Abstract
Objective:
To identify the role of miR-146a and tumor necrosis factor receptor-associated factor 6 (TRAF6) for improving the apoptosis of hippocampal neurons induced by microglia activation.
Methods:
Mouse microglial cell line (BV2 cell) was employed and treated with lipopolysaccharide. Mouse hippocampal nerve cell line (HT22 cell) was then grown in BV2 conditioned medium, and miR-146a overexpression and silencing cell lines were constructed. CCK8 and clone formation test were utilized to evaluate the proliferation ability of the transfected cells, and the level of inflammatory factors was measured by ELISA. Apoptosis was determined extensively by flow cytometry. The apoptosis-related protein and TRAF6 protein expressions were verified by Western blot. TRAF6 was identified to be the target gene of miR-146a based on double Luciferase Report. Finally, both TRAF6 and miR-146a were used to treat HT22 cells and the above indexes were detected repeatedly.
Results:
Interleukin (IL)-1β, tumor necrosis factor-α, and IL-6 expressions in BV2 cells increased significantly. miR-146a overexpression distinctly increased the cell proliferation ability and B-cell lymphoma-2 expression ((Bcl-2, p < 0.05); meanwhile, the apoptosis rate of cells, apoptosis-related proteins (Bcl-2 associated X and cleaved caspase-3), and TRAF6 gene and protein expressions were significantly decreased (p < 0.05). However, these above results were reversed for miR-146a silence. There is a targeting relationship between miR-146a and TRAF6. Silencing TRAF6 gene can promote HT22 cells’ proliferation and inhibit apoptosis. The effect of miR-146a on HT22 cells was reversed by adding TRAF6 mimics to miR-146a overexpression cells.
Conclusion:
miR-146a can inhibit the apoptosis of hippocampal neurons caused by microglia activation via targeting TRAF6 and down-regulating its expression.
Introduction
Microglia is an innate immune cell similar to macrophages in the central nervous system, which has two states of resting and activating. 1 Activated microglia is of great significance to monitor the microenvironment, clean up the damaged nerves, and prune neural process in healthy central nervous system so as to maintain synaptic homeostasis. 2 However, the continuously over-activated or out of control microglia may induce the release of many cytotoxic factors and lead to some inflammations, which can worsen neuronal damages, thereby resulting in neurodegenerative diseases. 3 Microglia cells are found to be activated and proliferated locally in the pathological process of central nervous system injury, inflammation, depression, Alzheimer’s disease, and even epilepsy. 4,5
Depression is a chronic and recurrent disease characterized by persistent depression and lack of interest in life. 5 There were some evidences suggested that microglia cells activation was closely associated with its occurrence and progression. 6,7 Microglia cells are activated immediately after stimulation, and the factors that cause their activations are in a wide range, such as lipopolysaccharide (LPS), adenine nucleoside triphosphate, pre-inflammatory cytokines, and so on. 8 –10 Similarly, patients with depression showed poor therapeutic effect with high expression inflammatory cytokines in serum. In the early stage of depression, inhibition of inflammatory biomarkers can relieve the depressive symptoms. 11 However, the mechanism of microglia cells activation in the depression process has not yet been understood and is still a focus of current research. Therefore, it is of great significance to explore the mechanism of inhibiting microglia activation in the treatment of neurological diseases.
MicroRNAs (miRNAs) are one group of small non-protein-coding RNA molecules of 20–22 nucleotides, which regulate the expression of target genes. They are considered as potential biomarkers and key regulators of many cellular processes in various diseases. 12 Many miRNAs are involved in inflammation and autoimmune process. 13,14 Of which, miR-146a has been found to regulate innate immune responses, 15 and the upregulation of its expression is associated with tissue inflammation. 16 A study has demonstrated that increasing miR-146a expression can reduce the inflammatory response mediated by microglia cells, while decreasing the expression of miR-146a can play an opposite role. 17,18 miR-146a can inhibit the transcriptional activity of nuclear factor kappa-B (NK-κB) by inhibiting the expression of the molecule tumor necrosis factor receptor-associated factor 6 (TRAF6) between toll-like receptor-4 (TLR-4) and NK-κB and can reduce the release of its downstream inflammatory cytokines, such as interleukin (IL)-1β, IL-6, and tumor necrosis factor (TNF)-α, ultimately inhibiting the inflammatory response. 17 In view of this, it is a new intervention idea that regulation of miR-146a expression is to inhibit the inflammatory response and promote the repair of nerve injury in the central nervous system.
In this study, we hypothesized that miR-146a improved the cell apoptosis of hippocampal nerve cell HT22 induced by mouse microglial cell BV2. Further, the targeting regulatory relation between miR-146a and TRAF6 was analyzed to explore the mechanism of miR-146a on improving the apoptosis of hippocampal neurons induced by microglia activation.
Materials, reagents, and methods
Cell culture
The mouse microglial cell BV2 and hippocampal nerve cell HT22 were purchased from Shanghai Institute of Biochemistry and Cell Biology that belongs to Chinese Academy of Sciences. Both cell lines were cultured in DMEM/F12 medium (GIBCO; Invitrogen, Grand Island, New York, USA) supplemented with 10% fetal bovine serum and 1% penicillin streptomycin in a constant temperature incubator (Thermo, New York, USA) containing 5% CO2 at 37°C. When the cells grew to about 80%, the logarithmic growth phase cells were collected for experiments. All procedures were approved as required in the ethics committee of Binzhou People’s Hospital.
LPS-treated BV2 cells
BV2 cells were divided into two groups: the blank control group (BC, cells were not treated) and the LPS group (cells were stimulated with 1 μg/mL of LPS; Sigma, St. Louis, Missouri, USA). 17
Detection of inflammatory factors in BV2 cells by ELISA
After BV2 cells were stimulated for 24 h, the supernatant was collected and centrifuged by 3000 r/min for 20 min. The content of IL-1β (orb79117; Biorbyt, Cambridge, UK), TNF-α (orb79138; Biorbyt), and IL-6 (orb79123; Biorbyt) in serum of BV2 was detected in strict accordance with the operation instructions of ELISA kit, and the data were recorded at 450 nm of enzyme marker (RT-6100, Lei Du).
Preparation of conditioned medium and cell grouping
BV2 cells, as adherent cells, were inoculated into six-well plates with 2 × 105/mL, 2 mL for each well. When the cells adhered to the wall, they were stimulated with LPS for 24 h, and likewise, the cell culture supernatant was achieved. The microglia cells medium without LPS was used as the blank conditioned medium (C-CM), and the conditioned medium after LPS treatment for 24 h was used as the conditioned medium (L-CM). The HT22 cell lines were subcultured one day before transfection and cultured in six-well plates. The transfection was carried out when the cell density was up to 90%. Five microliters of Lipofectamine TM2000 (Invitrogen, Carlsbad, California, USA) and 5 µL of plasmids (0.5 μg/μL; GenePharma, Shanghai, China) were respectively added into sterile centrifuge tubes containing 200 µL DMEM/F12 and were left for 5 min under the room temperature. The plasmids mixture was subsequently added into the Lipofectamine TM2000 mixture, mixed well and maintained for 20 min at room temperature. Finally, the mixture was dripped into the six-well plate, gently mixed and cultured for another 6 h. The plate was shaken every 1 h, so that liposomes and plasmids could fully interact with cells. According to different treatment methods, HT22 cells were divided into six groups: blank medium group (C), conditioned medium group (L), L + miR-146a mimic negative control group (L + NC1), L + miR-146a mimic (5′-UGAGAACUGAAUUCCAUGGGUU-3′) group (L + miR), L + miR-146a inhibitor control group (L + NC2), and L + miR-146a inhibitor (5′-AACCCAUGGAAUUCAGUUCUCA-3′) group (L + si-miR). The plasmids were purchased from GenePharma (Shanghai, China). The cells of C group were cultured in blank medium. In other groups, HT22 cells were cultured in BV2 activation conditioned medium. After transfection for 24 h, the cells were harvested and used for subsequent experiments.
Reverse Transcription Polymerase Chain Reaction (RT-PCR)
After cells were collected, they were centrifuged at 4°C (12,000 r/min, 15 min). Total RNA was extracted and isolated by TRIzol Kit (Takara, Dalian, China). OD260/OD280 between 1.8 and 2.0 indicates that RNA purity is qualified. The reverse transcription of miRNA was carried out by using miScript Reverse Transcription Kit (Qiagen, Germany). mRNA reverse transcription was performed using PrimeScriptTM RT Master Mix (Takara, Japan). Conditions are as follows: 95°C for 15 s, 60°C for 60 s, 72°C for 40 s (35 cycles). The data were processed with 2−ΔΔCt method, and the relative expression level was calculated with U6 and GAPDH as the internal parameter. The sequences of primers (Shanghai Shenggong Bioengineering Technology Service Co., Ltd, China) used for RT-PCR were as follows: miR-146a Forward: 5′-CCTGAGAAGTGAATTCCATGGG-3′ Reverse: 5′-TGGTGTCGTGGAGTCG-3′ U6 Forward: 5′-GACCTCTATGCCAACACAGT-3′ Reverse: 5′-AGTACTTGCGCTCAGGAGGA-3′ TRAF6 Forward: 5′-CCAATTCCCAGAATCCAGAAA-3′ Reverse: 5′-GACACAGAGGACCCACAGAGAA-3′ GAPDH Forward: 5′-TGACTTCAACAGCGACACCCA-3′ Reverse: 5′-CACCCTGTTGCTGTAGCCAAA-3′
CCK8 was used to detect the proliferation of transfected cells
The treated HT22 cells (2 × 104 cells/mL) were seeded into 96 well plates, and each well was 100 μL. Then, cells were cultured at 37°C with 5% CO2 for experiment. After 24, 48, 72, and 96 h, the cells were treated with 10 µL of CCK-8 solution (Tongren Institute of Chemistry, Japan) and continued incubation in 37°C incubator for 4 h. The absorbance value of each hole was quantified at 450 nm.
Clone formation experiment
HT22 cells were collected in logarithmic growth phase, digested with 0.25% trypsin and then they were blown into single cells. Two milliliters of cells (250 cells/mL) were seeded into each well in the six-well plate. After that, it was placed under the conditions of 37°C and 5% CO2 for 2–3 weeks and the fresh culture media was changed every 3 days. Then cells were fixed with methanol. Each hole was added into 1 mL Giemsa working fluid and dyed for 30 min. Finally, the plate was washed twice using the ultrapure water and photographed by camera after the water around the plate was sucked off with filter paper.
Detection of apoptosis by flow cytometry
After the above treatment, HT22 cells were incubated for 24 h. Cells were collected, resuspended at 1× precooled PBS (4°C), and centrifuged for 5–10 min at 2000 r/min. After washed, the cells were suspended with 1× Binding Buffer (300 μL). Five microliters of Annexin V-FITC were appended and mixed well, and then cells were kept away from light and incubated for 15 min under the room temperature. Five minutes before loading, propidium iodide (PI) dye (5 μL) was added, and 1 × binding buffer (200 μL) was added. Finally, the samples were detected by Beckman Coulter (Brea, California, USA) and assessed by CellQuest software (BD Bioscience, San Diego, California, USA).
Western blot detected the apoptosis-related protein and TRAF6 protein expression in HT22 cells
HT22 cells were lysed, centrifuged at 2000 r/min for 20 min, and the supernatant was obtained. Protein concentration was quantified by butyleyanoacrylate kit (Solarbio, Beijing, China). Four microliters of each protein sample were mixed using the 5× sample buffer and added into 10% SDS-PAGE. The protein was separated according to the molecular weight by electrophoresis at 80 V. The protein samples were transferred to the polyvinylidene fluoride (PVDF) membrane (Merck, Darmstadt, Germany) under the condition of half dry at 20 V for 30 min with electrical transfer instrument (Bio-Rad, New York, USA). Subsequently, the membrane was washed, sealed with 5% skimmed milk powder for 2 h, and subsequently the following primary antibodies were added: rabbit anti-mouse B-cell lymphoma-2 (Bcl-2) associated X (Bax, 1:500, orb224426; Biorbyt), Bcl-2 (1:500, orb226346; Biorbyt), cleaved caspase-3 (1:500, orb126608; Biorbyt), TRAF6 (1:500, orb228290; Biorbyt), and rabbit anti-GAPDH (1:500, orb67107; Biorbyt) polyclonal antibodies. After that, it was incubated overnight under 4°C. When it was washed and rewarmed, this membrane was incubated by using a goat anti-rabbit IgG-horseradish peroxidase (HRP) (1:1000, #7074; Cell Signaling Technology, USA) for 0.5 h. The results were recorded with the Roche Elecsys-2010 chemiluminescence instrument (Switzerland).
Dual-luciferase report assay
To determine whether TRAF6 is a direct target gene of miR-146a, the wild-type and mutant 3′UTRs of TRAF6 were amplified in pGL3/luciferase vector (Promega, Madison, Wisconsin, USA) and cloned to the downstream of luciferase gene. According to the manufacturer’s instructions, the HT22 cells were tested for luciferase activity on the basis of a dual luciferase reporter system (Promega, Shanghai, China) after transfection for 48 h. 19
To further verify the effect of miR-146a targeting TRAF6 on HT22 cells
The cells were divided into seven groups: (a) blank conditioned medium group (C): cells were cultured in blank conditioned medium group; (b) conditioned medium group (L): cells were cultured in conditioned medium; (c) L + miR-146a overexpression group (L + miR, cells that transfected by miR-146a mimics); (d) L + TRAF6 inhibitor negative control group (L + NC3, cells that transfected with TRAF6 inhibitor negative control); (e) L + TRAF6 inhibitor group (L + si-T): in which cells were transfected using the TRAF6 inhibitor; (f) L + miR-146a overexpression + TRAF6 overexpression negative control group (L + miR + NC4): cells were transfected using the miR-146a mimic and TRAF6 mimic negative control; and (g) L + miR-146a overexpression + TRAF6 overexpression (L + miR + T): transfected with miR-146a mimic and TRAF6 mimic. The above cells from (c) to (g) group were all cultured in conditioned medium. Cell proliferation and apoptosis were detected. The experimental design was shown in Figure 1.

The experimental design figure.
Statistical analyses
These experimental data were analyzed by using SPSS 19.0 statistical analysis software, which were expressed as mean ± standard deviation. T-test was exerted for data analysis between the two groups, and differences of data among multiple groups were calculated by one-way analysis of variance as well as subsequent Dunnett t-test. A value of p < 0.05 was regarded to be significant statistically.
Results
LPS acts on the expression of inflammatory factors in BV2 cells
Both IL-1β and IL-6 are extremely important mediators of the inflammatory response, which are participated in a series of cellular activities, such as cell proliferation and apoptosis. 20 TNF-α is also functioned by these roles of mediating systemic inflammation and stimulating the acute phase reaction. 21 As shown in Figure 2, when compared to BC group, the expression of IL-1β, TNF-α, and IL-6 increased significantly (p < 0.05) in the LPS group. And the levels of IL-1β, TNF-α, and IL-6 in BV2 cells were strongly boosted after LPS stimulation. To say, the above result means that LPS induced BV2 activation.

The effect of LPS on the expression of inflammatory factors in BV2 cells. (a) IL-1β. (b) TNF-α. (c) IL-6. Compared with BC group, @ p < 0.05. LPS: lipopolysaccharide; IL: interleukin; TNF-α: tumor necrosis factor-α.
miR-146a effects on proliferation and apoptosis of HT22 cells under BV2 activation
The HT22 cells were cultured in BV2 activation condition medium. The results demonstrated (Figure 3(a)) that miR-146a gene expression was significantly decreased in L group compared to C group (p < 0.05). Compared with the L group, the expression of miR-146a gene in the L + miR group cells was distinctly increased, while that in the L + si-miR group was markedly decreased (p < 0.05). In addition, as compared to C group, the proliferation ability (Figure 3(b) and (c)) and apoptosis rate (Figure 4(a)) of cells in other groups were significantly decreased, and the expression of apoptosis-related proteins (Figure 4(b)), Bax and cleaved caspase-3, in cells were overtly increased, while the expression of Bcl-2 was obviously decreased (p < 0.05). Compared with the L group, the results of the above indicators were contrary in L + miR group (p < 0.05). Nevertheless, the cells proliferation ability of L + si-miR group was decreased significantly (Figure 3(b) and (c)), and the apoptosis rate (Figure 4(a)) was obviously increased. The expression of Bax and cleaved caspase-3 in the cells was visibly increased, and the expression of Bcl-2 was distinctly decreased (Figure 4(b); p < 0.05).
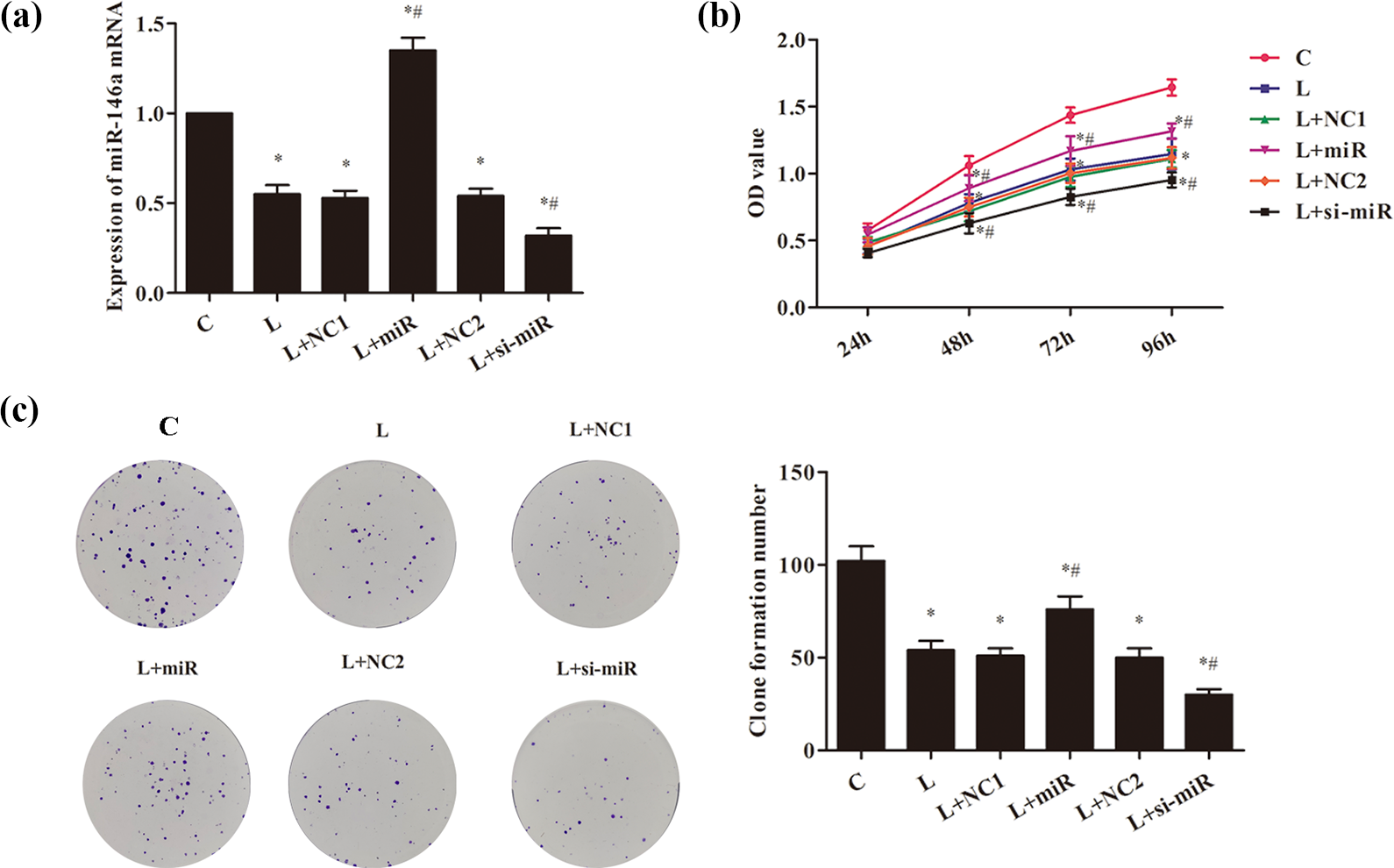
The effect of miR-146a on the proliferation of HT22 cells. (a) RT-PCR was used to detect the expression of miR-146a gene in the transfected cells. (b) CCK8 was utilized to detect the proliferation ability of transfected cells. (c) Clone formation experiment.

The effect of miR-146a on the apoptosis of HT22 cells. (a) Flow cytometry was utilized to evaluate the apoptosis of cells. (b) Western blot was exerted to detect the expression of apoptosis protein in the cells. Compared with C group, *p < 0.05; compared with L group, # p < 0.05.
Dual-luciferase reporter assay system
As shown in Figure 5(a) and (b), as compared with the C group, the expressions of TRAF6 gene and protein in other groups increased significantly with statistically significant difference (p < 0.05). When compared with L group, the expressions of TRAF6 gene and protein in L + miR group decreased significantly (p < 0.05), whereas those in L + si-miR group increased significantly (p < 0.05). Bioinformatics retrieval identified TRAF6 as the target of miR-146a (Figure 5(c)). In order to further verify whether miR-146a targets TRAF6, a dual luciferase reporting system was used (Figure 5(d)). The results demonstrated that miR-146a decreased the luciferase activity of TRAF6 with WT 3′UTR but not that of TRAF6 with mut 3′UTR.
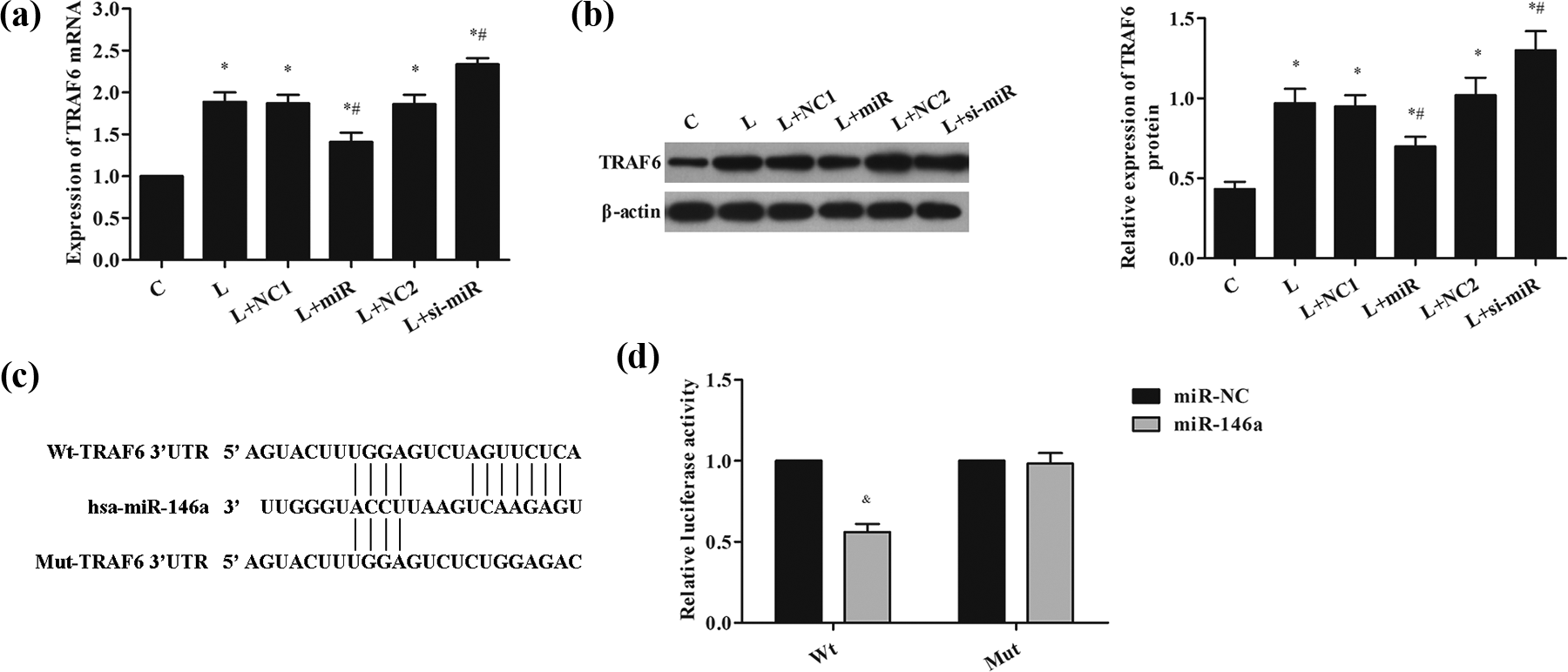
Dual-Luciferase Reporter Assay. (a) RT-PCR was used to detect the expression of TRAF6 mRNA in HT22 cells. (b) Western blot detected the expression of IRAK-1 protein in HT22 cells. (c) The predicted binding site of miR-146a and TRAF6 3′UTR region. (d) The dual luciferase expression of recombinant vector of miR-146a and its target gene TRAF6 was reported. Compared with C group, *p < 0.05; compared with L group, # p < 0.05; compared with miR-NC group, & p < 0.05. TRAF6: tumor necrosis factor receptor-associated factor 6; IRAL1: IL-1 receptor associated kinase 1.
miR-146a targeting TRAF6 acts on the proliferation and apoptosis of HT22 cells under BV2 activation
In order to verify the effect of miR-146a targeting TRAF6 on HT22 cells, TRAF6 inhibitor and mimic were transfected to HT22 cells. From Figure 6(a), the expression of TRAF6 mRNA in L + si-T group was obviously decreased, while significantly increased in L + miR + T group, compared with that in L group (p < 0.05). Further researches showed that the proliferation of HT22 cells was profoundly increased in L + si-T group, compared with L group (Figure 6(b) and (c); p < 0.05). However, the proliferation abilities were markedly lower in L + miR + T group than those in L + miR group (p < 0.05). The cell apoptosis rate and the expression of Bax and cleaved caspase-3 were also detected (Figure 7). The data showed, compared with L group, the expression of TRAF6 mRNA decreased significantly, the cell apoptosis rate was significantly decreased, the expression of Bax and cleaved caspase-3 was significantly decreased, but the expression of Bcl-2 was largely increased in L + miR, L + si-T, and L + miR + NC4 group (p < 0.05). Furthermore, in L + NC3 and L + miR + T groups, the expression of Bax and cleaved caspase-3 as well as apoptosis rate of cells were markedly increased, while the expression of Bcl-2 protein was significantly decreased in comparison with L + miR group (p < 0.05).

The effect of miR-146a targeting TRAF6 on the proliferation of HT22 cells. (a) RT-PCR was exerted to detect the expression of TRAF6 gene in the transfected cells. (b) CCK8 was applied to measure the proliferation ability of the transfected cells. (c) The clone formation experiment. TRAF6: tumor necrosis factor receptor-associated factor 6.

The effect of miR-146a targeting TRAF6 on the apoptosis of HT22 cells. (a) Flow cytometry was applied to assess the cell apoptosis. (b) Western blot was exerted to detect the expression of apoptosis protein in the cells. Compared with C group, *p < 0.05; compared with L group, # p < 0.05; compared with L + miR group, ^ p < 0.05. TRAF6: tumor necrosis factor receptor-associated factor 6.
Discussion
Microglia cells are activated immediately by LPS, pro-inflammatory cytokines, and other stimuli, which are often found in the pathological process of central nervous system injury and inflammation. 8 Once these cells are activated, they will release a large number of neuroactive substances, cytokines, and inflammatory mediators. The release of these substances can cause nerve inflammation and immune responses, which eventually lead to nerve dysfunction. 22 Consistent with the reports, we found that LPS induced pro-inflammatory cytokines secretion of BV2 microglia in our research. Furthermore, the HT22 cells activities were suppressed and cells apoptosis was increased after LPS induced BV2 treatment.
At present, miRNA therapy has attracted more attention on the treatment of neurological diseases and many experimental researches have been devoted to this. 23 –26 Interestingly, in vitro studies have shown that miR-146a may regulate the IL-1 receptor associated kinase 1 (IRAK-1) and TRAF6 on TLR-4 signaling pathway through negative feedback and may suppress NF-κB activity so as to further reduce the expression of proinflammatory factors, such as TNF-α, IL-6, and IL-1β, 23 –25 thereby inhibiting the activation of small keratinocytes and reducing the pathological damage caused by their activation. 27 Prior report has highlighted that induction of human miR-146a (has-miR-146a) regulates innate immune response by the mean of targeting TRAF6 and IRAK-1 genes 9. 24 Notably, the sequences of miR-146a, TRAF6, and IRAK-1 exhibit a high degree of nucleotide identity. Similar results were found in the present study; our study showed that miR-146a expression was increased in HT22 cells under BV2 activation. Further research identified transfection with miR-146a mimics in HT22 cells could decrease the apoptosis of cells and apoptosis-related proteins expression. However, the results were opposite when HT22 cells transfected with miR-146a inhibitor. According to previous reports, the expression of TRAF6 in HT22 cells was measured in this study. Expectedly, the levels of miR-146a regulated the expression of TRAF6, and their expression was opposite.
miRNA regulating gene expression occurs through specific binding to the target gene that contains the complementary nucleotide sequence. The TRAF6 functions as an adaptor molecule in the NF-κB pathway and acts as a crucial role in the initiation of the inflammatory response. 28 Bioinformatics retrieval identified TRAF6 as a target of miR-146a. Dual luciferase reporting demonstrated that miR-146a decreased the luciferase activity of TRAF6 with WT 3′UTR but not that of TRAF6 with mut 3′UTR. This finding is in agreement with some previous researches in humans and mice. 24,29,30 Therefore, in order to further verify the effect of miR-146a targeting TRAF6 on HT22 cells, TRAF6 inhibitor and TRAF6 mimic were added to treat cells. TRAF6 inhibitor transfection distinctly increased HT22 cells proliferation abilities and the expression of Bcl-2, meanwhile TRAF6 mimic transfection obtained opposite data. Contrary to TRAF6 mimic transfection, simultaneous transfection with miR-146a mimic and TRAF6 mimic weakened the effect of TRAF6 mimic. Based on this context, we believed that TRAF6-mediating miR-146a may emerge as a potential therapeutic target for rescuing apoptosis of hippocampal neurons caused by microglia activation.
The current study still exists many limitations. No insight is provided into the mechanisms about identification of the role of miR-146a and TRAF6 for improving the apoptosis of hippocampal neurons induced by microglia activation. Further work is needed to make clear the mechanisms and other molecules play a role in miR-146a mediated hippocampal neurons apoptosis and cytokine production.
Conclusion
Overall, this study revealed that miR-146a could inhibit the apoptosis of hippocampal neurons caused by microglia activation via targeting TRAF6 and down-regulating its expression.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
