure the cells were harvested by trypsinization, rinsed twice with DPBS solution and stained according to the manufacturer’s instructions. After staining, the cells were immediately analyzed via a ZE5 flow cytometer (Bio-Rad, USA). The raw data were analyzed using Floreada.io software (https://floreada.io/analysis).
Transcriptomic analysis
For transcriptomic analysis NHEK cells were seeded onto 100 mm Petri dishes (Corning, USA) at a density of 1.0 × 106 cells per plate. The cells were cultured for 24 h, then the culture medium was replaced with containing cadmium (25 µM). After 24 h, the cells were collected by trypsinization following RNA isolation. Total RNA was isolated according to the manufacturer’s protocol and quantified using NanoDrop-1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). Transcriptome data were obtained by high throughput paired-end sequencing using Illumina NovaSeq 6000 with a read length equal to 100 bp. The TruSeq Stranded mRNA Library Prep Kit was used to prepare RNA libraries. All steps were carried out in accordance with the manufacturer’s protocol. Transcriptome profiling was performed in three replicates with a separate process of RNA extraction for each replicate. Raw sequencing data were uploaded to the NCBI SRA (https://www.ncbi.nlm.nih.gov, accessed on 14 August 2023). The accession number is PRJNA1005459 for both control and treatment cells.
ELISA
NHEK were seeded in 12-well culture plates at a density of 5.0 × 104 cells/well and incubated under culture conditions. After 24 h, the culture medium was replaced with containing cadmium chloride (25 µM). The cells were treated with Cd for 24 h, after which the conditioned medium was collected for cytokine detection. Commercially available ELISA kits were used to measure the levels of IL-1, IL-6, IL-8, and PGE2 in the culture medium. The assays were performed according to the manufacturer’s protocol. Absorbance was measured using an iMark microplate reader (Bio-Rad, USA) at 450 nm.
Bioinformatic analysis
The R software environment was used for computations and visualization transcriptomic data (ver. 4.1) (R: The R Project for Statistical Computing https://www.r-project.org/). We used DESeq2 package 35 for differential expression analysis. For visualization we used EnhancedVolcano for volcano plot (https://github.com/kevinblighe/EnhancedVolcano, accessed on 25 July 2023) and ClusterProfiler for geneset enrichment analysis 36 as described in Poverennaya et al. 37 A source of biological knowledge, we utilized Wikipathways 38 and PROGENy databases. 39
Statistical analysis
All experiments were performed in at least 3 biological replicates. The graphs represent data as means ± SEM. Statistical analysis was carried out using one-way ANOVA with Tukey’s post-test. The analysis was performed using GraphPad Prism Version 8.0 software (GraphPad Software LLC, CA, USA). A p value of <0.05 was regarded as statistically significant.
Results
Evaluation of cadmium-induced cytotoxicity
The cytotoxic effects of cadmium chloride at different concentrations were evaluated using the MTT assay. The viability of pooled NHEK cells was analyzed after 24 h exposition to cadmium chloride at concentrations from 12.5 to 200 µM. A dose-dependent decrease in cell viability was observed within the selected range of concentrations. Significant differences compared to control were observed at concentrations of 25 µM and above (Figure 1(a)). Exposure to higher Cd concentrations (50–200 µM) resulted in severe cytotoxicity and drastically decreased cell viability. Treatment with Cd resulted in changes in cell morphology. No significant changes were observed after exposure to Cd at concentration of 12.5 µM. after treatment with 25 µM Cd, no visible changes in cell morphology were observed; however, a small number of floating cells appeared in the medium. Treatment with 50 µM Cd increased the ratio of floating cells compared to that in the control group, and some cells developed an irregular shape. Cytotoxicity of cadmium chloride in pooled NHEK after 24 h exposition at different concentrations (a) – MTT assay. Bars represent mean ± SEM. Asterisks indicate significant differences compared with the control group, **p < .01, ***p < .001 (b) – Morphology of NHEK after treatment with Cd at concentrations of 12.5–50 µM (scale bar = 50 µM).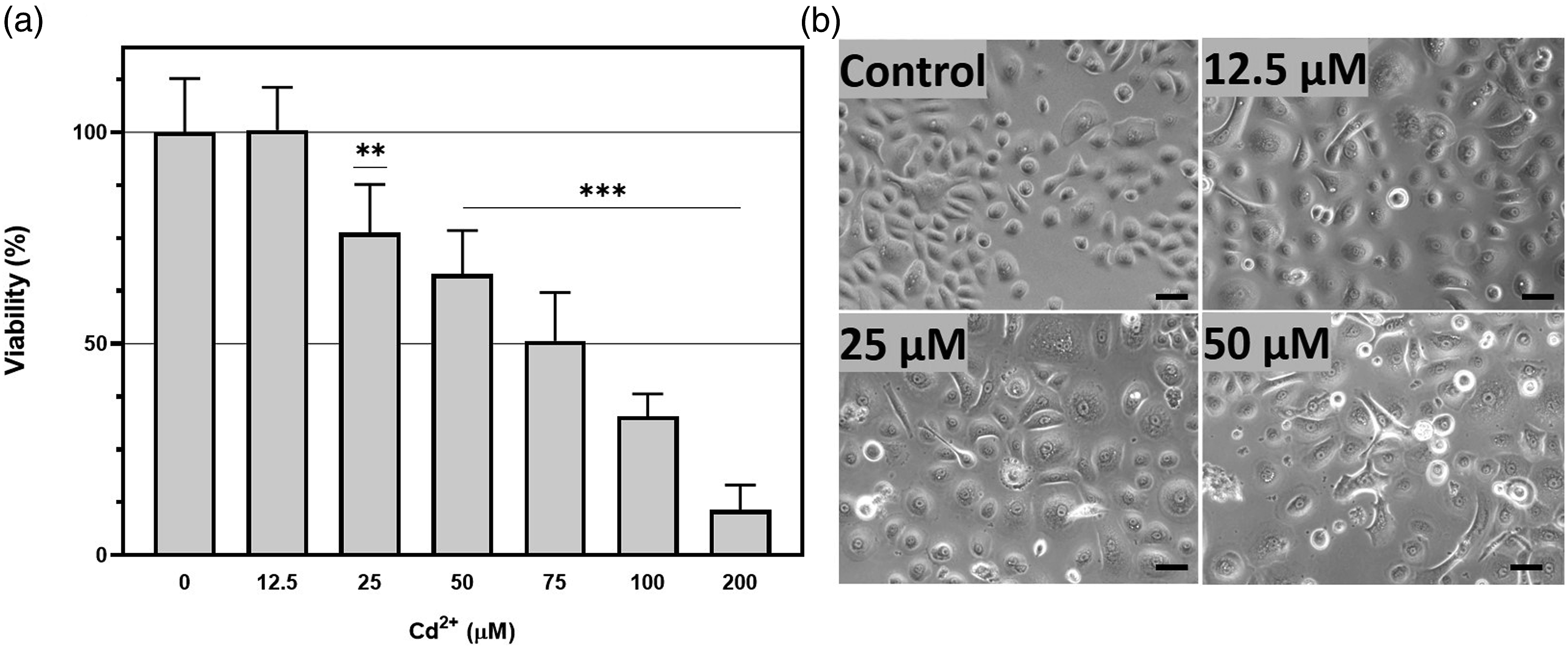
Exposure to cadmium chloride at a concentration of 25 µM resulted in moderate cytotoxicity; thus, this concentration was selected for further experiments, including transcriptomic analysis.
Apoptosis and cell cycle assays
The cytotoxicity of cadmium chloride at the selected concentration (25 µM) was further evaluated using the Annexin V-FITC/PI assay (Figure 2(a)). After 24 h exposition, the ratio of early apoptotic (EA, Annexin V-positive, Q3) cells increased from 0.68 to 4.41%, whereas the ratio of late apoptotic cells (LA, double-positive, Q2) increased to 5.85%. The overall ratio of apoptotic cells (EA + LA) was 10.84 ± 1.19%. In line with the MTT assay results, the ratio of viable cells (unstained, Q3) decreased to 89.59%. The cell cycle distribution analysis revealed that exposure to cadmium increased the ratio of cells in the G1 phase from 41.67 ± 1.15% to 55.5 ± 0.53%. In the S phase, the differences between the control and cadmium-treated groups were not significant. The proportion of cells in G2 decreased from 22.86 ± 1.17% to 14.35 ± 2.71% (Figure 2(b)). The results showed that exposure to cadmium chloride at the selected concentration resulted in cell cycle arrest at the G0/G1 phase. Evaluation of apoptosis and cell cycle analysis after 24 h exposition with 25 µM of cadmium chloride. (a) Annexin V-FITC/Propidium iodide assay. The statistical analysis graph represents cells in early apoptosis and late apoptosis/necrosis (Q2 + Q4) (b) Flow cytometry results of cell cycle analysis. Asterisks indicate significant differences between groups. *p < .05, ***p < .001.
Transcriptomic differences
In the result of RNAseq (Illumina) analysis 13257 and 13817 genes were identified in control and Cd-treated cells, respectively (TPM >0). Transcriptome profiling was employed to assess gene expression differences between the control cells and those exposed to Cd. Analysis of the differentially expressed genes revealed 1463 genes with significant alterations. Specifically, 547 genes were downregulated, while 916 genes were upregulated upon cadmium treatment (Figure 3(a)). To gain deeper insights into the impact of Cd on cellular pathways, we conducted gene set enrichment analysis using the list of differentially expressed genes (Figure 3(B), Supplementary Table SN). Remarkably, following Cd exposure, the number of activated genes was nearly twice as high as the number of suppressed genes. Genes with differential expression were sorted in decreasing order of the range of change. Transcriptomic profiling of intact and Cd-treated NHEK (a) Volcano plot showing differentially expressed genes between control and Cd-treated cells. X-axis: log2 fold change in gene expression. Y-axis: -log10 of the p-value, emphasizing statistical significance of gene expression changes. (b) Gene Set Enrichment Analysis of differentially expressed genes between control and Cd-treated cells. FDR—false discovery rate. The geneRatio value is the proportion of genes in a specific function/pathway from the total differentially expressed genes.
In response to cadmium exposure, the most pronounced activation was observed in eight metallothioneins (MT) genes: MT1E (log2FC = 4,84, FDR <0,0001), MT1F (log2FC = 7,79, FDR <0,0001), MT1G (log2FC = 12,26, FDR <0,0001), MT1H (log2FC = 14,11, FDR <0,0001), MT1L (log2FC = 7,79, FDR <0,0001), MT1X (log2FC = 5,85, FDR <0,0001), MT2A (log2FC = 1,81, FDR <0,0001) and MT1M (log2FC = 5,85, FDR <0,0001). MT genes encode proteins that form metal-thiolate bonds with cysteine residues, facilitating metal ion chelation. 40 Additionally, in cadmium-treated cells, there was an increase in the expression of the SLC30A1 gene (log2FC = 3,1, FDR <0,0001), which is involved in zinc transport. 41
As a result of treatment with cadmium chloride, there was a significant activation of the HMOX1 gene (log2FC = 8,71, FDR<0,0001). The induction of this gene represents an adaptive response that protects cells from oxidative damage during stress. 42 The expression of the PPP1R15 A gene (log2FC = 4,69, FDR <0.0001), was also increased, which is activated in response to genotoxic stress and plays an important role in cell recovery. 43 After cadmium exposure, the expression of certain cytokines was upregulated, such as IL1RL1 (log2FC = 2,92, FDR = 0.01), IL13RA2 (log2FC = 3,36, FDR <0.0001), IL36G (log2FC = 2,6, FDR = 0.04), IL6R (log2FC = 1,42, FDR = 0.003), IL7R (log2FC = 3,02, FDR = 0.01), IL20RB (log2FC = 1,78, FDR <0.0001), IL32 (log2FC = 2,52, FDR = 0.0002), IL11 (log2FC = 3,65, FDR <0.0001) and PTGS2 (log2FC = 1,97, FDR <0.0001). Cytokines are signaling molecules that play a key role in regulating immune response, inflammation, and cell growth and differentiation. 44 Data analysis revealed a decrease in the expression of genes encoding laminin-332 (LM) subunits: LAMC2 (log2FC = −1,29, FDR <0.0001), LAMB1 (log2FC = −2,29, FDR <0.0001) and LAMA3 (log2FC = −1,31, FDR <0.0001). LMs play an important structural role in maintaining epithelial adhesion and also have signaling roles in proliferation processes. 45 Furthermore, the expression of six genes from the "Minichromosome Maintenance” (MCM) family was reduced: MCM10 (log2FC = −1,87, FDR = 0.006), MCM6 (log2FC = −1,51, FDR <0.0001), MCM4 (log2FC = −1,77, FDR <0.0001), MCM3 (log2FC = −1,65, FDR<0.0001), MCM7 (log2FC = −1,63, FDR<0.0001) and MCMBP (log2FC = −1,41, FDR = 0.008). MCM proteins play a role in DNA replication and are involved in assembling and activating the replication fork during the S-phase of the cell cycle. 46 Genes MCM3, MCM4, and MCM7 are part of the eukaryotic MCM complex, which functions as a DNA helicase necessary for genome DNA replication. 46 Additionally, the expression of the FGFBP1 (log2FC = −2,75, FDR <0,0001) was notably reduced. The encoded protein plays a role in cell proliferation, differentiation, and migration by forming complexes with fibroblast growth factors, enhancing their biological activity on target cells. Importantly, the protein enhances the transmission of FGF2 signals during tissue regeneration, angiogenesis, and tumor growth. 47
During the enrichment analysis, the suppression of five DNA-related pathways was detected: «DNA repair pathways full network» (FDR = 1.22×
We also identified the activation of pathways associated with responses to oxidative stress and toxic effects, such as the «NRF2 pathway» (FDR = 0.0008), «Glucocorticoid receptor pathway» (FDR = 0.002), and «Nuclear receptors metapathway» (FDR = 2.46 × 10−5). Overall, the activation of these pathways following cadmium treatment indicates the development of an adaptive response. These pathways contribute to cell survival, reduce cellular damage, and restore cellular homeostasis.
For assessing intracellular signaling activity, we employed Pathway RespOnsive GENes (PROGENy)
39
— a model of consensus pathway signatures—to calculate a relative activity score for 14 distinct signaling pathways (Figure 4). Signal pathways activity of differentially expressed genes between control and Cd-treated cells using PROGENy database. NES - normalized enrichment score.
The «Hypoxia» pathway (NES = 14.09) is the most active pathway in the studied perturbation. In this pathway, the gene NDRG1 (log2FC = 2,97, FDR <0.0001) contributes significantly, being activated in cells treated with cadmium. NDRG1 is a stress-responsive protein whose expression is triggered by various stimuli, including hypoxia, oxidative stress, and exposure to heavy metals. In response to these stimuli, NDRG1 plays a role in regulating cellular proliferation, apoptosis, and angiogenesis. 48 Activation of the «p53» signaling pathway (NES = 4.23) was also notable, with the gene CDKN1A (log2FC = 1.18, FDR <0.0001) making a substantial contribution. The expression of the JAK-STAT signaling pathway was reduced after cadmium exposure (NES = −6.73). Within this pathway, the greatest contribution comes from genes of the Interferon-Inducible (IFI) family, such as IFIT1 (log2FC = −3,34, FDR = 0.002), IFIT3 (log2FC = −1,83, FDR = 0.001), IFITM1 (log2FC = −1,01, FDR <0.0001) and IFI44 (log2FC = −2,53, FDR <0.0001). These genes play a role in regulating responses to various pathogens and cellular stress.
Suppression of the MAPK signaling pathway (NES = −2.21) was also observed. The expression of the negative regulator of this pathway, the DUSP6 gene, was increased under the influence of cadmium (log2FC = −1.49, FDR <0.0001). This gene plays a role in controlling the duration and intensity of MAPK signal transmission.
Cadmium induces pro-inflammatory cytokines release in NHEK
In order to verify the results of transcriptomic analysis, we analyzed the levels of several pro- and anti-inflammatory cytokines in the NHEK condition medium after exposition with cadmium chloride at concentration of 25 µM using ELISA. The obtained results revealed that IL-1, IL-6, IL-10 and PGE2 were upregulated after cadmium treatment (Figure 5). Detection of cytokines in NHEK condition medium after 24 h exposition with subtoxic concentration of cadmium chloride (25 µM) via ELISA. *p < .5 compared to control group.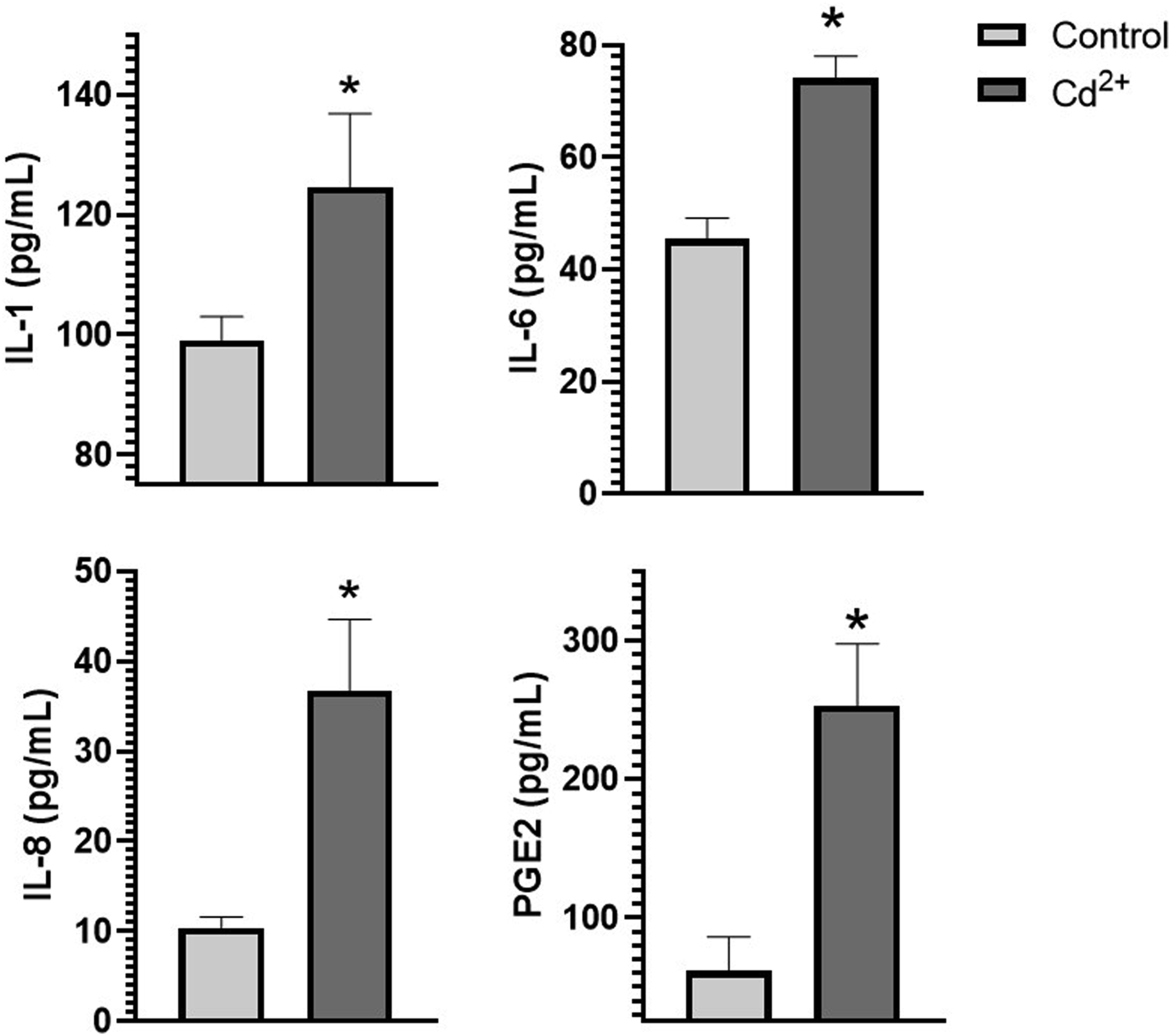
Discussion
This study aimed to comprehensively evaluate cytotoxicity in primary human keratinocytes after 24 h exposition to cadmium chloride with particular emphasis on transcriptomic analysis. First, the MTT assay was conducted to evaluate the toxicity of cadmium chloride in various concentration, which revealed a dose-dependent decrease in NHEK viability (Figure 1). The decrease of cell viability was statistically significant after exposure to Cd at concentrations of 25 µM and above. Based on the MTT assay, the concentration of 25 µM was selected for further experiments, including transcriptomic analysis. Annexin V/propidium iodide assay revealed, that Cd induces apoptosis in NHEK cells. In addition, cell cycle analysis revealed that exposure to Cd induced changes in cell cycle distribution, similar to the results obtained from HaCaT cell.20,49
Cadmium can substitute other metals like zinc, copper and magnesium in the metal-binding sites of metallothioneins (MTs), resulting in their dysfunction. 50 This, in turn, may result in alterations in the homeostasis of metals, carrying out the essential biological functions.50,51 Transcriptomic data revealed, that exposure to Cd causes significant alterations in zinc and copper homeostasis in NHEK. Also, exposition with cadmium results in significant upregulation of several MTs genes, as it has earlier been shown on different cells.52,53 Upregulation of MTs is a protective mechanism against metal-induced cytotoxicity.40,52,54 Cd has a high affinity to SH-groups in MTs, hence binding with MTs reduces cadmium toxicity.55,56
Cd cytotoxicity is heavily expressed through ROS formation and oxidative stress, reflected in our transcriptomic data.22,49,57–59 Enrichment analysis revealed that the Nrf2 pathway, known as the oxidative stress sensor, was activated after cadmium treatment. 42 Also, many of the Nrf2-downstream genes, associated with response to oxidative stress, were upregulated after exposure to Cd, including NQO1, NQO2, HMOX1, AKR1C1, AKR1C2, AKR1C3, SOD3).42,60–63 ROS can also induce various cytokines in keratinocytes, confirmed by transcriptomic analysis and ELISA.64,65 A recent study has shown that Cathepsin D preproprotein (CTSD), a protein responsible for intracellular protein degradation, was upregulated in HaCaT cells after treatment with cadmium. 66 According to our results, this gene was also upregulated in NHEK exposed to cadmium. Remarkably, two more autophagy-related genes (ATG9 and ATG12) were found to be upregulated after Cd2+ treatment.66,67
It is known that keratinocytes are a major source of cytokines in the epidermis. Pro-inflammatory cytokines are actively produced in response to damage and inflammation of the epidermis. Conversely, anti-inflammatory cytokines and growth factors are important regulators of the regenerative phase, exerting pleiotropic effects and influencing dermal cells (fibroblasts, macrophages). In the analysis of transcriptomic data from Cd-treated NHEK, elevated expression levels of several cytokine genes and/or their receptors were identified. These findings were further validated by determining the levels of individual cytokines in the culture medium (Figure 5). Our results align with those obtained in a rat model, which demonstrated that cadmium induces the expression of certain cytokines in epidermal keratinocytes. 15
These observations ensure coordinated behavior of both epidermal and dermal cells during the repair of damage of various origins. In our study, notably, an increase in FGFBP1 expression was observed, which enhances the transmission of the FGF2 signal—a fibroblast growth factor that exemplifies a pleiotropic cytokine of skin cells. Enhanced cytokine production by epidermal cells is also characteristic of several common chronic skin conditions, such as psoriasis, atopic dermatitis, and other diseases. In this context, the additional cytokine release (induced by cadmium) may have significant pathogenetic implications, influencing the severity of clinical manifestations, recurrence frequency, and resistance to therapy.
Our results indicate that cadmium triggers the activation of keratinocytes in response to damage. Indeed, under the influence of cadmium, the expression of KRT6B, KRT6C, KRT16, and KRT17. These keratins are important in damage regeneration.68–70 The induction of these keratins can be seen as the development of an adaptive response to the toxic effects of cadmium, along with cytokine secretion (Figure 5). The determination of the levels of certain cytokines in the culture medium after cadmium exposure confirms the development of an adaptive response. The expression of interleukin-1 β (IL-1β) is necessary to activate the T-cells producing IL-17. 71 The impact of IL-1β on keratinocytes leads to the activation of NFκB and/or MAPK (including ERK1/2 and p38)-AP1 signaling, along with IL-17, resulting in the activation of the expression of keratins 6/16/17. 72 The secretion of IL-6 by keratinocytes is characteristic of skin injuries and activates proliferation through STAT3 activation. 71 The expression of IL-10 in keratinocytes is associated with damage regeneration and prevention of fibrotic responses. 73 Increased expression of PGE2 is characteristic of atopic dermatitis. 74 Probably, PGE2 expression was altered to control the severity of skin injury and inflammation.
Our findings coincide with the data obtained from immortalized HaCaT keratinocytes, supporting the rationale for choosing HaCaT cells as an epidermal model to study Cd cytotoxicity and bioaccessibility.13,29 Specifically, we confirmed that Cd exposure leads to a decrease in proliferative activity and induction of apoptosis. 20 For the first time, we report of signaling pathways, affected by Cd the most, and adaptive mechanisms, which include induction of MTs, redox-sensitive genes and several cytokines in NHEK. The obtained results may provide a basis for further investigation of Cd cytotoxicity and screening of potential therapeutic targets.
Conclusions
In this study, Cd cytotoxicity in NHEK cells was evaluated. In addition to apoptosis and cell cycle arrest, exposure to Cd results in significant alterations at the transcriptomic level. The most prominent differences were associated with the adaptive response, which included the induction of MTs, several cytokines, and redox-sensitive genes.
Supplemental Material
Supplemental Material - Evaluation of Cd-induced cytotoxicity in primary human keratinocytes
Supplemental Material for Evaluation of Cd-induced cytotoxicity in primary human keratinocytes by Daniil Romashin, Viktoriia Arzumanian, Ekaterina Poverennaya, Alexandra Varshaver, Nataliya Luzgina and Alexander Rusanov in Human & Experimental Toxicology.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was performed within the framework of the Program for Basic Research in the Russian Federation for a long-term period (2021-2030) (No 122022800481-0).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
