Abstract
Ethephon, a member of the organophosphorus compounds, is one of the most widely used plant growth regulators for artificial ripening. Although million pounds of this chemical is being used annually, the knowledge regarding its molecular toxicity is yet not sufficient. The purpose of this study was to evaluate the potential developmental toxicity of ethephon using embryonic stem cell model. The mouse embryonic stem cells (mESCs) were exposed to various concentrations of ethephon and the viability, cell cycle alteration and changes in the gene expression profile were evaluated using high-throughput RNA sequencing. Further, the effect of ethephon on neural differentiation potential was examined. The results showed that ethephon at noncytotoxic doses induced cell cycle arrest in mESCs. Gene ontology enrichment analysis showed that terms related to cell fate and organismal development, including neuron fate commitment, embryo development and cardiac cell differentiation, were markedly enriched in ethephon-treated cells. Neural induction of mESCs in the presence of ethephon was inhibited and the expression of neural genes was decreased in differentiated cells. Results obtained from this work clearly demonstrate that ethephon affects the gene expression profile of undifferentiated mESCs and prevents neural differentiation. Therefore, more caution against the frequent application of ethephon is advised.
Keywords
Introduction
During the past century, the farming and crop production systems have changed speedily because of emerging demands for natural resources. The increasing rate of human population growth, changes in food consumption pattern and modifications in the ecosystem have led to new strategies, such as applying more pesticides, artificial fertilizer and ripening chemicals to fulfil the increasing demand for agricultural products. On the other hand, these strategies have increased human exposure to toxic chemicals and raised the urgent need for precise toxicology testing and more detailed studies on chemicals side effects. 1
Organophosphorus compounds (OPs) are widely used pesticides that primarily known for their neurotoxic effects during the prenatal and postnatal development period. 2,3 Exposure to OPs normally occurs through the breathing of vapours or via skin in the environment. 4 OPs could potentially pass through the placenta and blood–brain barrier. 5,6 They are potent to bind to cholinesterase enzyme and block the acetylcholine degradation. Accumulation of acetylcholine causes the overstimulation of the postsynaptic nervous system that affects the nervous message transmission. 3 The use of OPs has also been correlated with increased risk of multiple cancers. 7,8
Ethephon (C2H6ClO3P), as an OP, is presently used commonly as a plant growth regulator for artificial ripening of crops, vegetables and fruits. 9 Metabolic studies on rats showed that 78–84% of ethephon is promptly absorbed in the gastrointestinal tract within 120 h and is mainly distributed in the liver and kidney. However, the larger amount is primarily excreted in urine (50–60%) and faeces. 10
Experiment on rats showed some degenerative changes of hepatocytes, inflammatory and increment of Kupffer cells after consumption of 200 mg/kg/day of ethephon for 14 days. 11 Few in vitro studies have also demonstrated the cytotoxic and mutagenic effect of ethephon. 12 –14
It has been revealed that the early life stage and embryonic development are more sensitive to environmental toxicants than adult age. 15 –17 Considering the ability of environmental toxicants to pass through the placental barrier, maternal exposure could pose a great risk in embryo/fetal health, resulted in malformation or genetic deficiencies in adults. Developmental neurotoxicity (DNT) 12 of chemicals is one of the least tested health effects. Indeed, the guidelines for DNT studies mainly stand on in vivo experiments, which are expensive and prolonged. 18 Recent advances in cell biology have introduced newly in vitro models for screening the toxicity of chemicals that overcome the limitations of previous models, such as poor extrapolation of results to humans. In this regard, stem cells, particularly, embryonic stem cell (ESC) models emerged as new tools for drug discovery and predictive toxicology due to their exclusive characteristics relevant to stemness, self-renewal and differentiation ability to various cell types. 19 The advent of pluripotent stem cell technology, both in humans and rodents, allows scientists to generate neural cells and establish functional DNT evaluation methods. 20,21 So far, few studies have explored the effects of chemicals at low non-cytotoxic concentrations on the ESCs differentiation-associated gene expression. Such attempts lead to find the underlying mechanism of chemical toxicities. 22 –24
Studies on the developmental toxicity of ethephon are very limited. The goal of this study was to examine the cellular and molecular neurodevelopmental toxic effects of ethephon using an ESC model. We applied RNA sequencing (RNA-seq) next-generation sequencing technology to understand the gene expression changes of mouse ESCs (mESCs) at the whole-transcriptome level in response to ethephon. Furthermore, we assessed the neural differentiation of mESCs in the presence of ethephon.
Materials and methods
Chemicals and materials
Ethephon (2-chloroethylphosphonic acid, 96% titration) was from Sigma (Germany, CAS number: 16672-87-0). Also, 4,5 dimethylthiazol-2-yl-2,5-diphenyltetrazolium bromide (MTT), phosphate-buffered saline, potassium chloride, potassium ferricyanide (III), potassium hydroxide, propidium iodide, sodium citrate, sodium sulphate, sulphuric acid, trichloroacetic acid, Tris/HCl and Triton X-100 were from Sigma-Aldrich (Germany). 5-5-Dithiobis-2-nitro benzoic acid, bovine serum albumin, dichlorophenolindophenol, dipotassium phosphate, magnesium chloride, nicotinamide adenine dinucleotide and sodium bicarbonate were obtained from Merck Chemical Company (Merck, Germany). Mouse leukaemia inhibitory factor (LIF, 1000 U/ml) was purchased from Royan Institute (Tehran, Iran). Fetal bovine serum (FBS),
Cell culture and treatment
mESCs were purchased from Royan Stem Cell Bank (Tehran, Iran; www.royaninstitute.org) and cultured at 37°C, 5% carbon dioxide (CO2) in a flask coated with 0.1% gelatine. DMEM/F12 medium containing 20% ESCs FBS, (LIF, 1000 U/ml),
Cell viability assay
Cell viability was evaluated during short-term exposure of 24 h. For the determination of the cytotoxicity of ethephon on mESCs, MTT test was performed, as described previously. 14 In brief, mESCs were seeded on gelatin-coated-96 well plate and exposed to the various doses of ethephon (40, 80, 160, 320, 640, 1280 and 2560 µM). Then, 50 µl MTT solution (5 mg/ml) was added per well and incubated for 3 h at 37°C in dark. Then, dimethyl sulphoxide solvent was applied to the wells while shaking gently. After 15 min, the absorbance was measured at 593 nm by a microplate reader (BioTek, Winooski, VT, USA).
Cell cycle assay
After treatment with different concentrations of ethephon, cells were detached and collected by centrifugation followed by fixation in ice-cold 70% ethanol. Propidium iodide solution containing 50 and 8 μg/ml RNase was applied to stain the cells in dark for 30 min. Finally, cells were analyzed for cell cycle distribution using FACScan flow cytometer (Mindray, China) and FlowJo software (7.6.2 version).
RNA extraction
mESCs were treated with selected doses of ethephon (160 µM) for 24 h and the total RNAs were extracted from ethephon-treated and control groups using BioZol total RNA extraction kit (Bioer Technology, Hangzhou, China), according to the manufacturer’s instruction. Then, the isolated RNAs were assessed for purity and integrity by NanoDrop spectrometer as well as visualization on 2% agarose gel for ribosomal RNA band and measuring RNA integrity factor (RIN) values using the Agilent 2100 Bioanalyzer (Agilent, Palo Alto, California, USA). The A260/A230 ratio measurements between 1.8 and 2 and RIN factor more than 7 were included in the study (Supplemental Figure 1). The extracted RNAs were used for RNA-seq and quantitative real-time polymerase chain reaction (qRT-PCR) experiments.
Next-generation sequencing
Transcriptome analysis was carried out by Beijing Novogene Bioinformatics Technology Company (China). RNA samples from ethephon-treated and untreated mESCs were sent to the company to perform RNA-seq differential expression using HiSeq 4000 Illumina platform. The RNA-seq libraries were prepared with 150 bp paired-end sequences for each sample using the NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (New England BioLab, Ipswich, MA, US).
To calculate the quality control statistics for the generated data, FastQC was applied. Using Trimmomatic, the sequences were filtered to remove sequences with low Phred scores. The short reads fastq files were processed using Tophat (v2.0.1) and mapped to the mouse reference genome using default settings for paired reads. Cufflink program (version 1.3.0) was then applied to convert aligned short reads into fragments per kilobase of exon model per million mapped fragments (FPKM). After merging all transcripts made by cufflinks, cuffdiff was used to find differentially expressed genes with p value cut-off <0.05.
The list of downregulated and upregulated genes was subjected to DAVID gene ontology (GO) analysis. To identify enriched GO terms, functionally clustered genes were ranked according to the p value (<0.05) of Expression Analysis Systematic Explorer score. For pathway mapping, the Kyoto Encyclopaedia of Genes and Genomes (KEGG) was used.
Quantitative real-time PCR (qRT-PCR)
First-strand complementary DNAs (cDNAs) were synthesized using RevertAid cDNA synthesis kit (Thermo Scientific, Rockford, IL, USA), according to the provided instruction by the manufacturer. PCR reactions were administered duplicate in a total volume of 20 µl containing SYBR Green, PCR Master Mix (Applied Biosystems, Foster City, CA, USA) and 0.3 mM of each primer. The details of forward and reverse primers of all tested genes (including octamer-binding transcription factor 4 (OCT-4), NANOG, sex determing region Y HMG-box 2 (SOX-2), microtubule-associated protein 2 (MAP-2), β-tubulin III, nuclear factor erythroid 2-related factor 2 (NRF-2), Beta-actin (ACTB) and neuron-specific enolase (NSE)) are indicated in Supplemental Table 1. Amplification was performed on Roch 480 Light Cycler and data obtained as Ct values were normalized relative to β-actin. The fold changes in the expression of the targeted gene were studied compared to β-actin. The PCR products particularity were affirmed using melting curve analysis and consequent agarose gel electrophoresis. The melting curve and CT averages were shown in Supplemental Figure 2. Fold change was calculated based on 2−ΔΔCt to examine relative gene expression.
mESCs neural differentiation
Neuronal differentiation was performed according to the procedures reported previously with some modifications. 27 Briefly, cells were seeded in 3 cm dishes coated with 0.1% gelatine, containing neural differentiation medium (neurobasal medium supplemented with 1% N2, 2% B27, 40 ng/ml FGF-2, LIF, 0.1 mM 2ME and 20% ES FBS), and were incubated at 37°C and 5% CO2 incubator. After 24 h, the treated group was exposed to ethephon (160 µM) for 24 h. When the confluency reached about 80%, the cells were recultured at a concentration of 5 × 103 in the same medium containing no LIF and FGF. The medium was changed every other day. On the fifth day after first seeding, the medium was changed and supplemented with retinoic acid (4 µM). On the day 7, control and treated groups were evaluated for gene expression.
Statistical analysis
In this study, all the experiments, including MTT assay, cell cycle and qPCR gene expression, were performed in triplicate and repeated three times at three independent days. The achieved data were reported as mean ± standard error of the mean. One-way analysis of variance followed by Tukey’s multicomparison tests was completed by Stats Direct 3.2.10 to calculate the statistical difference (p < 0.05) between the treated and control groups.
Results and discussion
Cell viability
As shown in Figure 1, the viability of mESCs was not significantly affected at doses ≤640 µM ethephon.

Assessment of cytotoxicity of ethephon by MTT assay after 24 h. The results were demonstrated as mean ± SEM. *p < 0.05: significantly different from control. MTT: 4,5 dimethylthiazol-2-yl-2,5-diphenyltetrazolium bromide; SEM: standard error of the mean.
Cell cycle analysis
The control and ethephon-treated mESCs were assessed for analysis of cell cycle by flow cytometry. The results of the cell distribution in particular phases showed that cells mainly arrested at G 0/G 1 phase in response 160 µM ≥ ethephon, as compared to the untreated group. Accordingly, the distribution of cells in G 2/M phase was decreased (Figure 2).

Effect of ethephon on cell cycle distribution. Control (a) control untreated group; (b) cells treated with ethephon (80 µM); (c) cells treated with ethephon (160 µM); (d) cells treated with ethephon (320 µM). In the presence of ethephon, cell cycle shifted towards G 0/G 1 phase. *p < 0.05: significantly different from control; **p < 0.001: significantly different from control.
Differential gene expression profiling by RNA-seq
For RNA-seq analysis, the mESCs were treated with ethephon at 160 µM (approximately 1/20 of IC50). The number of clean reads in samples ranged between 39, 250, 713 and 32, 517, 701. Mapping output demonstrated that approximately 92% of the reads were aligned to the mouse genome reference (mm10). Of the total 50,616 genes in the database, 13,193 genes had at least FPKM >1 in one sample.
Of the 13,193 expressed genes, 327 had a p value <0.05 and 39 genes had p values <0.005. Data showed that 194 of the 327 expressed genes were downregulated, whereas 133 genes were upregulated after exposure to ethephon (Supplemental Tables 2 and 3). The p value cut-off of 0.05 was used to tabulate the differentially expressed genes. Based on this criterion, 327 genes were selected for pathway analysis and GO terms. To examine biological functions, GO analysis was done adopting David GO website on the 327 genes, which were expressed differentially. A selection of considerable GO terms for biological process and molecular functions was displayed in Figure 3. The results showed that GO terms related to cell fate and organismal development, including neuron fate commitment, embryo development and cardiac cell differentiation, were significantly enriched. There were also genes, such as Myc, Ep300, Gm 5741, Vegfc and Cblc that were related to cancer pathways and showed an increase in their expressions. The four significantly enriched pathways were identified in the KEGG pathway, as shown in Figure 3.
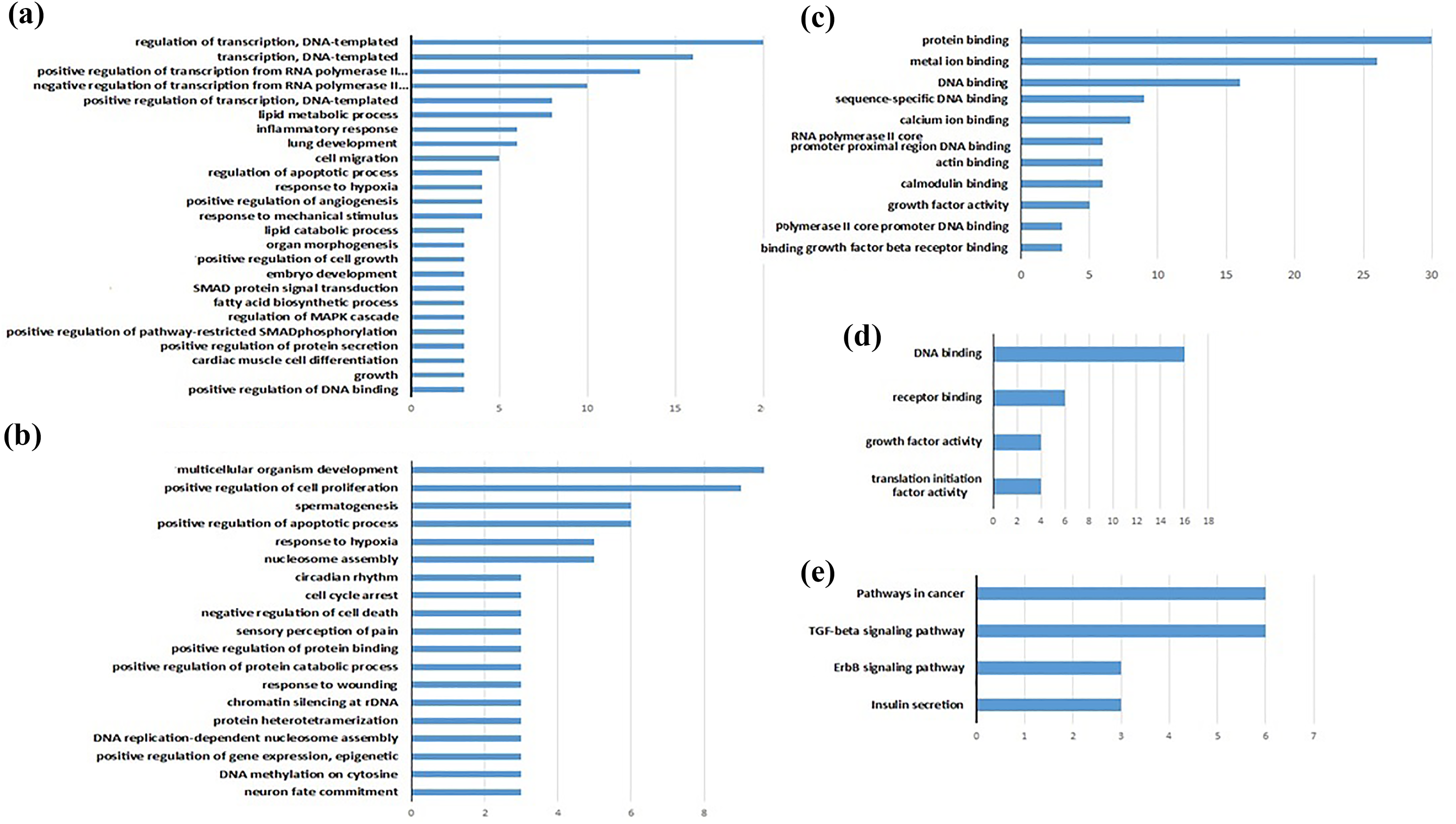
A selection of the enhanced GO biological process, GO molecular function, and KEGG pathways of differentially expressed genes in samples from ethephon treated-ESCs and control. (a) Biological process (upregulated genes), (b) biological process (downregulated genes), (c) molecular function (upregulated genes), (d) molecular function (downregulated genes) and (e) KEGG pathway. GO: gene ontology; KEGG: Kyoto Encyclopaedia of Genes and Genomes; ESC: embryonic stem cell.
Analysis of gene expression in mESCs by RT-PCR
Differential expression of stemness-related genes as identified by RNA-seq analysis was examined by real time RT-PCR assays between ethephon-treated and control group. The results showed a significant decrease in OCT-4 and SOX-2 expression in mESCs treated with 160 and 320 µM ethephon (p < 0.001), while there were increase in NANOG and NRF-2 expressions, as were shown in Figure 4.

Gene expression analysis of mESCs stemness markers in response to ethephon treatment after 24 h exposure. Ethephon affected the expression of stemness genes. *p < 0.05: significantly different from control. **p < 0.001: Significantly different from control; mESC: mouse embryonic stem cell.
Anteriorization of neural fate in the presence of ethephon
In the initial stage of neural induction, the rosette structures were formed in the control group after 4 days (96 h) differentiation. The neural shape filaments appeared at day 5–7 [Figure 5(a) and (b)]. There was a clear difference in the morphology of the differentiated mESCs exposed to 160 µM ethephon with the aberrant filaments.

mESCs neural differentiation in response to the ethephon treatment. (a-a′) Seven days after neural induction of ESC control and (a-b′) ethephon-treated group. The expression of neural markers 7 days after ESCs neural differentiation in the presence and absence of ethephon (160 µM). The black column represented the expression of neural genes in undifferentiated mESCs. Ethephon treatment attenuated the expression of neural markers. **p < 0.001: Significantly different from mESC control;. † p < 0.001: significantly different from differentiated control, # p < 0.01: significantly different from differentiated control. mESC: mouse embryonic stem cell; ESC: embryonic stem cell.
The expression of NSE (microtubule-associated protein), MAP-2 and β-tubulin III was assessed at the concentration of 160 µM of ethephon following neural induction of mESCs using RT-PCR. Obtained data showed that the expression of neural markers increased significantly in the differentiated cells while attenuated in ethephon-treated mESCs during neural differentiation (p < 0.001).
Discussion
Plant growth regulators are chemical substances that can affect the cellular growth or tissue differentiation. Among the plant growth regulators, ethephon is widely used in the field of agriculture notably in the induction of fast-growing and maturation of plants worldwide. Exposure to ethephon may occur directly through the consumption of residues within the fruits 28 or as a result of exposure to vapours or spray forms in the concentration of around 1000 ppm (1 mg/l). Based on the reported evidence in the rat model, between 0.1% and 0.5% of ethephon remains in tissues following oral administration of 50 mg/kg. 29 There are restricted certain data related to the adverse effects of ethephon in animal models, indicating its mutagenicity, teratogenicity and biochemical modulations at acute doses of 50–150 mg/kg. 12,30,31 Chromosomal aberrations were also detected in the foetus of pregnant mouse administered the same doses of ethephon. 30 While the acute reference dose of 0.05 mg/kg body weight per day was established as safe for human, still, concern remains over the molecular toxicity of this chemical that urges further exploration. In this regard, the present study attempted to investigate the developmental toxicity of ethephon using ESCs as an important in vitro model in toxicological studies. 19,32 To our knowledge, this is the first study evaluating the gene expression profile changes of mESCs in response to ethephon treatment. Our data on mESC cytotoxicity of ethephon showed a significant decrease in cell viability at doses higher than 640 µM of ethephon within 24 h. This was consistent with a previous study demonstrated the cytotoxic effect of ethephon after 24 h with IC50 value of 108.7 (747 µM), 126 (872 µM) and 137.2 µg/ml (948 µM) for Hep2, HepG2 and Vero cells, respectively. 13
Cell cycle distribution analysis at selected non-cytotoxic doses of ethephon showed an increased cell population in G 0/G 1 phase, indicating the role of ethephon in pathways influencing cell growth arrest. Studies on other mammalian cell types indicated ethephon-induced proliferative activity and induction of cell cycle G 2/M phase. 26,33
In this study, we applied the whole transcriptome sequencing analysis to investigate the global changes in gene expression profile of ESCs in response to non-toxic doses of ethephon. Non-cytotoxic dose of 160 µM ethephon was preferred on the basis of the results obtained in our previous study as well as the present data on viability assay and cell cycle analysis at concentrations with no effective changes.
Gene expression profile presented the complex signalling pathways and mechanisms affected by ethephon. Among 327 genes that significantly differentially expressed, ethephon could alter the expression of stemness-related markers, including SOX-2 and OCT4 in ESCs. In line with this observation, the RT-PCR data also revealed significant changes in SOX-2, OCT-4, NRF-2 and NANOG expression after 24 h exposure to ethephon. Indeed, mechanistic investigations have displayed modification in the expression of pluripotency-associated genes contributed to the altered phenotypic behaviour and impaired ability for mESCs multilineage cell differentiation. 34,35 The SOX-2 and OCT-4 transcription factors play key roles in the maintenance of pluripotency of ESCs that coregulate target genes involved in self-renewal capacity. 36 SOX-2 is also functioning as the inducer of neural differentiation in neural progenitors. 37,38 In the early embryonic stage, any deficiencies in the expression of these factors could impair embryo development at an early stage. 39 NANOG, which is associated with self-renewing in ESCs, involved in the maintenance of stem cells in the undifferentiated form. Reportedly, downregulation of OCT4 and NANOG could shift ESCs differentiation towards extraembryonic endoderm lineage. 40 Similar studies showed that manipulating the OCT4 and SOX-2 impacts on neuroectodermal commitment. 41 Taken the results obtained from the aforementioned studies, our results on RNA-seq analysis and RT-PCR suggested that ethephon treatment could affect cell fate decisions and mESCs response to the differentiation stimuli through affecting key pluripotency genes. Accordingly, the results of the GO analysis of genes with different expression clearly showed the enrichment of cell pathways involved in cell fates, including neuron fate commitment, embryo development and cardiac cell differentiation. Also, the expression of few genes with the regulatory function in transcription and replication, such as DNA-binding elements associated with polymerase II core promoter, that is the key components in the initiation of transcription by the RNA polymerase II machinery, was altered. 42
MYC is a global transcriptional factor that regulates different genes and controls multiple signalling cascades involved in the stem cell maintenance of pluripotency. 43 MYC is also known as a proto-oncogene that becomes activated in several tumours and recognized for its inducible role in cancer stem cell. 44,45 Our data from RNA-seq analysis showed an increased expression of MYC as well as other genes related to cancer pathways that might raise the possibility of ethephon oncogenicity. Although the data reported by the Environmental Protection Agency in 1995 showed no genotoxic activity at doses of approximately 500–2500 µg/ml of ethephon in mammalian cell, 46 our data suggested changes in the expression of oncogenic genes as well as genes involved in the developmental process at doses of 160 µM (23 µg/ml).
Furthermore, we evaluated the neural differentiation of ESCs in the presence of low doses of ethephon. We demonstrated that the exposure to the ethephon after 5-day treatment could alter the morphology of the differentiated cells and abrogate the formation of neural shape filaments. Accordingly, the expression of neural-specific markers, including NSE, MAP-2 and β-tubulin III, was decreased in response to the ethephon treatment. All evidence indicated the perturbation of ESCs neural differentiation in the presence of ethephon. The NSE, MAP-2 and β-tubulin III genes are expressed at ending steps of neural differentiation in terminally differentiated neurons. 27
Conclusion
Our study using ESC model provided strong evidence that in vitro generation of neurons from ESCs is affected by ethephon that suggests neurodevelopmental toxicity of this chemical. In addition, the comprehensive assessment of gene expression profiling of ESCs in response to ethephon showed significant modifications in the expression of the vast number of key regulatory genes including pluripotency genes. Considering the limited number of studies on ethephon embryonic toxicity, further in vivo studies are required to confirm the potent developmental neural toxicity of ethephon. Based on the data acquired from our study, more caution and greater control are advised for the application of this widely used compound.
Supplemental material
Supplemental Material, Supplementary_Fig_1 - Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon
Supplemental Material, Supplementary_Fig_1 for Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon by S Mohammadi Nejad, M Hodjat, SA Mousavi, M Baeeri, MA Rezvanfar, M Rahimifard, S Sabuncuoglu and M Abdollahi in Human & Experimental Toxicology
Supplemental material
Supplemental Material, Supplementary_Fig_2 - Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon
Supplemental Material, Supplementary_Fig_2 for Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon by S Mohammadi Nejad, M Hodjat, SA Mousavi, M Baeeri, MA Rezvanfar, M Rahimifard, S Sabuncuoglu and M Abdollahi in Human & Experimental Toxicology
Supplemental material
Supplemental Material, Supplementary_Table3 - Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon
Supplemental Material, Supplementary_Table3 for Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon by S Mohammadi Nejad, M Hodjat, SA Mousavi, M Baeeri, MA Rezvanfar, M Rahimifard, S Sabuncuoglu and M Abdollahi in Human & Experimental Toxicology
Supplemental material
Supplemental Material, Supplementary_Table_1_3 - Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon
Supplemental Material, Supplementary_Table_1_3 for Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon by S Mohammadi Nejad, M Hodjat, SA Mousavi, M Baeeri, MA Rezvanfar, M Rahimifard, S Sabuncuoglu and M Abdollahi in Human & Experimental Toxicology
Supplemental material
Supplemental Material, Supplementary_Table_2_1 - Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon
Supplemental Material, Supplementary_Table_2_1 for Alteration of gene expression profile in mouse embryonic stem cells and neural differentiation deficits by ethephon by S Mohammadi Nejad, M Hodjat, SA Mousavi, M Baeeri, MA Rezvanfar, M Rahimifard, S Sabuncuoglu and M Abdollahi in Human & Experimental Toxicology
Footnotes
Acknowledgment
The authors acknowledge Iran National Science Foundation (INSF) for the support of the second author (MH)'s postdoc program and also the Seat Award directed to MA.
Author contributions
SMN and MH contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
