Abstract
Keywords
Introduction
Hyperoxia-induced acute lung injury (HALI) is more common in clinical practice, which refers to damage to the lungs secondary to exposure to elevated oxygen partial pressure. 1 Oxygen therapy is one of the most vital methods for the clinical treatment of acute respiratory failure.2,3 Patients with hypoxemia often require inhalation of a high concentration of oxygen to maintain arterial oxygen partial pressure and improve oxygen supply to important organs, such as the brain.4,5 However, continuous inhalation of high concentration of oxygen can lead to damage to the lung tissue structure and functional damage due to the generation of oxidants, including reactive oxygen species (ROS), which can induce HALI, a serious complication of clinical oxygen therapy. 6 Hyperoxia-induced acute lung injury appeared after 24 h of oxygen therapy with FiO2 ≥ 0.75.7–9 Exposure to high concentration of hyperoxia leads to lung injury in experimental animal models and plays a major role in the pathogenesis of diseases such as Bronchopulmonary dysplasia (BPD) and acute respiratory distress syndrome (ARDS) in humans. 9 Oxygen toxicity became obvious with prolonged oxygen administration, and significant HALI was observed after 48 h of oxygen administration. Hyperoxia-induced acute lung injury is characterized by a large number of pro-inflammatory cytokines and pro-apoptotic signals, destruction of the pulmonary ventilation blood barrier, injury of endothelial and epithelial cells, and alveolar edema, which can lead to pulmonary ventilation dysfunction and pulmonary edema. A long period of exposure to hyperoxia may lead to acute inflammatory lung injury and chronic lung disease.10–12 Research on the incidence of HALI and related issues originated in the mid-20th century. Recent studies have indicated that the pathogenesis of HALI is complex, involving ROS/reactive nitrogen species (RNS), inflammation, cell apoptosis, and autophagy. 13 The pathogenesis of HALI is still unclear. Lung-protective ventilation and the appearance of PEEP adjunctive therapy for severe hypoxemia have greatly reduced the risk of HALI in the vast majority of patients requiring mechanical ventilation. 9 It is difficult to detect HALI early clinically, so it is particularly urgent to find possible markers.
There has been an increase in the number of studies focused on the functions of genes involved in HALI, particularly as technologies have developed and become more accessible.14–16 For instance, previous studies have identified Akap1, 17 P2x7, 18 and Socs-1 19 as key genes associated with HALI. Glycoprotein non-metastatic melanoma protein B (Gpnmb) is a protein-coding gene. The protein encoded by this gene is a type I transmembrane protein which has been reported to serve a significant role in tissue repair and angiogenesis.20,21 Glycoprotein non-metastatic melanoma protein B is believed to be involved in the development and progression of various diseases, including amyloidosis 22 and cutaneous melanoma. 23 The signaling pathways associated with GPNMB include PTK6 24 and G-Protein Coupled Receptors. 25 Interestingly, Gpnmb has been reported to be implicated in hyperoxia-induced neonatal lung injury.26,27 However, the potential functions of Gpnmb in HALI are still not fully understood.
Oxidative damage in endothelial cells is proposed to play an important role in endothelial dysfunction. 28 Numerous studies have shown that hydrogen peroxide (H2O2) induced epithelial cell dysfunction reflects a loss of the balance between pro- and anti-oxidant, and pro- and anti-inflammatory. 29 Stimulation of epithelial cells with H2O2 to construct cell models of HALI has been widely used. 30
The aim of the present study was to identify pivotal genes in hyperoxia-induced lung injury (HLI) via integrated bioinformatics analyses and validate the results in HALI via in vitro experiments. The flowchart is presented in Figure 1. Through a series of screening and verification tests, we identified Gpnmb as a pivotal gene in HLI. Functional experiments further demonstrated that Gpnmb silencing could ameliorate HALI in vitro by inhibiting mitochondrial-related apoptosis. This research provides a basis for guiding the development of therapeutics for HALI management. Flowchart used in the present study.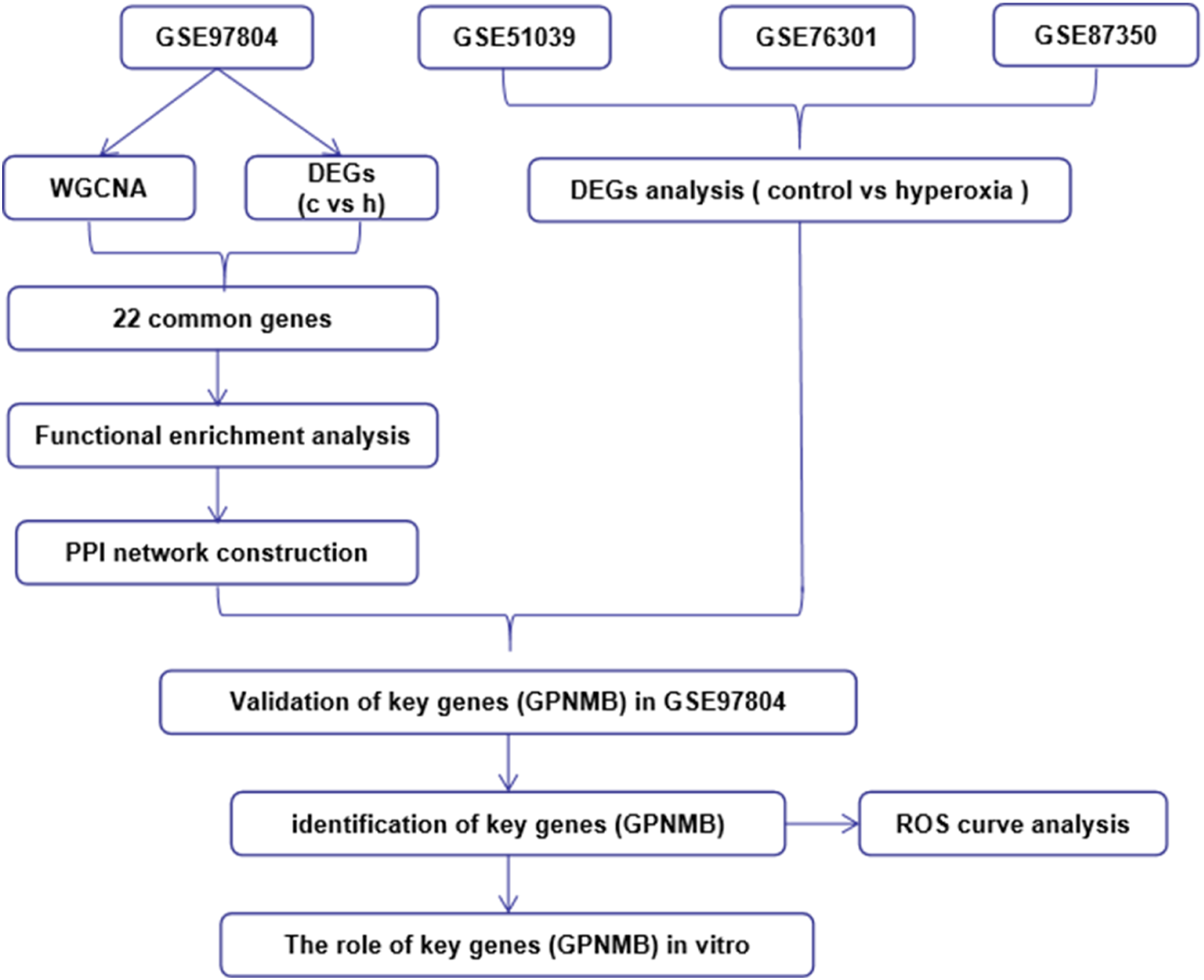
Materials and methods
Data acquisition
GEO datasets information.
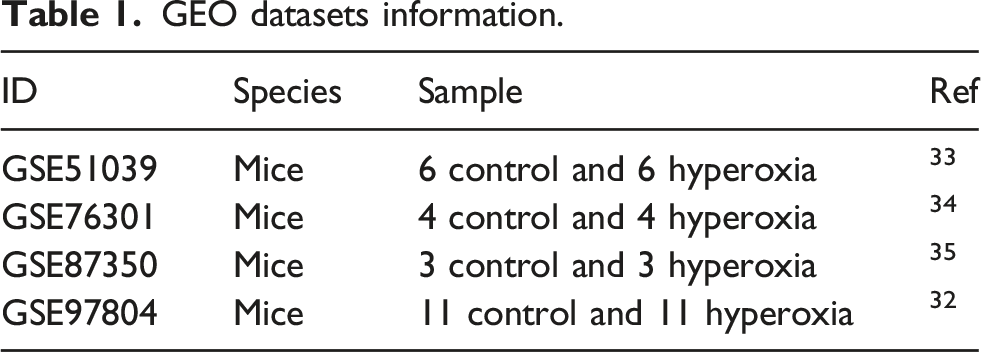
Identification of differentially expressed genes
Background correction, standardization, and expression value calculation were performed on the original dataset GSE97804, GSE51039, GSE76301, and GSE87350 from GEO using package Affy, Impute, and Limma of R software. Fold-change (FC) and adjusted p-values were used to screen DEGs. |log (FC)| ≥ 1 and adj. p-val <.05 were defined as the screening criteria for DEGs. Here adj. p-val <.05 was used in differential gene screening to control the false positive rate. 36 Heat and volcano maps of DEGs were constructed by using Pheatmap, Ggplot2, and other software packages. Sequence data was normalized and analyzed for differences.
Weighted gene co-expression network analysis of GSE97804 dataset
Weighted gene co-expression network analysis was performed according to the method in the literature. 37 The co-expression modules and key genes related to HLI in GSE97804 were established using the WGCNA package (version 3.6.0) in R. 38
Construction of the protein-protein interaction network
The identified genes were imported into the STRING online database (https://string-db.org), 39 and a protein-protein interaction (PPI) network was constructed. Cytoscape v3.6.1 was utilized to screen the top 30 genes in the network described previously. 40
Functional enrichment analysis
To assess the function of DEGs in HLI, Gene Ontology (GO) analysis41,42 and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis 43 was performed using the “clusterProfiler” package in R. 44 p < .05 was set as the cut-off criterion, as well as the p < .05 screening criteria for analysis of results.
Cell culture
The MLE-12 lung epithelial cells line was purchased from the American Type Culture Collection and incubated in DMEM supplemented with 10% FBS in a 5% CO2 humidified incubator at 37°C.
H2O2-stimulated MLE-12 cells
H2O2 has been widely used to induce an oxidative environment in vascular endothelial cell models in vitro. Oxidative injury was induced by exposure to H2O2 (500 μM) for 12 h at 37°C. Cells were plated in six-well plates and grown to confluence before adding H2O2 for up to 12 h.
Cell transfection
The MLE-12 cells were seeded into 6-well plates and divided into control group, H2O2 group, H2O2 + vector group, and H2O2 + siGpnmb group. Plasmids were purchased from Sangon Biotech, Co. Ltd. MLE-12 cells were transfected with the Gpnmb silencing plasmid using Lipofectamine™ 2000 (Invitrogen; Thermo Fisher Scientific, Inc.) according to the manufacturer’s instructions, whereas the control group was transfected with the empty plasmid. After 6 h, the cells were washed and maintained in culture for 12 h for further analysis. First, H2O2 + vector group and H2O2 + siGpnmb group were transfected with the vector or siGpnmb for 12 h, respectively. Then, H2O2 + vector group and H2O2 + siGpnmb group MLE-12 cells were treated with H2O2 for up to 12 h.
Cell Counting Kit-8 assay
Cellular viability was determined using a Cell Counting Kit-8 (CCK-8) assay (Beyotime Institute of Biotechnology). Briefly, 6000–8000 MLE-12 cells/well were plated into a 96-well plate. After treatment as mentioned above, 10 μL CCK-8 solution was added to the cell culture medium, and cells were incubated for 2 h. The absorbance of each well was detected at 450 nm using a Multiskan FC Microplate spectrophotometer (Invitrogen; Thermo Fisher Scientific, Inc.).
Apoptosis analysis
For apoptosis assessment, cultured MLE-12 cells were collected after transfection and rinsed with chilled PBS, followed by incubation with an apoptosis staining detection kit (Invitrogen; Thermo Fisher Scientific, Inc.) in the dark for 15 min. Then, the percentage of apoptotic cells was detected using a flow cytometer (Beckman Coulter, Inc.).
Western blotting
Total protein was extracted from MLE-12 cells using RIPA lysis buffer (Sigma-Aldrich; Merck KGaA). Total protein was quantified using the BCA protein assay kit (Sigma-Aldrich; Merck KGaA). A total of 40 μg protein was loaded per a lane of a 10% SDS-gel, resolved using SDS-PAGE, and transferred onto PVDF membranes (EMD Millipore). The membranes were blocked in 5% non-fat milk at room temperature for 1.5 h. Subsequently, the membranes were incubated overnight at 4°C with primary antibodies targeted against: β-actin (cat. no. ab8226; 1:1000; Abcam), Bcl-2 (cat. no. ab196495; 1:1000; Abcam), and Bcl-2 associated X (BAX) (cat. no. ab32503; 1:1000; Abcam). Subsequently, the membranes were incubated with a horseradish peroxidase-conjugated secondary antibody (cat. no. ab7090; 1:2000; Abcam) at room temperature for 2 h. Finally, the protein bands were observed by chemiluminescence using the ECL Advance Western Blotting Detection kit (Cytiva). Protein expression levels were quantified using ImageJ (version 1.52s; National Institutes of Health).
Intracellular ROS assay
A ROS Assay Kit (Beyotime, China) was applied to detect ROS content in MLE-12 cells. In brief, MLE-12 cells were plated into 6-well plates (1 × 105 cells/well) for 24 h. Then, MLE-12 cells were incubated with a DCFH-DA probe and washed. Finally, the ROS fluorescence was measured via an inverted fluorescence microscope (DMi8, Leica Microsystems, Germany).
JC-1 staining
Laser confocal scanning was used to detect JC-1 probe fluorescence intensity to determine mitochondrial membrane potential. JC-1 is a common fluorescent probe used to detect mitochondrial membrane potential. 45 MLE-12 cells were seeded in 6-well plates at a density of 5 × 104 cells/ml. After the cells had reached 60%–70% confluence, the cells were treated with 500 μM H2O2 for 12 h. The cells were collected and fixed with 4% ice-cold paraformaldehyde at 4°C for 10 min. After washing with PBS, MLE-12 cells were stained with JC-1 dye (Beyotime Institute of Biotechnology). Samples were observed using confocal microscopy (Zeiss, LSM 900, Germany).
Statistical analysis
All data are presented as the mean ± the standard error of the mean. Differences between three or more groups were compared using a one-way ANOVA, followed by a post hoc Tukey’s test. An unpaired Student’s t-test was used to compare the differences between two groups. p < .05 was considered to indicate a statistically significant difference.
Results
Identification of DEGs in GSE51039, GSE76301, GSE87350, and GSE97804 datasets
To investigate DEGs associated with HLI, the GSE51039, GSE76301, GSE87350, and GSE97804 datasets were obtained from GEO databases. As shown in Figure 2 the results showed that 299, 990, 2216, and 62 DEGs were identified by the limma package of R in HLI tissues compared with normal tissues based on |log (FC)|≥1 and adj. p-val <.05 on the GSE51039, GSE76301, GSE87350, and GSE97804 datasets, respectively. Identification of DEGs in GSE51039, GSE76301, GSE87350, and GSE97804 datasets. (a, c, e, and g) Volcano plot of all genes in HALI based on dataset GSE51039, GSE76301, GSE87350, and GSE97804, respectively. (b, d, f, and h) Heatmap plot of all DEGs based on datasets (GSE51039, GSE76301, GSE87350, and GSE97804), respectively.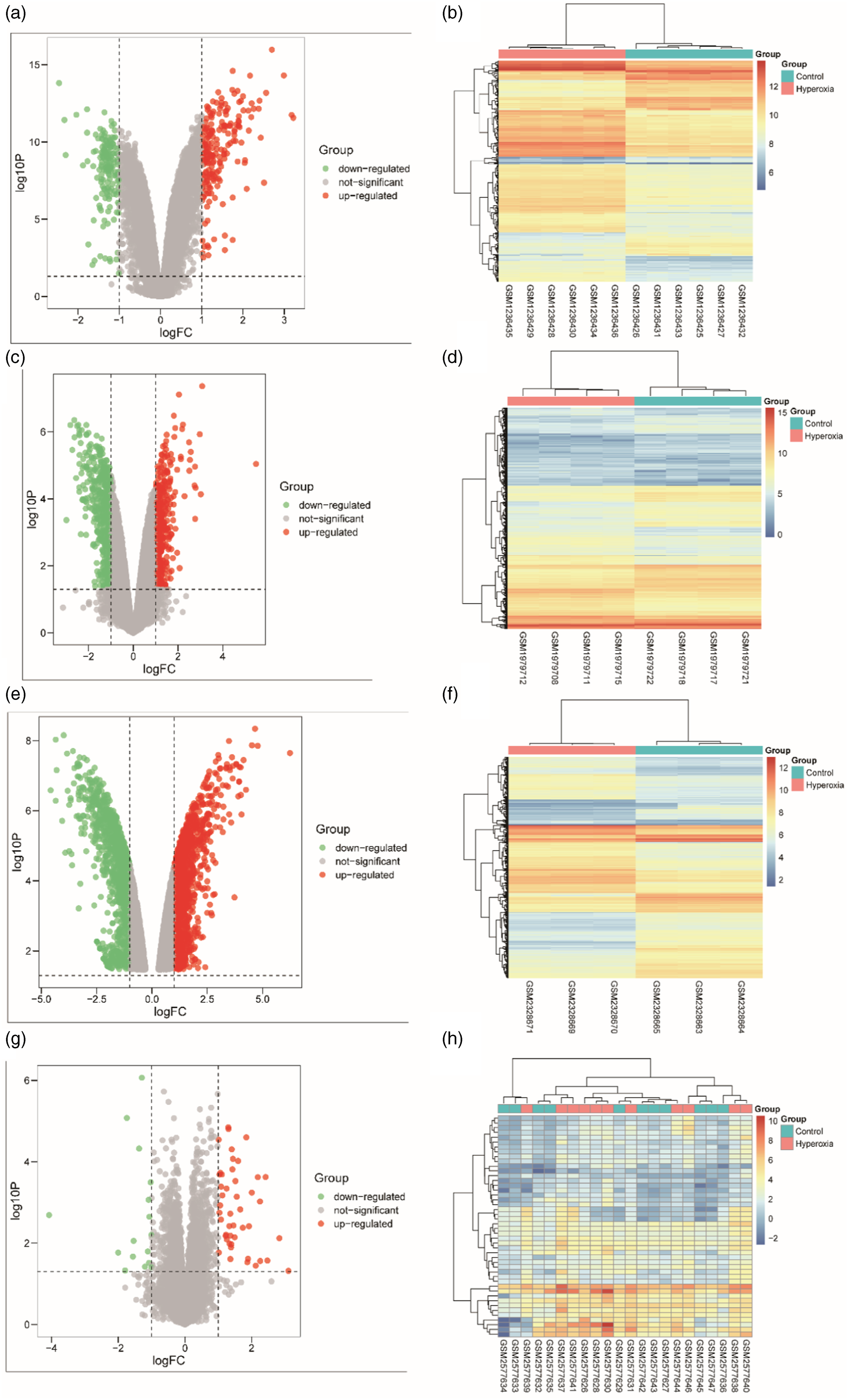
WGCNA of GSE97804 dataset
Next, WGCNA was performed to explore the biological correlation between co-expression modules and HLI. First, sample clustering was used to study the variations and outliers in all the datasets. As shown in Figure 3(a) WGCNA of GSE97804 dataset. (a) Sample clustering was conducted to detect outliers between the HLI and adjacent no tissues; (b) Analysis of the scale-free topology model fit index for soft threshold powers (β); (c) Cluster dendrogram was generated by hierarchical clustering to show the modules of highly interconnected groups of genes between HLI and normal tissues; (d) Interactive relationship analysis of co-expression genes. The light color indicates topological overlap, while the darker color indicates a high topological overlap. (e) The heatmap shows the eigengene adjacency. (f) Heatmap of the correlation between module eigengenes and clinical traits. p-value is shown in each color cell coded by the correlation between modules and traits (red indicates a positive correlation; blue indicates a negative correlation).
GO and KEGG analysis
A total of 22 common genes were identified between the DEGs analysis and the red module of WGCNA based on the GSE97804 dataset. Gene Ontology annotation and KEGG pathway enrichment analyses were used to explore the functions of the identified 22 common genes. As shown in Figure 4(a)–(c), the GO annotation results indicated that the common genes were primarily enriched in cell chemotaxis (biological process), external side of the plasma membrane (cellular component), and receptor-ligand activity (molecular function). Furthermore, the results of KEGG enrichment analysis revealed that common genes were primarily enriched in chemokine signaling pathway and cytokine-cytokine receptor interaction (Figure 4(d)). GO and KEGG pathways analysis of 22 common genes between DEGs and the red module of WGCNA based on GSE97804 dataset. (a) GO-biological processes (BP), (b) GO-cellular components (CC), (c) GO-molecular functions (MF), and (d) KEGG pathway.
Gpnmb was identified as a pivotal gene
To evaluate the potential association among the 22 common genes, a PPI network was constructed. As shown in Figure 5(a), The PPI network contained 16 nodes and 58 PPIs. Gpnmb was identified as one of the 16 nodes, indicating that Gpnmb may play an important role in HLI development. In addition, Gpnmb was also shown as an overlapping gene between GSE97804, GSE51039, GSE76301, GSE87350, and the red module of WGCNA based on GSE97804 dataset by Venn diagram (Figure 5(b)), indicating Gpnmb was a pivotal gene in HLI. Then, the abnormally high expression of Gpnmb in HLI was confirmed in the 4 datasets (Figure 5(c)). The receiver operating characteristic (ROC) curve of Gpnmb was shown in Figure 5(d), in which the area under the curve (AUC) was 0.8264, indicating it could be used as a potential biomarker for HLI diagnosis. Therefore, Gpnmb was identified as a pivotal gene. Gpnmb was identified as a pivotal gene. (a) Protein-protein interaction (PPI) network of 22 common genes. The topological degree was used as the criteria for node size. (b) Common genes between DEGs in GSE97804, GSE51039, GSE76301, GSE87350, and the red module. (c) Relative expression of Gpnmb between hyperoxia and control group. (d) ROC curve analysis of Gpnmb (95% CI: 0.717-0.93, **p < .01). *p < .05; **p < .01.
Gpnmb knockdown suppressed H2O2-induced MLE-12 cell apoptosis
To investigate whether Gpnmb was involved in HALI progression, an in vitro model was established by exposing MLE-12 cells to H2O2 treatment (500 μM) for the indicated time. As shown in Supplementary Figure 1, H2O2 treatment led to a significant reduction in MLE-12 cell viability in a time-dependent manner. As H2O2 treatment (500 μM) for 12 h inhibited MLE-12 cell viability by about 50%, H2O2 treatment (500 μM) for 12 h was selected for subsequent experiments. Flow cytometry results indicated that Gpnmb knockdown suppressed H2O2-induced apoptosis in MLE-12 cells (Figure 6(a)–(b)). CCK-8 results confirmed that Gpnmb knockdown enhanced the viability of H2O2-induced MLE-12 cells (Figure 6(c)). Therefore, Gpnmb might participate in HALI procession. Gpnmb knockdown suppressed H2O2-induced MLE-12 cell apoptosis. (a) Flow cytometry was used to detect the cell apoptosis. (b) The apoptosis rates. (c) The CCK-8 assay was used to detect the cell viability. n = 3, *p < .05, **p < .01, ***p < .001, ns for no significance.
Gpnmb knockdown reduced mitochondria-mediated apoptosis of H2O2-induced MLE-12 cells by regulating apoptosis-related proteins
To investigate whether Gpnmb knockdown reduced H2O2-induced MLE-12 cell apoptosis in a mitochondria-dependent pathway, the protein levels of GPNMB and key indicators of the mitochondrial apoptosis pathway (Bcl-2, BAX, and cleaved-caspase 3) were detected. First, MLE-12 cells were transfected with Gpnmb silencing plasmid, and the successful silencing of Gpnmb in MLE-12 cells was demonstrated by Western blotting (Figure 7(a)–(b)). As shown in Figure 7(c)–(g), H2O2 treatment increased GPNMB, BAX, and cleaved-caspase 3 protein levels and decreased Bcl-2 protein level in MLE-2 cells; however, Gpnmb silencing reversed the effects of H2O2 in MLE-12 cells. Collectively, Gpnmb silencing inhibited H2O2-induced MLE-12 cell apoptosis by regulating mitochondria-mediated cell apoptosis pathway. Gpnmb knockdown reduced mitochondria-mediated apoptosis of H2O2-induced MLE-12 cells by regulating apoptosis-related proteins. (a and b) The expression of Gpnmb after transfection with siGpnmb plasmid. (c–g) Protein levels of GPNMB, BAX, Bcl-2, and cleaved-caspase 3 in each group. n = 3, *p < .05, **p < .01, ***p < .001, ns for no significance.
Gpnmb knockdown inhibited intracellular ROS production and increased mitochondrial membrane potential in H2O2-induced MLE-12 cells
Next, ROS detection result showed that H2O2 increased the ROS generation in MLE-12 cells, and Gpnmb silencing reversed ROS generation in H2O2-induced MLE-12 cells (Figure 8(a)–(b)). Further, JC-1 staining revealed that H2O2 promoted depolarization of the mitochondrial membrane potential compared to the control group; Gpnmb silencing prevented depolarization of the mitochondrial membrane potential (Figure 8(c)–(d)). Therefore, Gpnmb deletion ameliorated H2O2-induced mitochondrial dysfunction in MLE-12 cells. Gpnmb knockdown inhibited intracellular ROS production and increased mitochondrial membrane potential in H2O2-induced MLE-12 cells. (a and b) ROS detection. (c and d) JC-1 staining. n = 3, *p < .05, **p < .01, ***p < .001, ns for no significance.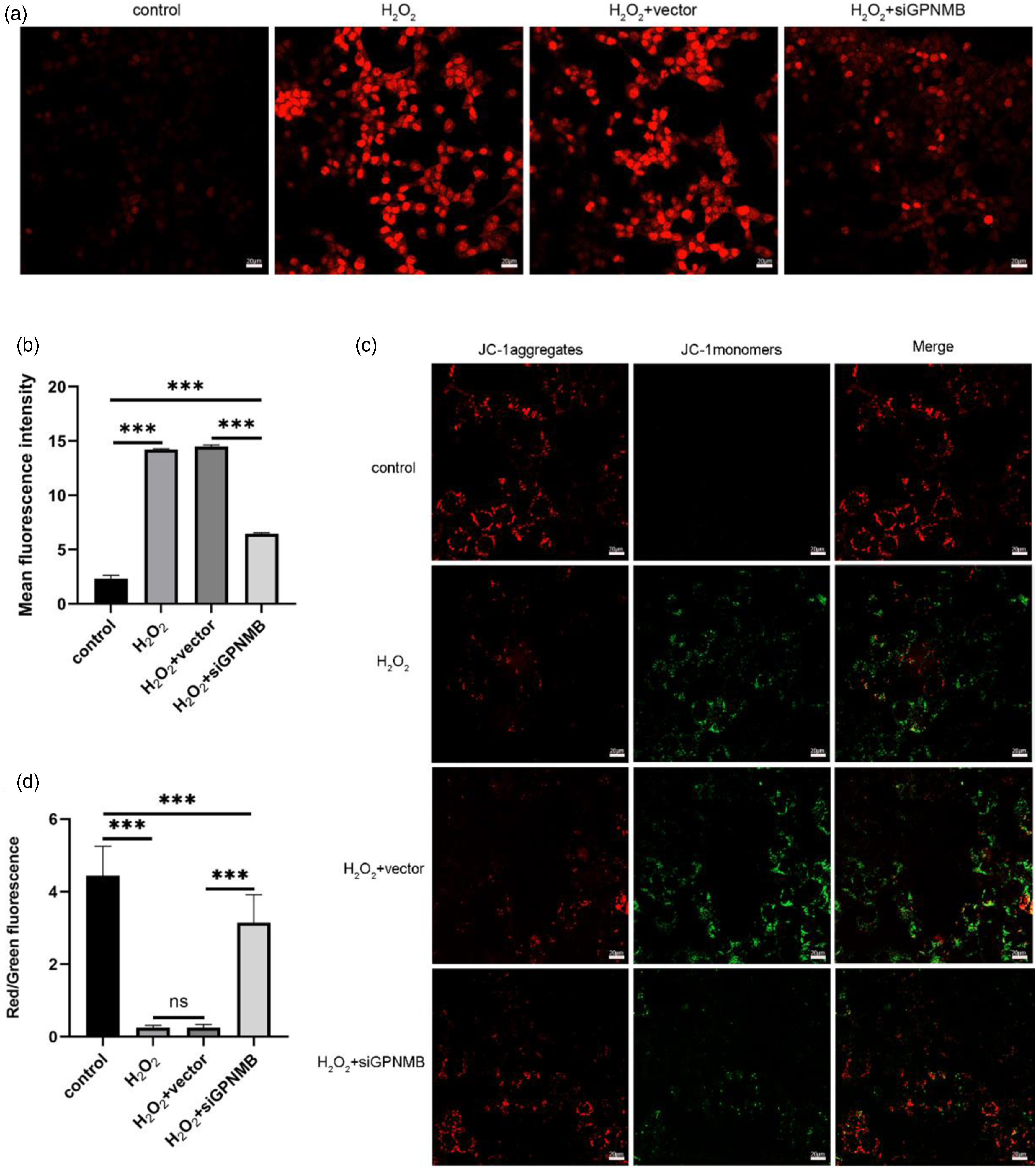
Discussion
Prolonged inhalation of high concentrations of oxygen can cause HALI. In clinical work, lung injury can only be detected by imaging after the disease is significantly aggravated, which seriously affects the treatment of HALI. Therefore, an increasing number of studies have focused on revealing HALI pathogenesis and searching for appropriate diagnostic markers and therapeutic targets.46,47 In this study, we found a relationship between Gpnmb and HALI, indicating that Gpnmb can be used as a new therapeutic target for HALI.
Initially, we analyzed raw data from GSE51039, GSE76301, GSE87350, and GSE97804 datasets to obtain DEGs associated with HLI (299, 990, 2216, and 62 DEGs). Then, WGCNA results provided 6 modules (yellow, blue, red, brown, green, and turquoise) based on the GSE97804 dataset. Among them, the red module (120 genes) showed strong positive correlations and was significantly co-expressed, revealing the important functions of these genes in HLI. Thereafter, GO annotation and KEGG pathway enrichment analysis were performed based on 22 common genes between the DEGs and the red module based on GSE97804 dataset. The results indicated these common genes were mainly associated with cell chemotaxis, the external side of the plasma membrane, receptor-ligand activity, chemokine signaling pathway, and cytokine-cytokine receptor interactions, suggesting that these genes played an important role in HLI. Next, Gpnmb was identified as an overlapping gene between DEGs of GSE97804, GSE51039, GSE76301, and GSE87350 datasets and the red module of WGCNA based on GSE97804 dataset. Then, abnormally high expression of Gpnmb in HLI was confirmed based on the 4 datasets. Also, the AUC of the Gpnmb ROC curve was 0.8264, indicating an acceptable diagnostic efficacy of Gpnmb in HLI. Therefore, Gpnmb might be a critical gene in HLI.
Glycoprotein non-metastatic melanoma protein B is a transmembrane glycoprotein expressed in several types of cells and may function as an adhesion molecule, or soluble ligand. 48 Glycoprotein non-metastatic melanoma protein B can regulate diverse cellular processes, including cell proliferation, apoptosis, and differentiation.49,50 Previous studies also demonstrated that Gpnmb plays an important role in hyperoxia-induced neonatal lung injury.26,27 Therefore, we speculated that Gpnmb may be also involved in HALI progression. Our experimental results confirmed that Gpnmb protein level was upregulated in H2O2-treated MLE-12 cells. Besides, Gpnmb silencing increased cell viability and decreased apoptosis in MLE-12 cells. This finding is consistent with a previous study of Parkinson’s disease in which Gpnmb is reported to inhibit cell proliferation and promote cell apoptosis. 51
Apoptosis is often accompanied by the destabilization of the mitochondrial membrane potential, which is widely considered to be one of the earliest events in the process of apoptosis. 52 The increase of ROS is accompanied by the decrease of mitochondrial membrane potential and the opening of mitochondrial membrane permeability conversion pore, making mitochondria the main executor of cell apoptosis signal. 53 Abnormally high levels of ROS lead to mitochondrial oxidative damage and accelerate mitochondria-dependent apoptosis. 54 To investigate whether Gpnmb knockdown reduced H2O2-induced MLE-12 cell apoptosis in a mitochondria-dependent pathway, the protein levels of key indicators of the mitochondrial apoptosis pathway (Bcl-2, BAX, and cleaved-caspase 3) were detected. The results showed that H2O2 treatment increased Bax, and cleaved-caspase 3 protein levels and decreased Bcl-2 protein level in H2O2-treated MLE-2 cells, which was reversed by Gpnmb silencing. In addition, Gpnmb silencing also inhibited the ROS generation and prevented depolarization of the mitochondrial membrane potential. Therefore, Gpnmb silencing may protect against apoptosis via inhibition of mitochondrial-mediated apoptosis.
The present study still has some limitations. In this study, Gpnmb was screened out via in silico analyses based on mice genome datasets. Its involvement in HALI was confirmed via in vitro experiments performed on MLE-12 (mouse pulmonary epithelial cell line) cells. Therefore, these findings cannot be directly extrapolated to humans because of species differences. In the future, these findings will be further verified using in vivo animal experiments and data from clinical patients.
In conclusion, this study identified Gpnmb as a pivotal gene in HLI and demonstrated that Gpnmb silencing exerted protective effects in HALI via inhibition of mitochondrial-mediated apoptosis in vitro (Figure 9), indicating Gpnmb may be a potential target for clinical prediction and treatment of HALI. This study provides a novel basis for guiding research into novel therapeutics for the management of HALI. Graphical abstract.
Supplemental Material
Supplemental Material - Gpnmb silencing protects against hyperoxia-induced acute lung injury by inhibition of mitochondrial-mediated apoptosis
Supplemental Material for Gpnmb silencing protects against hyperoxia-induced acute lung injury by inhibition of mitochondrial-mediated apoptosis by Xiao qin Wang, Song Qin, Yingcong Ren, Banghai Feng, Junya Liu, Kun Yu, Hong Yu, Zhenliang Liao, Hong Mei and Mei Tan in Human & Experimental Toxicology
Footnotes
Author contributions
Song Qin, Yingcong Ren, Banghai Feng, Xiaoqin Wang, Mei Tan and Hong Mei designed the research; Junya Liu, Song Qin, Hong Yu, Zhenliang Liao carried out the research; Kun Yu, Xiaoqin Wang carried out the analyses. Mei Tan and Song Qin wrote the manuscript. All authors reviewed and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (No. 81960362, No. 82160613), Natural Science Foundation of Guizhou Province (No: ZK [2022] general 660, No: ZK [2022] General 661), Science and Technology Program of Guizhou Province, CXTD [2021] 010; Project of Provincial and Ministerial Collaborative Innovation Center [2020] No.39 of the Ministry of Education and Science and Technology; Science and Technology Project of Zunyi City (ZunShiKeHe HZ No. (2021)13).
Data availability statement
All datasets generated for this study are included in the article/supplementary material.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
