Abstract
Sustained release nanoformulations of second line antitubercular drugs levofloxacin and ethionamide had shown promise in pharmacokinetics and acute and sub-acute toxicity studies. The present study evaluated the clastogenicity potential of the nanoformulations of these antitubercular agents. Clastogenicity was evaluated by (a) in vitro micronucleus assay (b) in vivo micronucleus assay in Swiss albino mice and (c) sister chromatid exchange (SCE) in CHO cell lines. Ethionamide and levofloxacin loaded nanoparticles were 312 ± 64 nm and 245 ± 24 nm in size respectively and drug encapsulation was 35.2 ± 3.1% w/w and 45.6 ± 9.4% w/w, respectively. The frequency of MN-NCE/1000 NCE and MN-PCE/1000 PCE were significantly reduced in mice treated with ethionamide nanoparticle (3.5 ± 0.9, 13.8 ± 16.68) and levofloxacin nanoparticles (5.6 ± 2.7, 16.7 ± 12.7) compared to the mice treated with free ethionamide (11.5 ± 4.1, p = 0.23 and 45.19 ± 19.21, p = 0.38) and free levofloxacin (14.7 ± 1.88, p < 0.0001 and 54.6 ± 18.1, p = 0.0017), respectively. For in vitro, micronucleus assay frequencies of micronuclei per thousand bi-nucleated cells (MN-BN/1000 BN) was 188.3 ± 20.20 and 148 ± 20.42 for ethionamide and levofloxacin nanoparticles as compared to 232.6 ± 16.04 (p = 0.52) and 175 ± 5.56 (p = 0.45) for free ethionamide and levofloxacin, respectively. The average number of SCE per cell for nanoformulation of ethionamide were not different from that of free drug (4.9 ± 0.51 vs 4.1 ± 0.55, p = 0.86). The SCE per cells were not significant difference for nanoformulation of levofloxacin (2.33 ± 1.36 vs 5.46 ± 0.25, p = 0.88). In vitro and in vivo assays have shown relatively less clastogenic potential of equivalent dose of ethionamide nanoparticles as compared to the conventional formulation.
Keywords
Introduction
Tuberculosis is the leading cause of death among infectious diseases. 1 Although existing anti-tubercular drugs (ATDs) can cure majority of tubercular patients, it claims around two million lives annually worldwide. 2 Drug compliance remains a major problem with long course of ATDs and the non-compliance to therapy could result in emergence of other advanced forms such as multidrug resistant tuberculosis (MDR-TB) and extensively drug resistant tuberculosis (XDR-TB). 3
Ethionamide and levofloxacin are second line ATDs widely used in the treatment of MDR TB. However, long treatment duration and daily dosing of these drugs are known to adversely affect patient compliance resulting in treatment failures and development of drug resistant strains of M. tuberculosis. Moreover, these second line anti-tubercular drugs are expensive, more toxic and less effective. 4 As very few classes of drugs have been developed in last several years for MDR-TB, formulating the already existing anti-tubercular drugs into sustained release formulations permitting lesser dosing frequency could be desirable strategy for improving patient compliance. 5
Previous work published from our laboratory had demonstrated sustained release profile both in-vitro and in-vivo of levofloxacin and ethionamide for nearly a week in drug loaded nanoparticles. We developed sustained delivery systems to increase compliance in TB patients. These nanoformulations were found safe in repeat dose toxicity studies. 6,7 However, clastogenicity potential of these formulations was not evaluated.
Evaluation of genotoxicity potential of nanoformulations has achieved only a limited much attention. 8 Genotoxicity potential of silver nanoparticles has been evaluated in various studies. 9,10 Although genotoxicity is a matter of concern because of the diverse size, structure, surface area, shape, charges and other characteristics of the formulation. Some of these characters, like size and charge may alter the toxicity at the genome level. 11 Levofloxacin has shown toxicity in vitro sister chromatid and micronucleus assay in human peripheral lymphocytes whereas ethionamide have shown in vitro clastogenic activity in sister chromatid exchange assay. 12,13,14 As stated above, both nanoformulations developed in our lab were found safe in preclinical toxicity. However, clastogenicity potential of these formulations was not evaluated. Evaluation of genotoxicity including clastogenicity is an important step required for translation of the product. In the current study, we aimed to evaluate the clastogenic potential of ethionamide and levofloxacin loaded poly-lactic-co-glycolic acid (PLGA) nanoparticles.
Materials and methods
Materials
Poly (DL-lactide-co-glycolide) (PLGA) (50:50 Resomer) was purchased from Birmingham Polymers, Inc (Birmingham, AL). Ethionamide (ETH) was a gift from Macleods Pharmaceutical (Mumbai, India). Levofloxacin (LF) was purchased from Sigma Aldrich (MO, USA). Poly-vinyl alcohol (PVA; MW 30,000 to 70,000 Da, 87–89% hydrolysed) and 5-bromodeoxyuridine were purchased from Sigma Chemicals, Sigma Aldrich (MO, USA). Acetonitrile (ACN; HPLC grade), dichloromethane (DCM) and methanol were purchased from Merck. Cyclophosphamide (CP), mitomycin C (MMC), cytochalsin B and colcemid were purchased from MP Biomedicals, LLC, France. Acridine orange, Giemsa stain and phosphate buffer solution (PBS) were purchased from Himedia, India. S9 metabolic fraction was purchased from BD Bioscience (USA).
Animals
Swiss albino mice of either sex weighing 20–30 g were obtained from central animal house of our institute. These were kept under controlled conditions and 12/12 h light/dark cycle. Animals were provided pellet diet and water ad libitum. The study was approved by Institutional Animal Ethics Committee and experiments were done in accordance with the CPCSEA guidelines.
Cell lines
Chinese Hamster ovary (CHO) cell lines were purchased from American Type Culture Collection (ATCC, Rockville, USA). This cell line was grown in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 6% FBS, 1% penicillin-streptomycin, 1% Hepes buffer 1 M, 1% non-essential amino acids and 1% l-glutamine. The maintenance medium for CHO cells was described above with 2% FBS.
Preparation of drug loaded PLGA nanoparticles
Both ethionamide and levofloxacin loaded PLGA nanoparticles were prepared by modified emulsion-diffusion-evaporation method and standardized in our lab. 6,7 In brief, ethionamide (20 mg) and PLGA (20 mg) were dissolved separately in methanol (800 µl) and DCM (7 ml), respectively. Drug solution was mixed in PLGA solution to make a single organic phase. The drug polymer solution was added slowly drop wise into aqueous phase (14 ml, 0.5% w/v PVA) and sonicated for 3 min at 20 kHz (20% amplitude) with the help of probe sonicator (Misonix, Farmingdale, NY; Model XL 2020). The resulting mixer was moderately stirred overnight to completely evaporate the volatile organic solvent. The pellet was isolated by centrifugation at 15,000 rpm for 20 min and washed with 5% v/v methanol once and twice with double distilled water (DDW). The suspension of nanoparticle in DDW were frozen at −20°C and lyophilized for 48 h in a lyophilizer (0.070 mbar vacuum; condenser temperature of −55°C) to obtain fine preparation of nanoparticles.
For levofloxacin loaded PLGA nanoparticles, levofloxacin and polymer were dissolved in DCM (1 ml) which was then drop wise added to aqueous phase (2 ml PVA) to form emulsion. After 10 min of magnetic stirring emulsion was stirred with probe sonicator (Misonix, Farmingdale, NY; Model XL 2020) set at level of 20% amplitude. Resulting emulsion was then added drop wise into 0.3% (w/v) PVA solution with constant stirring. The organic phase was evaporated off in 4–5 hrs and the nanoparticle pellet was isolated by centrifugation at 15,000 rpm for 20 minutes followed by three times washing with double-distilled-water. Prepared nanoparticles were characterized for the percentage of drug incorporated during nanoparticles preparation was determined using following procedure. In method, 100 µl of re-dispersed pellet was dissolved in 10 ml of 5% w/v Sodium Dodecyl Sulphate (SDS) in 0.1 M NaOH at 60°C for 10 min to accelerate hydrolysis of PLGA. The clear solution was assessed for drug content by HPLC.
Drug estimation
Ethionamide was analysed on a reversed phase C18 column (Cosmosil 5C18-MS-II from waters; 250 mm × 4.6 mm; 5 µm particles size) by using 0.02 M disodium phosphate buffer: acetonitrile (75:25) as the mobile phase (pH adjusted to 6.8) at a flow rate of 1.5 ml/ min, injection volume of 20 µl, retention time of 5.6 min and detection wavelength 254 nm. Levofloxacin was analysed on same column conditions with mobile phase consisting water and acetonitrile (80:20, v/v) with 0.3% of tri-ethylamine and pH adjusted to 3.3 with ortho-phosphoric acid at flow rate of 1 ml/min and injection volume of 20 µl. Size, charge and poly-dispersity index (PDI) was done as previously described. 6,7 Briefly, particle size, zeta potential and PDI was assessed by Delsa Nano C, Beckman Coulter, Inc. The lyophilized powder was diluted with double-distilled water (0.5 mg/mL). The size of nanoparticles was determined by dynamic light scattering technique (DLS). Zeta potential was estimated on the basis of electrophoretic mobility under an electric field. The PI of the particles was also obtained.
In-vitro studies
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) assay
The cytotoxicity of test compound was determined in vitro using CHO cells using the MTT test.
15
5 × 103 cells were seeded into each well of a 96-well plate. The medium was replaced after 24 h with 100 µl of fresh medium and 100 µl of test drug or PBS (for negative control). Plates were incubated for 24 h with the test compounds (10 µM to 1 nM) or control substances at 37°C in a humidified CO2 atmosphere. Then, 20 µl of 5 mg/ml of MTT was added to each well for 4 hr incubation. After that, the medium was discarded and replaced with 150 µl DMSO. Optical density (OD) data of converted dye was measured at a wavelength of 570 nm using 96-well plate reader. Percentage viability was calculated for treated cells using the following relationship:
In vitro micronucleus assay in CHO cell lines
The assay was done using the method described by He et al. 16 Briefly, for the micronucleus assay, 5 × 104 CHO cells were seeded into each well of a 6-well plate over a sterile glass cover-slip. One day after the seeding, the medium was replaced with 500 µl fresh complete medium containing PBS (negative control), test compound (with or without S9 fraction, Ethionamide 1 µM and levofloxacin 1.25 µM drug equivalent), mitomycin C (MMC, 0.5 μg/ml, positive control for the assay without S9) and Cyclophosphamide (CP, 12 μg/ml positive control for the assay with S9 fraction) for 4 hr. After treatment, the cells were rinsed with serum-free medium and replenished with complete medium containing 3 μg/ml of cytochalsin B. Then cells were further incubated for an additional 19 hrs. At the end of the incubation, the medium was removed and the cultures were replenished with complete medium and returned to the incubator for 1 h. Then cytochalsin B free medium was removed by gently inverting the culture slides and then blotting them on absorbent paper. The cells then washed with PBS and were incubated in 0.075 M hypotonic potassium chloride for 3–5 minutes. After the hypotonic treatment, the cells were fixed in methanol for approximately 20 minutes. The dishes were dried in air and stained for 20 min with fresh 5% Giemsa’s solution. Micronuclei (MN) were visualized under microscope at 100X magnification. Micronuclei in bi-nucleated (BN) cells were counted per 1000 bi-nucleated cells.
Sister chromatid exchange
The sister chromatid exchange (SCE) is designed to detect the clastogenic potential of the test agents by determining the number of exchanges that have occurred between the sister chromatid of a chromosome. The method of He et al. was used. 16 Briefly, in the assay, CHO cells were seeded into six-well plates at a cell density of 2 × 105 cells for 24 h, then cells were rinsed with fresh medium and cultured in the presence of the test compound (ethionamide 1 µM and levofloxacin 1.25 µM drug equivalent) and 5-bromodeoxyuridine (10 μg/ml) for a further period of 24 hours to allow two cell divisions. For the negative and positive controls, AAPBS and MMC were used instead of test samples, respectively. In the system with S9 metabolic activation, cells were treated with each nanoformulation for 2 h with S9 mix (10%). PBS and CP were used as the negative and positive control, respectively. After treatment, the medium containing nanoparticles and controls were removed. Cells were rinsed twice with PBS and cultured in fresh medium containing 5-bromodeoxyuridine (10 μg/ml) for 24 h. Colcemid (0.2 μg/ml) was added 2 h before chromosome preparation for metaphase arrest. Slides were prepared and stained with 4% Giemsa’s solution diluted with sorenson’s phosphate buffer.
In vivo micronucleus assay
Animal ethics committee approval was obtained for animal experiments. Experiments were performed on adult male Swiss albino mice, 5–6 weeks. Standard laboratory animal feed and water were given ad libitum. Animals were taken for the evaluation of clastogenicity in five groups: Ethionamide nanoformulation group (720 mg/kg, given every 7th day for 14 days) because of the sustained release profile observed in PK studies free ethionamide group (720 mg/kg, every day for 14 days), levofloxacin nanoformulation group (1728 mg/kg, given every 7th day, for 14 days) because of the sustained release profile observed in PK studies Free levofloxacin (1728 mg/kg, every day, for 14 days) and cyclophosphamide, as positive control group (50 mg/kg, every day, for 14 days). Six animals were taken in each group.
Peripheral blood was collected from the sub mandibular vein and smear was prepared on pre-cleaned slides. The smear was dried at room temperature and was fixed in absolute methanol for 5 minutes. After fixation slides were stained with acridine orange and washed twice with phosphate buffer (pH 6.8). Slides were observed under oil immersion objective using fluorescent microscope. The results were expressed ratio of micronucleated normochromatic erythrocytes (MNNCE) to 1000 normochromatic erythrocytes (NCE), ratio of micronucleated polychromatic erythrocytes (MNPCE) to 1000 polychromatic erythrocytes (PCE). Higher value of this ratio signifies the clastogenic potential of the compound. 17
Statistical analysis
The data from the in vitro MN and in vivo MN assay were expressed as mean ± SD. SCE assay were expressed as the mean ± SD of three independent experiments. At least 50 cells per experiment were examined. Results of the micronucleus assay and sister chromatid exchange between the groups was analysed by ANOVA followed by Tukey’s post hoc test. p value < 0.05 were considered statistically significant.
Results
Characterization of nanoparticles
The candidate drugs were evaluated for their characterization parameters like particle size, zeta potential, poly dispersity index, drug loading efficiency and drug entrapment efficiency. The optimized drug-polymer ratio was 1:1 with a PVA concentration 0.3% w/v for levofloxacin and 0.5% w/v for ethionamide nanoparticles. The average particle size for levofloxacin and ethionamide nanoparticles were 245 ± 24 nm and 312 ± 64 nm respectively. The encapsulation efficiencies for levofloxacin nanoparticle batches ranges from 32.4 to 45.6% w/w and for ethionamide nanoparticle batch encapsulation efficiency ranges from 25.7 to 37.12% w/w (Table 1).
Physiochemical characterization of candidate drugs nanoparticles.

Cytotoxicity determination for candidate drugs
The cytotoxicity of these test compounds was determined using MTT assay in CHO cells. The MTT assay was used to evaluate the effects of candidate drugs on cell proliferation and cell viability of CHO cell. The MTT is metabolized by mitochondrial dehydrogenase enzyme (NADPH) in metabolically active cell and convert it into insoluble formazan crystals which display purple colour. It was done to select the safe dose for further experiments. We have selected the concentration of each candidate drug at which at least 80% cells were viable. Based on the results of MTT assay of free ethionamide and free levofloxacin, 1.25 µM and 1 µM doses were considered safe for further experiments (Figure 1A and 1B). For the nanoformulation, the corresponding concentrations of free drugs were obtained based on drug loading of nanoparticles.

Cell viability assessment for candidate drugs; A: Levofloxacin (LF), B: Ethionamide (ETH); The cell viability data are presented as mean ± S.D.
In vitro micronucleus assay in CHO cells
A micronucleus is a whole lagging chromosome or acentric chromosomal fragment which is not integrated in any daughter nuclei during metaphase/anaphase of cell division. A clear and specific correlation between micronuclei and chromosomal damage has been reported. 17 For assessing chromosomal damage, number of micronucleated binucleated cells per 1000 binucleated cells (MN-BN/1000 BN) was calculated. Frequency of MN-BN cells (MN-BN/1000 BN) was highest in positive control group i.e. cyclophosphamide (260.6 ± 54.12) and mitomycin-C (266 ± 27) and lowest in negative control (79 ± 5.56). All the treatment groups showed a significant difference when compared to positive control of cyclophosphamide and mitomycin (p < 0.005). Frequency of MN-BN cells were higher in free ethionamide (p = 0.52) and free levofloxacin (p = 0.45) groups compared to the nanoformulations of these two drugs at same concentration in presence and absence of S9 fraction. But the difference was not statistically significant (Table 2 and Figure 2).
Number of micronuclei (MN) in 1000 bi-nucleated (BN) cells for positive control, negative control and test drugs with and without metabolic activation (S9 fraction); Values are presented as mean ± SD.
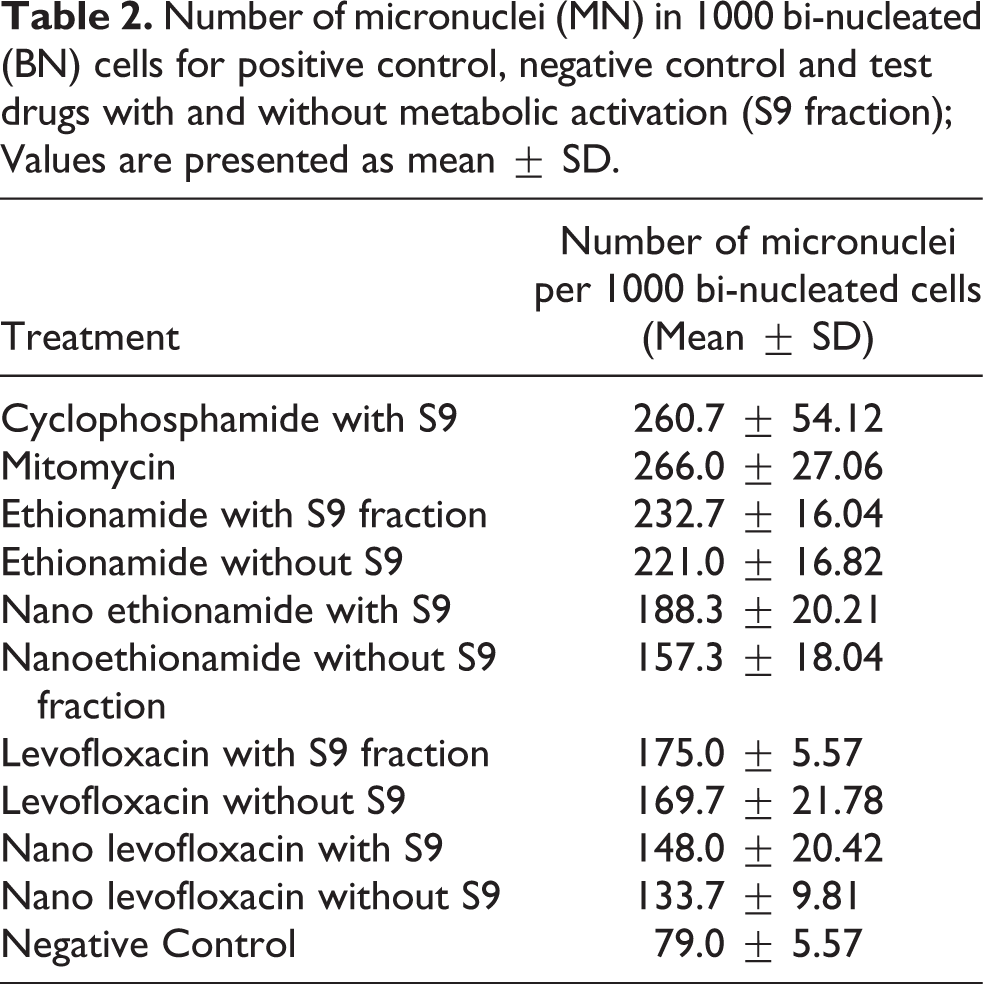

In vitro assessment of frequency of MN per 1000 bi-nucleated cells for different groups. Chinese hamster cells (CHO) were treated with free and nanoformulation of ETH and LF. DNA damage was evaluated by micronucleus assay. Treatment of free and nanoformulation of both drugs was given in presence (+S9) and in absence (−S9) fraction. Cyclophosphamide (CP) was used as positive control for the treatment groups containing S9 fraction. Mitomycin-C (MMC) was used as positive control for the treatment groups without S9 fraction. ANOVA was used to compare the means followed by multiple comparisons (Tukey’s post hoc test). A: Levofloxacin (LF); B: Ethionamide (ETH). #* p < 0.0001 for comparison with cyclophosphamide. $* p < 0.0001 for comparison with mitomycin. ns: no significant difference.
Mouse micronucleus assay
In mouse micronucleus assay, no deaths occurred prior to scheduled sacrifice. No significant drug related changes (body weight and hair colour) were observed. The frequency of MN-NCE/1000 NCE and MN-PCE/1000 PCE were significantly reduced in mice treated with ethionamide nanoparticle (3.5 ± 0.9 and 13.8 ± 16.68) and levofloxacin nanoparticles (5.6 ± 2.7 and 16.7 ± 12.7) compared to the mice treated with free ethionamide (11.5 ± 4.1, p = 0.23 and 45.19 ± 19.21, p = 0.38) and free levofloxacin (14.7 ± 1.88, p < 0.0001 and 54.6 ± 18.1, p = 0.0017), respectively (Table 3). The micronuclei frequency produced by the positive control, cyclophosphamide was significantly higher (p < 0.0001) than the respective treatment groups (22.9 ± 11.82) (Figure 3 and 4).
Ratio of MNPCE/1000 PCE and ratio of MNNCE/1000 NCE for positive control, free drugs and nanoformulation of levofloxacin and ethionamide; PCE: Polychromatic cells; MNPCE: Micronucleated polychromatic cells; MNNCE: Micronucleated normochromatic cells.


Microscopic analysis of micronucleus formation induced by free and nanoformulation of the levofloxacin in mice indicating MNPCE per 1000 PCE and MNNCE per 1000 NCE. ns: no statistically significant

Microscopic analysis of micronucleus formation induced by free and nanoformulation of the ethionamide in mice indicating MNPCE per 1000 PCE. ns: no statistically significant
Sister chromatid exchange assay
Sister chromatid exchange is used to measure the clastogenic properties of a test compound. Cells which divide in the presence of Bromodeoxy uridine (BrdU), replace thymidine in chromosomes and when active mutagens are present, an exchange between sister chromatid can be expected which can be visualized by comparing arms of chromatid. BrdU positive strands appear less intense than thymidine positive strands in the cells cultured for 2 cell cycle. Any event showing an exchange between dark and light intensity fluorescent plus Giemsa stained chromatid strands were counted as a positive clastogenic event. Clastogenic events were counted in 30 metaphases per treatment, and each treatment was performed in triplicate. Data are presented as a mean number of clastogenic event normalized on negative vehicle control. None of the nanoformulation showed any significant change in the number of the clastogenicevents as compared to respective free drug, i.e. ETH (p = 0.86) or LF (p = 0.88) (Table 4 and Figure 5). In the case of LF with and ETH without S9 fraction showed a lower number of such events. Overall, none of the nanoformulations showed any added clastogenic behaviour aside from native behaviour of the drug itself.
Numbers of sister chromatid exchange (SCE) per cell observed in the CHO cells treated with positive control or free or nanoformulation of candidate drugs with (+S9) or without (−S9) the addition of S9 mix. Data were shown as mean ± SD.

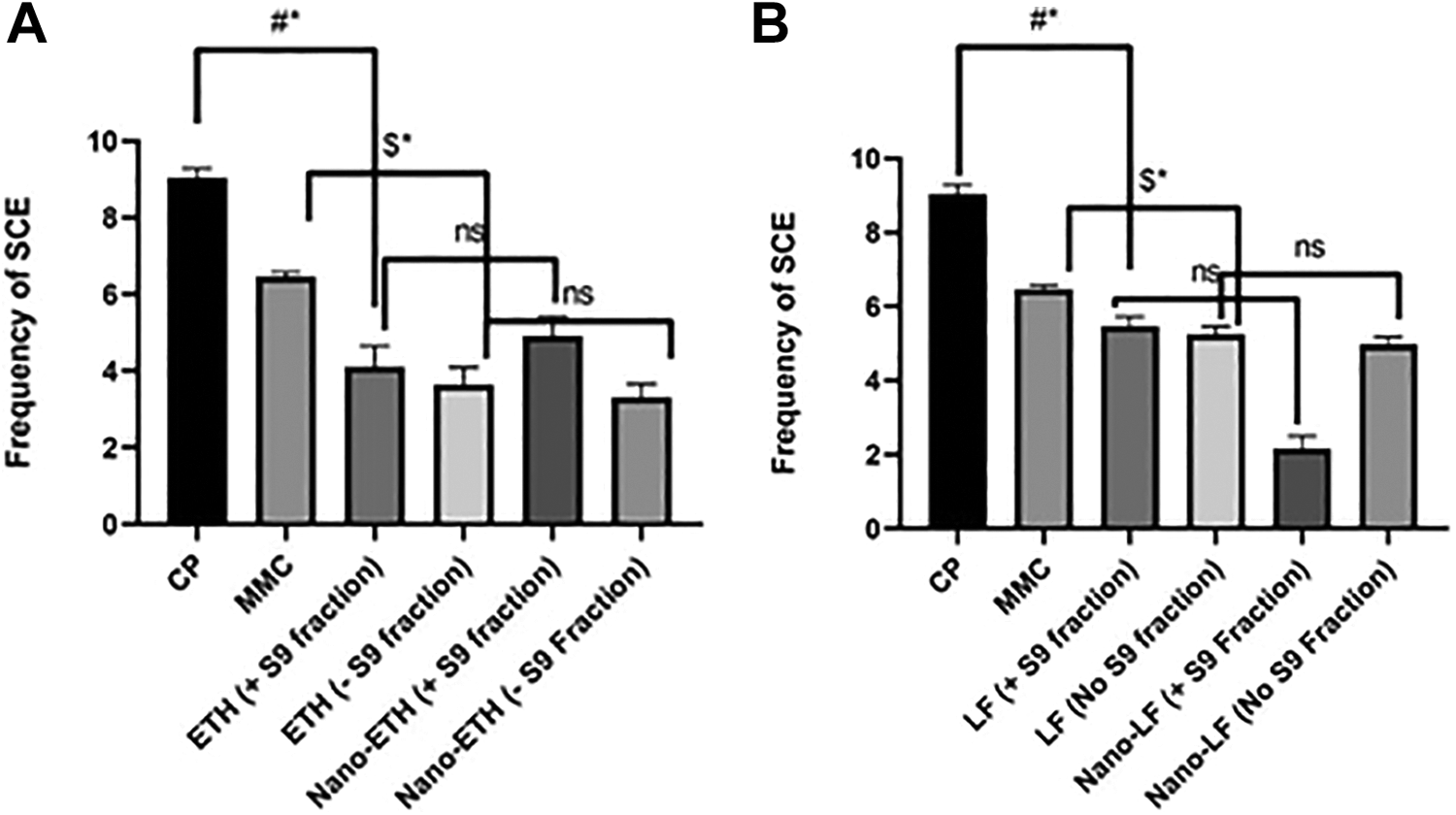
Numbers of sister chromatid exchange (SCE) per cell observed in the CHO cells treated with positive control or free or nanoformulation of candidate drugs with (+S9) or without (−S9) the addition of S9 mix (n = 3, mean ± SD). P < 0.05: Statistically significant difference. ns: no significant difference. A: Levofloxacin (LF); B: Ethionamide (ETH). #* p < 0.0001 for comparison with cyclophosphamide. $* p < 0.0001 for comparison with mitomycin. ns: no significant difference.
Discussion
In the present study, we were able to demonstrate that the PLGA based nanoformulation of levofloxacin and ethionamide did not have more clastogenic potential than the respective free drugs. There is dearth of studies assessing the genotoxic potential of drug encapsulated nanoparticles. Clastogenic potential as a component of genotoxicity is an important aspect in nanoformulation as with a reduction of particle size, its surface area increases, which can increase reactivity in the cells. 18 There are various assays which can test the genotoxic potential of carcinogens. Ames test (Salmonella reverse mutation assay), alkaline comet assay and micronucleus assay have been used in various studies on nanoformulations. 19 According to ICH guidelines, bacterial, in vitro and in vivo genotoxicity evaluation must be considered for any pharmaceuticals as part of its toxicity evaluation. In this study, we have reported in-vitro and in-vivo micronucleus assay and in vitro sister chromatid exchange. Micronucleus assay assesses both the chromosomal and genetic mutation in the dividing cells. Clastogenic effect can be evaluated by sister chromatid exchange. We have not reported evaluation in bacterial cells (Salmonella reverse mutation assay). There is evidence that states that due to differences in membrane structure and composition of bacteria compared to mammalian cell membranes, exact genotoxic potential of the nanoparticles cannot be evaluated. 20 Doak SK et al., mentioned that nanomaterials which have shown negativity for mutagenicity in the Ames test, have largely found to be positive in other genotoxic tests. 21 Other assays that have been used to evaluate genotoxic potential of an nanomaterial are Annexin V staining assay, Comet Assay, TdT mediate dUTP-biotin nick end labelling assay or interpretation of morphological changes. 22 However, these tests are not conventionally tested in genotoxic battery recommended by regulators for investigational new drug application. 23,24
For SCE and in vitro micronuclei test, no significant difference was noted. However, in the in vivo test a significant reduction in no. of micronuclei/ 1000 cells was noted. This would need further evaluation. Some possible explanation could be that incorporating the drugs in to the PLGA nanoparticles have decreased the clastogenic potential. Similar kind of finding have been observed in various other studies. 25,26 However, this observation would need to be examined with simultaneous measurement of drug concentrations in the systemic circulation. At the same time, it is reassuring that the number of chromosomal aberrations and number of micronuclei were not more than conventional treatment.
There is a debate among the scientific community and drug regulators regarding the biological effects and associated side effects of nanoparticles. 27 Nanoparticles having a dimension of size (1–100 nm) come in contact with the biological systems and have various chemical and physical interaction with cellular macromolecules like proteins. 28 Park et al., have shown the effect of silver (Au) nanoparticles at cellular level. 9 They have shown that smaller particles (20 nm) showed mutagenic potential by increasing frequency of mutation in MEF-LacZ cells but, larger particles (80 and 110 nm) did not have any significant changes in the DNA of the cells. In contrast, in a study done by Lebedova et al., had shown that silver nanoparticles induced DNA breakage in comet assay without any obvious size difference (5, 20, and 50 nm). The authors have also done genotoxic studies on gold (Au) and platinum (Pt) nanoparticles, where 5 nm Au NPs and 50 nm Pt NPs have shown breakage of DNA strands. 11 Evidence suggests that smaller nanoparticles account for genotoxic potential as they interact with cellular proteins and can induce conformational changes. 29 For our study, we used nanoparticles of size greater than 200 nm and these were not found to have clastogenic potential for the entire battery of tests evaluated. Moreover, in the studies mentioned above, metallic nanoparticles were evaluated. However, we have developed PLGA nanoparticles of drugs already in use for many years.
We have developed nanoparticles by using PLGA polymer (520 H). This polymer has been used widely for delivering drug into the biological systems. It is an US-FDA approved elastomeric copolymers for drug delivery. 16 Extensive toxicity evaluation of PLGA has not shown alteration in the histopathological evaluation or tissue damage. 10
The success of tuberculosis treatment is associated with compliance to drug therapy. Our formulations which give sustained levels of drugs for more than 5 days would be to be given for 18 months in order to meet the requirement of optimal duration of treatment. This extended duration of treatment may add to the potential of clastogenic changes. Therefore, doing clastogenic evaluation becomes a part of the new drug development process if delivering a drug through nanoparticles.
Previous studies 6,7 have demonstrated that the nanoparticles of ethionamide and levofloxacin were well distributed in the organ containing high number of phagocytosing cells, such as liver, spleen, and lung. Despite the distribution into these sites, an increase in clastogenic potential of the developed formulations was not evident.
Conclusion
In conclusion, we have evaluated our developed nanoformulation of levofloxacin and ethionamide for clastogenic potential through battery of test which evaluate chromosomal and genetic mutation. We found that encapsulating drug into nanoformulation did not account for any clastogenic potential, hence this PLGA based nanoformulation encapsulating second line anti TB drugs has a potential for evaluation in human studies.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Department of Biotechnology (DBT), New Delhi, India.

