Abstract
Lung adenocarcinoma (LUAD) is a malignant tumor that occurs in the lungs. Numerous reports have substantiated the participation of long non-coding RNAs (lncRNAs) in the tumorigenesis of LUAD. Previously, lncRNA alpha-2-macroglobulin antisense RNA 1 (A2M-AS1) was confirmed to be an important regulator in the biological processes of LUAD and dysregulation of A2M-AS1 was associated with non-small cell lung cancer (NSCLC) progression. However, the precise mechanism of A2M-AS1 in LUAD has not been elucidated. Therefore, our study was designed to investigate the detailed molecular mechanism of A2M-AS1 in LUAD. Herein, the expression of lncRNA A2M-AS1, microRNA (miRNA) miR-587, and bone morphogenetic protein 3 (BMP3) in LUAD cell lines and tissues were detected by real-time quantitative polymerase chain reaction (RT-qPCR) and western blotting. The viability, proliferation, migration and invasion of LUAD cells were tested by cell counting kit-8 (CCK-8), colony formation and Transwell assays. In vivo tumor growth was investigated by xenograft animal experiment. Interactions among A2M-AS1, miR-587 and BMP3 were measured by RNA pulldown and luciferase reporter assays. In this study, A2M-AS1 was downregulated in LUAD tissues and cells and related to poor prognosis in LUAD patients. A2M-AS1 overexpression suppressed LUAD cell proliferation, migration and invasion in vitro and inhibited tumor growth in vivo. Mechanistically, A2M-AS1 directly bound with miR-587 to promote BMP3 expression in LUAD cells. Low expression of BMP3 was found in LUAD tissues and cells and was closely correlated with poor prognosis in LUAD patients. BMP3 deficiency reserved the inhibitory influence of A2M-AS1 overexpression on LUAD cell behaviors. Overall, A2M-AS1 inhibits cell growth and aggressiveness via regulating the miR-587/BMP3 axis in LUAD.
Introduction
Lung adenocarcinoma (LUAD), the primary epithelial lung tumor, mostly originates from alveolar epithelium or bronchial mucosa epithelium. 1 LUAD is the most common subgroup of non-small cell lung cancer (NSCLC) which accounts for about 85% of lung cancers. 2 In 2018, new cases of patients with lung cancers reach 2,093,876, of which about 1,761,007 patients die. 3 Multi-modal therapies, such as radiotherapy, surgery, targeted therapy and chemotherapy, are the primary treatments for LUAD patients. 4 However, due to lack of effective diagnostic tools, most LUAD patients cannot be diagnosed with this cancer timely in the early stage and develop into the middle or even advanced clinical stage.5,6 The 5-years survival rate of LUAD patients is only about 15%. 7 Therefore, the exploration of reliable biomarkers is of great clinical importance to formulate effective treatment strategies and improve the accuracy of diagnosis of LUAD.
Noncoding RNAs (ncRNAs) are regarded as a type of transcripts without the ability to encode any protein, which are implicated in the modulation of various cellular activities. 8 According to sequence length of nucleotides, ncRNAs are classified into short ncRNAs (20–22 nucleotides) and long ncRNAs (lncRNAs) (over 200 nucleotides). 9 LncRNAs can express genes, modulate transcription and modify chromatin. 10 Differential expressed lncRNAs are associated with the progression of multiple carcinomas,9,11,12 including LUAD. 13 For example, lncRNA DPPA2 upstream binding RNA (DUBR) is dysregulated in LUAD and suppresses the mobility of LUAD cells by zinc finger and BTB domain containing 11 (ZBTB11)-mediated restriction of oxidative phosphorylation. 14 LncRNA LINC01006 regulates the miR-129-2-3p/catenin beta 1 (CTNNB1)/Wnt/β-catenin axis to facilitate the epithelial-mesenchymal transition (EMT), migration and proliferation of LUAD cells. 15 Knockdown of lncRNA family with sequence similarity 201 member A (FAM201 A) inhibits cell invasion and proliferation in LUAD via the miR-7515/glyoxalase I (GLO1) axis. 16 Moreover, accumulating studies reveal the participation of lncRNA alpha-2-macroglobulin antisense RNA 1 (A2M-AS1) in several cancers. A2M-AS1 sponges miR-146b to upregulate mucin 19 (MUC19), thus accelerating breast cancer cell proliferative and invasive properties, and suppressesing apoptosis. 17 A2M-AS1 is identified as a prognostic biomarker for pancreatic ductal adenocarcinoma. 18 Interestingly, A2M-AS1 was confirmed to be an important regulator in the biological processes of LUAD 19 and dysregulation of A2M-AS1 was associated with NSCLC progression. 20 However, the precies mechanism of A2M-AS1 in LUAD remains uncharacterized.
MicroRNAs (miRNAs) belong to a class of ncRNAs, which are single-stranded RNAs of 19–25 nucleotides that function by complementary base pairing with target messenger RNAs (mRNAs), degrading the corresponding mRNAs, or inhibiting mRNA translation.21,22 It is widely known that miRNAs are involved in regulating multiple biological processes as well as the development of various human diseases, including cancers. For example, miR-338–3p targets mitogen-activated protein kinase kinase kinase 2 (MAP3K2) to modulate the extracellular regulated protein kinase (ERK)1/2 signaling pathway, thereby suppressing the migration, invasion and proliferation of human LUAD cells. 23 MiR-126-5p represses cell migration and facilitates apoptosis to enhance the radiosensitivity of LUAD cells through modulating the enhancer of zeste two polycomb repressive complex two subunit (EZH2)/Krüppel-like factor 2 (KLF2)/baculoviral IAP repeat containing 5 (BIRC5) axis. 24 Our bioinformatics analysis showed that miR-587 was sponged by A2M-AS1 and targeted by bone morphogenetic protein 3 (BMP3). Notably, miR-587 was previously discovered to be upregulated in NSCLC samples and closely linked to tumor staging, and miR-587 ehnanced the proliferative and migratory capabilities of NSCLC cells by downregulating cylindromatosis (CYLD). 25 BMP3 is a member of the transforming growth factor β (TGF-β) superfamily which plays a pivotal role in embryonic development by inducing early skeletal formation. 26 BMP3 has been reported as a tumor suppressor in many types of human cancers, such as cholangiocarcinoma, 27 gastric carcinoma 28 and colorectal cancer. 29 Nevertheless, whether miR-587 affects LUAD progression by targeting BMP3 has not been investigated before.
Therefore, in this study, we aim to evaluate the effect of the A2M-AS1/miR-587/BMP3 axis on LUAD progression by confirmatory experiments in both LUAD cell lines and a xenograft tumour model. We for the first time hypothesize that A2M-AS1 might inhibit the pathogenesis of LUAD by regulating the miR-587/BMP3 axis. Our results may provide a theoretical basis for LUAD therapy and diagnosis.
Material and methods
Bioinformatic analysis
GEPIA database (https://gepia2.cancer-pku.cn/) was applied to analyze A2M-AS1 expression in LUAD tissue samples (N = 483) and normal tissue samples (N = 347), as well as in LUAD patients with different clinical stages. The relationship between LUAD patients prognosis and A2M-AS1 (or BMP3) expression was analyzed by Kaplan-Meier analysis (https://kmplot.com/analysis/). The potential miRNAs binding to A2M-AS1 (screening condition: Score ≥0.913) and the specific binding site between miR-587 and A2M-AS1 was predicted by DIANA database (https://carolina.imis.athena-innovation.gr/). The potential target mRNAs of miR-587 was predicted by miRDB database (https://mirdb.org/) (screening condition: Score Rank ≤5), and the specific binding site between miR-587 and BMP3 3′ untranslated region (3′UTR) was predicted by Targetscan database (https://www.targetscan.org/).
Cell culture
Lung/brunch normal epithelial cell line Beas-2B, LUAD cell lines H1975 and HCC827 were purchased from purchased from American Type Culture Collection (ATCC; Manassas, VA, USA). These cell lines were cultured in Dulbecco’s Modified Eagle Medium (DMEM; Gibco, NY, USA) added with 1% antibiotics and 10% fetal bovine serum (FBS; Gibco) in humidified incubator containing 5% CO2 at 37°C.
Clinical specimens
Characteristics of 23 LUAD patients used in the study.

Cell transfection
Before transfection, H1975 and HCC827 cells were cultured in 6-well plates (5 × 105 cells/well). Full-length A2M-AS1 was inserted into the pcDNA3.1 vector to overexpress A2M-AS1 with empty vector as a negative control. MiR-587 inhibitor was used to downregulate miR-587 with NC inhibitor as a negative control. Short hairpin RNA (shRNA) targeting BMP3 (sh-BMP3) was used to knock down BMP3 with sh-NC as a negative control. When the cell confluence reached 80%, a total of 50 nM pcDNA3.1/A2M-AS1, 50 nM sh-BMP3, 50 nM miR-587 inhibitor or 50 nM of their corresponding negative controls were transfected into LUAD cell lines using Lipofectamine 2000 (Invitrogen, CA, USA) at 37°C for 48 h based on supplier’s guidance.
Real-time quantitative polymerase chain reaction
Sequences of primers used for RT-qPCR.

Cell counting kit-8 assay
H1975 and HCC827 cell viability was detected using the Cell Counting Kit-8 (CCK-8; Dojindo, Japan). After H1975 and HCC827 cells were transfected with different vectors for 24, 48, 72 h, CCK-8 reagent (10 μl) was added to each well. Afterwards, the plate was incubated at 37°C for 4 h. The absorbance at 450 nm was measured by a microplate reader (Thermo Fisher, USA).
Colony formation assay
After transfection, H1975 and HCC827 cells were seeded into 6-well plates (800 cells/well). Two weeks later, colonies were treated with phosphate buffered saline (PBS; Solarbio) for cleaning, methanol (Solarbio) for fixation and Crystal Violet (Sigma–Aldrich) for staining. Then, the visible colonies were counted manually in five randomly chosen fields.
Transwell assay
Transwell assay was performed using 24-well transwell chamber which was precoated with or without Matrigel (BD, San Jose, USA). The upper chamber was added with treated H1975 and HCC827 cells (4 × 104 cells/well) in serum-free Roswell Park Memorial Institute-1640 (RPMI-1640) medium and the lower chamber was added with 10% fetal bovine serum medium. The cells were incubated with 5% CO2 at 37°C for 1 day. Then, those cells on the lower surface were treated with methanol (Sigma) for fixation and 1% crystal violet (Sigma) for staining. Finally, the migrated and invasive cells were photographed using a ×200 magnification microscope (Olympus, Tokyo, Japan) under three randomly selected fields.
Western blot analysis
H1975 and HCC827 cells were collected and lysed in Radio-Immunoprecipitation Assay (RIPA) lysis buffer (Beyotime, Shanghai, China). The lysates was treated with 12,000 g centrifugation for 10 min and then bicinchoninic acid (BCA) protein assay kit (Beyotime) was used to quantify total protein in supernatant. After bathing in the cold water, equal amounts of proteins were loaded on 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel and transferred to polyvinylidene fluoride (PVDF) membranes (Millipore). Following blocking the nonspecific binding sites with 5% defated milk, the membranes were probed with anti-BMP3 (1:200, ab134724, Abcam) and anti-GAPDH (1:1000, ab181602, Abcam) at 4°C overnight, along with horseradish peroxidase-labeled IgG (1:10,000, ab6721, Abcam) at room temperature for 2 h. The intensity of protein bands was visualized with enhanced chemiluminescence (ECL) assay kit (Millipore, USA) and quantified by densitometry using the Quantity One software (Bio-Rad).
RNA pulldown assay
A2M-AS1 probe with biotin (Bio-A2M-AS1) and negative control with biotin (Bio-NC) were purchased from GenePharma (Beijing, China). The biotinylated RNAs were incubated with the lysates of H1975 and HCC827 cells overnight, followed by pulldown by M-280 streptavidin magnetic beads (Invitrogen, USA) at 4°C overnight. After washing with wash buffer, the RNA mix bound to the beads was eluted and miRNA was extracted with an RNeasy Mini Kit (QIAGEN) for RT-qPCR.
Luciferase reporter assay
Full-length A2M-AS1 or BMP3 3′UTR sequence with wildtype (WT) or mutant-type (MUT) binding sites for miR-587 was subcloned into the pmirGLO luciferase vector (Promega, Madison, WI, USA) to construct A2M-AS1-WT/MUT or BMP3 3′UTR-WT/MUT vectors, separately. All plasmids were synthesized by GenePharma (Shanghai, China), and transfected into LUAD cells using Lipofectamine 2000 (Invitrogen). Renilla luciferase activity was used as a standardized control.
Generation of A2M-AS1-overexpressing cell line
Recombinant lentiviruses containing full-length A2M-AS1 (LV-A2M-AS1) and its negative control sequence (LV-NC) were obtained from GENECHEM (Shanghai, China). H1975 cells were infected with LV-A2M-AS1 or LV-NC plus 5 μg/mL polybrene (GENECHEM). After 48 h, infection efficiency was examined by RT-qPCR, and H1975 cells that can stably overexpress A2M-AS1 were selected with 2 μg/mL puromycin.
Xenograft assay in vivo
The in vivo assays were carried out under the guidelines and approval of the Animal Ethics Committee of Tumor Hospital Affiliated to Nantong University (Jiangsu, China). BALB/c nude mice (6 weeks old; approximately 20 g) were purchased from Vital River Co, Ltd (Beijing, China) to establish xenograft models. H1975 cells (1×106 per injection) carrying lentiviral vector LV-NC or LV-A2M-AS1 were subcutaneously inoculated into the flanks of the nude mice. From week one to week 5, callipers (Shoufeng, Shanghai, China) were used to measure tumor size every 7 days after tumor engraftment. The formula: volume = π/6 × a × b2 (a = longest diameter, b = shortest diameter perpendicular to a).The tumor volume was monitored every 7 days. After 5 weeks, the mice were sacrificed by cervical dislocation and tumors were excised. The tumor weight and tumor size in LV-NC and LV-A2M-AS1 groups were measured and recorded.
Statistical analysis
All the experimental data were analyzed using GraphPad Prism software 6.0 and represented as the means ± standard deviation (SD). Student’s t-test was utilized for comparison between two groups, and one-way analysis of variance (ANOVA) followed by Bonferroni’ second post hoc test was required for comparisons among multiple groups. p < 0.05 was considered statistically significant.
Results
A2M-AS1 is upregulated in LUAD and predicts poor prognosis
First, the expression pattern of A2M-AS1 in LUAD was analyzed. The data from GEPIA reveal that A2M-AS1 is downregulated in LUAD tissue samples compared to normal tissue samples (*p < 0.05, Figure 1(a)) and patients with LUAD at advanced stage exhibits downregulation of A2M-AS1 (**p < 0.01, Figure 1(b)). Additionally, A2M-AS1 was remarkably decreased in LUAD tissues in contrast to the corresponding normal tissues (***p < 0.001, Figure 1(c)). Kaplan-Meier survival curves revealed that low A2M-AS1 expression was associated with a low overall survival rate of LUAD patients (**p < 0.01, Figure 1(d)). Furthermore, A2M-AS1 was underexpressed in LUAD cell lines compared with the normal lung/brunch epithelial cell line Beas-2B (***p < 0.001, Figure 1(e)). Overall, A2M-AS1 is decreased in LUAD and may be related to advanced clinical stages and poor prognosis of LUAD patients. A2M-AS1 is downregulated in LUAD. (a) A2M-AS1 expression in LUAD tissue samples and normal tissue samples was analyzed by GEPIA. (b) A2M-AS1 expression in different clinical stages of LUAD patients was analyzed by GEPIA. (c) Detection of A2M-AS1 expression in adjacent non-tumor tissues and tumor tissues via RT-qPCR. (d) Kaplan-Meier survival analysis was required for predicting the association of A2M-AS1 expression with prognosis of LUAD patients. (e) Detection of A2M-AS1 expression in LUAD cell lines and Beas-2B cells using RT-qPCR ***p < 0.001.
A2M-AS1 overexpression inhibits LUAD cell proliferation, migration and invasion
Next, the biological function of A2M-AS1 in LUAD cells was explored. A2M-AS1 was overexpressed in H1975 and HCC827 cells by transfection with pcDNA3.1/A2M-AS1 (***p < 0.001, Figure 2(a)). The growth curves obtained from CCK-8 assay demonstrated that A2M-AS1 elevation remarkably inhibited the viability of H1975 and HCC827 cells (**p < 0.01, Figure 2(b)). Furthermore, A2M-AS1 upregulation suppressed LUAD cell proliferation by decreasing the colony number of LUAD cell line, as suggested by colony formation assay (**p < 0.01, Figure 2(c)). Additionally, Transwell assay confirmed that the elevation of A2M-AS1 resulted in reduced number of migrated and invaded LUAD cells, showing that A2M-AS1 enhancement restrains LUAD cell migration (**p < 0.01, Figure 2(d)) and invasion (***p < 0.001, Figure 2(e)). In summary, A2M-AS1 overexpression inhibits LUAD cell proliferative, invasive and migratory abilities. A2M-AS1 overexpression inhibits LUAD cell proliferation, migration and invasion. (a) Detection of A2M-AS1 expression in H1975 and HCC827 cells after overexpressing A2M-AS1 using RT-qPCR. (b) Evaluation of H1975 and HCC827 cell viability after overexpressing A2M-AS1 by CCK-8 assay. (c) Evaluation of H1975 and HCC827 cell proliferation after overexpressing A2M-AS1 by colony formation assay. (d-e) Evaluation of H1975 and HCC827 cell migration and invasion after overexpressing A2M-AS1 using transwell assays **p < 0.01, ***p < 0.001.
A2M-AS1 upregulation inhibits tumor growth in vivo
To explore the bioeffects of A2M-AS1 on tumor growth, an xenograft tumor model was constructed. H1975 cells infected with LV-A2M-AS1 or LV-NC were subcutaneously injected into nude mice. As Figures 3(a)–(c) indicated, tumors from the LV-A2M-AS1 group had smaller size, volume (***p < 0.001) and weight (**p < 0.01) than those in the LV-NC group. The expression of A2M-AS1 was upregulated in tumors after infection of LV-A2M-AS1 (**p < 0.01, Figure 3(d)). Overall, A2M-AS1 overexpression suppresses tumor growth in vivo. A2M-AS1 upregulation inhibits tumor growth in vivo. (a–c) Tumor size, tumor volume and tumor weight in nude mice after A2M-AS1 overexpression were imaged and recorded. (d) The of A2M-AS1 expression in tumors after treated with LV-A2M-AS1 or LV-NC **p < 0.01, ***p < 0.001.
A2M-AS1 sponges miR-587
To investigate the possible molecular mechanism of the anti-tumor effects of A2M-AS1 in LUAD, DIANA was utilized to predict 10 miRNAs containing binding site for A2M-AS1, namely miR-95-5p, miR-6733-5p, miR-7161-5p, miR-5093, miR-6739-5p, miR-1266-3p, miR-548u, miR-587, miR-4775 and miR-1273e (Figure 4(a)). Among these candidate miRNAs, miR-587 displayed the highest enrichment in the Bio-A2M-AS1 group in H1975 and HCC827 cells, as demonstrated by RNA pulldown assay (***p < 0.001, Figure 4(b) and (c)). Thus, miR-587 was selected for subsequent study. Then, the possible binding site between A2M-AS1 and miR-587 was obtained from DIANA (Figure 4(d)). RT-qPCR revealed that miR-587 level in LUAD tissues was higher than in normal tissues (***p < 0.001, Figure 4(e)). Furthermore, compared to Beas-2B cell line, LUAD cell lines exhibited higher level of miR-587 (***p < 0.001, Figure 4(f)). The expression of miR-587 was downregulated in H1975 and HCC827 cells after transfection with miR-587 inhibitor (***p < 0.001, Figure 4(g)). The outcome of luciferase reporter assay presented that the luciferase activity of A2M-AS1-WT was markedly increased by miR-587 inhibitor rather than that of A2M-AS1-MUT in LUAD cell lines (***p < 0.001, Figure 4(h) and (i)), indicating the interaction between A2M-AS1 and miR-587 in LUAD cells. Taken together, A2M-AS1 binds to miR-587 in LUAD cells. A2M-AS1 sponges miR-587. (a) DIANA was used to predict the miRNAs containing binding site for A2M-AS1. (b-c) The enrichment of potential miRNAs from the bio-A2M-AS1 probe was measured by RNA pulldown assay. (d) The binding site between A2M-AS1 and miR-587 was predicted by DIANA. (e) Detection of miR-587 expression in adjacent non-tumor tissues and tumor tissues using RT-qPCR. (f) Detection of miR-587 expression in LUAD cell lines and beas-2B cells using RT-qPCR. (g) Detection of miR-587 expression in H1975 and HCC827 cells after inhibiting miR-587 using RT-qPCR. (h-i) A luciferase reporter assay was used to validate the interaction between A2M-AS1 and miR-587 in H1975 and HCC827 cells after inhibition of miR-587 **p < 0.01, ***p < 0.001.
BMP3 is a target of miR-587
Subsequently, the target genes of miR-587 were further investigated. Based on the prediction from miRDB database, five possible mRNAs (BMP3, SLC35F1, GLI3, RAD54B and COL11A1) possibly binding to miR-587 were identified (Figure 5(a)). After LUAD cell lines were transfected with miR-587 inhibitor, only BMP3 expression was significantly increased among these candidates (***p < 0.001, Figure 5(b)). Hence, BMP3 was selected for further experiments. The protein level of BMP3 was upregulated after miR-587 deficiency in LUAD cells (Figure 5(c)). The potential binding site between miR-587 and the BMP3 3′UTR was predicted by Targetscan (Figure 5(d)). A luciferase reporter assay further revealed that knockdown of miR-587 elevated the luciferase activity of BMP3-WT, but that of BMP3-MUT was not affected in both H1975 and HCC827 cells (**p < 0.01, Figure 5(e)), which confirmed the binding between miR-587 and BMP3. To sum up, BMP3 is targeted by miR-587 in LUAD cells. BMP3 is targeted by miR-587. (a) MiRDB was applied to predict the mRNAs containing binding site for miR-587. (b) Detection of candidate mRNA expression in H1975 and HCC827 cells after miR-587 inhibitor using RT-qPCR. (c) Measurement of BMP3 protein expression in H1975 and HCC827 cells after inhibiting miR-587 by western blotting. (d) The binding site between BMP3 and miR-587 was obtained from targetscan. (e) H1975 and HCC827 cells cotransfected with miR-587 inhibitor and plasmids containing WT or MUT 3′UTR of BMP3 were subjected to luciferase activity measurement **p < 0.01, ***p < 0.001.
BMP3 is downregulated in LUAD and predicts a poor prognosis
First, the aberrant expression of BMP3 in LUAD was evaluated by RT-qPCR. The results manifested that BMP3 expression was markedly downregulated in LUAD tissue samples compared to normal tissue samples (**p < 0.01, Figure 6(a)). Similarly, BMP3 expression was significantly lower in two LUAD cell lines than in Beas-2B cells (***p < 0.001, Figure 6(b)). Next, the performance of BMP3 expression for LUAD diagnosis was validated by Kaplan-Meier survival analysis, which showed that low BMP3 expression was associated with a poor prognosis in LUAD patients (**p < 0.01, Figure 6(c)). Collectively, BMP3 is downregulated in LUAD and predicts a poor clinical outcome. BMP3 is downregulated in LUAD. (a) BMP3 expression in tumor tissues and adjacent non-tumor tissues was detected by RT-qPCR. (b) Detection of BMP3 expression in LUAD cell lines and a lung/brunch normal epithelial cell line beas-2B using RT-qPCR. (c) Kaplan-Meier plotter was applied to predict the relationship between BMP3 expression and prognosis of LUAD patients **p < 0.01, ***p < 0.001.
BMP3 knockdown attenuates the suppression of A2M-AS1 elevation on LUAD cell proliferation, migration and invasion
In rescue assays, the regulatoty role of the A2M-AS1/BMP3 axis in LUAD cells was explored. The mRNA and protein levels of BMP3 were reduced in H1975 cells after transfection with sh-BMP3 (***p < 0.001, Figure 7(a)). CCK-8 assay indicated that BMP3 knockdown offset the restriction of A2M-AS1 overexpression on LUAD cell viability (**p < 0.01, Figure 7(b)). The outcome of colony formation assay revealed that decreased colony number caused by elevating A2M-AS1 was recovered by BMP3 deficiency in H1975 cells (***p < 0.001, Figure 7(c)). In addition, A2M-AS1 upregulation suppressed the number of migrated LUAD cells and invaded LUAD cells, which was restored by BMP3 silencing (***p < 0.001, Figure 7(d) and (e)). Collectively, BMP3 knockdown attenuates the suppressive effects of A2M-AS1 overexpression on LUAD cell proliferation, migration and invasion. BMP3 knockdown attenuates the suppressive effect of A2M-AS1 overexpression on LUAD cell proliferation, migration and invasion. H1975 cells were transfected with vector, A2M-AS1 or A2M-AS1 + sh-BMP3. (a) Measurement of BMP3 mRNA and protein levels in H1975 cells after knocking down BMP3 using RT-qPCR and western blotting. (b) Evaluation of H1975 cell viability in each group by CCK-8 assay. (c) Evaluation of H1975 cell proliferation in each group via colony formation assay. (d-e) Detection of H1975 cell migration and invasion in each group via transwell assays *p < 0.05, **p < 0.01, ***p < 0.001.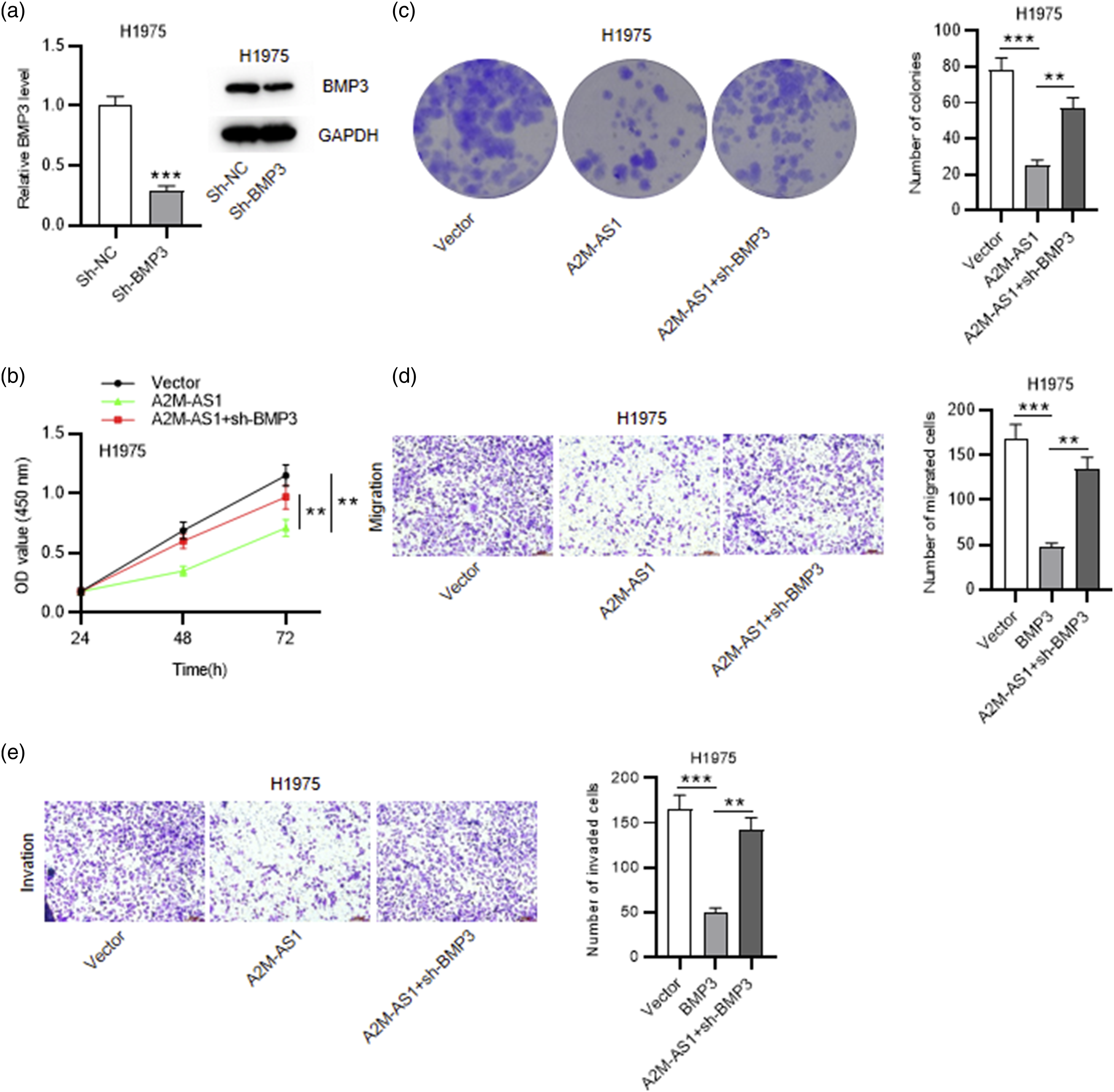
Discussion
LUAD is one of the most prevalent malignant cancers around the world. 30 So far, various lncRNAs have been found to be involved in the pathophysiological processes of LUAD.31,32 For example, lncRNA LINC00520 upregulates forkhead box R2 (FOXR2) expression by sponging miR-1252-5p to accelerate LUAD progression. 33 LncRNA H19 faciliates the viability and epithelial-mesenchymal transition (EMT) of LUAD cells via binding with miR-29b-3p to regulate signal transducer and activator of transcription 3 (STAT3). 34 LncRNA titin-antisense RNA 1 (TTN-AS1) acts as a oncogene to promote cell mobility and EMT process via regulating the miR-142-5p/cyclin dependent kinase 5 (CDK5) axis in LUAD. 35 In this study, A2M-AS1 is downregulated in LUAD tissues and cells. Additionally, low expression of A2M-AS1 was related to advanced clinical stage and poor prognosis in LUAD patients. A2M-AS1 upregulation inhibited the proliferation, invasion and migration of LUAD cells. Our results indicated that A2M-AS1 is a tumor suppressor in LUAD. Limited studies have reported that A2M-AS1 also participates in pathogenesis of breast carcinoma, pancreatic ductal adenocarcinoma and NSCLC.18,20,36 Especially, A2M-AS1 facilitates breast cancer cell invasion, proliferation and suppressed apoptosis via sponging miR-146b to upregulate MUC19 37 , suggesting that A2M-AS1 acts as an carcinogenic gene in breast cancer development. Therefore, A2M-AS1 plays different roles in cancers, which may depend on the cellular and molecular context.
Since the competing endogenous RNA (ceRNA) hypothesis was put forword, increasing evidence indicated that lncRNA can affect miRNA expression and thus reduce its restriction on target genes as a ceRNA. 38 Therefore, we predicted that A2M-AS1 may be involved in LUAD regulation via the ceRNA mechanisms. In this study, according to the prediction of bioinformatics websites and the results of RNA pulldown and luciferase reporter assays, miR-587 was identified to be a direct target of A2M-AS1 and was overexpressed in LUAD tissues and cells. Previously, miR-587 was demonstrated to be abnormally expressed and participate in the carcinogenesis of several cancers. For example, miR-587 expression downregulates in prostate cancer. 39 MiR-587 interacts with Ribosomal Protein SA to suppress hepatocellular carcinoma progression 40 and in cervical cancer by repressing IRF6. 41 Notably, miR-587 was reported to play an oncogenic role in NSCLC by reducing CYLD expression. 25 However, in this study, the precise effect of miR-587 on tumorigenesis of LUAD remains unclear. Therefore, for further understanding of the ceRNA mechanism of A2M-AS1 in LUAD, the biological function of miR-587 should be explored in future researches.
Methylated BMP3 was identified to be a target gene of miR-587. Recent studies have reported that BMP3 exerts important functions in the tumorigenesis of various types of cancers. 42 BMP3 is hypermethylated and inactive in gastric cancer 28 and biliary cancer. 27 Additionally, for those patients suffering from abnormal breast mass, hypermethylated BMP3 may lead to increasing risk of breast cancer. 43 Furthermore, BMP3 is characterized by low activity in the early stages of most colorectal carcinoma cases 44 and shows hypermethylation in colorectal carcinoma. 45 These findings indicate that BMP3 serves as a tumor suppressor in several kinds of cancers. In our study, LUAD tissues and cells exhibited downregulated BMP3 expression level and low expression of BMP3 was related to a poor prognosis in LUAD patients. MiR-587 inhibitor contributed to increased mRNA and protein levels of BMP3. Additionally, BMP3 knockdown reserved the suppressive effects of A2M-AS1 on the proliferative, migratory and invasive behaviors of LUAD cells, suggesting that BMP3 also played an antioncogenic role in LUAD. Of interest, as an indispensable component of TGF-β superfamily, BMPs play important roles in LUAD progression. 46 BMP2 and BMP7 are upregulated in LUAD and they both promote tumor cell proliferation and migration.47,48 However, like BMP3, BMP5 is also downredulated in LUAD and acts as a tumor suppressor. 49
In conclusion, A2M-AS1 sponges miR-587 to regulate BMP3 expression, thus inhibiting LUAD cell proliferation, migration and invasion. Our results highlighted that the A2M-AS1/miR-587/BMP3 axis might be potential therapeutic targets for LUAD. However, to figure out the mechanism of A2M-AS1 in LUAD more precisely, functional experiments of miR-587 in LUAD should be carried out in our future work.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Nantong Science and Technology Bureau (No. MS12020046).
