Abstract
Background
Nasopharyngeal carcinoma (NPC) is cancer with high mortality and poor prognosis. Circular RNAs (circRNAs) have been identified in a wide variety of cancers. But the functional mechanism of circ_000285 in NPC remains unclear.
Purpose
To decipher the biological function and molecular mechanism of circ_000285 in NPC.
Methods
Quantitative PCR (RT-qPCR) was applied for detecting the expression of circ_0000285, miR-1278, and FNDC3B. Western blot was used to measure the protein levels of Fibronectin type III domain containing 3B (FNDC3B), Bcl2 associated X (Bax), and B cell leukemia/lymphoma 2 (Bcl2). Cell proliferation, migration, and invasion were analyzed by colony formation, 5-ethynyl-2′-deoxyuridine (EdU), and transwell assays. Cell apoptosis was detected by flow cytometry assays. ELISA assay was used to analyze Caspase-3 activity. Bioinformatics was used to predict, and the target relationship between miR-1278 and circ_0000285 or FNDC3B was verified by luciferase reporter assay. Tumor xenograft models were established to examine how circ_0000285 functions during the mediation of NPC tumor growth in vivo.
Results
Increased circ_0000285 and FNDC3B expressions, and a decreased miR-1278 expression were observed in NPC tissues and cell lines. Knockdown of circ_0000285 inhibited NPC cell proliferation, migration, invasion, and while promoting NPC cell apoptosis in vitro. Circ_0000285 knockdown-mediated anti-tumor effects in NPC cells could be largely reversed by silencing of miR-1278 or overexpression of FNDC3B. Circ_0000285 could up-regulate FNDC3B expression by sponging miR-1278 in NPC cells. Knockdown of circ_0000285 could inhibit tumor growth in vivo.
Conclusion
Circ_0000285 upregulates FNDC3B expression by adsorbing miR-1278 to promote NPC development.
Keywords
Introduction
Nasopharyngeal carcinoma (also known as NPC) is a rare tumor originating in the nasopharynx and occurring in the head and neck.1,2 Environment and diet are important to influence factors in the development of NPC.3,4 In the treatment of NPC, the improvement of intensive radiotherapy has significantly improved the survival rate of patients with NPC.5,6 The purpose of this study was to investigate the molecular mechanism of NPC progression and to further improve the prognosis of NPC patients.
As a kind of noncoding RNA, circular RNAs (circRNAs) lack ployA tails and are highly stable and not easily degraded. 7 With the development of scientific research, more and more evidence has been found that there are thousands of endogenous circRNAs in mammalian cells, and play significant roles in a variety of diseases.8–10 In NPCs, multiple circRNAs have been found to participate in the regulation of tumorigenesis and development. For instance, circRNF13 can regulate the development of NPC by mediating SUMO2 expression. 11 Circ_ABCB10 promoted the growth and metastasis of NPC. 12 Here, we aimed to explore the molecular mechanism of circ_0000285 in NPC progression.
MiRNAs are short RNAs with a length of 19–25 nucleotides.13,14 It has been found that a single miRNA can target multiple mRNAs to regulate gene expression. 15 Of course, miRNAs also regulate disease occurrence and development through mRNA.16,17 MiR-653-5p regulates the progression of gastric cancer by targeting TART3. 18 In addition, miR-1278 regulates inflammatory responses by targeting IL-22 and CXCL14. 19 Whereas, there was not yet known which mRNA was targeted by miR-1278 in NPC and its specific mechanism is not yet known, so it is of great interest to us.
Fibronectin type III domain containing 3B (FNDC3B) is a regulator of adipocyte differentiation.20,21 Meanwhile, it is also a member of the FN family and has been found to regulate cell invasion and metastasis in many types of tumors. 22 FNDC3B can participate in the regulation of endoplasmic reticulum stress and adverse prognosis of cervical cancer. 21 It was found that FNDC3B plays a carcinogenic function in liver cancer. 23 However, the specific regulatory mechanism of FNDC3B in nasopharyngeal carcinoma remains unclear.
Currently, increasing studies are focusing on the circRNA-miRNA-mRNA regulatory mechanism that circRNAs might function as competing endogenous RNAs (ceRNAs) for miRNAs to indirectly affect protein synthesis.24,25 Here, our bioinformatics analysis indicates miR-1278 is a potential downstream target of circ_0000285. Therefore, we supposed that circ_0000285 might exert oncogenic functions by modulating miR-1278/FNDC3B. And this research was carried out to verify this hypothesis. Our findings will aid in the identification of a potential biomarker for predicting NPC progression and an underlying therapeutic target for its therapy.
Materials and methods
Patient sample collection
Fifty-one NPC tissues and matched adjacent normal tissues were collected at Xingtai People’s Hospital, and the patients were informed before the operation, and all patients signed informed consent. All experiment was approved by the ethics committee of Xingtai People’s Hospital. After the tissues were removed, it was stored in liquid nitrogen immediately and then transferred to a −80°C refrigerator.
Cell culture and transfection
Human normal nasopharyngeal epithelial cells (NP69) and NPC cell lines (HK1, C666-1, 5–8F, and SUNE-1) were gained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). All NPC cells were maintained routinely in RPMI-1640 medium (Invitrogen, Carlsbad, CA, USA) containing 10% fetal bovine serum (FBS, GIBCO, Carlsbad, CA, USA) at 37°C in a humidified incubator with 5% CO2. Meanwhile, NP69 cells were cultured in keratinocyte medium (Invitrogen, Carlsbad, CA, USA).
Small interfering RNA (siRNA) against circ_0000285 (si-circ_0000285) and negative control (si-NC), miR-1278 and miR-NC, miR-1278 inhibitor (anti-miR-1278) and anti-NC, pcDNA3.0 and pcDNA3.0-FNDC3B were synthesized and purchased from Sangon Biotech (Shanghai, China). According to instructions, Lipofectamine 3000 reagent (Thermo Fisher Scientific, Inc., Waltham, MA, USA) was used for cell transfection.
RNA extraction and real-time quantitative PCR (RT-qPCR)
Primers sequences used for PCR.

Colony formation assay
The transfected NPC cells were cultured for 24 h and prepared into cell suspension. Then, an appropriate amount of cell suspension was inoculated into the 6-well plate for further culture. When visible clones appeared on the culture plate, the culture was terminated. Then, the culture medium in the culture plate was fixed with 4% paraformaldehyde for 15 min. Finally, the colonies were stained with 0.1% crystal violet (Sangon Biotech, Shanghai, China) for 30 min. At last, visible colonies were observed under a microscope (Olympus, Tokyo, Japan) and counted manually.
5-ethynyl-29-deoxyuridine (EdU) assay
The EdU Apollo In Vitro ImagingKit (RiboBio, Guangzhou, China) was applied to analyze cell proliferation. Firstly, the transfected cells were cultured with 10 μL EdU reagent for 24 h, then fixed with 4% paraformaldehyde. The cells were then stained with the Apollo staining solution for 30 min. The cells were then stained with DAPI. The staining solution was observed and photographed under a microscope.
Transwell assay
The upper transwell chambers pre-coated with or without Matrigel (BD Biosciences, San Jose, CA, USA) were used to analyze cell invasion ability or migration ability, respectively. NPC cells suspended in serum-free medium were seeded onto the upper chambers. DMEM medium containing 10% FBS was added to the lower chamber. After 24 h of incubation, the cells were removed from the upper compartment using a cotton swab. Subsequently, cells attached to the lower surface were fixed with paraformaldehyde (Sangon Biotech, Shanghai, China) and stained with 5% crystal violet (Sangon Biotech, Shanghai, China) for 20 min. The cells were photographed under a microscope.
Flow cytometry assay
Propidium iodide (PI)/Annexin V Cell Apoptosis Kit (Invitrogen, Carlsbad, CA, USA) was used to analyze cell apoptosis to according instructions. In short, the transfected NPC cells were cultured for 48 h and stained with FITC-Annexin V and PI. Finally, cell apoptosis was detected and analyzed by a flow cytometer (BD Biosciences, San Jose, CA, USA).
Enzyme-linked immunosorbent assay (ELISA)
Human Active Caspase-3 (Ser29) SimpleStep ELISA® Kit (Abcam, Cambridge, MA, USA) was applied to analyze caspase-3 activity. The transfected NPC cells were cultured for 24 h and then were collected for protein extraction. The wells of the 96-well plate were added with 50 μL standard samples or experimental samples. Then, 50 μL Antibody Cocktail was added to each well to incubate with cells at room temperature for 1 h. 350 μL wash buffer was added to each well three times, and then 100 μL TMB development solution was added to the well for incubation for 10 min. The absorbance at 450 nm was read on the microplate reader (Invitrogen, Carlsbad, CA, USA).
Western blot
Protease inhibitor PMSF lysis buffer (Abcam, Cambridge, MA, USA) was used to extract proteins from cell lines and tissue samples. BCA reagents (Beyotime, Shanghai, China) were used to determine protein concentrations. Proteins were isolated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) gel and transferred onto the polyvinylidene fluoride (PVDF) membrane. After being blocked in 5% skim milk, the membrane was incubated with primary antibodies (Sigma, St Louis, MO, USA) overnight at 4°C. After the membrane was cleaned, the horse radish peroxidase-conjugated secondary antibody (Easybio, Beijing, China) was incubated with the membrane at room temperature for 1 h, and ECL chemiluminescence reagent (Beyotime, Shanghai, China) was used to develop and analyze the protein bands. Primary antibodies include anti-Bax (ab32503; 1:8000; Abcam, Cambridge, MA, USA), anti-Bcl2 (ab32124; 1:5000; Abcam, Cambridge, MA, USA), anti-GAPDH (ab8245; 1:20000; Abcam, Cambridge, MA, USA) and anti-FNDC3B (AV49753; 1:8000; Sigma, St Louis, MO, USA).
Luciferase reporter assay
The interaction between miR-1278 and circ_0000285 or FNDC3B was verified by dual-luciferase reporter assay. The partial sequence of circ_0000285 or FNDC3B 3′ untranslated region (3′UTR), including the wild-type or mutant complementary sequence with miR-1278, was inserted into pmirGLO vector (Promega, Madison, WI, USA), and the constructed reporter plasmids were termed as circ_0000285 WT/MUT and FNDC3B 3′UTR WT/MUT. NPC cells were co-transfected with miR-1278 or miR-NC and the luciferase reporter plasmids for 48 h. The luciferase activities in different transfected groups were measured by a Luciferase Assay System Kit (Promega, Madison, WI, USA).
Animal experiments
6 weeks old female BALB/c nude mice (Vital River Laboratory Animal Technology Co., Ltd., Beijing, China) were selected for this experiment, and the experiment was approved by the animal ethics committee of Xingtai People’s Hospital. C666-1 cell lines transfected with short hairpin RNA (shRNA) targeting NC (sh-NC, Sangon Biotech, Shanghai, China) or circ_0000285 (sh-circ_0000285, Sangon Biotech, Shanghai, China) were prepared into cell suspension. Then the cell suspension (1.5 × 106 cells/mouse) was injected into the mice, and the tumor volume was recorded at 5, 10, 15, 20, 25, and 30 days. The mice were then euthanized and the tumor weight was measured. Then fresh tumor tissue was used for immunohistochemistry (IHC) and expression analysis.
IHC assay
The fresh tumor tissue was sectioned, roasted, dewaxed and rehydrated, and then repaired. The tissues were incubated with the primary antibody (Ki-67, 1:100, Abcam, Cambridge, MA, USA) and secondary antibody. After incubation, the membrane was cleaned and DAB was added for staining. Finally, the film was sealed and a microscopic examination was performed.
Statistical analysis
Student’s t-test or one-way ANOVA was performed for difference analysis. The results were expressed as the mean ± standard deviation (SD). Pearson correlation analysis was used to analyze the expression association. All the assays were repeated three times. p < 0.05 was deemed as a significant difference.
Results
Up-regulation of circ_0000285 was detected in NPC tissues and NPC cell lines
RT-qPCR was performed to measure the expression of circ_0000285, the results showed that circ_0000285 expression was significantly highly expressed in 51 NPC tissues rather than 51 normal tissues (Figure 1(a)). Consistently, the expression of circ_0000285 was increased in NPC cell lines (HK-1, C666-1, 5–8F, and SUNE-1) compared with that in nasopharyngeal epithelial cell line (NP69) (Figure 1(b)). In short, circ_0000285 RNA level was higher in NPC tissues and cell lines. Circ_0000285 expression was up-regulated in NPC tissues and cells. (a) RT-qPCR was performed to analyze circ_0000285 expression in 51 NPC tissues and matched normal tissues. (b) RT-qPCR was performed to analyze circ_0000285 expression in NPC cell lines (HK-1, C666-1, 5–8F, and SUNE-1) and human normal nasopharyngeal epithelial cells NP69. **p < 0.01, ***p < 0.001.
Silencing of circ_0000285 inhibited cell proliferation, migration, invasion, and while promoting cell apoptosis in NPC cell lines
In our study, we found that circ_0000285 expression was increased in NPC cell lines, and was more significantly expressed in C666-1 and 5–8F cell lines. So we chose these two cell lines for the next experiment. At first, RT-qPCR was performed to analyze the transfection efficiency of circ_0000285 siRNA, and the results suggested that the expression of circ_0000285 was reduced in C666-1 and 5–8F cells transfected with si-circ_0000285 (Figure 2(a)). Colony formation assay proved that the number of colonies decreased after the knockdown of circ_0000285 (Figure 2(b)). EdU assay confirmed that down-regulation of circ_0000285 could suppress cell proliferation (Figure 2(c)). Cell migration and invasion of C666-1 and 5–8F cells declined after the knockdown of circ_0000285 (Figures 2(d) and (e)). Flow cytometry assay exposed that circ_0000285 knockdown promoted cell apoptosis (Figure 2(e)). Caspase-3 is the most important terminal shearing enzyme during apoptosis.
26
ELISA assay suggested that caspase-3 activity was augmented after transfection of si-circ_0000285 (Figure 2(g)). Bax and Bcl-2 are promoters and inhibitors of apoptosis, respectively.
27
We found that circ_0000285 knockdown up-regulated the protein level of Bax and reduced the protein level of Bcl2 in NPC cells (Figure 2(h)). All in all, our data strongly meant that knockdown of circ-0000285 could restrain cell migration, migration, and invasion, but facilitate cell apoptosis in NPC cell lines. Knockdown of circ_0000285 could inhibit cell proliferation, migration, invasion, and while promoting cell apoptosis in NPC cells. (a) RT-qPCR was performed to analyze circ_0000285 expression in C666-1 and 5–8F cells after knockdown of circ_0000285. (b) Colony formation assay was conducted to detect the colonies number. (c) EdU assay was conducted to analyze cell proliferation. (d–e) Transwell assay was conducted to analyze cell migration and invasion. (f) Flow cytometry assay was conducted to analyze cell apoptosis. (g) ELISA assay was conducted to detect Caspase-3 activity. (h) Western blot was performed to analyze Bax and Bcl2 protein levels. ***p < 0.001.
Circ_0000285 could act as a sponge of miR-1278
RT-qPCR analysis proved miR-1278 expression was lower in NPC tissues and NPC cell lines (HK-1, C666-1, 5–8F, SUNE-1) compared with their respective controls (Figures 3(a) and (b)). According to starbase prediction, we found the target binding sites between circ_0000285 and miR-1278 (Figure 3(c)). Besides, miR-1278 overexpression could inhibit the luciferase activity of C666-1 and 5–8F cells transfected with circ_0000285 WT, while there was no change in cells transfected with circ_0000285 MUT (Figures 3(d) and (e)). Furthermore, miR-1278 expression was up-regulated after the knockdown of circ_0000285 in C666-1 and 5–8F cells (Figure 3(f)). Ultimately, Pearson correlation coefficient analysis found that there was a negative correlation between circ_0000285 and miR-1278 in 51 NPC tissue samples (Figure 3(g)). To sum up, circ_0000285 could target miR-1278. Circ_0000285 could target miR-1278. (a) RT-qPCR was performed to analyze miR-1278 expression in NPC and normal tissues. (b) RT-qPCR was performed to analyze miR-1278 expression in NPC cell lines (HK-1, C666-1, 5–8F and SUNE-1) and human normal nasopharyngeal epithelial cells NP69. (c) The target binding sites between circ_0000285 and miR-1278 were predicted by Starbase. (d–e) The luciferase activities of C666-1 and 5–8F cells co-transfected with circ_0000285 WT or circ_0000285 MUT and miR-NC or miR-1278 were detected. (f) RT-qPCR was performed to analyze the expression of miR-1278 in C666-1 and 5–8F cells when transfection of si-NC or si-circ_0000285. (g) Pearson correlation coefficient was utilized to analyze the relationship between circ_0000285 and miR-1278 in NPC tissue samples. ***p < 0.001.
Circ_0000285 could regulate the progression of NPC by targeting miR-1278
In order to further study whether circ_0000285 regulates NPC progression through miR-1278, we carried out a series of experiments. The RNA level of miR-1278 was impeded in C666-1 and 5–8F cells with the transfection of anti-miR-1278 (Figure 4(a)). Both colony formation assay and EdU experiments confirmed that anti-miR-1278 could weaken the inhibitory effect of si-circ_0000285 on cell proliferation of C666-1 and 5–8F cells (Figure 4(b)–(e)). Meanwhile, transwell assay analysis demonstrated that the inhibitory function of si-circ_0000285 on cell migration and invasion could be restored with anti-miR-1278 (Figure 4(f)–(i)). Flow cytometry assay analysis meant that anti-miR-1278 could recover the promoting effect of circ_0000285 silence on cell apoptosis (Figure 4(j) and (k)). ELIAS assay and western blot assay strongly meant that the promoting effect of si-circ_0000285 on cell apoptosis was reversed after co-transfection of anti-miR-1278 in C666-1 and circ_0000285 knockdown reduced the malignant progression of NPC cells by up-regulating miR-1278. Downregulation of miR-1278 could restore the effect of circ_0000285 knockdown on the progression of NPC. (a) RT-qPCR was performed to analyze the expression of miR-1278 after transfection of anti-NC or anti-miR-1278 in C666-1 and 5–8F cells. (b–c) Colony formation assay was performed to analyze the number of colonies after transfection of si-NC, si-circ_0000285, si-circ_0000285+anti-NC or si-circ_0000285+anti-miR-1278. (d-e) EdU assay was performed to analyze cell proliferation of C666-1 and 5–8F cells. (f–i) Transwell assay was performed to analyze cell migration and invasion; (j) and (k): Flow cytometry assay was performed to analyze cell apoptosis; L and M: ELISA assay was performed to analyze Caspase-3 activity. (n–o) Western blot was performed to analyze the protein levels of Bax and Bcl2. **p < 0.01, ***p < 0.001.
There was a target relationship between miR-1278 and FNDC3B
Primitively, RT-qPCR and western blot assays presented that FNDC3B mRNA and protein levels were upregulated in NPC tissues and cell lines (HK-1, C666-1, 5–8F, SUNE-1) compared to normal tissues and NP69 cells (Figure 5(a)–(d)). Starbase prediction found that miR-1278 could target FNDC3B (Figure 5(e)). Luciferase reporter assay also confirmed that miR-1278 could target FNDC3B in C666-1 and 5–8F cells (Figure 5(f) and (g)). Moreover, miR-1278 expression was significantly raised after transfection miR-1278 mimic in C666-1 and 5–8F cells (Figure 5(h)). Besides, the protein level of FNDC3B was detected by western blot, and the results disposed of that miR-1278 overexpression down-regulated the protein level of FNDC3B in NPC cells (Figure 5(i)). Beyond that, western blot results displayed that knockdown of circ_0000285 limited FNDC3B protein expression, and anti-miR-1278 could restore the inhibitory influence of si-circ_0000285 on FNDC3B protein level in C666-1 and 5–8F cells (Figure 5(j) and (k)). Pearson correlation coefficient analysis confirmed that there was a negative correlation between miR-1278 and FNDC3B mRNA in 51 NPC tissues (Figure 5(l)). Therefore, miR-1278 could target FNDC3B, and circ_0000285 could regulate FNDC3B expression by targeting miR-1278 in C666-1 and 5–8F cells. MiR-1278 could target FNDC3B. (a) RT-qPCR was performed to analyze the mRNA level of FNDC3B in NPC and normal tissues. (b) Western blot was conducted to measure FNDC3B protein level in NPC and normal tissues. (c–d) RT-qPCR and western blot assays were performed to analyze FNDC3B mRNA and protein levels in NP69, HK-1, C666-1, 5–8F and SUNE-1 cell lines. (e) The target relationship between miR-1278 and FNDC3B was predicted by Starbase. (f–g) The luciferase activities of C666-1 and 5–8F cells after co-transfection of FNDC3B 3′UTR WT or FNDC3B 3′UTR MUT and miR-NC or miR-1278 were detected. (h) RT-qPCR was performed to analyze miR-1278 expression in C666-1 and 5–8F cells after transfection of miR-NC or miR-1278. (i) Western blot was performed to analyze the level of FNDC3B protein in C666-1 and 5–8F cells after transfection of miR-NC or miR-1278. (j–k) Western blot was performed to analyze FNDC3B protein level in C666-1 and 5–8F cells after transfection of si-NC, si-circ_0000285, si-circ_0000285 + anti-NC, or si-circ-0000285 + anti-miR-1278. (l) Pearson correlation coefficient was used to analyze the relationship between miR-1278 and FNDC3B in NPC tissue samples. ***p < 0.001.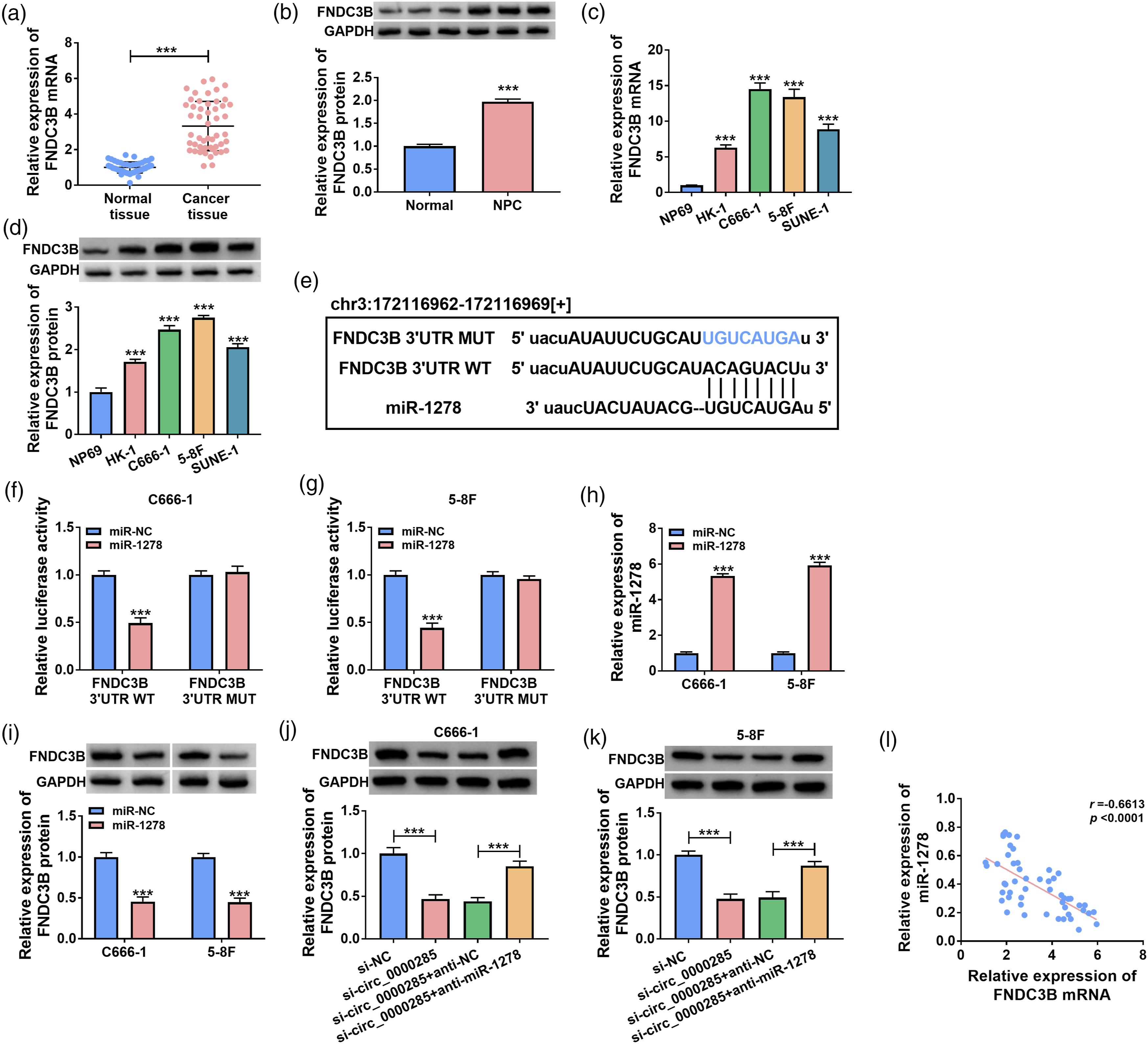
FNDC3B overexpression could reverse the anti-tumor effects of circ_0000285 silencing in NPC cells
Finally, western blot analysis proved that the protein level of FNDC3B was higher after transfection of pcDNA-FNDC3B in C666-1 and 5–8F cells (Figure 6(a)). On this basis, colony formation and EdU assay indicated that overexpression of FNDC3B could overturn the inhibiting effect of si-cric_0000285 on NPC cell proliferation (Figure 6(b)–(e)). Circ_0000285 knockdown-induced suppressive effects on the migration and invasion of NPC cells were largely counteracted by FNDC3B overexpression (Figure 6(f)–(i)). Meanwhile, the results of flow cytometry and ELISA assays showed that circ_0000285 knockdown-induced apoptosis of NPC cells was attenuated by the overexpression of FNDC3B (Figure 6(j)–(m)). Western blot assay showed that circ_0000285 knockdown-induced up-regulation of Bax and down-regulation of Bcl2 were reversed by the overexpression of FNDC3B (Figure 6(n) and (o)). In summary, all data proved that circ_0000285 could regulate the progression of NPC through FNDC3B. Overexpression FNDC3B could recover the function of si-circ_0000285 on NPC progression. (a) Western blot was performed to analyze the protein level of FNDC3B after the transfection of pcDNA3.0 or pcDNA3.0-FNDC3B in C666-1 and 5–8F cells. (b–o) C666-1 and 5–8F cells were transfected with si-NC, si-circ_0000285, si-circ_0000285 + pcDNA3.0, or si-circ_0000285 + pcDNA3.0-FNDC3B. (b–c) Colony formation was performed to analyze colony formation. (d–e) EdU assay was performed to analyze cell proliferation. (f–i) Transwell was performed to analyze cell migration and invasion; (j) and (k): Cell apoptosis was analyzed by flow cytometry. (l–m) Caspase-3 activity was analyzed by ELISA; (n) and (o): Western blot was performed to analyze Bax and Bcl2 protein levels. **p < 0.01, ***p < 0.001.
Silencing of circ_0000285 could prevent the growth of NPC tumor in vivo
To verify the regulatory mechanism of circ_0000285 in vivo, we first constructed the xenograft mice model. In this assay, C666-1 cells with the stable knockdown of circ_0000285 were established, and the C666-1 cell line stably transfected with sh-NC was used as the control. C666-1 cell suspension (1.5 × 106 cells/mouse) was injected into the nude mice. The results showed that the tumor volume was significantly reduced by the knockdown of circ_0000285 (Figure 7(a)). The tumor weights of the mice were weighed after euthanasia, and it was revealed that the low expression of circ_0000285 inhibited the growth of tumor weight (Figure 7(b)). Beyond that, IHC analysis suggested that knockdown of circ_0000285 could restrain Ki-67 expression (Figure 7(c)). Circ_0000285 expression was reduced in tumor tissues in the sh-circ_0000285 group compared with the sh-NC group (Figure 7(d)), suggesting the successful knockdown. Circ_0000285 knockdown up-regulated miR-1278 expression and reduced FNDC3B protein expression in tumor tissues in vivo (Figure 7(e) and (f)), which was consistent with the results in vitro. In a word, knockdown of circ_0000285 could inhibit tumor growth in vivo. Downregulation of circ_0000285 could inhibit the growth of NPC tumor in vivo. (a) The tumor volume was analyzed; (b): The tumor weight was analyzed. (c) IHC was conducted to analyze the expression of Ki-67 in xenograft tumor tissues. (d-e) RT-qPCR was performed to analyze the expression of circ_0000285 and miR-1278 in xenograft tumor tissues. (f) Western blot was performed to analyze the protein of FNDC3B level in xenograft tumor tissues. ***p < 0.001.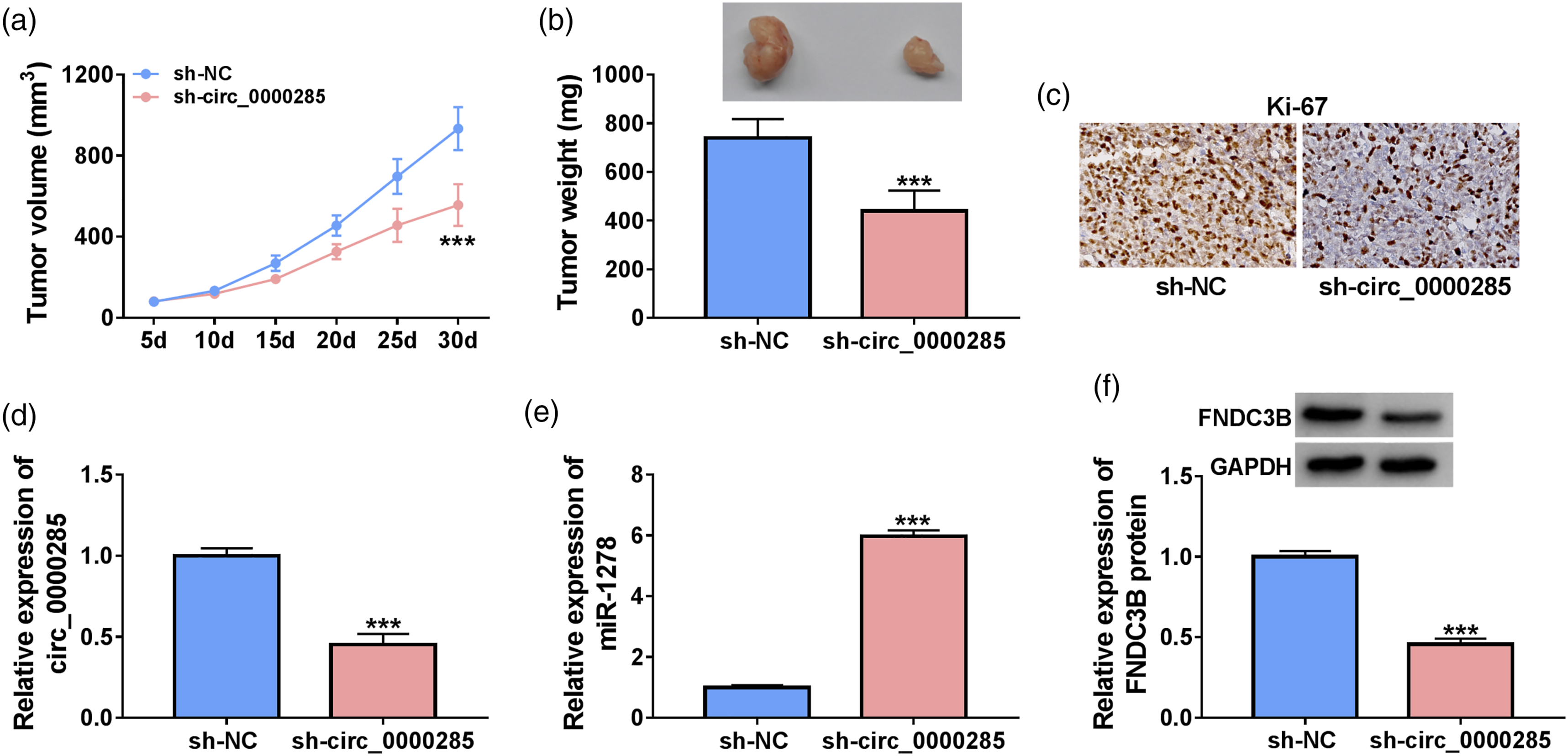
Discussion
NPC is the head and neck cancer with the highest metastatic potential. 28 CircRNAs have been reported to regulate cancer progression by acting as sponges for miRNAs and thereby regulating gene expression.29,30 The mechanism of circRNAs in NPC has also been studied a lot.31,32 In our study, the function and mechanism of circ_0000285 in NPC were investigated. Initially, we analyzed that circ_0000285 expression was significantly upregulated in NPC tissues and cells. In addition, knockdown of circ_0000285 could inhibit NPC cell proliferation, migration and invasion, and promote cell apoptosis. In summary, the above findings confirmed the tumor-promoting role of circ_0000285 in nasopharyngeal carcinoma.
Based on the ceRNA hypothesis, the interaction between circ_0000285 and miR-1278 was predicted by Starbase database. We further verified their targeting relationship by luciferase report experiments. Previous studies have suggested that miR-1278 acted as a tumor suppressor in multiple cancers, including NPC. For instance, Yang et al. found that circ_0003221 facilitated the progression of laryngeal squamous cell carcinoma by sponging miR-1278 and up-regulating YAP1. 33 Du et al. demonstrated that circ_0101675 contributed to the progression of non-small cell lung cancer by acting as a molecular sponge for miR-1278. 34 In NPC, miR-1278 was reported to elevate the cisplatin sensitivity and suppress the autophagy of NPC cells by targeting ATG2B. 35 We found that miR-1278 expression was significantly decreased in NPC tissues and cells. Moreover, circ_0000285 was negatively correlated with miR-1278. Then, miR-1278 inhibitor was co-transfected with si-circ_0000285 to explore cell proliferation and other related processes, and the results showed that circ_0000285 could play a role in the regulation of NPC progression through sponging miR-1278. Our study further investigated the specific regulatory mechanism of circ_0000285 in NPC.
In addition, we further found that miR-1278 can target FNDC3B, and FNDC3B was up-regulated in NPC tissues and cells. Accumulating evidence has identified FNDC3B as an oncogene in a variety of human cancers, including NPC. For example, Li et al. found that FNDC3B facilitated the proliferation and invasion abilities of colorectal cancer cells by targeting PI3K/mTOR signaling. 36 Lin et al. demonstrated that FNDC3B contributed to the metastasis of hepatocellular carcinoma cells. 37 Zhong et al. found that FNDC3B promoted the epithelial-mesenchymal transition of tongue squamous cell carcinoma cells. 38 In terms of NPC, a previous study reported that FNDC3B 3′UTR shortening could escape from miRNA-induced gene silencing and contribute to NPC progression. 39 We found that FNDC3B was negatively regulated by miR-1278 in NPC cells. FNDC3B overexpression reversed circ_0000285 knockdown-mediated anti-tumor effects in NPC cells, suggesting that circ_0000285 knockdown suppressed NPC progression largely by down-regulating FNDC3B. Pearson correlation analysis also found a negative correlation between FNDC3B and miR-1278, and a significant positive correlation between FNDC3B and circ_0000285. We found that circ_0000285 could positively regulate FNDC3B expression by sponging miR-1278 in NPC cells. Xenograft tumor model confirmed that circ_0000285 knockdown suppressed xenograft tumor growth in vivo.
In conclusion, our results validated that circ_0000285 is expected to be a novel diagnostic biomarker and therapeutic target for NPC, and these findings reported that the ceRNA network consisting of circ_0000285, miR-1278, and FNDC3B is a new mechanism partook in NPC development. However, this project is limited by the small sample size, and our future research will try to recruit more respondents.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The authors declare that they have no conflicts of interest.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
