Abstract
Coconut oil (CO) is enriched with medium chain saturated fatty acids like lauric acid (LA), capric acid and caprylic acid, which are known to have several health benefits. LA, the predominant fatty acid in CO, is reported to possess anticancer activity mediated through oxidative stress-induced apoptosis; however, there is no clear information on its cellular signalling mechanism. The present study screened the anticancer potential of various fatty acids present in CO (capric acid, caprylic acid and LA) using in silico tools such as CDOCKER in Accelrys Discovery Studio by targeting proteins like epidermal growth factor receptor (EGFR), cyclin-dependent kinase and thymidine synthase (TS). The results were further confirmed using cell culture-based studies and quantitative PCR. Among the tested compounds, LA was found to be the most active and showed a higher affinity towards EGFR and TS. Corroborating with these results, LA-induced dose-dependent cytotoxicity towards HCT-15 (human colon cancer), HepG2 (human hepatocellular carcinoma) and Raw 264.7 (murine macrophages) cells exhibiting morphological characteristics of apoptosis. Further, in HCT-15 cells exposed to LA (30 and 50 µg/mL), the expression of EGFR was found to be downregulated by 1.33- and 1.58-fold. The study thus concludes that the anticancer activity of LA may be partially mediated by the downregulation of EGFR signalling and consequent reduction in cell viability through apoptosis. Since EGFR signalling is crucial in cancer cell survival and is a prime target in drug development, the present study has pharmacological significance.
Introduction
Coconut oil (CO) has been used for edible and nonedible purposes worldwide. Different forms of CO exist, which include the virgin CO, copra oil as well as refined bleached and deodorized oil. 1 –3 Fatty acid profile of these forms is similar, with higher amounts of medium chain saturated fatty acids (MCFAs), such as lauric acid (LA; 48–52%). CO also contains higher levels of polyphenols such as ferulic, coumaric, gallic acids, catechin, kaempferol or their sugar derivatives. Studies have reported the health beneficial effects of CO under different conditions of hyperlipidemia, systemic and hepatic damages. 4 –7 However, limited studies have been conducted on the anticancer potential of CO. Craig-Schmidt et al. 8 observed a significant reduction in DMBA-induced mammary tumorigenesis in Balb/c mice treated with a combination of CO and menhaden oil. Later studies by Enos et al. 9 observed a significant reduction in the ulcerative colitis incidence in an animal fed with CO, which is mediated by downregulating the chronic inflammatory cytokine expressions as well as preventing apoptotic changes in the colon mucosa.
Studies on the individual fatty acid components of oil revealed the potential role of MCFAs, mainly LA. LA is shown to have different pharmacological properties including anti-diabetic, 10 hypotensive in spontaneously hypertensive rats 11 and known to be protective of neuronal cells by promoting ketone body formation. 12 In addition, anticancer activities of LA have been reported previously in colon and breast cancer cells. In colon cancer cells (Caco2), apoptotic changes and cell cycle arrest in G0/G1 and G2/M phases have been induced by LA treatment at a dose of 0.5 mM (about 100 µg/mL); the treatment also increased intracellular reactive oxygen species with a concomitant decrease in the intracellular reduced glutathione levels. 13,14 Apart from these reports, there is no information on the role of cellular signalling pathways with respect to the anti-colon cancer potential of LA. Similarly, in breast cancer cells, 100 µM LA treatment (about 20 µg/mL) induced apoptosis mediated through the phosphorylation of epidermal growth factor receptor (EGFR) and also Rho-associated kinase pathway. 15 Although these reports indicated that LA exerts its anticancer activity mediated by apoptosis, there is no available information on its role in various pro-survival pathways in carcinogenesis and progression.
There are several different pathways associated with cancer progression and cell survival, among these EGFR, cyclin-dependent kinase (CDK), 16 dihydrofolate reductase (DHFR), 17 thymidylate synthase (TS), 18 vascular endothelial growth factor receptor-2 kinase (VEGFR), 19 estrogen receptor (ER) 20 and B-cell lymphoma-extra-large (Bcl-xl) 21 are the important. Among these different pathways, the EGFR axis is the important one. 22 –24 Several of the natural products are known to interfere with these pathways and induce apoptosis in cancer cells. 25 –27 Specifically, the EGFR pathway downregulation induces cytotoxic and anti-metastatic effects 27 –29 in various cancers and also showed significant improvement in the survival rate of cancer patients. 30
Considering the roles of EGFR pathway on cancer progression and survival, the present study aims to evaluate the role of LA in inhibiting EGFR expression and induction of cytotoxicity. In addition, the interaction of LA with other cellular pathways such as CDK, VEGFR, TS and DHFR is also analysed using in silico and in vitro techniques.
Materials and methods
Chemicals and cell lines
LA and MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) were procured from Sigma-Aldrich (St. Louis, Missouri, USA). Cell culture media and other reagents were obtained from Invitrogen (Carlsbad, California, USA). Kits and reagents for real-time quantitative PCR (qPCR) were purchased from Promega (Madison, Wisconsin, USA). All other reagents and chemicals used in the study were of reagent grade. Cell lines such as HCT-15, HepG2 and Raw 264.7 were obtained from National Centre for Cell Sciences, Pune, Maharashtra, India.
Molecular docking studies
The molecular targets chosen in this study are EGFR, CDK, 16 DHFR, 17 TS, 18 VEGFR, 19 ER 20 and Bcl-xl, 21 which are reported to be involved in the initiation, progression and cachexia events of cancers, especially in colon cancers. In addition, these molecules are the active targets for cancer prevention and drug discovery. The molecular targets such as for cancer were selected for the study. The X-ray crystallographic structures of these target proteins were retrieved from the protein data bank (PDB) (Online supplemental material 1). The 3D structures of LA, capric acid, caprylic acid, myristic acid, gallic acid and caffeic acid taken as ligands were retrieved from PubChem database. The downloaded 3D structures of proteins and ligands were cleaned and processed in Discovery Studio 4.0. The protein structures were then energy minimized by applying the CHARMM force field to remove the steric clashes between the atoms in order to get a stable conformation as per the previous reports of Kim et al. 31 The ligands were filtered using Veber and Lipinski’s rule as previously defined by Lipinski and colleagues. 32,33 The binding sites of the target receptors were predicted based on the ‘receptor cavity method’. Using this protocol, active sites of the target receptor were identified based upon the inhibitory property of the amino acid residues present in the binding site.
In the docking analysis, each ligand was docked with the target proteins. Strategies of Discovery Studio 4.0 exhaustively docks or score possible positions of the ligand binding site of the target protein. A molecular dynamics (MD) simulated-annealing-based algorithm, namely, CDOCKER, was used to score the interacting compounds as defined by Puratchikody et al. 34 The final poses were then scored based on the total docking energy, which is composed of intramolecular energy of ligand and the ligand–protein interactions. The lowest energy structure was taken as the best fit. Interpretation of the values was done using standards provided by Discovery Studio such as CDOCKER energy, CDOCKER interaction energy, hydrogen bonds and binding energy.
The MCFAs, which gave better dock results, were further filtered through ADMET descriptors as per previous reports of Lagorce et al. 35 This includes evaluation of molecular properties that influence absorption, distribution, metabolism, excretion and toxicity which are recognized as a long side therapeutic potency as key determinants of whether a molecule can be successfully developed as a drug. 36
In vitro toxicity study using cell lines
The cells were seeded at a density of 1 × 105 cells/mL on the flat bottom of a 96-well plate and allowed to reach sub-confluency. Different concentrations of LA (0–80 µg/mL) were added and incubated for 48 h. The cell viability was determined using MTT assay according to the method described by Mosmann. 37 The morphological changes were analysed at 200× magnification under a phase contrast objective of an inverted microscope (Magnus INVI, Bangalore, Karnataka, India). The images were captured using Magnus Pro software (Magnus, Bangalore, Karnataka, India)
The real-time qPCR analysis
The HCT-15 cells (1 × 105 cells/mL) were plated in 12-well plates and allowed to adhere overnight. At sub-confluency, the cells were treated with high (50 µg/mL) and low dose (30 µg/mL) of LA for 12 h. Total cDNA was synthesized using SuperScript™ III CellsDirect cDNA Synthesis System (Invitrogen, Carlsbad, California, USA). The qPCR amplification was carried out in Applied Biosystems 7300 to study the gene expression profile of EGFR using β-actin as the reference gene. The master mix used in PCR consisted of 12.5 µL SYBR Premix Ex Taq (2X), 1 µL of each forward and reverse primers, 0.5 µL ROX Reference Dye and 5 µL template.
Custom-designed primer sequence of EGFR were: Forward: 5′GAGACGAGAACTGCCAGAA3′. Reverse: 5′GTAGCATTTATGGAGAGTC3′.
Ct values obtained from the melting curve of qPCR were used to calculate the ΔCt values and ΔΔCt values were represented as fold change. 38
Statistical analysis
Cell culture studies were conducted in three independent sets of the experiment, each in triplicate. The values are represented as mean ± SD of the three experiments.
Results
Molecular docking
The results of the present study showed more than 30 amino acids in the active site of protein-DHFR, whereas only two in VEGFR. The number of amino acids varied according to proteins and the strength of interactions. Among the six tested ligands, five passed the Veber and Lipinski’s rule. Myristic acid failed the Veber and Lipinski’s rule since its log P value exceeded 6.
MCFAs in coconut recorded binding energy ranging between −78.91 and −145.8 kcal/mol with the target EGFR. The binding energy of LA recorded was −102.75 kcal/mol with hydrogen bond distance of 2.1 Å. The difference in CDOCKER energy and CDOCKER interaction energy was 4.32 kcal/mol. Also, it binds with MET at 769th position in the identified active site (Table 1). In the case of CDK, the least binding energy was recorded for capric acid (−113.86 kcal/mol) compared to other ligands. The CDOCKER energy and CDOCKER interaction energy of capric acid were also high with minimum energy difference (Online supplemental material 2). From the results, it is clear that LA scored the least binding energy with a value of −185.42 kcal/mol against the target DHFR. Almost all the ligands had significant interaction with the TS. The LA formed three hydrogen bonds with the two amino acids in the active sites of TS, and also recorded the least binding energy (−213.1 kcal/mol), higher CDOCKER energy and CDOCKER interaction energy. Further, LA showed significant interactions with the other target proteins VEGFR, ER and CDK (Table 1).
Different dock scores obtained for cancer targets against LA as the ligand.

LA: lauric acid; EGFR: epidermal growth factor receptor; CDK: cyclin-dependent kinase; DHFR: dihydrofolate reductase; TS: thymidylate synthase; VEGFR: vascular endothelial growth factor receptor-2 kinase; ER: estrogen receptor; Bcl-xl: B-cell lymphoma-extra-large. *p < 0.05.
ADME Toxicity
The ADMET solubility levels were represented as ‘0–4’, where ‘4’ being the highly soluble and ‘0’ the least soluble. The ADMET of blood–brain barrier crossing (BBB level) and absorptivity also ranges between 0 and 4, where ‘0’ being the highly absorptive and ‘4’ the least absorbable. The three MCFAs (LA, capric acid and caprylic acid) and the phenolic compound, caffeic acid, recorded good pharmacokinetic descriptors. These compounds passed the ADMET filters by exhibiting scores that fall within the optimum level (Table 2). Moreover, the BBB level of gallic acid was 3, exceeding the optimum range. Nine FDA-approved therapeutic drugs used in docking analysis were also filtered through ADMET descriptors. Pharmacokinetic properties of these drugs obtained from ADMET analysis are presented in Table 2. The solubility level of Sorafenib and Tamoxifen was found to be very low. Letrozole, Gefitinib and Tamoxifen had the BBB level in the preferred range; however, for other drugs, the BBB level was found undefined or too low. In order to achieve effective absorption in the body, the absorption level should be 0. Drugs like Fluorouracil, Raltitrexed, Methotrexate, Ponatinib and Tamoxifen had a pharmacokinetic absorption level ≥1, indicating the reduced absorption in the body; whereas, the tested MCFAs and polyphenols of CO had higher pharmacokinetic absorptivity. The hepatotoxicity scores were calculated by ADMET programme, where those obtained a score below −4.0 are considered as false and those above −4.0 are considered as true, where true being hepatotoxic and false being non-toxic. All the standard chemotherapeutic drugs used in this were found to be hepatotoxic; on contrary, the tested coconut phytocompounds, except gallic acid, were least toxic towards the liver.
ADME toxicity scores of different coconut phytocompounds and commercial anticancer drugs.a

BBB: blood–brain barrier.
aThe ADMET solubility levels have been represented as ‘0–4’, where ‘4’ being the highly soluble and ‘0’ the least soluble. The ADMET of BBB crossing and absorptivity also ranges between 0 and 4, where ‘0’ being the highly absorptive and ‘4’ the least absorbable. The hepatotoxicity scores were calculated by ADMET programme, where those obtained a score below −4.0 are considered as false and those above −4.0 are considered as true, where true being hepatotoxic and false being non-toxic.
Cytotoxicity analysis
The in vitro cytotoxicity study indicated that the LA had a dose-dependent effect on the cell growth of different cell lines such as HCT-15, HepG2 and Raw 264.7. The MTT assay showed a gradual increase in cytotoxicity with the escalating doses of LA from 0 to 80 µg/mL, where untreated cells considered as 0% cell death. In HCT-15 cells, the percentage of cell death at doses of 10, 30, 40, 60 and 80 µg/mL was 13.7 ± 2.46, 45.3 ± 2.11, 58.8 ± 1.46, 75.1 ± 4.01 and 97.5 ± 3.22%, respectively (Figure 1). Similarly, in HepG2 cells, the respective doses of 10, 30, 40, 60 and 80 µg/mL LA induced 15.1 ± 1.68, 28.6 ± 1.91, 45.7 ± 2.48, 65.7 ± 1.09 and 86.5 ± 3.41% cell death. In murine macrophage cell, Raw 264.7, the same doses of LA induced 18.2 ± 2.93, 48.1 ± 2.01, 64.3 ± 4.08, 90.9 ± 1.06 and 98.1 ± 5.01% cell death at 48 h. The IC50 values were found to be 39.0 ± 1.29, 46.0 ± 2.08 and 36.0 ± 1.87 µg/mL, respectively. These morphological alterations that are characteristics of apoptosis were visible in the LA-treated cells (Figure 2). The results emphasize the anticancerous activity of LA in inhibiting the mass multiplication of cancer cells.
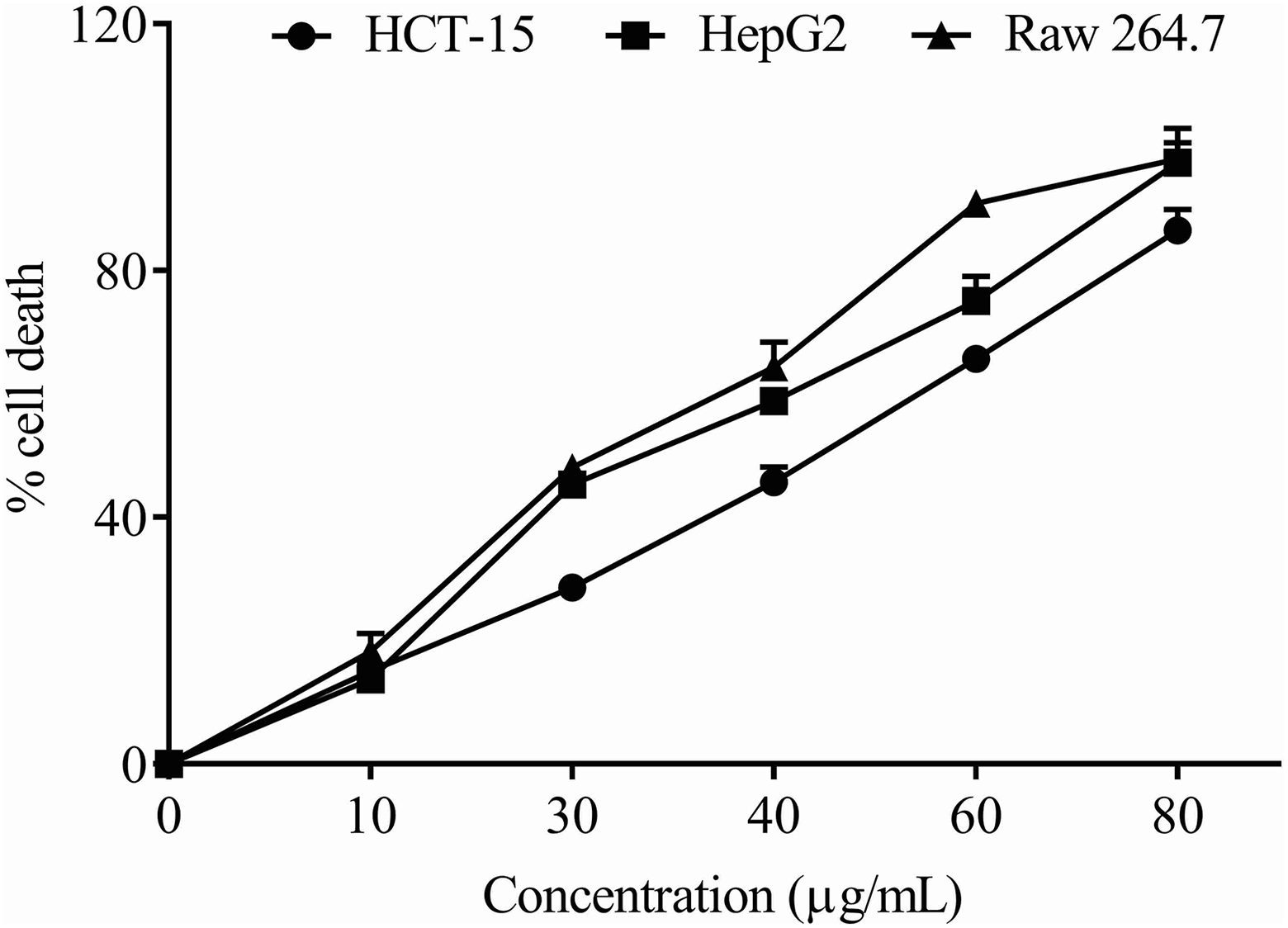
Cytotoxicity analysis of LA on HCT-15 (human colorectal cancer), HepG2 (human hepatocellular carcinoma) and Raw 264.7 (murine macrophage) using MTT assay. MTT: 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide; LA: lauric acid.
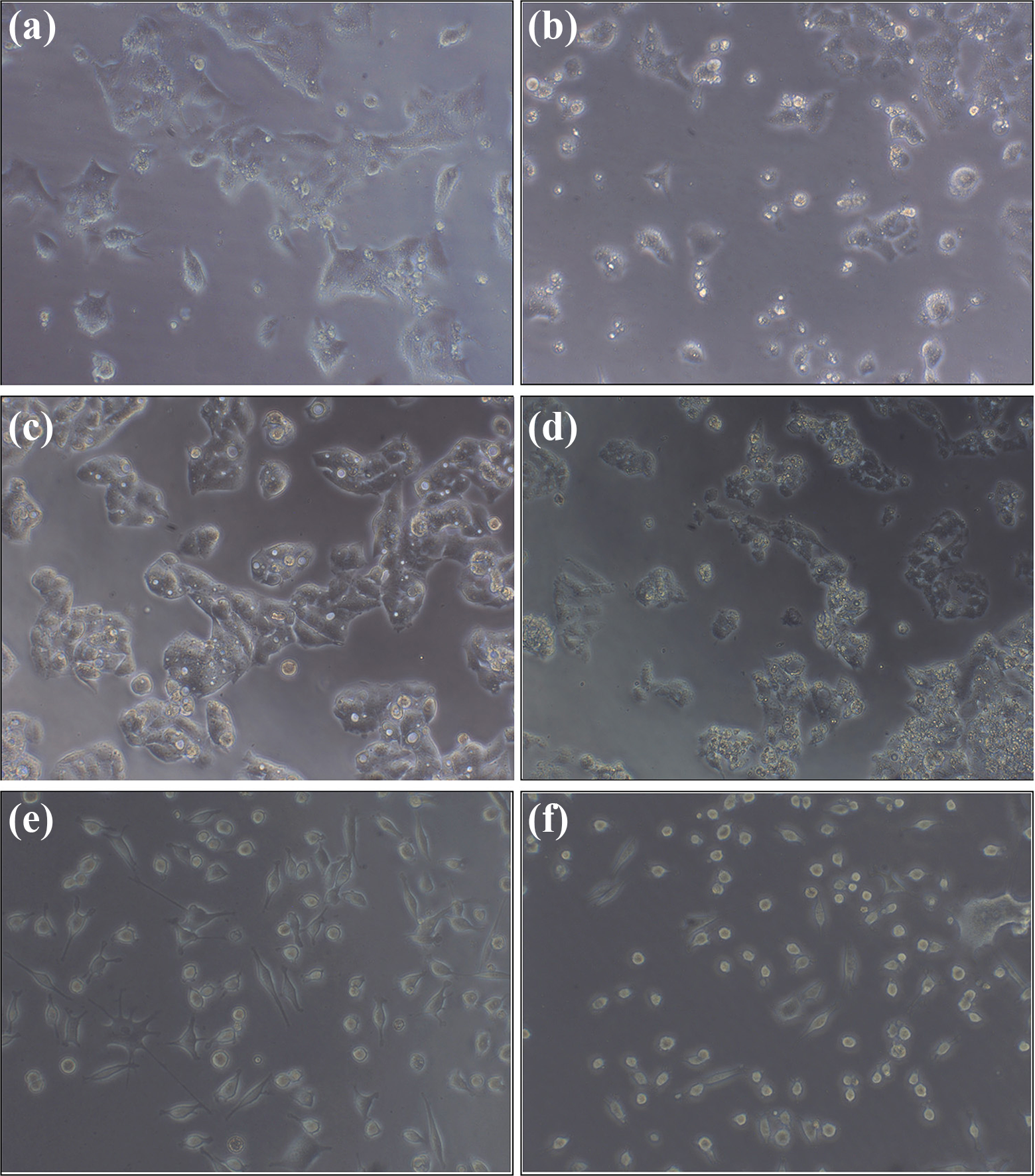
The effect of LA on the morphology of various cells: (a, c and e) untreated HCT-15, HepG2, and Raw 264.7 cells; (b, d, and f) LA (at its IC50 values)-treated HCT-15, HepG2 and Raw 264.7 cells. The cells treated with LA show significant alterations in their cell membrane structure and nuclear condensation is also evident in these cells. In addition, shrinking of the cells treated with the LA (at its IC50 concentration) has been observed. The images were captured using Magnus INVI microscope (Magnus, Chennai, Tamil Nadu, India) at 20× phase contrast objective. LA: lauric acid.
Morphological changes in cells induced by LA treatment in cells
Untreated HCT-15, HepG2 and Raw 264.7 cells showed normal morphology with uniformly spread cells throughout the culture plate. In the lower doses of LA treatment (10 and 30 µg/mL), there observed a minor lipid accumulation as well as cell membrane bulging. In the medium doses (40 and 60 µg/mL), the membrane blebbing and nuclear condensation was visible (Figure 2), with the mild incidence of necrotic cells. However, in the high-dose LA (80 µg/mL)-treated cells, the cells were visualized to be shrunk, entire nuclear materials were condensed and cell membrane integrity was lost.
Real-time qPCR analysis
The gene expression was quantified using the 2−ΔΔCt method and accordingly, a positive ΔΔCt indicates downregulation and a negative ΔΔCt indicates upregulation. The ΔΔCt values observed for the gene EGFR are presented in Figure 3(b). When the cells were treated with 30 µg/mL of LA, a 1.33-fold decrease in the EGFR gene expression was noticed. In the cells treated with 50 µg/mL of LA, the expression of EGFR was decreased by 1.58-fold.

(a) Hydrogen bond interaction between LA and EGFR. (b) Real-time qPCR analysis of EGFR expression in HCT-15 cells treated with different doses of LA (30 and 50 µg/mL). The results are expressed as the fold change with respect to the normal untreated cells. LA: lauric acid; EGFR: epidermal growth factor receptor; qPCR: quantitative PCR.
Discussion
The results indicate that the MCFAs of CO (LA, capric acid, caprylic acid, caffeic acid and gallic acid) interact with the selected targets identified for cancers, of which the most notable has been observed as TS and EGFR. TS is a crucial enzyme in the folate pathway which plays significant roles in the DNA synthesis and cell division. 39 The inhibition of TS activity has been known to prevent the abnormal synthesis of DNA and thereby reducing the severity of colorectal, pancreatic, ovarian, gastric and breast cancers. 40 On TS, the binding energy of LA recorded the lowest followed by caffeic acid. All the MCFAs studied recorded good CDOCKER energy and CDOCKER interaction energy with minimum energy difference in all. Hydrogen bond interactions between the lead ARG residues (in the 50th position) of TS and the MCFAs, especially with LA, indicate a strong and stable complex formation. Besides TS, LA shows inhibition of EGFR; it has been reported that EGFR signalling is actively involved in the regulation of apoptosis resistance and cell proliferation. 22 –24 Mutations that lead to EGFR overexpression or upregulation have been associated with a number of cancers, especially those of lung and colon. 41
Further, studies using cancer cells in vitro confirmed the observations of in silico analysis. The LA treatment induced a dose-dependent toxicity in various cancer cells such as HCT 15 (human colorectal cancer), HepG2 (human hepatocellular carcinoma) and Raw 264.7 (murine macrophage cells). These results are in line with the previous reports of Fauser et al., 13 where they observed apoptosis induction in LA-treated colorectal cancer cells mediated by reactive oxygen species. It is thus expected that the cytotoxicity induced by LA in HCT-15 cells in this study may also be mediated by apoptotic cascade; further detailed studies are needed for the confirmation of these aspects. Similarly, Lappano et al. 15 have also shown that LA treatment induces apoptotic events in breast cancer cells, where morphological changes of apoptosis mediated by the activation of a Rho-associated kinase pathway are observed. Similarly, our studies have also observed morphological alterations associated with apoptosis in LA-treated cells.
In previous studies, LA induced apoptosis in breast cancer cells at a dose of 100 µM (about 20 µg/mL). On contrary, in colon cancer cells (Caco2), apoptotic changes were induced by a dose of 0.5 mM (about 100 µg/mL). 15 In the current study, a dose of 50 µg/mL has been found to be the median lethal concentration for HCT-15 cells. It is thus possible that the dose and response to LA vary with the origin and nature of cells.
Results of the present study emphasize a reduction in EGFR expression in LA-treated cells. Previous studies have also indicated that EGFR downregulation corresponds to a reduction in cell viability in colon cancer cells. 29,42 –44 In contrast, the previous study by Lappano et al. 15 had shown that LA triggers apoptosis in breast cancer cells mediated through the phosphorylation events of EGFR. It is therefore assumed that the disparity observed in the EGFR signalling in the two studies may be dependent on the origin of cancer cells; however, further studies are necessary to ascertain this assumption.
In conclusion, the present study highlights the interaction of MCFAs present in CO with various protein targets involved in cancer, especially EGFR and TS. Among these MCFAs, LA, the predominant fatty acid of CO, has been found to have higher efficacy. It is thus possible that by virtue of these MCFAs and other antioxidant polyphenols, CO may be a promising nutraceutical for cancer prevention.
Supplemental Material
Supplementary_Material_1 - Lauric acid induce cell death in colon cancer cells mediated by the epidermal growth factor receptor downregulation: An in silico and in vitro study
Supplementary_Material_1 for Lauric acid induce cell death in colon cancer cells mediated by the epidermal growth factor receptor downregulation: An in silico and in vitro study by DL Sheela, A Narayanankutty, PA Nazeem, AC Raghavamenon and SR Muthangaparambil in Human & Experimental Toxicology
Supplemental Material
Supplementary_material_2 - Lauric acid induce cell death in colon cancer cells mediated by the epidermal growth factor receptor downregulation: An in silico and in vitro study
Supplementary_material_2 for Lauric acid induce cell death in colon cancer cells mediated by the epidermal growth factor receptor downregulation: An in silico and in vitro study by DL Sheela, A Narayanankutty, PA Nazeem, AC Raghavamenon and SR Muthangaparambil in Human & Experimental Toxicology
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This work was financially supported by DBT, Government of India and Kerala Agricultural University, Vellanikkara, Thrissur (Order no. R7/66460/2013 ii). A Narayanankutty is a recipient of the Council of Scientific and Industrial Research-Senior Research Fellowship (09/869(0012)/2012 EMR-I).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
