Abstract
Introduction
Gastric carcinoma (GC) is the second most common malignant tumor and the fourth most prevalent cancer globally. 1 Although radiotherapy and chemotherapy can significantly increase the survival rate of GC patients, the 5-year survival rate of GC patients remains only 25–30%. 2 The occurrence and development of GC is a multi-stage, complex evolutionary process involving multiple factors, multiple genes, and multiple intracellular and extracellular signal pathways; 3 however, its specific mechanism has not been fully clarified. Therefore, elucidation of the key factors causing GC and their molecular mechanisms is particularly important for improving patient quality of life and reducing patient mortality.
Long non-coding RNAs (lncRNAs) are molecules over 200 nucleotides in length that are localized in the nucleus or cytoplasm 4 and are involved in epigenetic, transcriptional, and post-transcriptional gene expression regulation. 5 Accumulating evidence has indicated that lncRNAs are further involved in tumor proliferation, invasion, and metastasis and have potential as tumor biomarkers and therapeutic targets for human cancers.6,7 Down syndrome cell adhesion molecule-anti-sense 1 (DSCAM-AS1) is a recently discovered lncRNA located on 21q22.2 8 that may have important roles in many human cancers. 9 High-level expression of DSCAM-AS1 was found to promote cervical cancer progression via oncogenesis. 10 The oncogenetic role of DSCAM-AS1 has also been documented in hepatocellular carcinoma. 11 After a significant increase in DSCAM-AS1 expression was observed in melanoma samples, silencing DSCAM-AS1 inhibited cell proliferation and migration, as well as colony formation. 9 However, the role of DSCAM-AS1 in GC requires clarification. In this study, we investigated DSCAM-AS1 expression in GC tissues and cell lines to elucidate its role in GC regulation. The findings of this study may be useful for the development of novel therapeutic approaches in GC.
Materials and methods
Patient tissues
We collected 62 GC tissues and adjacent normal tissues from patients of Tangdu Hospital, Air Force Medical University, between 2013 and 2019. This study was approved by the Tangdu Hospital, Air Force Medical University Ethics Committee (TDLL-KY-202105-01). All patients signed an informed consent form prior to the study.
Cell culture and transfection
Four human GC cell lines (BGC-823, NCI-N87, SGC-7901, and MGC-803) and one human normal gastric epithelial cell line (GES-1) were cultured in Delbecco’s modified Eagle medium (DMEM; Gibco C11885500BT, Thermo Fisher, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA) followed by incubation at 37°C humidified with 5% CO2. Transfection was performed using Lipofectamine 2000 reagent to prepare an siRNA against DSCAM-AS1 (si-DSCAM-AS1), miR-204 mimic (miR-204), and miR-204 inhibitor and their respective controls.
RNA extraction and quantitative reverse transcription-polymerase chain reaction (qRT-PCR)
Total RNA was isolated from the samples using Trizol reagent (Invitrogen) and the concentration and quality of RNA was assessed using a spectrophotometer (Thermo Fisher). Total RNA was reverse-transcribed into cDNA using a High Capacity cDNA Reverse Transcription Kit (Thermo Fisher) or microRNA Reverse Transcription Synthesis Kit (Thermo Fisher). Next, qPCR analysis was performed using Maxima SYBR Green qPCR Master Mix (2×) (Thermo Fisher) and specific primers. Fold changes were calculated using the 2–ΔΔCt method, followed by statistical analysis.
MTT assay
Cell viability was detected using an MTT assay. The treated cells (∼104 cells per 96-well plate) were incubated for 24, 48, or 72 h. Sample absorbance was read at 490 nm under a microplate reader (Bio-Rad, Hercules, CA, USA).
Cell apoptosis assay
An Annexin V–fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis detection kit (Keygen, Nanjing, China) was used to measure apoptosis. Briefly, cells were collected after centrifugation and re-suspended in 400 μL of binding buffer, and then incubation with Annexin V–FITC and PI in the dark for 0.5 h, followed by apoptotic cell detection via flow cytometry (Partec AG, Arlesheim, Switzerland).
Transwell migration assay
A Transwell assay was performed to detect cell migration. Briefly, BGC-823 or SGC-7901 cells were cultured and seeded into the upper chamber of an incubator containing Matrigel at a density of 2 × 104 cells per chamber. The lower chamber was supplemented with RPMI-1640 medium containing 10% FBS. After 48 h of incubation, cells adhering to the lower surface were fixed in methanol for 15 min, followed by 20% methanol and 0.2% crystal violet for staining. Inserts were counted in images obtained using a light microscope (Olympus, Tokyo, Japan).
Luciferase reporter assay
Wild-type DSCAM-AS1-3′-UTR (WT-DSCAM-AS1-3′-UTR) or TPT1-WT (WT-TPT1-3′-UTR) and mutant DSCAM-AS1-3′-UTR (MUT-DSCAM-AS1-3′-UTR) or TPT1-MUT (MUT-TPT1-AS1-3′-UTR) containing the miR-204 binding sites were inserted into pmiR-RB-REPORT (Ribobio) for luciferase reporter tests. Each of these vectors was then transfected into BGC-823 or SGC-7901 cells. Luciferase activity was measured using a dual luciferase assay system (Promega, Madison, WI, USA).
Western blot assay
Total protein was extracted from the collected cells, and a bicinchoninic acid (BCA) assay was used to quantify the protein. Using 10% sodium dodecyl–sulphate polyacrylamide gel electrophoresis (SDS–PAGE), 20 μg of protein was separated and transferred onto polyvinylidene fluoride or polyvinylidene difluoride (PVDF) membranes. After blocking with non-fat milk (5%; Yili, Beijing, China), the membranes were probed with primary antibodies against TPT1 (1:1000) and GAPDH (1:1000) for 12–16 h at 4°C, followed by secondary antibody incubation. Finally, an enhanced chemiluminescence reagent (Applygen, Beijing, China) was used to visualize all blots. TPT1 protein levels were quantified using ImageJ software and normalized with GAPDH.
Animals and xenograft tumor model establishment
10 BALB/c nude mice were purchased from the Shanghai Animal Center (Shanghai, China). SGC-7901 cells with and without si-DSCAM-AS1 (2 × 106 cells in 200 μL of phosphate-buffered saline [PBS], n = 5/group) were subcutaneously injected into 4-week-old male BALB/c nude mice. After 24 days, the nude mice were anesthetized with pentobarbital sodium, and the tumor tissues were weighed and photographed. All animal experimental procedures were approved by the Ethics Committee for Animal Studies of Tangdu Hospital, Air Force Medical University (TD.20190102).
Statistical analysis
The monitoring data are analyzed by SPSS19.0 statistical software. The results of data analysis are shown as mean ± standard deviation (mean ± SD). Multigroup data analysis is founded on one-way ANOVA. LSD test was employed for subsequent analysis. p < 0.05 indicated the difference was significant.
Results
DSCAM-AS1 was upregulated in GC tissues and cell lines
DSCAM-AS1 expression in GC tissues and cell lines was detected using qRT-PCR. DSCAM-AS1 expression was significantly higher in GC tissues (Figure 1(A)) and cell lines (Figure 1(B)) than in paracancerous tissues, suggesting that DSCAM-AS1 plays a central role in GC. The expression of DSCAM-AS1 was upregulated in GC tissues and cells. A: The expression level of DSCAM-AS1 was determined in 62 GC and matched adjacent normal tissues using qRT-PCR. B: The relative DSCAM-AS1 expression in 4 GC cell lines and the human◆ normal◆ gastric◆ epithelial◆ cell◆ line◆ GES-1. Data are mean ± SD. *p < 0.05, **p < 0.01. GC: gastric carcinoma, DSCAM-AS1: Down syndrome cell adhesion molecule-anti-sense 1.
Downregulation of DSCAM-AS1 inhibited GC cell proliferation and migration
We transfected siRNAs specifically against DSCAM-AS1 into BGC-823 and SGC-7901 cells (Figure 2(A)) to explore the role of DSCAM-AS1 in GC proliferation and migration. MTT results suggested cell viability suppression after transfection with si-DSCAM-AS1 (Figure 2(B)). The flow cytometry data showed the opposite results (Figure 2(C)). Silencing DSCAM-AS1 expression significantly inhibited GC cell migration (Figure 2(D)). DSCAM-AS1 knockdown suppressed GC cell proliferation and migration. A: DSCAM-AS1 siRNA was transfected into GC cell lines BGC-823 and SGC-7901. The expression of DSCAM-AS1 was detected by qRT-PCR following transfection. B: GC cell proliferation was determined by MTT assay after DSCAM-AS1 knockdown. C: Cell apoptosis of BGC-823 and SGC-7901 cells following DSCAM-AS1 knockdown. D: The migration capacity of GC cells after effective knockdown of DSCAM-AS1. Data are mean ± SD. *p < 0.05, **p < 0.01. GE: gastric carcinoma, DSCAM-AS1: Down syndrome cell adhesion molecule-anti-sense 1.
MiR-204 downregulation in GC tissues and cells was regulated by DSCAM-AS1
The StarBase v2.0 online database predicted that miR-204 had substantial complementary binding sites with DSCAM-AS1 (Figure 3(A)). The luciferase assay results confirmed the existence of binding sites between DSCAM-AS1 and miR-204, and fluorescence intensity was replenished after binding sites between DSCAM-AS1 and miR-204 had mutated (Figure 3(B)), and miR-204 levels decreased significantly in GC tissues and cell lines (Figures 3(C) and (D)). DSCAM-AS1 acted as a molecular sponge for miR-204 in GC cells. A: miR-204 contains the potential binding site for DSCAM-AS1 predicted by Starbase2.0. B: The luciferase activity was detected in GC cells after co-transfected with wild-type (WT) or mutant (MUT) DSCAM-AS1 reporter and miR-204 mimics or miR-NC. C: The relative expression of miR-204 in GC tissues and adjacent normal tissues detected by qRT-PCR. D: miR-204 expression in 4 GC cell lines and the human◆ normal◆ gastric◆ epithelial◆ cell◆ line◆ GES-1 determined by qRT-PCR. Data are presented as mean ± SD. *p < 0.05, **p < 0.01. DSCAM-AS1: Down syndrome cell adhesion molecule-anti-sense 1.
TPT1 was a target of miR-204
The downstream targets of TPT1 were predicted by the StarBase v2.0 database. Binding sites were detected between miR-204 and TPT1 (Figure 4(A)). To verify the targeting relationship between miR-204 and TPT1, we conducted a dual-luciferase reporter assay. The results show that the luciferase activity of TPT1-WT was sharply suppressed in miR-204-transfected BGC-823 and SGC-7901 cells, whereas the luciferase activity of TPT1-MUT showed little change (Figure 4(B)). The mRNA (Figure 4(C)) and protein (Figure 4(D)) levels of TPT1 were notably decreased in BGC-823 and SGC-7901 cells transfected with miR-204. TPT1 was a direct target of miR-204 in GC cells. A: TPT1 was a candidate target of miR-204 predicted by TargetScan. B: Luciferase reporter assay was conducted in BGC-823 and SGC-7901 cells after transfected with wild-type (WT) or mutant (Mut) TPT1 vectors and miR-204 mimics or miR-NC. C: The relative mRNA levels of TPT1 in GC cells BGC-823 and SGC-7901 after miR-204 overexpression. D: The protein expression of TPT1 in GC cells BGC-823 and SGC-7901 after miR-204 overexpression. Data are presented as mean ± SD. *p < 0.05, **p < 0.01.
DSCAM-AS1 regulates the miR-204/TPT1 axis to exert its effect in GC
Next, we investigated whether DSCAM-AS1 mediates the miR-204/TPT1 axis to influence GC progression. The miR-204 inhibitor group showed decreased expression of miR-204 after transfection (Figure 5(A)), and the results of the MTT assay showed that silencing miR-204 partially reversed its suppressive effects on GC cell proliferation induced by the DSCAM-AS1 knockdown (Figure 5(B)). We also observed a significant decrease in TPT1 protein levels following transfection with si-DSCAM-AS1, which was restored by miR-204 inhibition in BGC-823 and SGC-7901 cells (Figure 5(C)). DSCAM-AS1 exerted its effects on GC cells via miR-204/TPT1 axis. A: The expression of miR-204 was inhibited in GC cells after miR-204 inhibitors transfection. B: miR-204 inhibition partially reversed the suppressive effects of DSCAM-AS1 knockdown on GC cells proliferation. C: The protein expression of TPT1 in BGC-823 and SGC-7901 cells after DSCAM-AS1 knockdown and/or miR-204 inhibition. Data are mean ± SD. *p < 0.05, **p < 0.01. DSCAM-AS1: Down syndrome cell adhesion molecule-anti-sense 1.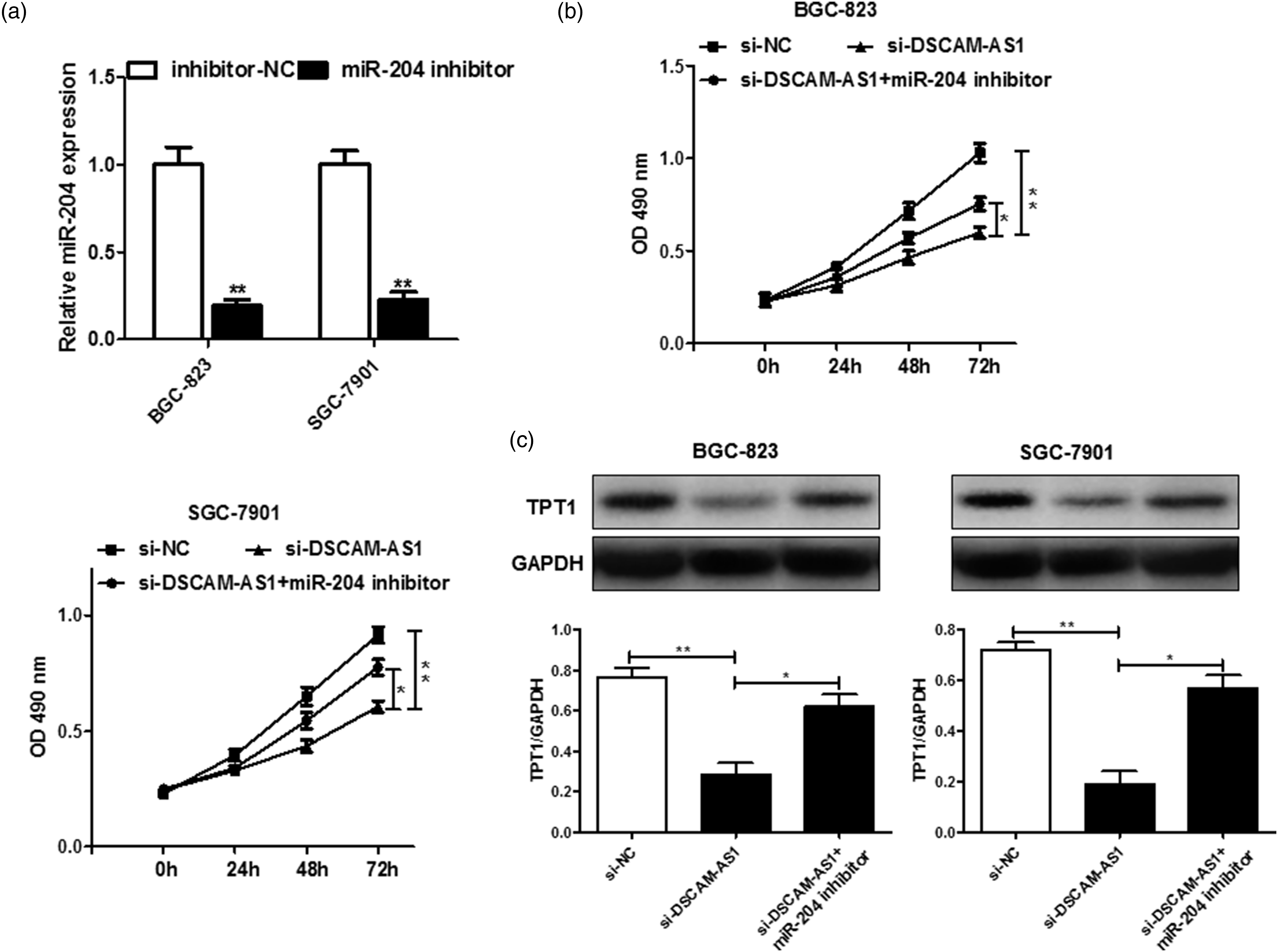
DSCAM-AS1 deletion hindered GC tumor growth in vivo
To further explore the role of DSCAM-AS1 in vivo, SGC-7901 cells pre-treated with si-DSCAM-AS1 were injected into nude mice. The tumor volume was markedly increased in the si-DSCAM-AS1 group (Figure 6(A)), consistent with the tumor weight results (Figure 6(B)). Our qRT-PCR analysis results showed that the DSCAM-AS1 and TPT1 levels were dramatically downregulated, whereas miR-204 expression was upregulated, in the si-DSCAM-AS1 group (Figure 6(C)). Cells stably transfected with si-DSCAM-AS1 were used to establish xenograft models. A: Tumor volume was calculated every 4 days. B: Mice were killed and tumor weight was examined in each group. C: The expressions of DSCAM-AS1, miR-204, and TPT1 were detected in two groups by quantitative reverse transcription polymerase chain reaction. *p < 0.05, **p < 0.01. DSCAM-AS1: Down syndrome cell adhesion molecule-anti-sense 1.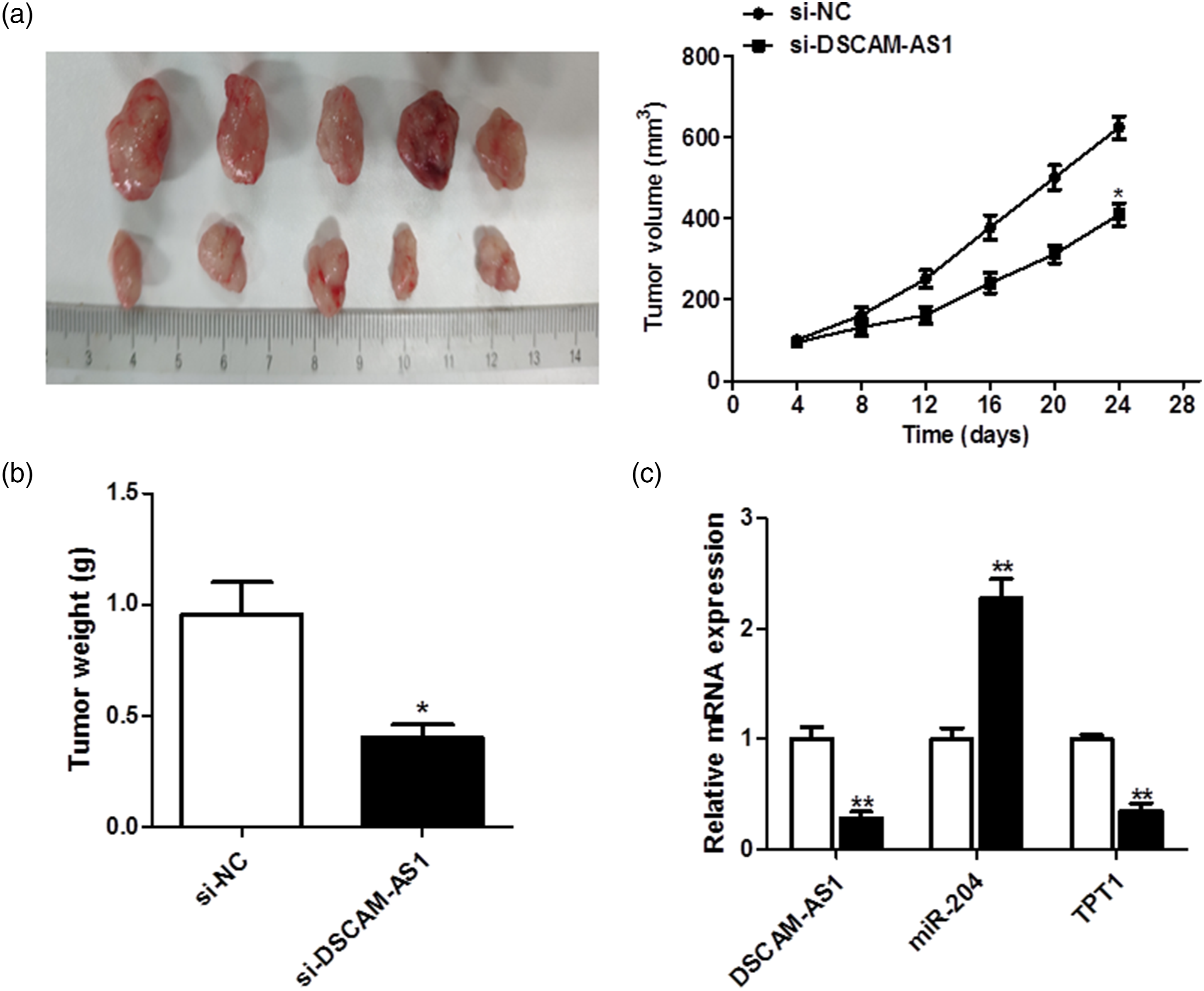
Discussion
Since its discovery, increasing numbers of studies have reported that lncRNAs play key roles in the occurrence and development of various cancers, including GC. 12 Therefore, the relationships between lncRNA and human diseases, especially tumors, have received increasing research attention. Multiple studies have shown that lncRNA is abnormally expressed in human cancer, in association with genes related to cancer promotion or tumor suppression. For example, increased lncRNA SNHG8 expression in GC tissues was shown to be significantly associated with shorter survival times, with higher SNHG8 expression in non-survivors than in survivors (p = 0.002). 13 LncRNA MEG-3 was downregulated in GC and lncRNA-MEG3 transfection suppressed tumor growth mainly by decreasing the expression of vascular endothelial growth factor and increasing the expression of Bcl2. 12 Also, lncRNA GHET1 was determined to be upregulated in GC. The over-expression of this lncRNA correlates with tumor size, tumor invasion, and poor survival and GHET1 enhances the c-Myc mRNA stability and its expression to regulate the proliferation in GC cells. 14 The carcinogenic effect of lncRNA DSCAM-AS1 in tumors was first discovered and reported by Niknafs et al. 15 : In breast cancer, lncRNA DSCAM-AS1 is related to estrogen receptors, and it interacts with hnRNPL to participate in the occurrence and development of breast cancer and mediates resistance to tamoxifen. Later, studies have also proven through bioinformatics analysis and experimental verification that the abnormal expression of lncRNA DSCAM-AS1 in breast cancer can promote its abnormal proliferation and avoid apoptosis, 16 and affect the prognosis of breast cancer patients by participating in cell cycle regulation. 17 Liao et al. 18 reported on the carcinogenic effect of lncRNA “DSCAM-AS1 in non-small cell lung cancer. lncRNA DSCAM-AS1 promotes its invasion and metastasis by upregulating the expression of BCL11A. lncRNA DSCAM-AS1 can also participate in the occurrence and development of melanoma, liver cancer and colon cancer through an endogenous competitive RNA mechanism.18-20 The research results of Li et al. 21 suggest that lncRNA DSCAM-AS1 can participate in the invasion and metastasis of ovarian cancer by upregulating SOX4. The biological role of lncRNA and DSCAM-AS1 in cervical cancer was reported by Liang et al. 22 in 2020: The expression of lncRNA and DSCAM-AS1 in cervical cancer cells was significantly upregulated. Inhibiting the expression of lncRNA DSCAM-AS1 can prevent cell growth, invasion, and migration, suggesting that lncRNA DSCAM-AS1 can promote the development of cervical cancer. However, the clinical significance, biological role, and effects of DSCAM-AS1 in GC remain to be elucidated.
In the present study, we determined that DSCAM-AS1 expression levels were increased in clinical GC tissue specimens and GC cells, suggesting that DSCAM-AS1 is a molecular marker of GC. Silencing DSCAM-AS1 inhibited the migration and proliferation of GC cells, suggesting that it plays a key role in GC tumor suppression, as shown in our MTT, flow cytometry, and Transwell assay results. Silencing DSCAM-AS1 inhibited GC growth through miR-204 upregulation and TPT1 signal inactivation in vivo. Thus, DSCAM-AS1 was involved in the progression of GC. Accumulating evidence indicates that lncRNAs mediate the development of many cancers through competition for miRNA sponges.23,24 In the present study, we used a dual-luciferase reporter gene assay to demonstrate that DSCAM-AS1 targeted the 3′-UTR of miR-204 in GC cells, and that miR-204 expression was negatively correlated with DSCAM-AS1. We also showed that miR-204 was reduced and that TPT1 was targeted by miR-204 in GC cells. Furthermore, our rescue assay results verified that DSCAM-AS1 mediated the miR-204/TPT1 axis to affect GC. Mechanistically, the loss of miR-204 reversed the inhibitory effects on GC cell proliferation induced by silencing DSCAM-AS1 expression. TPT1 protein levels were significantly decreased when transfected with si-DSCAM-AS1, and this decrease was partly abolished after the loss of miR-204. Thus, our results demonstrate that DSCAM-AS1 influences GC through regulation of the miR-204/TPT1 axis.
In summary, we confirmed that DSCAM-AS1 and TPT1 were markedly increased in GC tissues and cells. Our assay results demonstrated that DSCAM-AS1 was involved in GC progression via mediation of the miR-204/TPT1 axis. Together, our findings may offer a promising and novel therapeutic target for the prevention of GC.
Footnotes
Author’s Contribution
Nan Wang, and Ying Yang authors equally worked this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by Social Development Projects of Shaanxi Province (No. 2019SF-004) and National Natural Science Foundation of China (82072722).
Ethical approval
All experiments were preformed according to the Ethics Committee of Tangdu Hospital, the Air Force Medical University.
Informed consent
The consent to participate was obtained from patients.
Consent for publication
The informed consent obtained from study participants.
