Abstract
This study examined the association between clock gene expression and the effect of methamphetamine (MA) on drug-metabolizing enzymes from the perspective of drug metabolism. The relationship between expression of the clock genes BMAL1 and PER2 and the drug-metabolizing enzymes CYP3A4 and CYP2D6 was investigated using livers from autopsy cases of MA-intoxication deaths. Additionally, the effect of MA exposure on various genes was examined in HepG2 human hepatocellular carcinoma cells. Comparisons of the expression of various genes in MA users according to blood MA concentration revealed that CYP3A4 expression was similar to that of PER2, and CYP2D6 expression was similar to that of BMAL1. In cultured cell experiments, BMAL1 and CYP2D6 expression decreased depending on the time elapsed after MA addition, and PER2 and CYP3A4 expression increased slightly in a concentration-dependent manner. These results were consistent with the findings of autopsy examinations. Expression of CYP3A4 and CYP2D6 under BMAL1 and PER2 suppression, but not CYP2D6 under PER2 suppression alone, was upregulated in response to MA. These results suggest that CYPs are regulated via the clock genes BMAL1 and PER2 during MA metabolism.
Introduction
Generally, the movement and action of drugs in the body vary greatly depending on the time of administration, and clock genes play an important role in drug disposition.1-4 Clock genes affect drug metabolism and excretion by inducing fluctuations in the activity of the drug-metabolizing enzyme cytochrome P450 (CYP) and in the expression of transporters in a 24-h cycle. 1 Studies in mice have reported that the clock gene Bmal1 regulates CYP3A4 expression and drug metabolism. 2
Methamphetamine (MA) and amphetamine (AMP) are widely abused psychostimulants. 5 MA is partially metabolized by CYP2D6 and CYP3A4.6,7 Certain genetic factors, such as the CYP2D6 phenotype, reportedly contribute to individual differences in sensitivity to stimulants. 8 In cases of death due to MA intoxication, fatal side effects may occur even in the therapeutic concentration range, and it can be difficult to determine the cause of death and understand the pathophysiologic process that led to death. However, the relationship between CYP activity and drug metabolism in actual human tissues remains largely unknown. As the expression of clock genes affects drug metabolism, differences in clock gene expression or the timing of drug administration may also affect individual differences in MA sensitivity.
To date, there are no comprehensive reports describing the effects of MA on CYP2D6 and CYP3A4 in the human liver or the role of clock genes in MA metabolism. Cathinone, the major active compound of the natural amphetamine known as khat, is structurally similar to AMP. Several studies have reported that cathinone inhibits several CYP isoforms in vitro.9,10 Similarly, structurally similar MA may also inhibit CYP expression. In this study, the expression of the clock genes BMAL1 and PER2 was examined using livers from cases of MA-intoxication death from the perspective of drug metabolism in order to clarify the relationship between clock gene expression and the effect of MA on drug-metabolizing enzymes.
Methods
Ethics statement
This study was evaluated by the Independent Ethics Committee of the Osaka Metropolitan University Graduate School of Medicine, which approved opt-out of informed consent for the use of autopsy data for analysis (authorization no. 4404).
Human samples
Case profiles.

Measurement of MA
AMP and MA were extracted using a Gilson ASPEC XL-274 automated SPE solid/liquid-phase extraction instrument (Middleton, WI, USA), as follows. As an internal standard, a 50-μL solution containing 3-phenylpropylamine was added to 0.5 mL of each sample. The pH of the samples was adjusted to approximately 7.0 by adding 6 mL of 0.1
Total RNA extraction and qRT-PCR analysis
Total RNA was isolated using an Isogen kit (Nippon Gene, Toyama, Japan) following the manufacturer’s instructions. cDNA was synthesized from total RNA using a High-Capacity RNA-to-cDNA kit (Applied Biosystems, Foster City, CA, USA). Quantitative reverse transcription (qRT)-polymerase chain reaction (PCR) was performed by adding 20.0 μL of the reaction mixture, consisting of 10.0 μL of TaqMan gene expression master mix (2×), 1.0 μL of TaqMan gene expression assay (20×), 4 μL of cDNA, and 5 μL of H2O, to each well of a 96-well reaction plate (0.1 mL/well). Primers for the transcripts encoding aryl hydrocarbon receptor nuclear translocator-like protein (ARNTL/BMAL1) (TaqMan assay ID: Hs00154147_m1), period circadian clock protein (PER2) (TaqMan assay ID: Hs00256143_m1), cytochrome P450 family three subfamily A member 4 (CYP3A4) (TaqMan assay ID: Hs00604506_m1), and cytochrome P450 family two subfamily D member 6 (CYP2D6) (TaqMan assay ID: Hs04931916_gH) were used with a StepOnePlus real-time PCR system (Applied Biosystems). A threshold of 0.2 was used, and the software automatically calculated the threshold cycle (Ct) value. As a preliminary step, the stability of several endogenous reference genes (Actinβ, β2M, GAPDH, HMBS, HPRT1, PPIA, and TBP) in the liver was tested. As a result, HMBS (TaqMan assay ID: Hs00609297_m1) was selected as the most stable endogenous reference gene. 15
Effects of MA
Effect of MA on HepG2 cells
The effects of MA exposure were evaluated using HepG2 cells (Cellular Engineering Technologies, Coralville, IA, USA). Cells were cultured in HepG2 hepatocellular carcinoma expansion medium (Cellular Engineering Technologies) supplemented with 10% fetal bovine serum (Biological Industries Ltd, Kibbutz Beit-Haemek, Israel, Product No. 04-121-1A) and adjusted to 1.5 × 105 cells/well in 24-well plates. MA-addition experiments were performed the next day. Cells were cultured in an incubator with 4.7% CO2 at 37°C. Cells were counted using a Cell Counter model R1 (Olympus Optical Co, Ltd, Tokyo, Japan). MA was added to the cell cultures at final concentrations of 0.2, 2, 5, and 10 μmol/dL per well (n = 6 for each). At 1 and 3 h after addition of MA, cells were collected, and gene expression was analyzed using RT-PCR. 16
Effect of MA under suppression of BMAL1 and PER2 expression
To confirm the effect of MA under suppression of BMAL1 and PER2 expression, Silencer® Select pre-designed siRNAs (Thermo Fisher Scientific) were used to suppress expression of these genes. The IDs of the siRNA assays used were s1616 for BMAL1 and s16932 for PER2. Silencer® Select Negative Control #1 siRNA was used as the negative control (NC). On the day prior to the assay, cells were cultured in a 24-well plate at 1.5 × 105 cells per well in 1 mL of serum-containing, antibiotic-free medium. Next, 1.5 μL of Lipofectamine RNAiMAX reagent was added to 25 μL of Opti-MEM medium and mixed. Additionally, 1.0 μL of 5 μ
Statistical analysis
Pearson correlation coefficients were used to compare differences between groups. The nonparametric Mann-Whitney U test was used for comparisons of two groups, and the Kruskal–Wallis test was used for analyses involving multiple comparisons. Graphs of test results for the autopsy samples are shown as box and whisker plots, with the horizontal line in each box indicating the median. Box whiskers indicate 90% confidence intervals. Circles and asterisks indicate outliers and unexplained outliers, respectively. Maximum gene expression values were log-transformed for graphical presentation. The results of cell culture experiments are presented as bar graphs, in which the bar indicates the mean, and the whiskers indicate the standard deviation. All analyses were performed using the SPSS 9.0 statistical package (SPSS, Inc., Chicago, IL, USA). Significance was defined as p < 0.05. 17
Results
Time of death and expression of various genes in MA users
There was no statistically significant difference between the time of death and expression of the BMAL1, PER2, CYP3A4, and CYP2D6 genes (Figures 1(a) to (d)). Gene expression in liver tissue of autopsy cases, categorized by time of death. Expression of BMAL1 (a), PER2 (b), CYP3A4 (c), and CYP2D6 (d) in liver samples recovered at autopsy. The data are presented as box and whisker plots, in which the central horizontal line in each box represents the median; the boxes span the interquartile range, and the whiskers represent the 90% confidence interval. Circles represent outliers, and asterisks represent abnormal values.
Expression of various genes according to blood MA concentration
BMAL1 expression tended to be increased at moderate and severe blood MA concentrations but tended to decrease as MA concentration increased further (Figure 2(a)). Conversely, PER2 expression increased with increasing blood MA concentration and was highest at the fatal level of MA (Figure 2(b)). CYP3A4 expression was similar to that of PER2 and tended to increase with increasing blood MA concentration (Figure 2(c)). CYP2D6 expression was similar to that of BMAL1, which was highest at the severe blood MA concentration but decreased at the fatal level (Figure 2(d)). Gene expression in liver tissues of autopsy cases, categorized by blood methamphetamine concentration. Expression of BMAL1 (a), PER2 (b), CYP3A4 (c), and CYP2D6 (d) in liver samples collected at autopsy. The data are presented as box and whisker plots, in which the central horizontal line in each box represents the median; the boxes span the interquartile range, and the whiskers represent the 90% confidence interval. Circles represent outliers, and asterisks represent abnormal values.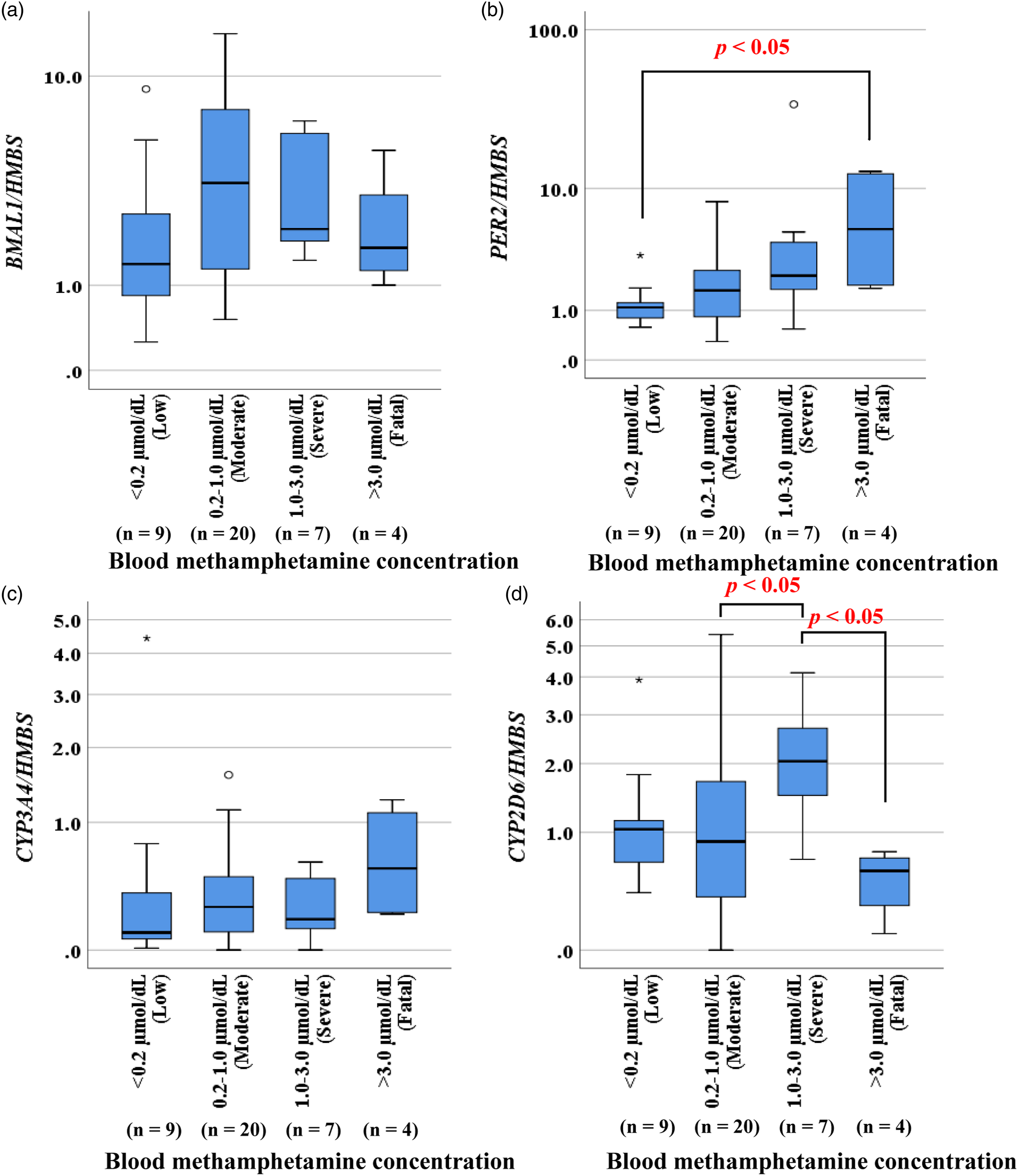
Effect of MA on gene expression in HepG2 cells
The expression of BMAL1 and CYP2D6 in HepG2 cells tended to be higher at 1 h than 3 h after addition of MA at various concentrations (Figures 3(a) and (d)). Although the range increased slightly after MA addition, the expression of PER2 and CYP3A4 tended to increase slightly with increasing MA concentration (Figures 3(b) and (c)). Expression of the BMAL1 (a), PER2 (b), CYP3A4 (c), and CYP2D6 (d) genes in HepG2 cells after methamphetamine addition. The data are presented as bar charts, in which the bars show the means, and the whiskers show the standard deviations.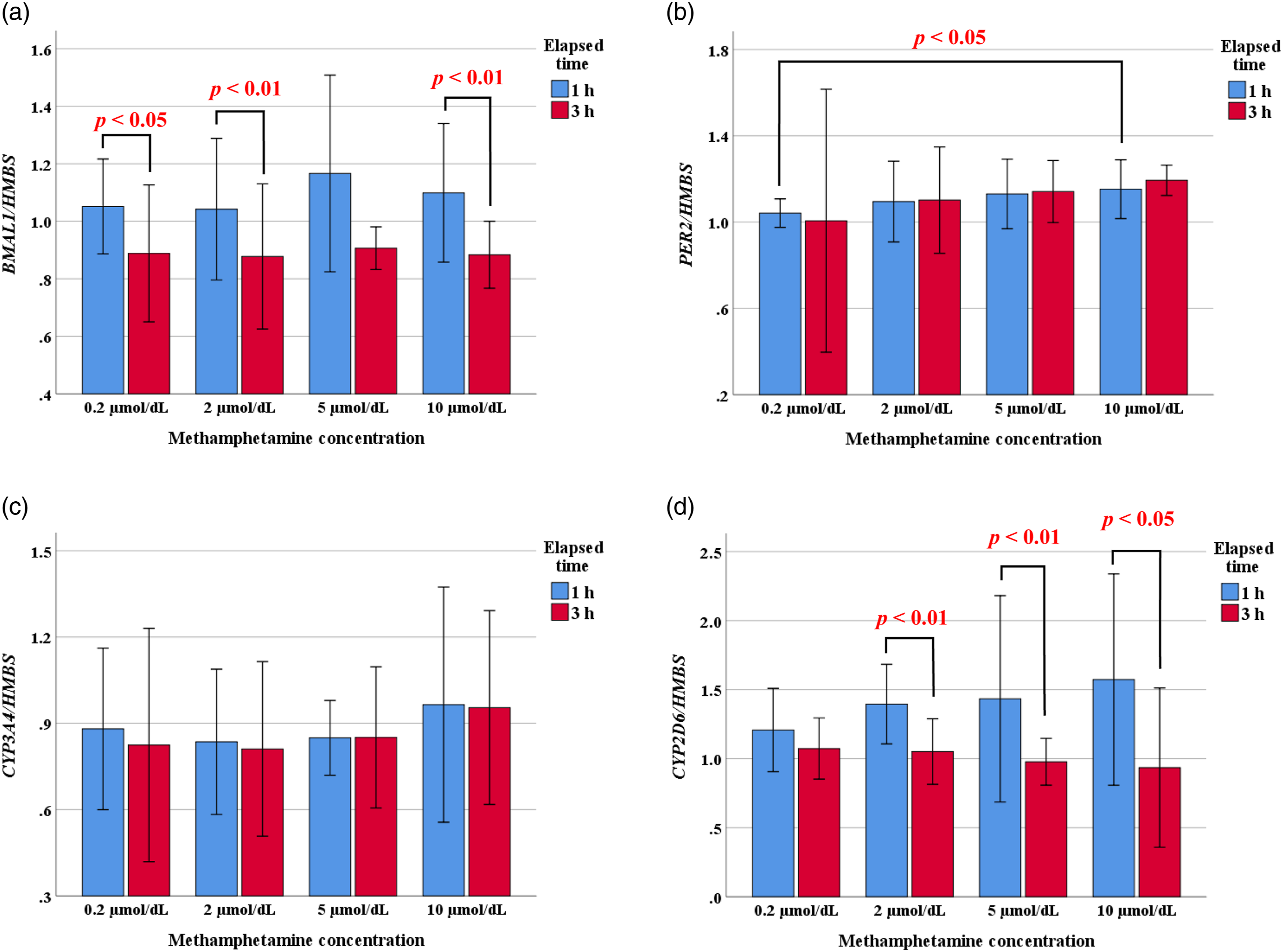
Effect of MA under suppression of BMAL1 and PER2 expression
Introduction of siRNA suppressed BMAL1 expression (Figure 4(a)). Following suppression of BMAL1, the expression of CYP3A4 and CYP2D6 increased compared with NC cells. Furthermore, MA addition under BMAL1 suppression led to increased CYP3A4 expression and decreased CYP2D6 expression over time (Figures 4(b) and (c)). PER2 expression was also suppressed following introduction of siRNA (Figure 4(d)). PER2 suppression increased CYP3A4 expression compared with NC cells; however, only a minimal increase in CYP2D6 expression was observed under PER2 suppression. Although CYP3A4 expression increased over time following addition of MA under PER2 suppression, CYP2D6 expression was not changed over time following addition of MA under PER2 suppression (Figures 4(e) and (f)). Expression of the BMAL1 (a), CYP3A4 (b), and CYP2D6 (c) genes in HepG2 cells after methamphetamine addition under BMAL1 suppression. Expression of the PER2 (d), CYP3A4 (e), and CYP2D6 (f) genes in HepG2 cells after methamphetamine addition under PER2 suppression. The data are presented as bar charts, in which the bars show the means, and the whiskers show the standard deviations.
Discussion
This study examined the association between clock gene expression and the effect of MA on drug-metabolizing enzymes from the perspective of drug metabolism. The expression of various genes was compared in livers of autopsy cases of individuals who were MA users and in HepG2 human hepatocellular carcinoma cells treated with MA.
No association was observed between time of death and expression of the BMAL1, PER2, CYP3A4, and CYP2D6 genes. The expression of BMAL1 and CYP2D6 tended to increase at moderate or severe levels of blood MA and decrease at fatal levels. Conversely, the expression of PER2 and CYP3A4 increased with increasing blood MA concentration and was the highest at fatal blood MA concentrations. To complement the results obtained using human tissue samples, experiments were performed using HepG2 cells. The expression of BMAL1 and CYP2D6 decreased over time after MA addition, whereas the expression of PER2 and CYP3A4 increased slightly in a concentration-dependent manner after MA addition, although the extent of the increase was small. These results suggest that MA affects the expression of clock genes and CYPs.
To further investigate the relationship between the clock genes BMAL1 and PER2 and CYP expression, cells were treated with MA under siRNA-mediated suppression of gene expression. The expression of CYP3A4 and CYP2D6 increased under BMAL1 suppression. Similarly, CYP3A4 expression was upregulated under PER2 suppression. Conversely, CYP2D6 expression did not increase over time under PER2 suppression and was thus unaffected by MA. With the exception of CYP2D6 expression under PER2 suppression, MA affected the expression of all of the examined CYPs under BMAL1 and PER2 suppression. These results show that CYPs are regulated by a clock gene feedback group and suggest that CYPs respond to MA under the control of clock genes.
Lim et al. reported that cathinones, which are similar in structure to amphetamines, inhibit the activity of CYP2D6 and CYP3A4.9,10 However, the present study showed that the expression of only the CYP2D6 gene was suppressed after MA addition. Additionally, previous studies have reported that MA does not affect CYP3A4. 18 The difference between cathinone and MA in terms of the effect on CYP3A4 is unknown, but it may be due to structural differences. 10 The results of the present research lead us to hypothesize that the clock genes inhibit the effect of MA on CYP3A4 expression. In addition, MA may affect not only CYP2D6 and CYP3A4 but also various other CYP isoforms via clock genes.
A limitation of the present study was that clock gene expression in stimulant users did not exhibit a diurnal expression pattern, and thus, it was not possible to assess individual differences in MA sensitivity among the deceased in relation to differences in clock gene expression. The results of the present study showed that clock genes affect the expression of CYP3A4 and CYP2D6. These data suggest that clock genes mediate individual differences in MA sensitivity. In order to investigate this hypothesis in more detail, it will be necessary to conduct a large-scale investigation including more cases of stimulant-poisoning death and investigate the relationship between MA concentration and clock genes in such cases.
Conclusion
In conclusion, the results of analyses of autopsy samples and cell culture experiments indicate that expression of the BMAL1 and PER2 genes modulates the expression of CYP3A4 and CYP2D6. Furthermore, the results of this study suggest that MA intake affects the expression of clock genes and drug-metabolizing enzymes.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Data availability statement
All data generated or analyzed during this study are included in this published article and its supplementary information files.
