Abstract
The promise of precision medicine is now a clinical reality. Advances in our understanding of the molecular genetics of colorectal cancer (CRC) are leading to the development of a variety of biomarkers that are being used as early detection markers, prognostic markers, and markers for predicting treatment responses. This is no more evident than in the recent advances in testing CRCs for specific molecular alterations in order to guide treatment with the monoclonal antibody therapies cetuximab and panitumumab, which target the epidermal growth factor receptor. In this review, we update a prior review published in 2010 and describe our current understanding of the molecular pathogenesis of CRC and how these alterations relate to emerging biomarkers for early detection and risk stratification (diagnostic markers), prognosis (prognostic markers), and the prediction of treatment responses (predictive markers).
Keywords
Introduction
The promise of precision medicine is becoming a clinical reality, with colorectal cancer (CRC) molecular genetics at the forefront of this major advance in clinical medicine. The most clear example of the role that molecular alterations can have on improving clinical care is the testing of CRCs for specific molecular alterations in order to guide treatment with the monoclonal antibody (mAb) therapies cetuximab and panitumumab, which target the epidermal growth factor receptor (EGFR; Allegra et al. 2009). Indeed, the discovery that acquired Kirsten rat sarcoma gene (KRAS) mutations are a robust predictive marker of resistance to cetuximab and panitumumab (Karapetis et al. 2008; Siena et al. 2009) has led to clinically validated and cost-effective testing strategies to direct these drugs to patients who have the best chance of responding to these agents. This discovery resulted from a detailed understanding of the molecular pathology of CRC, including the role of KRAS mutations in colorectal carcinogenesis, as well as knowledge of the EGFR signaling pathways (Vogelstein et al. 1988). The success of KRAS mutation testing in predicting treatment response is just the beginning of the use of genetic markers for directing the care of CRC patients. Many other molecular markers in CRC show promise for their use in treatment selection, prognosis, and early cancer detection. In this context, knowledge of the underlying genetic and epigenetic alterations of colorectal tumorigenesis and the potential of specific molecular alterations for clinical decision making is expected to become part of the working knowledge of care providers managing CRC patients. However, despite the promising advances in the molecular pathology of CRC that are highlighted in this review, it is important to emphasize that clinicopathological staging and histologic assessment of tumor tissue is still the cornerstone of prognostication and treatment selection. The modern tumor node metastasis (TNM) classification system is recommended, although the original Dukes staging system is still used by some clinicians and is taught to pathologists in training (Shia et al. 2012). The pathologic features with greatest prognostic power are depth of tumor invasion, burden of lymphovascular invasion (estimated by the number of lymph nodes infiltrated by cancer), and presence of distant metastases. Efforts to correlate molecular alterations with histologic features have had limited success, although microsatellite instability (MSI) is a molecular feature that shows modest correlation with certain histologic features such as cribriform architecture and medullary histology (Alexander et al. 2001). Thus, molecular testing is usually required for accurate assessment of specific gene mutations, epigenetic alterations, or genomic instability that provides prognostic and predictive information beyond clinicopathologic features.
In this symposium review, we have updated a review published in 2011 (Pritchard and Grady 2011). We examine genetic and epigenetic mechanisms associated with CRC and discuss how these alterations relate to emerging biomarkers for early detection and risk stratification (diagnostic markers), prognosis (prognostic markers), and the prediction of treatment responses (predictive markers; Table 1). The molecular features of CRC that are currently most clinically useful will be emphasized in this review, and a detailed description of the molecular genetics and molecular biology of the germane genetic and epigenetic alterations will be provided. We conclude by reviewing the potential role for molecular markers in the selection of targeted CRC therapies that are in preclinical development or in phase I and II trials.
Selected biomarkers that have been evaluated in colorectal cancer.
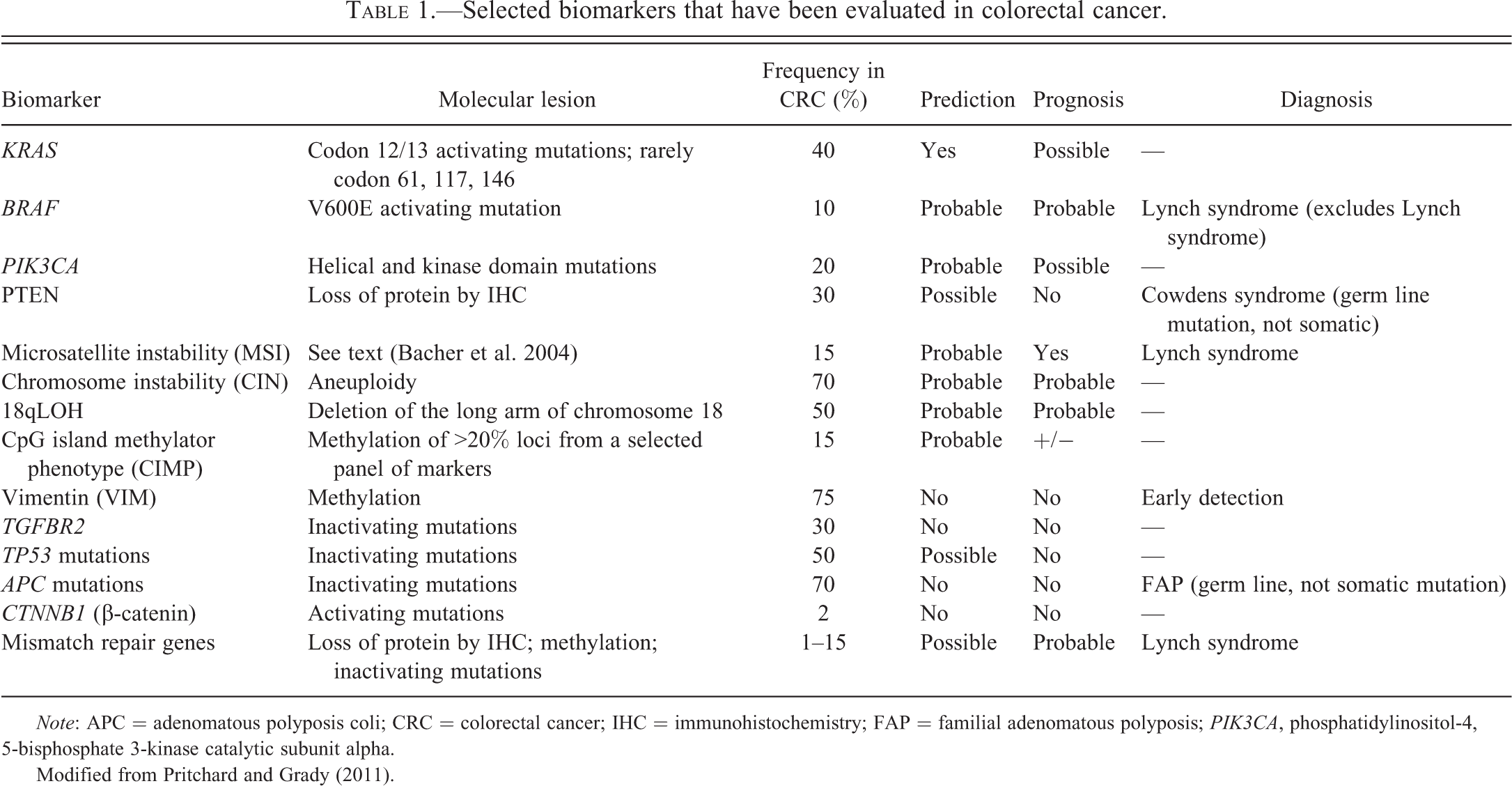
Note: APC = adenomatous polyposis coli; CRC = colorectal cancer; IHC = immunohistochemistry; FAP = familial adenomatous polyposis; PIK3CA, phosphatidylinositol-4, 5-bisphosphate 3-kinase catalytic subunit alpha.
Modified from Pritchard and Grady (2011).
Molecular Mechanisms of Colorectal Carcinogenesis
The Polyp/Carcinoma Progression Sequence
CRC arises as the result of the accumulation of acquired genetic and epigenetic changes that transform normal glandular epithelial cells into invasive adenocarcinomas. Steps that transform normal epithelium to benign neoplasms (adenomas and sessile serrated polyps [SSP]), followed by invasive carcinoma, and eventually metastatic cancer are described in the classic tumor progression model proposed by Fearon and Vogelstein (Figure 1; Vogelstein et al. 1988). Since this model was originally proposed, our understanding of the molecular pathogenesis of CRC has advanced considerably and led to numerous revisions of the Vogelstein and Fearon model. For instance, the original model proposed that only tubular and tubulovillous adenomas had the potential to progress to invasive adenocarcinoma. It is now recognized that serrated polyps including sessile serrated adenomas/polyps (SSA/P) and traditional serrated adenomas (TSA) also have the potential for malignant transformation (Goldstein 2006; Jass 2004). These polyps are an alternative pathway to malignancy whereby a subset of hyperplastic polyps progress to serrated neoplasms (SSP or TSA) and a fraction of these serrated neoplasms progress to CRC. Premalignant serrated polyps more frequently arise in the proximal colon (Baker et al. 2004) and are associated with MSI and aberrant DNA methylation at CpG islands, whereas conventional tubular adenomas appear to arise most commonly via biallelic inactivation of the adenomatous polyposis coli (APC) tumor-suppressor gene and display chromosome instability (Noffsinger 2009). Furthermore, other molecular alterations, such as BRAF V600E mutations, are characteristically found more often in tumors arising via the serrated neoplasia pathway (Noffsinger 2009).
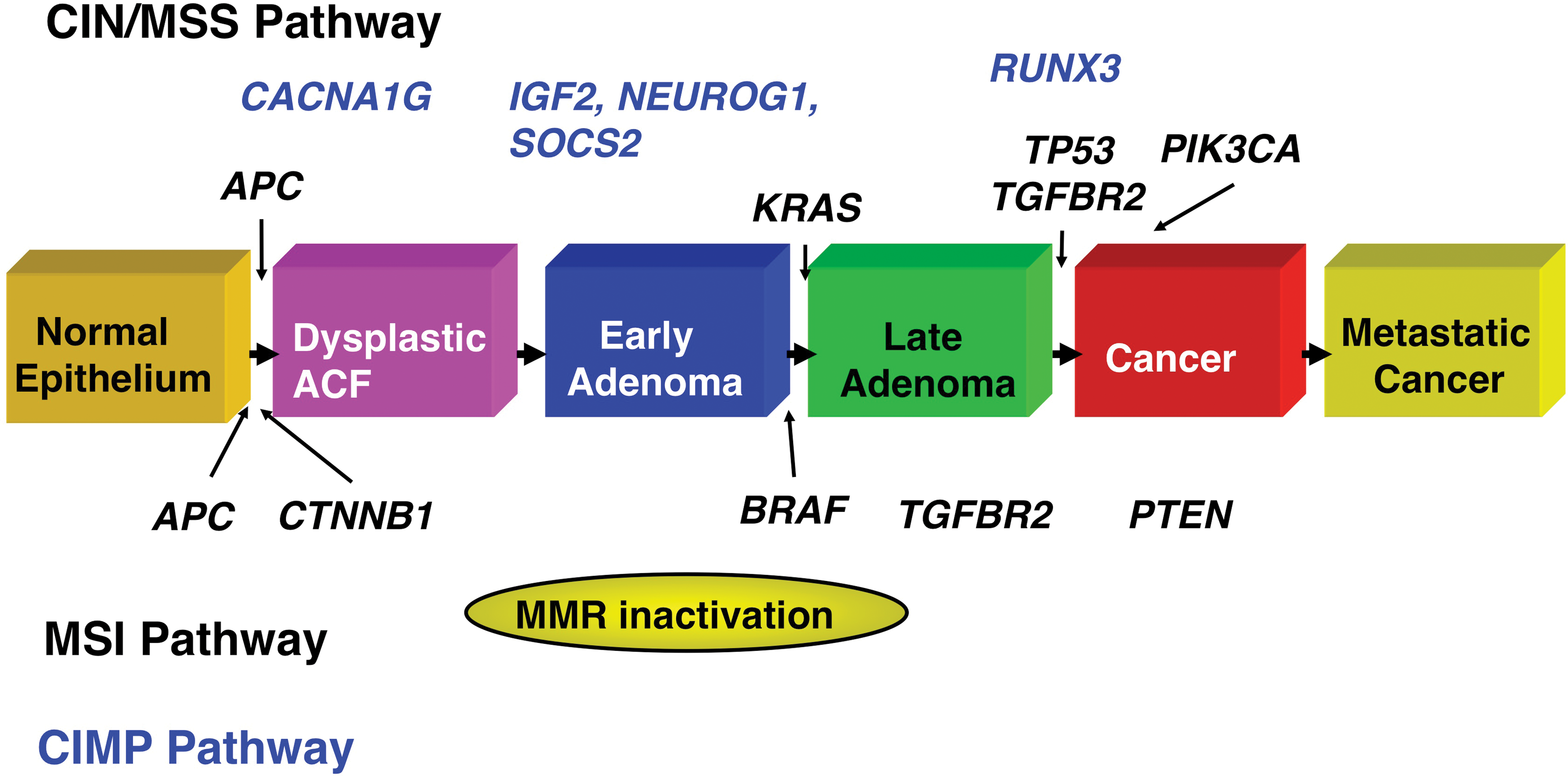
The polyp-to-carcinoma progression sequence. Colorectal carcinogenesis progresses by at least two well-recognized pathways. The chromosome instability (CIN) pathway is characterized by classic tubular adenoma histology and the early acquisition of adenomatous polyposis coli (APC) mutations that lead to deregulated wingless type MMTV integration site (WNT) signaling, frequent activating mutations of the KRAS oncogene at the early adenoma stage, loss of heterozygosity at chromosome 18q (18qLOH) in late adenomas, and TP53 mutations that facilitate the transition to frank malignancy. By contrast, tumors that harbor microsatellite instability (MSI) frequently acquire BRAF mutations and are not associated with 18qLOH or TP53 mutations. Sporadic MSI cancers appear to commonly arise via the serrated neoplasia pathway, in which SSAs are the most frequently observed precancerous lesions; SSAs appear to progress to CRCs via the CpG Island Methylator Phenotype (CIMP) pathway. (Modified from Pritchard and Grady 2011)
A Fundamental Feature of CRC: Genomic and Epigenomic Instability
Genomic and epigenomic instability distinguishes neoplastic from normal colonic epithelium and is a hallmark feature of colorectal carcinogenesis (Hanahan and Weinberg 2000; Hanahan and Weinberg 2011; Little, Vineis, and Li 2008). At least 4 kinds of genomic or epigenetic instability have been described in CRCs: (1) chromosomal instability (CIN), (2) MSI, (3) CpG island methylator phenotype (CIMP), and (4) global DNA hypomethylation. Overlap between these categories and imprecise use of these terms have led to confusion and confound interpretation of the literature (Walther et al. 2009). Thus, in this section, we will first define the different types of genomic and epigenetic instability in CRC and then will delineate in general terms how these mechanisms are clinically relevant.
CIN
The most common form of genomic instability is chromosome instability, which is found in as many as 85% of CRCs (Grady and Carethers 2008). Chromosome instability, which can be recognized by the presence of aneuploidy, is defined as the presence of numerical chromosome changes or multiple structural aberrations and can be assessed by DNA flow cytometry (Walther, Houlston, and Tomlinson 2008). Despite the frequent occurrence of CIN in CRC, the mechanisms that give rise to this form of genomic stability and the role of aneuploidy in tumor progression remain poorly understood. However, there is some evidence that CIN promotes cancer progression by increasing clonal diversity (Grady 2004; Hermsen et al. 2002; Maley et al. 2006). Importantly, from a clinical perspective, large meta-analyses have demonstrated that CIN is a marker of poor prognosis in CRCs, but technical aspects of assays that can determine the CIN status of CRCs have limited its use in clinical laboratories (Popat, Hubner, and Houlston 2005; Walther, Houlston, and Tomlinson 2008).
MSI
MSI tumors, which account for approximately 15% of CRCs, are generally regarded as being mutually exclusive of CIN tumors because they display a normal karyotype and exhibit unique gene mutations compared to CIN CRCs, although there does appear to be a subset of tumors that show both CIN and MSI (Walther et al. 2009). MSI CRC has been defined by the presence of at least 30% unstable loci in a panel of 5 to 10 loci consisting of mono- and dinucleotide tracts selected at a National Cancer Institute consensus conference (Boland et al. 1998). Currently, many clinical laboratories assess MSI using a panel of 5 mononucleotide markers (BAT-25, BAT-26, NR-21, NR-24, and MONO-27) that were selected for high sensitivity and specificity (Bacher et al. 2004). A subset of tumors with only 10 to 29% unstable loci has been designated as a form of microsatellite tumors designated “MSI-low.” Although there is evidence that MSI-low cancers have distinct features compared with MSI (also referred to as MSI-High, or MSI-H) and microsatellite stable (MSS) tumors, there is considerable controversy regarding whether MSI-low is a unique molecular subclass of CRC (Baron et al. 2003; Grady and Carethers 2008; Walther et al. 2009). CRC patients with MSI tumors have been shown to have a better prognosis compared to patients with CIN cancers (Popat, Hubner, and Houlston 2005; Walther, Houlston, and Tomlinson 2008), and probably respond differently to adjuvant chemotherapy compared to patients with MSS (also called CIN) CRCs (Fallik et al. 2003; Jo and Carethers 2006; Sargent et al. 2008).
In contrast to CIN, the mechanisms underlying MSI are relatively well understood and involve inactivation of genes in the DNA mismatch repair (MMR) family either by aberrant DNA methylation or by somatic mutation (Grady 2004). Furthermore, individuals with the hereditary cancer syndrome, Lynch syndrome (also called hereditary nonpolyposis colorectal cancer [HNPCC]), almost exclusively develop MSI CRCs because they have germ line mutations in 1 of the MMR genes, which include mutL homolog 1 (MLH1), mutS homolog 2 (MSH2), mutS homolog 6 (MSH6), and Postmeiotic segregration increased 2 (PMS2). In contrast, sporadic MSI CRCs most often have loss of MMR activity as the result of silencing of MLH1 by aberrant DNA methylation (Grady 2004; Kane et al. 1997). It is also now recognized that sporadic MSI tumors are associated with the serrated neoplasia pathway and frequently carry BRAF V600E mutations, while cancers resulting from germ line mutations in MMR genes (Lynch syndrome) do not have mutated BRAF (Domingo et al. 2004; Wang et al. 2003). Thus, the presence of a BRAF mutation in an MSI CRC effectively excludes the possibility that the tumor arose as the consequence of Lynch syndrome (Figure 2).
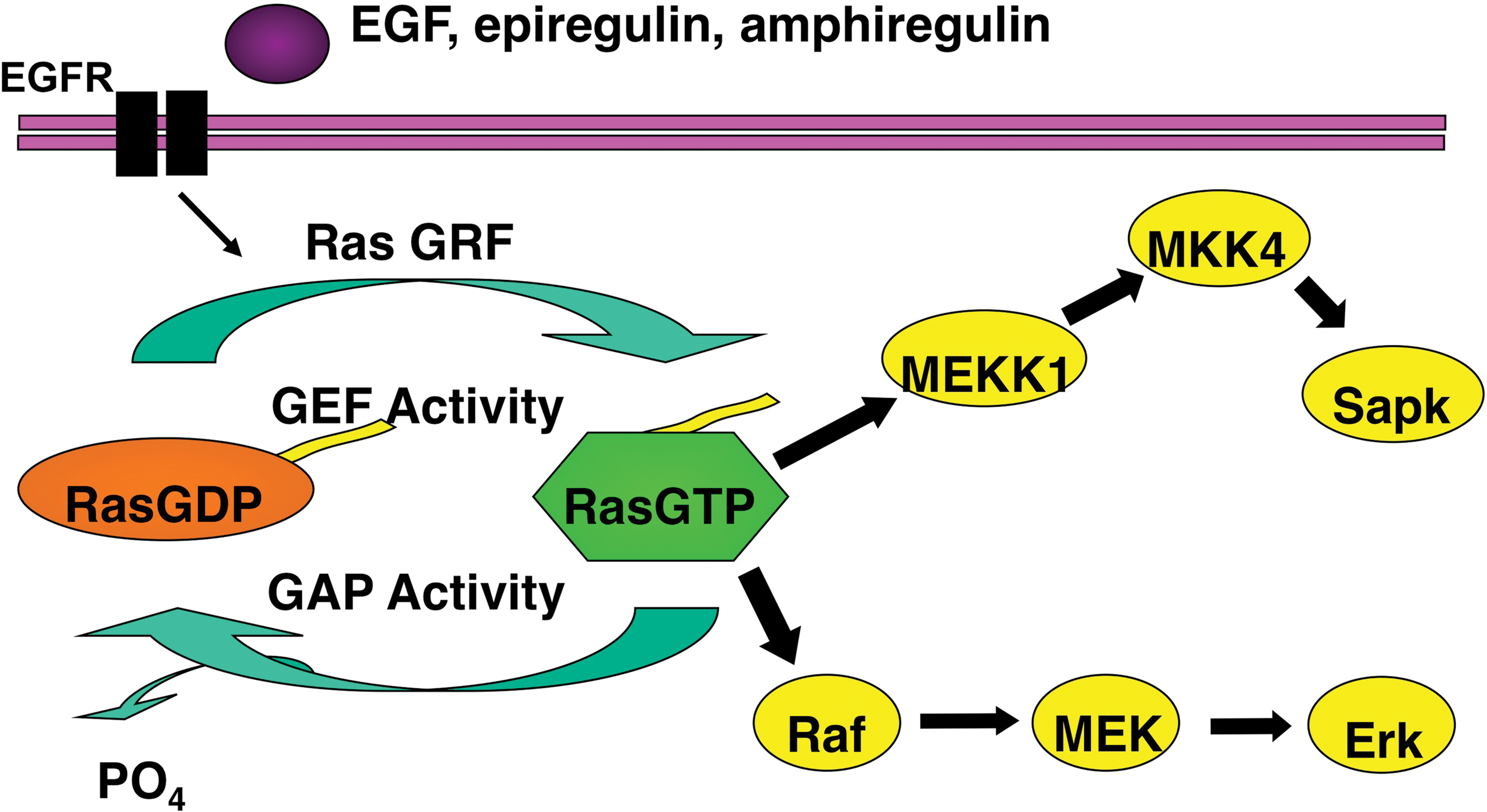
Mediators of epidermal growth factor receptor (EGFR) signaling. EGFR forms a homodimer after ligand activation, which results in phosphorylation/activation of the intracellular kinase domain and a cascade of downstream signaling including activation of the Ras/Raf/MAPK and phosphoinositol-3-kinase (PI3K) pathways that are associated with cell growth, differentiation, survival, and invasion. Monoclonal antibodies used to treat patients with metastatic colorectal cancer including cetuximab and panitumumab bind to the extracellular portion of EGFR and inhibit signaling in some patients. Activating mutations in KRAS occur in ∼40% of colorectal cancers and are thought to confer resistance to these drugs by bypassing the need for upstream EGFR signals. Activating mutations in BRAF—the direct downstream effector of KRAS—occur in ∼10% of colorectal cancers and also probably confer resistance to anti-EGFR monoclonal antibodies. Emerging evidence supports an additional role of oncogenic aberrations in the PI3K pathway in cetuximab and panitumumab resistance. Modified from Pritchard and Grady (2011).
CIMP
Epigenetic instability in CRC is manifested as both hypermethylation of gene promoters that contain CpG islands and global DNA hypomethylation. Aberrant DNA methylation is present in essentially all CRCs; however, there is a subset of CRCs (approximately 20%) that have an extremely high proportion of aberrantly methylated CpG loci. This class of CRCs has been characterized as having a CIMP. The mechanism/mechanisms that give rise to CIMP are still unknown, although the strong association between BRAF V600E mutations and CIMP CRC suggests a role for activated BRAF in the pathogenesis of CIMP CRCs and a link between sporadic MSI and CIMP (Barault et al. 2008; Weisenberger et al. 2006). However, in vitro studies of mutant BRAF in CRC cell lines have not demonstrated a direct cause and effect relationship between BRAF and CIMP (Hinoue et al. 2009). Furthermore, although CIMP tumors do appear to represent a distinct subset of CRC, the clinical utility of this designation is hindered by lack of a universally accepted criteria for determining CIMP. CIMP is usually defined as methylation of at least 3 loci from a selected panel of 5 gene-associated CpG islands. Because this panel is not always the same across studies, attempts are being made to facilitate standardization of CIMP markers for clinical use (Nosho et al. 2008; Weisenberger et al. 2006). Some authors have proposed 2 classes of CIMP, CIMP-low and CIMP-high, depending on the number of methylated marker loci detected (Barault et al. 2008). Another group suggested that CIMP CRCs be divided into 2 distinct classes (called CIMP1 and CIMP2) based on the results of unsupervised cluster analysis of a large panel of methylation markers (Shen et al. 2007). Finally, considerable overlap between CIMP and sporadic MSI tumors adds to the challenge of incorporating CIMP status into clinical trials and clinical decision making (Weisenberger et al. 2006). Retrospective studies suggest CIMP will ultimately be shown to be a predictive marker for CRC, but the data are not adequate at this time to recommend its clinical use (Iacopetta, Kawakami, and Watanabe 2008; Shen et al. 2007). Thus, the discovery and classification of CIMP tumors have advanced our understanding of the molecular pathology of CRC, but has not yet impacted clinical care.
In addition to aberrant gene methylation, a global decrease in methylation has been identified in many CRCs and is tightly associated with CIN CRCs (Matsuzaki et al. 2005; Rodriguez et al. 2006). Further research is needed to determine if the measurement of global DNA hypomethylation in CRC has any clinical utility.
Gene Mutations and Epigenetic Alterations: Impact on Signaling Pathways
Just as important as genomic and epigenomic instability for the pathogenesis of CRC is the accumulation of mutations in specific genes and the resulting deregulation of specific signaling pathways that control the hallmark behaviors of cancer: cell proliferation, differentiation, apoptosis, immortalization, angiogenesis, and invasion (Hanahan and Weinberg 2000; Hanahan and Weinberg, 2011). The best-studied pathways that are deregulated in CRC are the WNT-β-catenin signaling pathway, the transforming growth factor-beta (TGF-β) signaling pathway, the EGFR–mitogen-activated protein kinase (MAPK) pathway, and the phosphatidylinositol 3-kinase (PI3K) pathway (Siena et al. 2009; Walther et al. 2009). Selected deregulated pathways in CRC and targeted therapies in clinical use or in clinical trials are summarized in Table 2.
Pathways commonly deregulated in colorectal cancer and targeted drugs in clinical use (boldface) or under investigation.
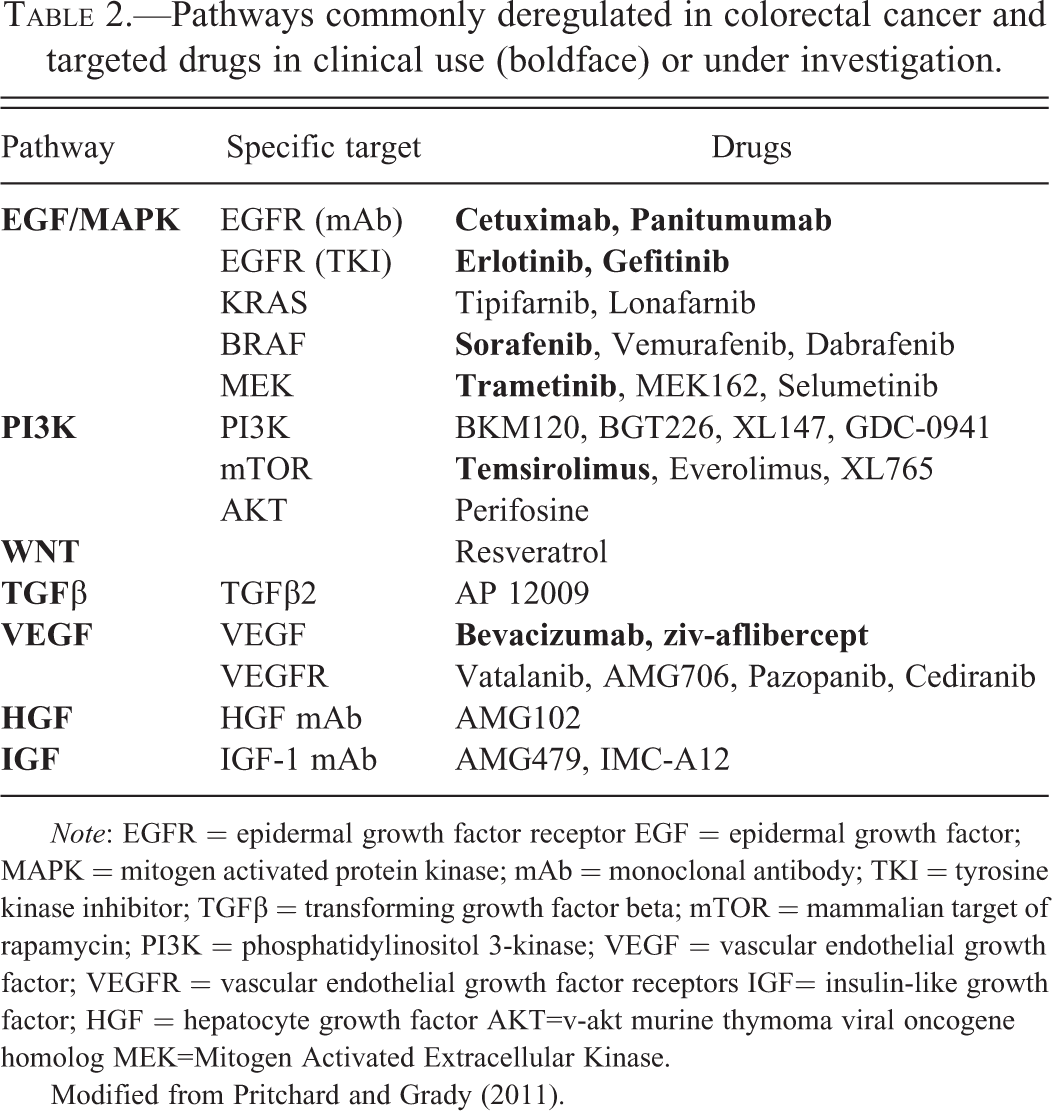

Note: EGFR = epidermal growth factor receptor EGF = epidermal growth factor; MAPK = mitogen activated protein kinase; mAb = monoclonal antibody; TKI = tyrosine kinase inhibitor; TGFβ = transforming growth factor beta; mTOR = mammalian target of rapamycin; PI3K = phosphatidylinositol 3-kinase; VEGF = vascular endothelial growth factor; VEGFR = vascular endothelial growth factor receptors IGF= insulin-like growth factor; HGF = hepatocyte growth factor AKT=v-akt murine thymoma viral oncogene homolog MEK=Mitogen Activated Extracellular Kinase.
Modified from Pritchard and Grady (2011).
Key tumor suppressor genes that do not necessarily mediate their effects through signal pathway deregulation, such as TP53, and recurrent cytogenetic aberrations, such as 18q loss of heterozygosity (LOH), are also well studied in CRC and affect the malignant transformation of colon epithelial cells through specific effects on the behavior of the cells (Figure 1). The use of these molecular alterations in the management of patients with CRC is discussed in more detail below.
APC/β-Catenin/WNT Pathway
Mutations in the APC gene occur in up to 70% of sporadic CRCs and are the cause of the familial adenomatous polyposis (FAP) cancer predisposition syndrome. APC mutations can be found at the earliest stages of neoplasia and are predominantly associated with the classic tubular adenoma pathway and CIN cancers (Figure 1; Chung 2000; Miyaki et al. 1999; Vogelstein et al. 1988). The APC protein negatively regulates WNT signaling by facilitating the targeting of the transcription factor β-catenin for ubiquitin-mediated proteasomal degradation. Disruption of the APC protein results in increased WNT signaling through stabilization of nuclear β-catenin, which increases the transcription of WNT target genes. Activating mutations in the gene for β-catenin (CTNNB1) that inhibit APC-mediated degradation are also observed in colorectal neoplasia, although they are found more frequently in adenomas (12.5%) than invasive cancer (1.4%), suggesting that CTNNB1-mutant neoplasms do not frequently progress to carcinoma (Samowitz et al. 1999). Despite the critical and nearly universal role of WNT pathway activation in colorectal carcinogenesis, there is currently no clinical use for APC or CTNNB1 mutations for treatment selection, prognosis, or early cancer detection. There has been intense effort to develop small molecule inhibitors of this pathway, but these efforts are still confined to the preclinical arena (Anastas and Moon 2013). If these agents eventually reach the clinic, the assessment of APC mutations, activated β-catenin (by the detection of nuclear localization of β-catenin by immunostaining), and epigenetic alterations that can affect WNT ligand availability (e.g., methylation of Secreted Frizzled-related Protein 2 (SFRP2), etc.) is likely to have a role in directing the selection of patients who will respond to these agents.
TGF-β Pathway
Deregulation of TGF-β signaling, which is generally considered a tumor-suppressor pathway in the colon, occurs in the majority of CRCs (Chittenden et al. 2008). Inactivating mutations have been observed in receptor genes (Transforming Growth Factor-β Receptor Type 2 (TGFBR2) and Transforming Growth Factor-β Receptor Type 1 (TGFBR1)), postreceptor signaling pathway genes (SMAD2, SMAD4), and TGF-β superfamily members (Activin Receptor type 2 (ACVR2); Deacu et al. 2004; Eppert et al. 1996; Grady et al. 1999; Grady and Markowitz 2008). Functionally significant mutations in TGFBR2 have been detected in as many as 30% of all CRC and are associated with the malignant transformation of late adenomas. TGFBR2 mutations are most common in MSI tumors, but also occur in approximately 15% of MSS tumors (Figure 1; Grady et al. 1999, 1998; Markowitz et al. 1995). SMAD4 is located on 18q in a region commonly deleted in CRCs and is associated with adenoma formation and adenoma–carcinoma progression in mouse models, supporting a role for SMAD4 as a tumor suppressor gene (Takaku et al. 1998). Furthermore, loss of SMAD4 expression as detected by immunostaining has been reported in >50% of CRCs and is associated with lymph node metastases (Tanaka et al. 2008). There is still not any definite clinical role for any genetic markers in the TGF-β signaling pathway; however, there is some evidence that SMAD4 expression levels may be associated with prognosis and response to 5-fluorouracil (5-FU), and there is ongoing investigation of 18qLOH as a predictive marker, which is discussed further in the next section (Alazzouzi et al. 2005; Boulay et al. 2002).
Loss of the long arm of chromosome 18 (18q LOH) is the most frequent cytogenetic alteration in CRC and is observed in up to 70% of CRCs, and it may influence CRC behavior by deregulating TGF-β signaling (Popat, Hubner, and Houlston 2005; Vogelstein et al. 1988). Two genes implicated as tumor suppressor genes that are affected by this cytogenetic alteration are deleted in colorectal carcinoma Delected in Colon Cancer (DCC) and SMAD4. Additional mediators of the TGF-β pathway, SMAD2 and SMAD7, are also in the 18qLOH region, suggesting that 18qLOH may promote tumorigenesis at least in part through deregulation of TGF-β signaling. It appears that deletion at 18q is associated with a worse prognosis; however, efforts to definitively link 18qLOH to prognosis are limited by a lack of consistent results across studies and by heterogeneous detection methods (Popat, Hubner, and Houlston 2005). Ongoing clinical trials (e.g., NCT00217737, also designated ECOG 5202) are assessing the utility of 18qLOH for directing the use of specific adjuvant therapies.
TP53
Mutations in the tumor-suppressor gene TP53 occur in about half of all CRCs and promote the malignant transformation of adenomas (Figure 1; Vogelstein et al. 1988). Like APC, TP53 is a key tumor suppressor that has been extensively studied in CRC but currently has no predictive or prognostic role in the clinical setting (Walther et al. 2009).
EGFR/RAS/RAF/RAF/MAPK Signaling
KRAS, a member of the RAS family of proto-oncogenes, is the most frequently mutated gene in all of human cancer and an important oncogene in CRC. The KRAS protein is a downstream effector of EGFR that signals through BRAF to activate the MAPK pathway and promote cell growth and survival (Figure 2). Mutations in KRAS codons 12 or 13 occur in approximately 40% of CRCs and lead to constitutive signaling by impairing the ability of guanosine triphosphate hydrolase (GTPase)-activating proteins to hydrolyze KRAS-bound guanosine triphosphate (GTP; Downward 2003). KRAS mutations occur after APC mutations in the adenoma-to-carcinoma progression sequence, but are still a relatively early event in tumorigenesis (Figure 1; Vogelstein et al. 1988). Acquired KRAS mutations are maintained throughout carcinogenesis, as evidenced by the nearly perfect concordance of KRAS-mutation status in primary and metastatic CRC (Artale et al. 2008; Zauber et al. 2003). This fact is critical to the utility of KRAS mutational analysis on archived primary tumor specimens in patients with metastatic disease and usually eliminates the need for additional biopsy tissue from a metastatic lesion.
The BRAF gene, mutated in ∼10% to 15% of CRCs, encodes a protein kinase that is the direct downstream effector of KRAS in the Ras/Raf/MAPK signaling pathway. The majority of BRAF mutations are a single base change resulting in the substitution of glutamic acid for valine at codon 600 (V600E; sometimes referred to as V599E; Siena et al. 2009). KRAS and BRAF mutations are mutually exclusive, supporting the hypothesis that an activating mutation in either gene is sufficient to promote tumorigenesis via increased MAPK signaling (Rajagopalan et al. 2002). As discussed above, BRAF mutations are much more frequent in MSI tumors (∼35%) compared to MSS tumors (∼5%) and are very tightly linked to CIMP cancers and the serrated neoplasia pathway (Lubomierski et al. 2005; Rajagopalan et al. 2002). Emerging evidence supports a role for BRAF as a genetic marker for prognosis and possibly for predicting response to therapy.
Alterations in EGFR ligands and the EGFR gene itself are also observed in a subset of CRCs. There is some data to support that upregulation of the EGFR ligands epigregulin and amphiregulin are associated with an anti-EGFR drug response (Jacobs et al. 2009; Lievre, Blons, and Laurent-Puig 2010).
PI3K/AKT Signaling Pathway
Mutations in PI3K pathway genes are observed in up to 40% of CRC and are nearly mutually exclusive of one another (Parsons et al. 2005). The most frequent mutations of the PI3K pathway occur in the p110α catalytic subunit phosphatidylinositol-4, 5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA), which are reported in up to 32% of CRCs and may promote the transition from adenoma to carcinoma (Figure 1; Samuels et al. 2004). Mutations are also observed in Phosphatase and Tensin homolog (PTEN), a tumor suppressor gene that negatively regulates PI3K signaling in as many as 30% of MSI tumors and 9% of CIN tumors (Danielsen et al. 2008). In addition, loss of PTEN expression (as assessed by immunostaining) occurs in 30 to 40% of CRCs (Yu et al. 2013). The PI3K pathway is modulated by EGFR signaling in part via KRAS activation, and there is a plausible role for both PIK3CA and PTEN mutations as predictive markers of anti-EGFR therapy (Figure 2; Razis et al. 2008; Sartore-Bianchi, Di Nicolantonio, et al. 2009). Recent evidence suggests that mutations in PIK3CA are predictive biomarkers of response to aspirin therapy (Liao et al. 2012). Mutations of genes in this pathway also have the potential to be used as predictive biomarkers for therapies that target the PI3K pathway, mammalian target of rapamycin (mTORC) pathway, as well as the MAPK pathway.
Risk Stratification and Early Detection
One use of molecular markers in the management of CRC is in risk stratification for identifying individuals at high risk for developing CRC and for the early detection of colon adenomas and early-stage CRCs. With regard to risk stratification, the most robust molecular markers to date are germ line mutations in genes that cause the hereditary colon cancer syndromes (e.g., APC mutations and FAP, bone morphogenetic protein receptor, type IA [BMPR1A], and Juvenile polyposis, etc.) and MSI tumor status, which is an indicator of the possibility of having Lynch syndrome. The use of MSI tumor testing in the diagnosis of Lynch syndrome will be discussed below in the context of MSI testing being a risk stratification marker because of its association with Lynch syndrome.
Lynch Syndrome (aka HNPCC)
Identifying individuals with Lynch syndrome (also known as HNPCC) dramatically alters their clinical management and can lead to effective CRC prevention programs for these individuals and their family members. However, currently the definitive molecular diagnosis of Lynch syndrome requires expensive germ line DNA mutation analysis of multiple DNA MMR genes (MLH1, MSH2, MSH6, epithelial cell adhesion molecule [EPCAM], and PMS2). To facilitate the most cost-effective strategies for identifying patients at high risk for Lynch syndrome who are candidates for genetic testing, the evaluation of molecular features of CRCs that have occurred in these individuals or other family members can be used to predict the likelihood of identifying a germ line mutation in one of the MMR genes. It is now common practice for molecular diagnostics labs to offer a stepwise series of molecular tests that are used to identify CRCs that arose secondary to Lynch syndrome. These tests are based on the molecular pathology of CRC (Figure 2; Baudhuin et al. 2005). A common approach is to initially test the tumors for loss of MMR gene products (MLH1, MSH2, MSH6, and PMS2) by immunohistochemistry (IHC) and for MSI by polymerase chain reaction (PCR) as the first-tier screening test (see e.g., http://www.mayomedicallaboratories.com/test-catalog/Clinical+and+Interpretive/17073), although there is support for the use of IHC alone as a first-line test (Shia 2008). Tumors that display MSI and loss of MLH1 protein expression by IHC are then subjected to reflex testing for BRAF V600E mutation status and MLH1 promoter hypermethylation to help distinguish sporadic MSI tumors (∼35% BRAF-mutant and 99% MLH1-methylated) from Lynch syndrome MSI tumors (BRAF-wild-type [WT], infrequent MLH1-methylation; Figure 2, Table 3; Bettstetter et al. 2007; Domingo et al. 2004; Zhang 2008). This strategy is most effective in excluding individuals who are unlikely to have a MMR gene mutation from undergoing germ line mutation testing. It is notable that in those tumors that have MSI and loss of MSH2, MSH6, or PMS2 the likelihood of having a germ line mutation is extremely high. Also of interest, it is now recognized that a strategy that relies on clinical criteria alone for the diagnosis of individuals at risk for Lynch syndrome underdiagnose this syndrome (Hampel et al. 2005). In light of the substantial effect of a missed diagnosis on an individual’s likelihood of developing cancer in the future, a strategy that employs universal MSI testing of all CRC is being advocated by some experts in this area (South et al. 2009, [NCT01850654]). It remains to be determined if this strategy is cost effective and if the benefits outweigh the risks.
Biomarkers used in the diagnosis of lynch syndrome (HNPCC).
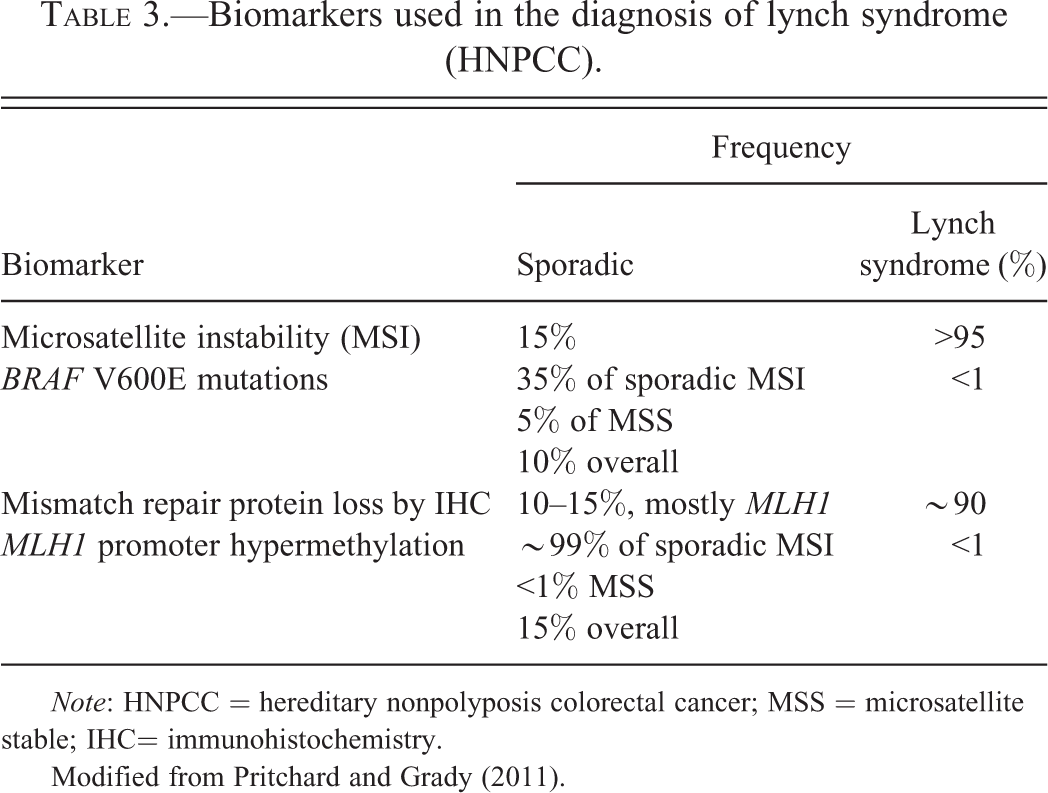
Note: HNPCC = hereditary nonpolyposis colorectal cancer; MSS = microsatellite stable; IHC= immunohistochemistry.
Modified from Pritchard and Grady (2011).
Molecular Markers and CRC Early Detection
Colonoscopy is the most accurate test currently used for CRC screening; however, it is expensive and associated with procedure-related complications and poor patient compliance (Ahlquist, Moertel, and McGill 1993). In contrast, another commonly used CRC screening test, fecal occult blood testing (FOBT), is inexpensive and simple to perform, but has a relatively low sensitivity and specificity (Smith et al. 2013). Advances in our understanding of the molecular pathology of CRC has led to the identification of promising early detection molecular markers for use in noninvasive CRC screening assays (Ahlquist et al. 2008; Osborn and Ahlquist 2005). Stool-based methylated VIMENTIN (mVim) is a clinically validated marker for early CRC detection that was previously available for clinical use in the United States (Table 1; Itzkowitz et al. 2007). The test relies on the fact that a majority of CRCs (53–84%) carry an aberrantly methylated vimentin (VIM) gene. A PCR-based assay that simultaneously measured mVim and DNA integrity reported a sensitivity of 83% and a specificity of 82%, with approximately equal sensitivity in stage I to III CRC patients (Itzkowitz et al. 2008). At this time, methods are under development to enhance the performance of stool- and plasma-based methylation assays for clinical purposes (Li et al. 2009; Zou et al. 2009). The use of molecular assays, such as the fecal-mVIM assay, in the clinical care of patients is an area that is likely to undergo rapid advances in the near future.
Prognostic Biomarkers
Genomic Instability and Prognosis: MSI versus CIN
Meta-analyses across a diverse range of patients have firmly established that MSI CRC have a better prognosis, which is independent of stage, and that CIN tumors have an unfavorable prognosis (Table 4; (Popat, Hubner, and Houlston 2005; Walther, Houlston, and Tomlinson 2008). The combined hazard ratio (HR) for MSI CRCs for overall survival (OS) was estimated to be 0.65 (0.59–0.71, 95% confidence interval [CI]) with only 1 of the 32 included studies reporting an HR >1.0 (Popat, Hubner, and Houlston 2005). Conversely, the overall HR associated with CIN CRC was determined to be 1.45 (1.27–1.45, 95% CI) based on 63 eligible studies and over 10,000 patients (Walther, Houlston, and Tomlinson 2008). Despite the clear association of MSI and CIN with prognosis, these markers have not yet been adopted into routine clinical decision making. It is most likely that MSI testing will be adopted into clinical practice before CIN testing because of the availability of a reliable assay for assessing MSI status.
Prognostic biomarkers in colorectal cancer.
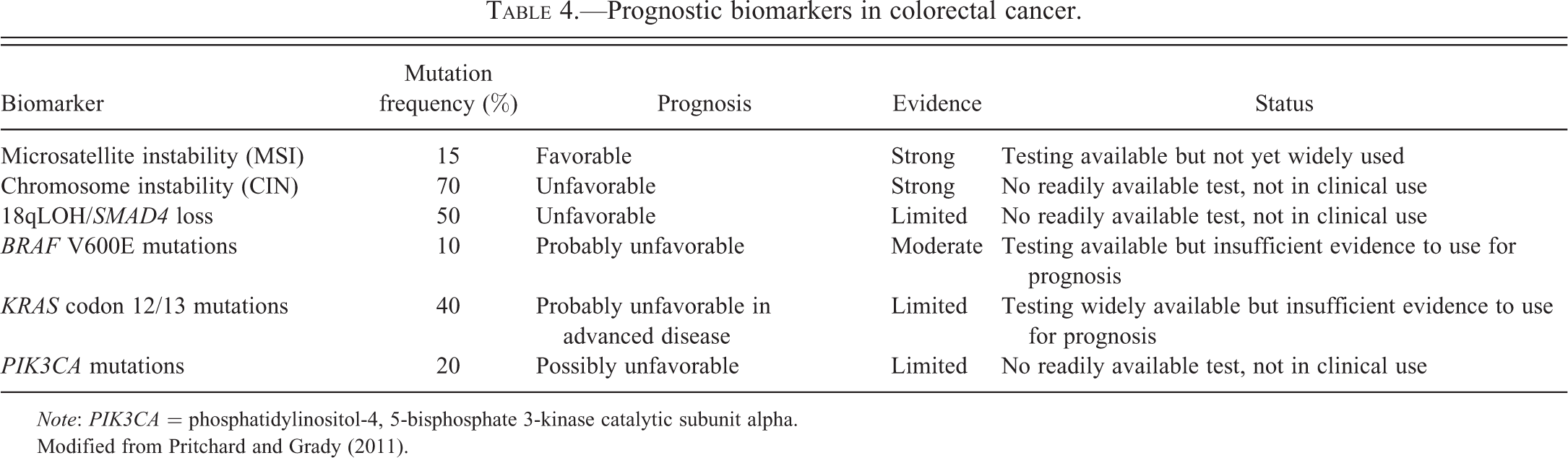
Note: PIK3CA = phosphatidylinositol-4, 5-bisphosphate 3-kinase catalytic subunit alpha.
Modified from Pritchard and Grady (2011).
18qLOH and Prognosis
CRC patients with 18qLOH appear to have a worse prognosis compared to patients with tumors that do not carry 18qLOH. A meta-analysis of 17 independent studies that was limited by evidence of publication bias found an overall HR of 2.00 (1.49–2.69 95% CI) for 18qLOH across all patients, and an HR of 1.69 (1.13–2.54 95% CI) in the adjuvant setting (Popat, Hubner, and Houlston 2005). Candidate genes in the 18q region, including DCC and SMAD4, have been studied individually for prognostic roles, with inconsistent results (Carethers et al. 1998). The independent prognostic contribution of 18q deletion in CRC has been called into question due to the tight association between 18qLOH and CIN, and the inverse association of 18qLOH and MSI (Walther et al. 2009). This assertion is supported by a recent study that found no difference in prognosis attributable to 18qLOH in a prospectively collected cohort of 555 non-MSI tumors stage I to IV (Ogino, Nosho, Kirkner, Kawasaki, et al. 2009). Thus, it is unclear at this time whether 18qLOH represents an independent prognostic marker or is merely a surrogate marker for CIN/MSS CRCs.
Mediators of EGFR Signaling and Prognosis
Several recent studies have assessed the prognostic significance of KRAS, BRAF, and PIK3CA mutations in CRC (Fuchs, Ogino, and Meyerhardt 2009; Ogino, Nosho, Kirkner, Shima, et al. 2009; Richman et al. 2009; Roth et al. 2010; Tol, Nagtegaal, and Punt 2009; Zlobec et al. 2010). In the vast majority of studies, mutant KRAS has not been independently associated with differences in relapse-free or OS in stage II or III CRC, but mutant BRAF has been shown to be prognostic for OS in this group of patients (Fuchs, Ogino, and Meyerhardt 2009; Roth et al. 2010; Yokota 2012). In contrast, mutant KRAS and BRAF have been reported as markers of poor prognosis in advanced CRC. In the largest study that has addressed the prognostic role of KRAS mutations in advanced CRC to date, patients with mutant KRAS cancers had a worse OS (HR = 1.40; 1.20–1.65 95%CI) but similar progression free survival (PFS) compared to patients with tumors bearing WT KRAS (Richman et al. 2009). The potential prognostic value of KRAS mutations is of particular interest in advanced CRCs because the KRAS mutational status of tumors is now being routinely collected in this setting in order to assess for eligibility for treatment with cetuximab or panitumumab. At this time, the use of KRAS mutation status for prognosis in CRC is still premature but appears to have significant potential to be adopted into clinical use in the near future. Similarly for mutant PIK3CA, most studies to date have not demonstrated a role for PIK3CA mutations in either exon 9 (helical domain) or exon 20 (kinase domain) as prognostic markers, although one observational cohort study did find that patients with concurrent exon 9 and 20 had a worse prognosis (Liao et al. 2012).
Predictive Biomarkers
Although the treatment of CRC still primarily relies on the surgical resection of the primary tumor to achieve a cure, considerable progress in the medical treatment of stage III and IV CRC has occurred over the last 15 years. The adjuvant therapy of stage III CRC has become more effective as the standard regimen has advanced from 5FU and leucovorin to 5FU and oxaliplatin or irinotecan (Meyerhardt and Mayer 2005). Furthermore, the treatment of stage IV CRC patients has expanded to include targeted therapies (cetuximab, panitumumab, bevacizumab; see Table 2) in addition to 5FU, oxaliplatin, and irinotecan. With the identification of multiple effective agents for the treatment of CRC has come a need for predictive markers for selecting optimal treatment regimens for patients. This is particularly applicable to CRC because of the heterogeneity in response among colon cancers and because of the toxicity and cost of the medical treatments. The potential of genetic and epigenetic alterations to be effective predictive molecular markers has received considerable attention lately and has led to the use of some of these markers in the routine care of patients with CRC (Table 5).
Colorectal cancer biomarkers as predictors for drug selection.
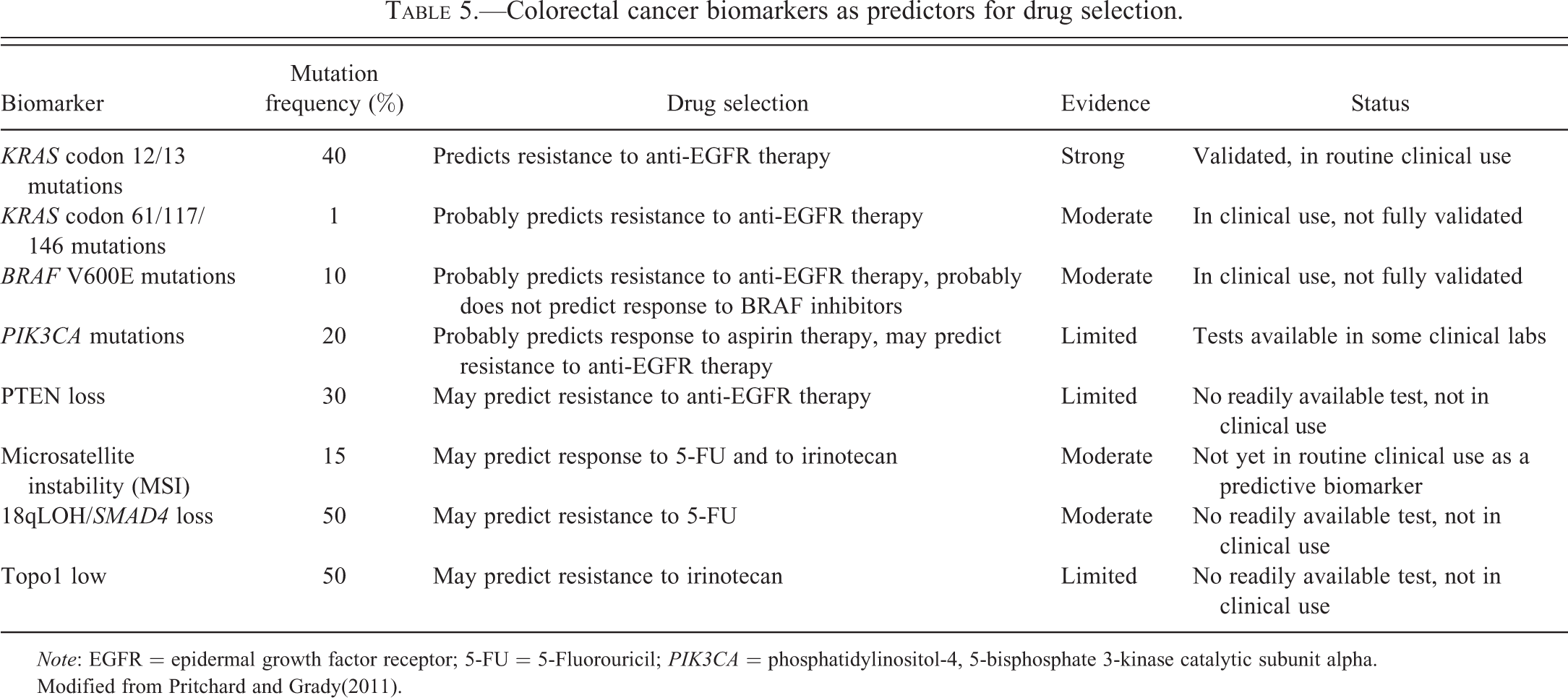
Note: EGFR = epidermal growth factor receptor; 5-FU = 5-Fluorouricil; PIK3CA = phosphatidylinositol-4, 5-bisphosphate 3-kinase catalytic subunit alpha.
Modified from Pritchard and Grady(2011).
The advent of cancer therapeutics that target specific molecules and pathways highlights the potential for underlying genetic and epigenetic lesions in CRC to guide personalized treatment decisions. A clear demonstration of the potential of mutant genes to direct therapy is that of mutant KRAS and treatment with cetuximab. Only ∼15% of patients with metastatic CRC respond to mAb therapies targeting the EGFR, which prompted intense research into resistance mechanisms that could be secondary to alterations in the EGFR gene and/or mutations in downstream effectors. These studies have produced one well-validated and exceedingly robust predictive marker (mutant KRAS) and several more promising biomarkers that require further validation (mutant BRAF, PIK3CA, PTEN; Bardelli et al. 2003). Research efforts are also focused on identifying molecular features of CRC that predict response to adjuvant chemotherapy with cytoxic agents: 5-FU, irinotecan, and oxaliplatin (Walther et al. 2009). In this section, we will discuss genetic features of CRC that have been evaluated for a role in guiding treatment selection. We have focused primarily on acquired tumor mutations as predictive markers, but it is important to note that inherited (germ line) polymorphisms also influence the effects of chemotherapy on cancers and the risk for drug toxicity, particularly in the case of 5-FU and irinotecan (reviewed in Walther et al. 2009).
Predictors of Response to Anti-EGFR mAb Therapies
EGFR-targeted mAbs cetuximab (Erbitux®) and the fully humanized mAb panitumumab (Vectibix®) have proven to be effective in patients with metastatic CRC both as single agents and in combination with traditional chemotherapy (Ciardiello and Tortora 2008; Cunningham et al. 2004; Saltz et al. 2004). However, while these therapies improve both PFS and OS, they are effective in only a minority of metastatic CRC patients (Bardelli et al. 2003). These drugs are generally well tolerated but are still associated with treatment-related morbidity, including skin rash, diarrhea, and nausea, and are also expensive. To better target anti-EGFR mAb therapy to patients most likely to benefit, KRAS mutation status and additional molecular markers of cetuximab and panitumumab resistance have been extensively evaluated (Siena et al. 2009).
KRAS
Results of 4 large phase III randomized have established unequivocally that metastatic CRC patients with KRAS mutations in codon 12 or 13 do not benefit from cetuximab or panitumumab therapy (Amado et al. 2008; Bokemeyer et al. 2009; Karapetis et al. 2008; Van Cutsem et al. 2007). Prior to the publication of these pivotal trials, the link between KRAS mutation status and anti-EGFR mAb response was already firmly supported by several smaller studies (Benvenuti et al. 2007; Di Fiore et al. 2007; Lievre et al. 2006), but the data were not sufficient to warrant routine clinical testing. The recently published randomized trials have established the use of KRAS mutational analysis as a predictive marker for anti-EGFR mAb resistance in patients with metastatic CRC in most of the relevant clinical settings. These settings include the use of cetuximab or pantumimab in combination with conventional cytotoxic chemotherapy (e.g., 5-FU, FOLFOX, FOLFIRI) as first-line treatment of metastatic disease (Bokemeyer et al. 2009; Douillard, Siena, and Cassidy 2009; Van Cutsem et al. 2009) and as monotherapy in relapsed/refractory patients (Amado et al. 2008; Karapetis et al. 2008; Van Cutsem et al. 2007).
A second relevant question related to anti-EGFR mAb therapy is whether mutant KRAS predicts an adverse outcome in the setting of these treatments. The reported HRs were almost exactly 1.0 in a total of 348 KRAS-mutant chemotherapy-resistant or refractory cancers treated with either panitumumab (Amado et al. 2008) or cetuximab (Karapetis et al. 2008) as monotherapy, confirming lack of benefit, but also suggesting no harm from anti-EGFR mAb treatment related to PFS or OS in this population. In contrast, the reported HRs were usually greater than 1.0 in studies of cetuximab or panitumumab as first-line treatment in combination with FOLFOX4 (FU, leucovorin, and oxaliplatin) or FOLFIRI (FU, leucovorin, and irinotecan) chemotherapy (Bardelli et al. 2003). The results of the OPUS trial (oxaliplatin and cetuximab in first-line treatment of metastatic CRC) in particular suggest it may be harmful to add anti-EGFR mAb treatments to 5FU, leukovorin, and oxaliplatin in patients with KRAS-mutant metastatic CRC (Bokemeyer et al. 2009).
Based on the evidence from large trials, European and U.S. practice guidelines either recommend or require KRAS mutational analysis on CRC tumor tissue prior to the initiation of cetuximab or panitumumab treatment (European Medicines Agency 2008; NCCN 2010; Allegra et al. 2009). The European health authority confines use of panitumumab monotherapy, and cetuximab as mono- or combination therapy, to metastatic CRC patients who are found to carry nonmutated (WT) KRAS in the primary tumors (European Medicines Agency 2008). The American Society for Clinical Oncology recently published a provisional opinion stating that “All patients with metastatic colorectal carcinoma who are candidates for anti-EGFR antibody therapy should have their tumors tested for KRAS [codon 12 and 13] mutations…and [KRAS-mutant] patients should not receive anti-EGFR antibody therapy” (Allegra et al. 2009). Similarly, the National Comprehensive Cancer Network (NCCN) guidelines require evidence of WT KRAS prior to cetuximab or panitumumab therapy in all metastatic CRC settings (NCCN 2010).
Despite the nearly perfect negative predictive value of mutant KRAS, it is still only a minority (∼30%) of KRAS codon 12/13 WT patients who respond to anti-EGFR mAb therapy (Bardelli et al. 2003). This has led to research into additional biomarkers that might predict lack of benefit in those individuals with tumors that have WT KRAS. There is evidence that rare KRAS mutations in codons 61 or 146 (∼2% of CRC) behave similarly to codon 12/13 mutations (Loupakis et al. 2009), but incorporating these mutations into routine clinical practice will require analysis of a larger group of patients. Other promising markers of anti-EGFR mAb resistance are BRAF V600E mutations, PIK3CA mutations, and loss of PTEN protein expression (Siena et al. 2009).
BRAF
The biological rationale for BRAF V600E mutations as an additional biomarker of anti-EGFR mAb resistance is strong: (1) BRAF is the immediate downstream effector of KRAS in the Ras/Raf/MAPK signaling pathway (Figure 2) and (2) BRAF V600E activating mutations are 100% mutually exclusive of KRAS mutations in CRC, implying that the activation of either protein is sufficient for colon tumorigenesis. Prior studies support the use of BRAF V600E as a negative predictor of response to anti-EGFR mAb therapy, leading to the evolving use of BRAF mutation testing in KRAS-WT patients prior to treatment as a means to further stratify patients into responders and nonresponders. A retrospective analysis showed that 0/11 tumors with mutant BRAF responded to cetuximab or panitumumab compared to 22/68 (32%) of BRAF-WT/KRAS-WT patients (Di Nicolantonio et al. 2008). Similar results were observed for patients treated with cetuximab plus irinotecan. None of the patients with tumors with mutant BRAF (N = 13) responded compared to 24/74 (32%) patients with tumors with BRAF-WT/KRAS-WT (Loupakis et al. 2009), although not all studies have found as robust a relationship between BRAF V600E mutation status and anti-EGFR antibody response (Laurent-Puig et al. 2009; Tol, Nagtegaal, and Punt 2009). BRAF mutations also appear to be associated with worse prognosis independent of treatment, which can confound the assessment of its role as a predictive marker for response to EGFR-directed therapies (Laurent-Puig et al. 2009; Tol, Nagtegaal, and Punt 2009). Despite the currently limited data, and lack of complete consensus, it is likely that BRAF mutation status has a role in anti-EGFR mAb treatment decisions and soon will be adopted into the planning for treatment with cetuximab and panitumumab.
PI3K Pathway Members
Molecular lesions in the PI3K pathway, which in CRC are primarily mutations in PIK3CA and loss of PTEN protein expression (Yu and Grady 2012), have been proposed as additional anti-EGFR mAb resistance markers because EGFR mediates its effects through the PI3K pathway as well as the MAPK pathway (Bardelli et al. 2003). However, the relationship of oncogenic alterations in PI3K signaling with cetuximab or panitumumab response is much less clear than that of KRAS and BRAF mutations. In several small studies published to date, PIK3CA mutations or PTEN loss have been associated with lack of response to cetuximab (Jhawer et al. 2008; Mao et al. 2012; Prenen et al. 2009; Sartore-Bianchi, Di Nicolantonio, et al. 2009; Sartore-Bianchi, Martini, et al. 2009). Both PIK3CA mutations and PTEN loss may coexist with KRAS or BRAF mutations, which weakens the biological rationale of the activation of this pathway as an absolute predictor of anti-EGFR mAb therapeutic response. Nonetheless, the balance of evidence points toward a probable predictive role of molecular events that activate the PI3K pathway for being negative predictive markers for EGFR mAb-based therapy. In fact, there is modest data demonstrating that when PIK3CA mutations and PTEN loss of expression are combined with KRAS and BRAF mutational analysis, up to 70% of patients unlikely to respond to cetuximab or panitumumab may be identified (Bardelli and Siena 2010; Sartore-Bianchi, Di Nicolantonio, et al. 2009). This observation has led to the idea that colon cancer may be able to be classified like breast cancers (e.g., triple negative breast cancers), and these cancers have been termed quadruple negative for patients who do not have alterations in any of these 4 biomarkers (Bardelli and Siena 2010; Sartore-Bianchi, Di Nicolantonio, et al. 2009). However, at this time, further studies are needed to determine whether mutant PIK3CA or PTEN loss should be incorporated into clinical practice (Bittoni et al. 2012).
EGFR Mutations and Amplification
The most obvious candidate biomarker for resistance to antibodies that target EGFR is the EGFR gene itself. Early studies that focused on EGFR overexpression failed to show a consistent relationship with treatment response because of lack of standardization of the assays used to assess EGFR expression, which were based on immunostaining, fluorescent in situ hybridization (FISH), or quantitative reverse transcriptase (RT)-PCR, and because of interobserver variability inherent in some of these techniques (Shia et al. 2005). EGFR gene amplification appears to be a more promising predictive biomarker, but it has also been fraught with technical challenges that have limited its use in the clinic (e.g., false negative results secondary to dilution of tumor DNA with WT DNA in PCR-based assays, and lack of consistent tissue processing and scoring systems in FISH assays; Siena et al. 2009). Activating mutations in the EGFR catalytic domain are seen frequently in lung cancer and are associated with sensitivity to anti-EGFR tyrosine-kinase inhibitors, but these mutations are quite rare in CRC (Siena et al. 2009). Thus, EGFR is unlikely to be a clinically useful predictive marker for anti-EGFR mAb therapy. Furthermore, although preliminary studies have shown that the EGFR ligands amphiregulin and epiregulin are overexpressed in CRC and may predict response to cetuximab, lack of standardization of the assays, and studies that reproducibly demonstrate the same effect have prevented amphiregulin and epiregulin expression levels from being used as clinical biomarkers for directing therapy with EGFR mAbs (Jacobs et al. 2009).
Predictive Molecular Markers for Cytotoxic Chemotherapy: 5-FU, Irinotecan, and Oxaliplatin
Currently, the tumor biomarkers that demonstrate the greatest promise for guiding adjuvant chemotherapy with conventional cytotoxic drugs in CRC patients are MSI and 18qLOH.
MSI
5-FU-based regimens have been shown to be ineffective or even detrimental to patients with MSI tumors (French et al. 2008; Ribic et al. 2003). Evidence that a functioning MMR system is required for the cytotoxic effect of FU provides a plausible biological rationale for 5-FU resistance in MSI tumors (Grady and Carethers 2008; Jo and Carethers 2006). However, the finding of 5-FU resistance in MSI CRC is not uniform and may vary with tumor stage and may differ between sporadic MSI cancers versus Lynch syndrome cancers (Kim et al. 2007; Liang et al. 2002; Sinicrope and Shi 2012). An ongoing phase III trial enrolling stage II and III CRCs (NCT00217737) will prospectively assess the role of MSI in predicting response to adjuvant chemotherapy (Walther et al. 2009). At this time, the mixed outcomes from published studies make it unclear if MSI is a clinically useful predictive marker for 5-FU responsiveness.
MSI tumors appear to be more responsive to irinotecan-based adjuvant chemotherapy (Bertagnolli et al. 2009; Fallik et al. 2003). Recently published results from a large randomized trial of stage III CRC demonstrated improved outcomes (disease-free survival [DFS]) in MSI patients treated with an irinotecan-containing regimen that included 5-FU compared to 5-FU/leukovorin alone (Bertagnolli et al. 2009). In light of the prior results of the cancer and leukemia group B (CALGB) 89803 study showing no benefit of adding irinotecan to 5FU as adjuvant therapy in unselected stage III CRC patients, the finding that MSI is a predictive biomarker for irinotecan suggests MSI could be useful for adjusting adjuvant therapy for CRC patients (Saltz et al. 2007). However, it should be noted that these results were not confirmed in a similar study (Pan-European trials in adjuvant colon cancer [PETACC]; Tejpar et al. 2010). Thus, at this time, neither the European Group on Tumor Markers nor the American Society of Clinical Oncology recommend using MSI to direct adjuvant chemotherapy for stage II or stage III CRC patients.
An important issue to consider with regard to MSI is that the majority of CRCs that have MSI are sporadic CRCs that have inactivated the MLH1 gene through aberrant promoter methylation. The majority of these sporadic MSI tumors also can be classified as CIMP cancers as well. It is not known whether the associations seen between 5FU and irinotecan effects in sporadic MSI tumors also apply to MSI tumors that arise in the setting of Lynch syndrome.
18q LOH
Loss of 18q has been associated with an adverse response to 5-FU-based adjuvant chemotherapy (Boulay et al. 2002; Watanabe et al. 2001). There is some evidence that this effect is due to loss of the SMAD4 gene located in the 18q21-deleted region, although this remains to be determined with more definitive studies (Alhopuro et al. 2005; Boulay et al. 2002). A number of ongoing clinical trials are assessing the predictive value of 18qLOH and MSI status for the treatment of colon cancer. These include an Eastern Cooperative Oncology Group (ECOG) trial of stage II CRC patients being treated with 5-FU, oxaliplatin, and bevacizumab (NCT00217737), a trial in patients being treated with olaparib for metastatic disease (NCT00912743), as well as a retrospective analysis assessing MSI and 18qLOH in patients with CRC (stage II or III) treated with 5-FU or 5-FU and irinotecan (CLB-9581 or CLB-89803).
Topoisomerase 1 Expression
In a large randomized trial that compared 5-FU alone to 5-FU with irinotecan and 5-FU with oxaliplatin in advanced CRC, high expression of topoisomerase 1 (Topo1) measured by IHC correlated with responsiveness to irinotecan (Braun et al. 2008). Conversely, cancers with low Topo1 expression (602/1,269; 47%) did not appear to benefit from the addition of irinotecan (HR 0.98; 95% CI, 0.78–1.22). Irinotecan is a Topo1 inhibitor, thus the level of Topo1 expression has a clear biological rationale as a biomarker for predicting irinotecan response. Replication of these initial results in multiple independent studies is required before Topo1 should be considered for use as a predictive marker.
Polymorphisms: Biomarkers for CRC
In addition to somatic molecular alterations in CRCs being used as biomarkers for informing the prognosis and care of patients with this common form of cancer, germ line polymorphisms that alter the pharmacokinetics and pharmacodynamics of adjuvant chemotherapy have potential to be biomarkers for guiding treatment selection. For example, alterations in thymidylate synthetase and dehydropyrimidine dehydrogenase have been extensively studied in relation to 5-FU response and appear to have potential to identify patients that are at increased risk of adverse effects from 5-FU. However, very few of these polymorphisms have been adequately validated for clinical use, so the majority are not ready to be used to direct patient care (Ezzeldin and Diasio 2008; Schwab et al. 2008). One exception to this generalization is a homozygous polymorphism that reduces the activity of UDP-glucuronosyltransferase (UGT1A1, an enzyme that detoxifies irinotecan), which is associated with a dose-related increased incidence of irinotecan toxicity (Hoskins et al. 2007; Palomaki et al. 2009). This has led to a commercial UGT1A1 genotyping test that was approved by the Food and Drug Administration in 2005 to help guide irinotecan dosing.
Conclusion and Future Directions
Over 4 decades of investigation into the molecular mechanisms of CRC behavior have now culminated in biomarkers that are sufficiently validated for routine clinical use. KRAS-mutation analysis to guide anti-EGFR treatment stands as one of the first successes in the era of precision medicine. MSI and BRAF mutations already have a clear role in triaging molecular genetic testing in Lynch syndrome, and these markers are poised to take on a much greater role in prognostication and prediction of therapeutic responses for sporadic CRCs.
The use of assays for mutant KRAS, mutant BRAF, and MSI demonstrates how the molecular testing of CRC tissue can reduce medical costs and improve patient outcomes by targeting therapies to the appropriate patient populations. Thus, it is anticipated that the use of molecular genetic markers in clinical decision making is likely to expand as more markers are identified and validated. For example, studies are in progress for assessing the efficacy of the multikinase/BRAF-inhibitor sorafinib, and specific inhibitors of the PI3K signaling pathway in the treatment of CRC (Siena et al. 2009). There is evidence that sorafinib may only be effective in the setting of concurrent EGFR inhibitors, which has prompted an ongoing phase II National Cancer Institute sponsored clinical trial of sorafinib plus cetuximab in metastatic CRC patients (NCT00343772; Di Nicolantonio et al. 2008; Wagle et al. 2011). If these initial findings are validated, the indications for mutational analysis of BRAF and KRAS would expand. Furthermore, CRC patients with tumors carrying mutant BRAF might also benefit from newer selective mutant BRAF inhibitors such as plexin (PLX)-4032 combined with anti-EGFR mAb therapy. PIK3CA mutations or PTEN loss are likely to become clinically relevant for the treatment of CRC patients as specific PI3K pathway inhibitors (such as XL147, BGT226, GDC0941, XL765, and NVP-BEZ325) are assessed in clinical trials (Yu and Grady 2012; Yuan and Cantley 2008). The expanding repertoire of drugs designed to inhibit specific oncogenes and oncogenic signaling pathways again highlights that molecular mechanisms of CRC will increasingly play a role in the clinical care of patients with CRC. The use of molecular markers for risk stratification and early detection of CRC is also showing promise and will be part of the era of molecular medicine that is rapidly emerging.
Footnotes
Abbreviations
Acknowledgment
We wish to thank Jonathan Tait for helpful suggestions.
Authors’ Note
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Work on this publication was supported by the National Cancer Institute of the National Institutes of Health under award number P30CA15704, UO1CA152756, 5U01HG006507, U54CA143862, and P01CA077852, (WMG). Support for these studies was also provided by a Burroughs Wellcome Fund Translational Research Award for Clinician Scientist (WMG).
