Abstract
Arsenic has been associated with significant effects on human health. Exposure to inorganic arsenic has been associated with the changes in gene expression. Promoter of CDKN1A antisense DNA damage activated RNA (PANDAR) expression is induced by p53 protein and DNA damage response. Here, we investigated whether the ability of arsenic metabolism in individuals affected the expression of PANDAR, DNA damage, and DNA methylation. Levels of gene expression and DNA damage were examined by the quantitative polymerase chain reaction and DNA methylation was measured by the methylation-sensitive high-resolution melting curve. In our study, we demonstrated that arsenic exposure increased PANDAR expression and DNA damage among arsenic smelting plant laborers. The PANDAR expression and DNA damage were positively linked to monomethylarsonic acid % (R = 0.25, p < 0.05 and R = 0.32, p < 0.01) and negatively linked to dimethylarsinic acid % (R = −0.21, p < 0.05 and R = −0.31, p < 0.01). Subjects with low primary methylation index had increased levels of DNA damage (51.62 ± 2.96 vs. 60.93 ± 3.10, p < 0.05) and methylation (17.14 (15.88–18.51) vs. 15.83 (14.82–18.00), p < 0.05). Subjects with low secondary methylation index had increased levels of PANDAR expression (4.88 ± 0.29 vs. 4.07 ± 0.23, p < 0.01) and DNA damage (17.38 (15.88–19.29) vs. 15.83 (14.82–17.26), p < 0.01). DNA methylation of PANDAR gene was linked to the regulation of its expression in peripheral blood lymphocytes among laborers (Y = −2.08 × X + 5.64, p < 0.05). These findings suggested arsenic metabolism ability and exposure affected the expression of PANDAR, DNA damage, and DNA methylation.
Introduction
Arsenic, a ubiquitous element in nature, has been associated with significant effects on human health. 1 –6 Arsenous oxide (As2O3) was a recognized human carcinogen. Exposure to inorganic arsenic (iAs) has been associated with malignant tumors in several organs. 5 Recent findings highlighted that chronic exposure to arsenic increased risks, including hypopigmentation, chronic obstructive pulmonary disease, type 2 diabetes, ischemic heart disease, and peripheral arterial disease. 1 –8 However, the molecular mechanism of arsenic pathogenicity is still unclear.
In general, As2O3 entered the body via the respiratory tract. 9 Liver was the major site for arsenic metabolism, but the metabolic pathways of iAs in human body were still not fully understood. The most generally well-accepted methylation pathway involved two steps: conversion to monomethylarsonic acid (MMA) and further methylation to dimethylarsinic acid (DMA). 10,11 This methylation of iAs to MMA and DMA had a shorter biological half-value period than iAs and could be quickly discharged from the body. Urine iAs, MMA, and DMA were considered as metabolic markers. 1,2,6,9,12 The percentages of urinary arsenic metabolites and methylation index reflected the ability of arsenic metabolism in individuals. 13
S-Adenosylmethionine (SAM) was consumed by arsenite methyltransferase (AS3MT) protein to metabolize arsenic resulting in exhaustion of SAM. 5,14 DNA methylation required SAM to provide methylation donor, and the decrease of SAM in arsenic metabolism could lead to a shortage of methyl donor. Arsenic exposure could reduce the level of DNA methylation. iAs seems to reduce DNA methyltransferase (DNMT) activity by inhibiting transcription or inactivating proteins. 2,15 Previous studies have proved that DNA methylation is an important way of modulation of genetic epigenetics. Thus, iAs might alter the methylation status of genomic DNA and increase the transcription of some genes. 16 –21
The metabolism of iAs could cause oxidative stress and damage through generating free radicals and active oxygen species, which finally brought about a broad variety of base modification and DNA break. DNA damage was a key initial event in the carcinogenic potential of arsenic. 5,22 –24 Elevated DNA damage levels were observed in the laborers occupationally are exposed to iAs. It had been shown that arsenic concentration in laborers’ urine was correlated with DNA damage levels or skin lesion risks. 9,25,26
Long noncoding RNA (lncRNA) was a new discovery member in non-coded RNA. The promoter of cyclin dependent kinase inhibitor 1A (CDKN1A) antisense DNA damage activated RNA (PANDAR) was first discovered in 2003 as an lncRNA and started from about 1500 bp upstream of CDKN1A transcription initiation site. PANDAR was induced by p53 protein and its knockdown increased apoptosis. PANDAR interacts with nuclear transcription factor Y subunit alpha (NF-YA) or BMI1 proto-oncogene, polycomb ring finger (BMI1) to limit the transcription of apoptosis-related genes. 27,28 The previous studies strongly indicated that DNA methylation in the regulatory sequence was related to the alteration of lncRNA gene expression. 29 Genomic sequences of the promoter region were analyzed by the Methprimer program revealed that there was a high-frequency CpG island between the base sequence sites +1281 to +1404. 30 It was thus likely that PANDAR methylation was linked to the regulation of PANDAR messenger RNA expression.
Arsenic caused DNA damage and PANDAR was induced in DNA damage response. Therefore, in our current research, we investigated whether the ability of arsenic metabolism in individuals affected the expression of lncRNA PANDAR, DNA damage, or DNA methylation.
Materials and methods
Study population
In our case–control study, the exposed group consisted of arsenic-exposed laborers who were employed in the As2O3 smelter, and the control group consisted of agricultural laborers who settled in villages. The farm in the control group was about 10 km away from the arsenic smelter. Exposure group and control group laborers have similar living habits and environment. The arsenic-exposed laborer had at least 3-month exposure to the As2O3. If laborers had been exposed to pesticide, ultraviolet, or alkylating agents in the past 3 months, they would be excluded. We used questionnaires to record demographic characteristics such as age, smoking, education, drinking, and years of service. We used anticoagulant tubes to collect blood. We used plastic bottles to collect urine and each laborer donated 10–20 mL. The research design was agreed by the Theory Committee of Kunming Medical University.
Measurement of urinary arsenic metabolites
The DMA, MMA, and iAs in laborers urine were analyzed as described. 8,9 One milliliter of urine was digested with sodium hydroxide solutions at 95°C for 4 h. Speciation analysis of arsenic was based on the low-temperature separation of liquid nitrogen and hydride generation of volatile arsenic. Measurement of urinary arsenic was performed by an atomic absorption spectrophotometer with an arsenic speciation pretreatment system. The minimum detection limit of the method was 2 ng and the urine arsenic concentration was showed as µg/g creatinine. total arsenic concentrations (tAs), the proportions of urinary arsenic metabolites, and two methylation indices were calculated in accordance with the formula given below.
RNA and DNA extraction
Peripheral blood was stored in the centrifuge tube containing ethylenediaminetetraacetic acid (EDTA). Lymphocytes were separated using Lymphocyte Separation Medium (Solarbio Biotech Company, China). After isolation, we extracted DNA by using the TIANamp Blood Clot DNA Kit (Tiangen Biotech Company, China). We extracted total RNA from lymphocytes using TRNzol (Tiangen Biotech Company) and reversed transcription to complementary DNA using the TIANScriptⅡRT Kit (Tiangen Biotech Company) according to the method of the instructions.
PANDAR expression
Quantitative polymerase chain reaction (Q-PCR) used FastStart Essential DNA Green Master (Roche, Germany) and run on LightCycler (LC)® 96. The sequence of primers are as follows. PANADAR forward: TGCACACATTTAACCCGAAG; PANADAR reverse: CCCCAAAGCTACATCTATGACA and beta-actin (β-actin; internal control) forward: ATTGCCGACAGGATGCAGAA; reverse: GCTGATCCACATCTGCTGGAA. Each polymerase chain reaction (PCR) was repeated three times. PANDAR expressions were normalized to internal control. The amount of gene expression was showed by
Determination of DNA damage level of PANDAR gene
The method was developed by Sikorsky and colleagues.
31,32
DNA damage reduced PCR amplification efficiency and increased Ct value. Q-PCR primers were synthetized to analyze the exon of PANDAR gene and β-actin sequence (control fragment). The following primers were used. PANADAR forward: ATTCTTTCAGGAATGCCGCAGAT; reverse: TCTTTACCAATTCATCCTGGCTCTG and β-actin forward: CGGGAAATCGTGCGTGACAT; reverse: GAAGGAAGGCTGGAAGAGTG. Total DNA were extracted from lymphocytes. Q-PCR was performed on the LC 96. The reaction mixture (15 µL) was composed of 2× FastStart Essential DNA Green Master (Roche), 0.5 µL of 20 µM forward primers, 0.5 µL of 20 µM reverse primers, 1 µL DNA, and 4.5 µL of water. DNA damage was reflected by
DNA methylation analysis
DNA methylation was determined by high-resolution melting (HRM) analysis. 33,34 DNA was modified with EZ DNA Methylation-Gold™ Kits which were produced by the Zymo Research. Determination of DNA methylation has been described previously. 34 MethPrimer was used to design primers. 30 A 182-bp PCR product was obtained from 10 ng bisulfite-treated DNA with outside primers: TGAATTGTGGTATATTATTTTTTTT and ACCTAACCAACATAAAAAAACCCTC.
PCRs contained 0.1 μL of the 182-bp PCR product. We diluted the PCR products, and then DNA methylation tested by the HRM Analysis Kit. HRM analysis was performed using an LC 96 Instrument. The PCR and HRM program run in accordance with the standard procedure of the instrument. The following primers were used: inside forward: TGGAATAGATTTTTAAAATAAGGTTTTT and inside reverse: CCCTCTCTCTACTAAAATACAAAC.
The standard curve was made from the standard products of Zymo Research. The methylation ratio of the standard curve was 0%, 25%, 50%, 75%, and 100%. Standard curves with known methylation ratios were included in each assay and were used to calculate the methylation rate of each reaction. We calculated the percentage methylation of samples by the standard curve in HRM assays. The calculation method was carried out as described previously. 34
Statistical analysis
The R3.4.1 software package was used to calculate the data of the article. PANDAR expression and three arsenic species levels were transformed by logarithm. Student’s t test, Mann–Whitney U test, χ2 test, Spearman correlation coefficient, Pearson correlation coefficient, and multiple linear regression were used as data analysis in our study. The standard of statistical difference was p < 0.05.
Results
Demography of laborers and urinary arsenic species levels
The general information and urinary arsenic species of the subjects were listed in Table 1. Three arsenic species levels, iAs%, and MMA% increased in the exposed group, whereas DMA%, PMI, and SMI were lower. The difference between the two groups was statistically significant.
Demographics of subjects and urinary arsenic levels.

iAs: inorganic arsenic; MMA: monomethylarsonic acid; DMA: dimethylarsinic acid; tAs: total arsenic; PMI: primary methylation index; SMI: secondary methylation index.
a Compared with the control group, p < 0.05.
b Compared with the control group, p < 0.01.
PANDAR expression, DNA damage levels, and DNA methylation among exposure group and control group
PANDAR expression and DNA damage levels were measured by Q-PCR. DNA methylation in the promoter region of the PANDAR gene was measured by methylation-sensitive high-resolution melting (MS-HRM). DNA damage, PANDAR expression, and DNA methylation were shown in Figure 1. Student’s t test was used to analyze gene expression and DNA methylation. Mann–Whitney U test was used to analyze DNA damage. PANDAR expression (4.79 ± 0.22 vs. 3.52 ± 0.0.31, p < 0.01) and DNA damage levels [17.09 (15.70–19.09) vs. 15.78 (14.27–16.34), p < 0.05] were elevated in the exposed group, but DNA methylation (56.33 ± 2.48% vs. 55.95 ± 4.68%, p > 0.05) was not change.

(a) DNA damage levels, (b) PANDAR expression, and (c) DNA methylation in peripheral blood lymphocytes among exposure group and control group. PANDAR expression and DNA damage in the exposure group were higher than those in the control group. The line inside each box represents the median; the upper and lower limits of the box were the 75th and 25th percentiles. PANDAR expression and DNA methylation were expressed as
The interrelationship between arsenic exposure and the level of PANDAR expression, DNA damage, and DNA methylation
The interrelationship between arsenic exposure and level of PANADR expression, DNA damage, and DNA methylation in laborers are presented in Tables 2 and 3. The level of DNA damage and PANDAR expression were positively linked to the level of arsenic in urine. There was no interrelationship between DNA methylation and the level of arsenic in urine. The PANDAR expression and DNA damage were positively linked to MMA% and negatively linked to DMA%.
Association between arsenic species and level of PANDAR expression, DNA damage, and DNA methylation in all subjects.
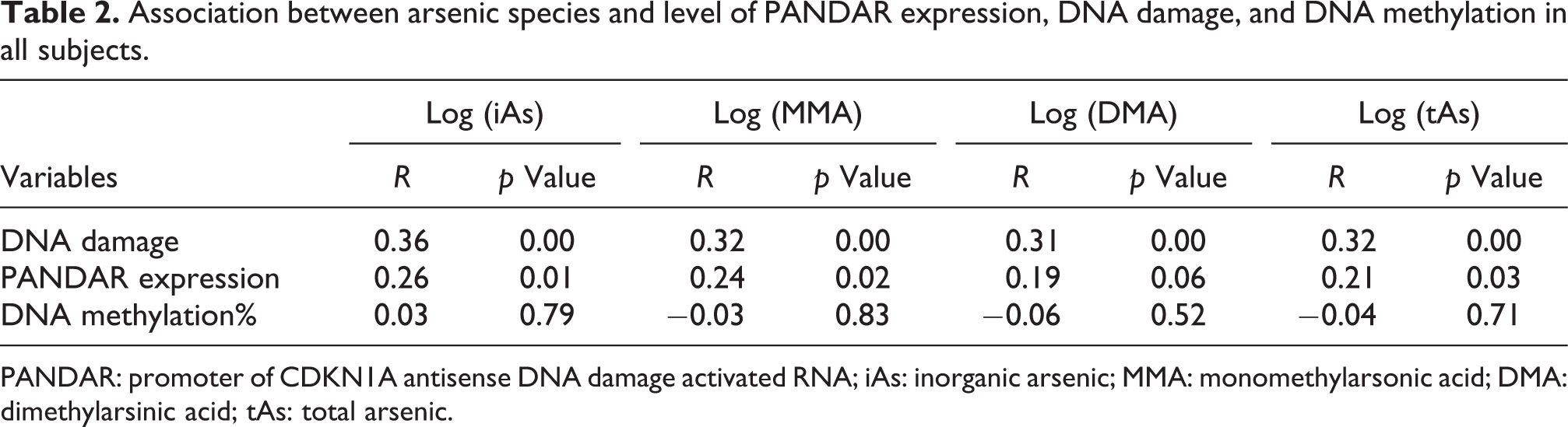
PANDAR: promoter of CDKN1A antisense DNA damage activated RNA; iAs: inorganic arsenic; MMA: monomethylarsonic acid; DMA: dimethylarsinic acid; tAs: total arsenic.
Association between the percentages of urinary arsenic and level of PANDAR expression, DNA damage, and DNA methylation in all subjects.
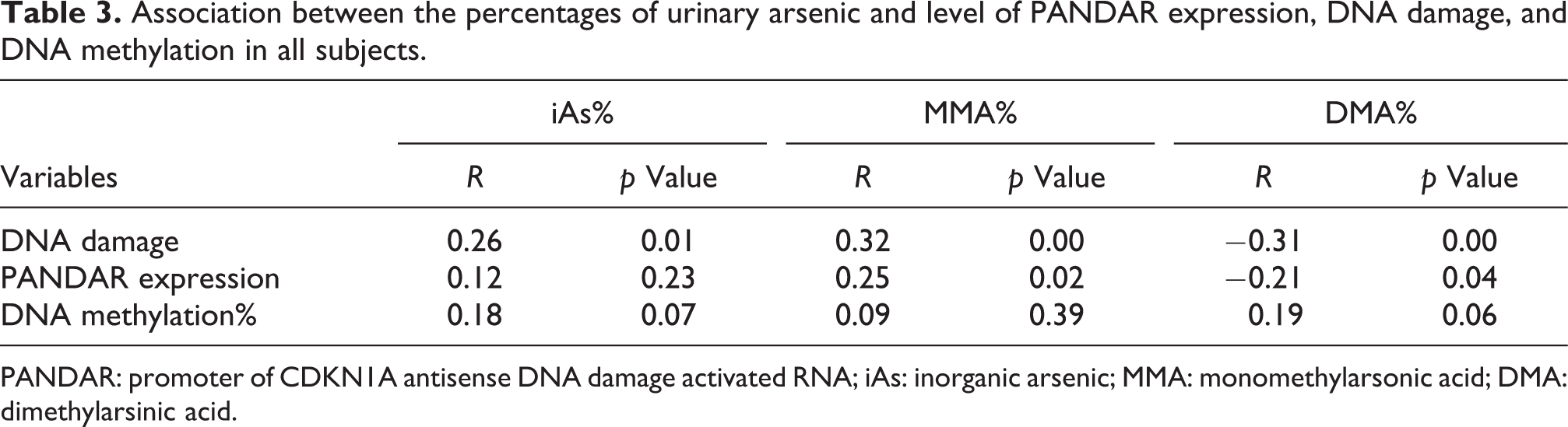
PANDAR: promoter of CDKN1A antisense DNA damage activated RNA; iAs: inorganic arsenic; MMA: monomethylarsonic acid; DMA: dimethylarsinic acid.
Association of PMI and SMI with the level of DNA damage, PANDAR expression, and DNA methylation in subjects
All the laborers were divided into two groups according to PMI or SMI. The results were summarized in Tables 4 and 5. Levels of DNA methylation and DNA damage in the high PMI laborers were lower than that in the low PMI laborers. Levels of PANDAR expression and DNA damage in high SMI laborers were lower than that in low SMI laborers.
Comparisons of DNA methylation, gene expression, and DNA damage stratified by PMI levels in all subjects.

PMI: primary methylation index; PANDAR: promoter of CDKN1A antisense DNA damage activated RNA.
a Compared with the low PMI group, p < 0.05.
Comparisons of DNA methylation, gene expression, and DNA damage stratified by SMI levels in all subjects.

SMI: secondary methylation index; PANDAR: promoter of CDKN1A antisense DNA damage activated RNA.
a Compared with the low SMI group, p < 0.05.
b Compared with the low SMI group, p < 0.01.
The interrelationship between gene expression and DNA methylation or DNA damage in subjects
Linear regression analysis was used to analyze the relationship between gene expression and DNA methylation. There was inversely correlated between the methylation and PANDAR expression in peripheral blood lymphocytes. The results were shown in Figure 2. We did not observe correlations between gene expression and DNA damage.
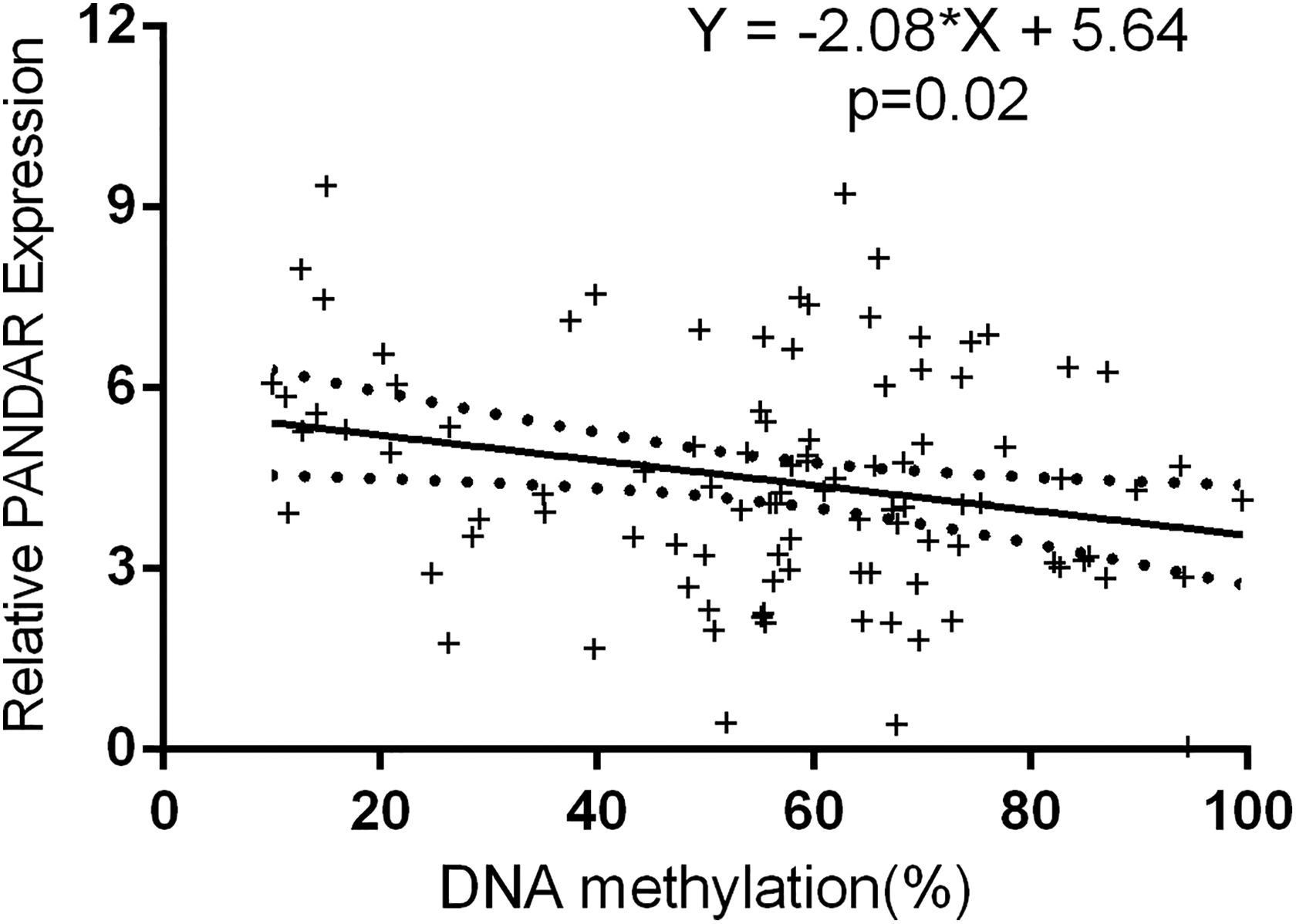
Relationship between DNA methylation and PANDAR expression in all subjects. The X-axis shows the DNA methylation and the Y-axis shows the level of PANDAR expression in peripheral blood lymphocytes. Black dots represent a 95% confidence interval. PANDAR: promoter of CDKN1A antisense DNA damage activated RNA.
Discussion
The arsenic smelting plant laborer inhaled high levels of As2O3 through the air. Human metabolism of As2O3 led to the production of the methyl metabolites, MMA and DMA. 1 –3 In our study, we had measured the metabolites of arsenic in urine. The results clearly showed that the contents of iAs, MMA, and DMA in the exposed group were much higher than those in the control group. The urinary arsenic level of workers in our study was higher than that of people exposed to the environment, 4,21,24,35 which was consistent with previous occupational exposure studies. 9 Our study showed that MMA% and iA% were higher and DMA% was lower in the exposed group. The results also suggested that increased exposure to iAs alters the proportion of metabolites in urine. 9,24,36,37
Arsenic exposure linked to changes in gene expression patterns in cells or populations. 17,19,20,38 Studies had shown that iAs could change the expression of apoptosis-related genes. 19 PANDAR interacted with NF-YA to limit the transcription of apoptosis-related genes. 27,28 In our study, arsenic exposure increased PANADR expression and the PANDAR expression was positively linked to the concentration of urinary tAs, DMA, and MMA among arsenic smelting plant laborers. iAs would be transformed to MMA and DMA. DMA could be rapidly discharged by the kidneys, but iAs and MMA were not easily discharged. 39 MMA, DMA, and iAs had different effects on gene expression. 19 Our research indicated that the PANDAR expression was positively linked to MMA% and negatively linked to DMA%. The possible reason was that the effect on gene expression was weakened after iAs is metabolized.
Arsenic caused DNA damage by producing active oxygen species, which finally brought about a broad variety of base modifications and DNA breaks. 9,18,24,25 The experiments showed that PANDAR was induced by DNA oxidative damage. 27 Our results suggested that the level of DNA damage was positively correlated with arsenic exposure and MMA% and negatively correlated with DMA%. We further investigated whether DNA damage was associated with gene expression. The final results suggested that there was no interrelationship between DNA damage and PANDAR expression. Our data also showed that individual metabolic capacity was very important in arsenic exposure-induced injury. 9,40
It has been proved that the disorder of gene transcription caused by arsenic can be controlled by epigenetics. 17 DNA methylation was an important way of modulation of genetic epigenetics. Metabolism of iAs reduced SAM which is required to DNA methylation as a methyl donor and arsenic appeared to restrain DNMT activity. 14 Chronic arsenic exposure could lead to changes in genome methylation and has been linked to altered gene expression patterns. 5,16,17,20,35 To further identify PANDAR expression with methylation changes, the MS-HRM technique was applied to determine methylation levels in the promoter region of PANDAR gene. The result showed that the methylation of PANDAR was significantly associated with the suppression of its RNA expression. Abnormal DNA methylation was related to the transcriptional changes of genes. 29,38
Recent findings highlight that the methylation metabolism of arsenic was related to various health effects. PMI and SMI reflect the ability of arsenic methylation in vivo. Some studies had shown that individuals with low methylation ability were distinguished by lower urine SMI, were more likely to suffer from arsenic-related diseases. 1 –3,21,26,37 Our data suggested that subjects with low methylation ability had significantly higher levels of gene expression, DNA damage, and DNA methylation than those with high metabolic efficiency. Our results are consistent with previous findings that arsenic metabolism affects gene expression levels, DNA damage, and DNA methylation. 3,21,26,37 The mechanism responsible for the outcome that exposure to iAs would lead to an increased level of gene expression, DNA damage, and methylation might attribute to various health hazards induced by arsenic.
Conclusion
In summary, our results suggested that arsenic metabolism ability affected the expression of lncRNA PANDAR, DNA damage, and DNA methylation. PANDAR expression and DNA damage in peripheral blood lymphocytes of laborers were potential biomarkers of effects. These facts provided clues for elucidating the health hazards mechanism of arsenic.
Footnotes
Acknowledgment
The authors acknowledge Weihua Weng for his technical input.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China [Grant Number 81460491] and Yunnan Applied Basic Research projects [Grant Number 2017FB123].

