Abstract
ADAM33 represents an important gene of susceptibility for lung function impairment. This work aimed to evaluate the association between genetic polymorphism of ADAM33 at four single nucleotide polymorphisms (T1, T2, S1, and Q1) and arginase activity with respiratory functions impairment in wood workers. The study was done to compare ventilatory functions and arginase activity of 82 wood workers and 81 controls. Genotyping was determined by using the polymerase chain restriction fragment length polymorphism method. Forced expiratory volume in the first second (FEV1), forced vital capacity (FVC), and peak expiratory flow rate (PEF) of the workers were significantly reduced compared with the controls. T1 single nucleotide polymorphism (SNP) was associated with obvious decline in the FEV1, FVC, and PEF in wood workers, while T2 SNP was associated with decline in FEV1 and PEF. A significant increase in arginase activity was found in T2 and S1 SNPs of the exposed workers. Increase in duration of exposure was correlated with the decline in ventilatory functions. This inverse correlation was significant for pulmonary function indices in AA and GG genotypes of T1 and T2, respectively. Moreover, significance was detected for FVC and FEV1 in AA and GA genotypes of S1 and Q1. A positive correlation between arginase activity and duration of exposure was found to be significant in GG genotype of S1 SNP. An association between ADAM33 gene polymorphism and impaired lung functions was detected in wood dust-exposed workers. Arginase activity may play an associated important role in increasing this impairment in wood workers.
Introduction
Wood processing in furniture industry generates wood dust. 1 When wood dust becomes airborne from processes such as sanding, cutting, drilling, chipping, sawing, or turning to shape, it causes potential health problem. 2 Occupational exposure to wood dust has long been associated with cross-shift decline in forced expiratory volume in the first second (FEV1) 3 and respiratory symptoms, including asthma, chronic bronchitis, bronchial hyper reactivity, 4 and overall decrease in lung functions. 5,6
However, there is growing evidence that interactions between susceptibility genes and environmental factors are important determinants for respiratory health problems, the causes of respiratory function impairment are complex and not totally understood. 7,8
Garlisi and his colleagues 9 demonstrated that ADAM33 is an active proteinase that is able to cleave α2-macroglobulin, an important member of pulmonary defense system. Studies have shown that ADAM33 is expressed in asthmatic subepithelial fibroblasts and smooth muscle but not in respiratory epithelium. 8,10 These results suggested that ADAM33 is involved in the pathogenesis of airway obstruction affecting tissue remodeling, a physiological process intricately related to airway inflammation. 11 Additional studies have demonstrated that single nucleotide polymorphisms (SNPs) within ADAM33 were associated with accelerated decline of lung function in the general population and in asthmatic patients. 12
Arginase exists as two distinct isoenzymes, arginase I and II, which are encoded by different genes. In the airways, both arginase I and II are constitutively expressed in bronchial epithelial cells, endothelial cells, (myo)fibroblasts, and alveolar macrophages,
13
while arginase II is also expressed in parenchymal epithelial cells.
14
A number of studies have reported that arginase expression in airway smooth muscle was below detection limit,
14
while other studies have indicated that either isoform may be (conditionally) expressed in these cells.
15
Arginase may play an important role in the pathogenesis of various pulmonary disorders.
16
Increased expression and/or activity of arginase has been observed in airways and lung tissue obtained from various animal models of allergic asthma
16
and from asthmatic patients.
17
Increased consumption of
The aim of this work was evaluation of arginase activity and ventilatory functions among workers exposed to wood dust and their association with SNPs (T1, T2, S1, and Q1) of ADAM33 gene in those workers.
Subjects and methods
This study was carried out in a factory for furniture manufacture in Cairo governorate. Eighty-two workers exposed to wood dust (workers) were enrolled in the study, and 81 apparently healthy subjects not occupationally exposed to wood dust (controls) were also recruited. Complete history was obtained from all the included persons in the form of a questionnaire including personal, medical, and detailed environmental and occupational histories. Based on this questionnaire, persons with history of recent or previous exposure to wood dust were excluded from the control group. Wood workers employed for less than 5 years were excluded from workers’ group. Smokers were also excluded from the two groups, as there was an association between ventilatory abnormalities and the cytogenetic changes produced by smoking. 21 A written consent was signed by all the participants before beginning the study, which was approved by the National Research Centre Ethical Committee, and withdrawal from the study was allowed any time.
Ventilatory function tests
Spirometric measurements were performed in sitting position using a portable spirometer, according to the criteria of the American Thoracic Society. 22 All ventilatory function parameters in the form of FEV1, forced vital capacity (FVC), and peak expiratory flow rate (PEF) were expressed as percent of the predicted value for each person after adjustment for age, gender, race, and height.
Laboratory studies
Venous blood samples were withdrawn from all subjects under complete sterile conditions, part of the blood was stored in ethylenediaminetetraacetic acid tubes at −70°C until used in DNA extraction, the rest of the blood samples were collected in plain tubes, centrifuged, and the separated sera were frozen at −20°C. Samples were thawed and mixed thoroughly just prior to the assay to avoid erroneous results of repeated freeze/thaw cycles, hemolyzed, and lipemic samples were avoided.
– Arginase assay was based on colorimetric determination of urea by condensation with diacetylmonoxime in an acid medium in the presence of ferric chloride (oxidant) and carbazide (accelerator), according to Mellerup.
23
– ADAM33 gene polymorphism analyses (T1, T2, S1, and Q-1) were done through isolation of genomic DNA from peripheral blood leukocytes of the studied groups using Qiagen Kit, Germany. Genotypes were assayed with polymerase chain reaction (PCR) restriction fragment length polymorphism method. The PCR primers were shown in Table 1. The PCR was performed with a 25-μL reaction mixture containing 100 ng of genomic DNA, 0.5 μmol/L of each primer, 200 μmol/L of each deoxynucleotide triphosphate (dNTP), 2.5 U of Taq DNA polymerase (Omega, Doraville, Georgia, USA), 10× PCR buffer supplied by Invitrogen Corporation (10 mmol/L Tris-hydrochloric acid, pH 8.8, 50 mmol/L potassium chloride), and 2.0 mmol/L magnesium chloride. The PCR profile consisted of an initial melting step of 5 min at 94°C, followed by 35 cycles of 1 min at 94°C, 1 min at 58°C for T2&Q-1 A/G, 1 min at 60°C for T1 A/G, 1 min 59°C for S1 G/A, followed by1 min at 72°C and a final elongation at 72°C for 5 min.
Primer sequences and reaction conditions of ADAM33 genotyping.

Statistical analysis
The collected data and the laboratory results were computerized. Statistical analysis was done through Statistical Package for Social Science (SPSS) software computer program version 18 (SPSS, Inc., Chicago, Illinois, USA). The quantitative results were expressed as means ± standard deviation (SD) and qualitative results as number (No.) and percent (%). Independent t-test, Pearson’s χ2, and analysis of variance (ANOVA) with the least significant difference (LSD) as a post hoc test were used in the analysis of the results. Level of significance was adjusted at p ≤ 0.05.
Results
Significant reductions were observed in the mean values of predicted FVC%, predicted FEV1%, and predicted PEF% in wood workers compared with the controls (Table 2). Serum arginase showed significant increase in exposed workers compared with the controls.
Age, ventilatory functions, and arginase activity in the exposed and control groups.

FVC: forced vital capacity; FEV1: forced expiratory volume in the first second; PEF: peak expiratory flow rate; NS: not significant.
Figure 1 illustrates the distributions of the ADAM33 gene loci T1, S1, and Q-1 genotypes among the exposed workers and their controls. For T1, the genotype GA was the highest genotype among the exposed workers, while the homozygous AA was the highest in the controls, with significant difference (χ2 = 6.625, p < 0.05). In T2, homozygous GG showed the highest percentage in the exposed and control groups, without significant difference (χ2 = 2.963, p > 0.05). For S1, there was a significant difference between the two groups (χ2 = 16.98, p < 0.001), as the genotype GA was the most common in the exposed group, and the homozygous AA was the highest among the control group. The homozygous GG for Q-1 was the most common among the exposed group, and the genotype GA showed the highest percentage among the controls, with significant difference (χ2 = 22.51, p < 0.001).
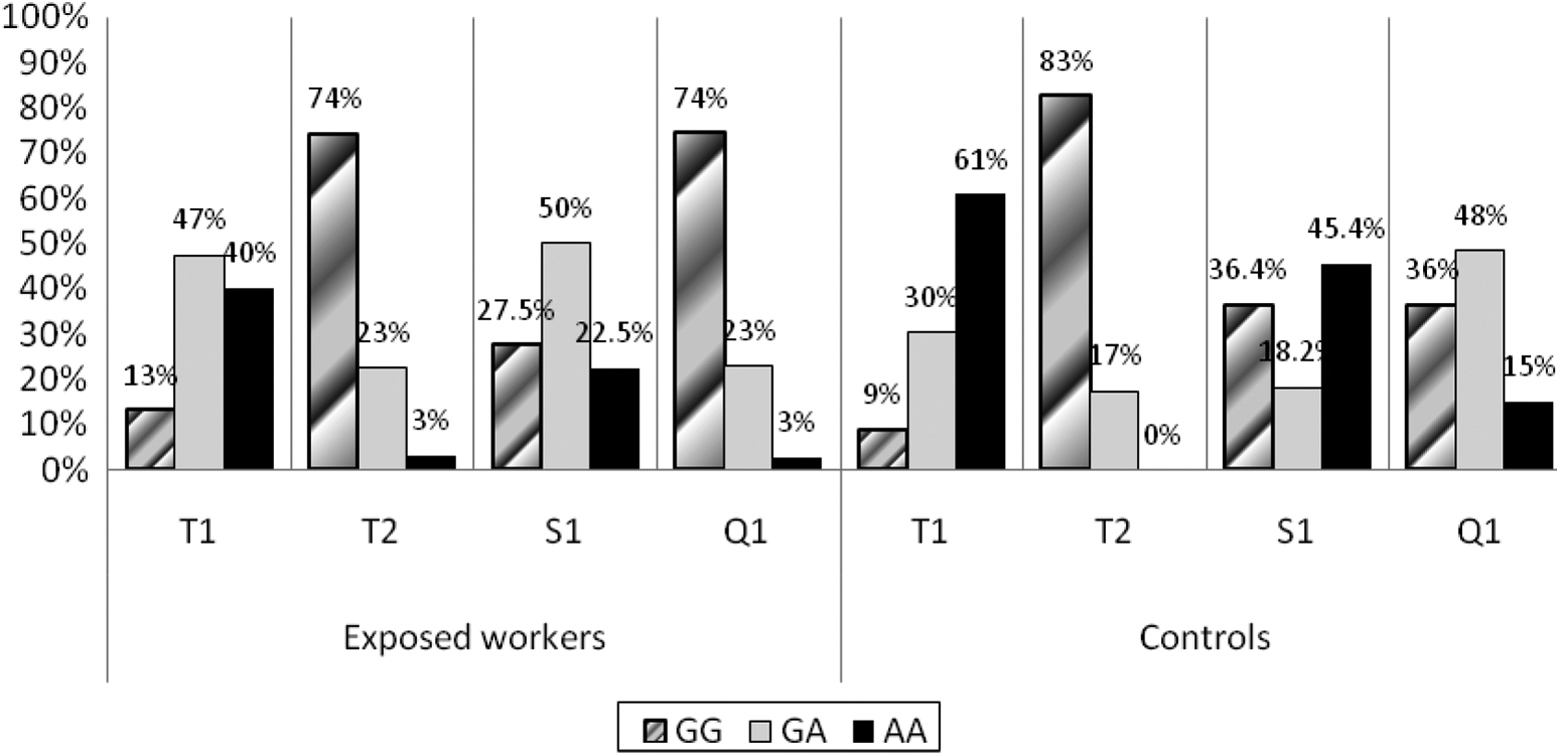
Distribution of different genotypes of T1, T2, S1, and Q-1 SNPs of ADAM33 gene among exposed workers and control groups. SNP: single nucleotide genotype.
Table 3 revealed that in T1 SNP, FVC, and FEV1 were significantly decreased in the workers with GG and AA genotypes compared with the workers with GA, and PEF was significantly decreased in AA genotype compared with the GG and GA. In T2 SNP, PEF in GG and GA was significantly decreased compared with AA genotype. There was no significant difference in the ventilatory functions between the different genotypes of S1 and Q1 loci.
The respiratory functions in different genotypes of T1, T2, S1, and Q1 of the exposed group.a

FVC: forced vital capacity; FEV1: forced expiratory volume in the first second; PEF: peak expiratory flow rate; LSD: least significant difference.
aIndependent t-test was used in comparisons, as zero% of the controls had AA in T2.
Serum arginase was significantly elevated in T2 AA compared to GG and GA, and in S1 GG SNPs compared to GA (Table 4). There was no significant difference in arginase activity between the different genotypes in T1 and Q1.
Comparisons of the arginase activity of different genotypes in T1, T2, S1, and Q1 SNPs of the exposed workers.a

aIndependent t-test was used in comparisons, as zero% of the controls had AA in T2.
A negative correlation was observed between the duration of exposure and measured ventilatory functions in the four studied SNPs, which were obviously significant in AA of T1 and S1, in GG of T2, and in GA of S1 and Q1 genotypes of SNPs (Table 5). PEF was also inversely correlated with the duration of exposure in GG of S1 and Q1. While a positive significant correlation was detected between duration of exposure and arginase activity in AA genotype of T1 and in GG of S1 SNP.
Association between duration of exposure in different genotypes of T1, T2, S1, and Q1 of the workers with their respiratory functions and arginase activity.

FVC: forced vital capacity; FEV1: forced expiratory volume in the first second; PEF: peak expiratory flow rate.
aCannot be calculated as only two workers in the group.
Discussion
Wood dust is a known occupational allergen that may induce respiratory diseases including asthma and allergic rhinitis in exposed workers. 24 An association between fresh wood dust exposure and asthma, coughing, bronchitis, and acute and chronic impairment of lung function was proven. 5 The present study detected significant reduction in the predicted percentages of FEV1, FVC, and PEF in wood workers compared with their controls. These findings were in agreement with Sriproed et al., 25 they observed a significant decrease in the ventilatory functions (FEV1/FVC and PEF) in high dust level-exposed workers across work shift compared with low dust level-exposed workers. Also Jacobsen et al. 6 detected a dose–response relationship between wood dust exposure and cross-shift decline in FEV1 in wood workers.
In the present study, there was a significant elevation in the serum arginase levels in the wood workers compared to their controls. Serum arginase was significantly different in T2 GA and S1 GA SNPs, being highest in AA and GG genotypes, respectively, compared to the other genotypes. Hyseni et al. 20 mentioned that exposure to particulate matter (PM) might induce elevation in arginase, as PM upregulates arginase II activity and expression in human bronchial epithelial cells, in part via epidermal growth factor-dependent mechanisms independent of oxidative stress. Another study reported that increased arginase activity may be involved in lung fibrotic disorders, including idiopathic pulmonary fibrosis. 16 The increase in arginase activity may be contributed in AHR allergic asthma to the stimulation of peroxynitrite formation. 16 Therefore, the current study highlights the association between elevation in the arginase levels in workers exposed to wood dust and the significant decline in the ventilatory functions observed in the wood workers compared to the nonexposed subjects.
The current results were in accordance with Wang et al., 12 they confirmed that T1 and T2 ADAM33 SNPs were significantly associated with decreased FEV1 and FEV1/FVC. ADAM33 is selectively expressed in mesenchymal cells. Alterations in its activity may alter the function of bronchial smooth muscle cells and fibroblasts, ultimately leading to airway remodeling. Airway remodeling can lead to bronchial hyperreactivity and airway obstruction, which is related to asthma. 8 Polymorphisms of the ADAM33 gene could predict excess decline in lung function. 26
In the present study, ADAM33 polymorphism was different between the included exposed workers and their controls. This difference was attributed to that the included subjects in the two groups were selected randomly, and their genotypes were defined at the end of the study. In T1 and S1 genotype, GA was the commonest in the exposed group, while the homozygous AA was the commonest in the control one. For T2, the homozygous GG was the most detectable polymorphism in both groups. Moreover, GG was the most common SNPs in Q1 genotype of the exposed group, and the heterozygous GA was the predominant in the controls. Therefore, statistical analysis comparing the SNPs in ADAM33 and its association with respiratory functions and the arginase levels was done in the workers.
PEF is the diagnostic ventilatory function for asthma. 27 In the present study, the PEF was significantly declined among the ADAM33 T2 GG genotype in wood workers, while a meta-analysis study revealed an association between the prevalence of asthma and the ADAM33 T1 GG genotype. 27
The ADAM33 protein is expressed in airway smooth muscle cells and fibroblasts, and it has been proposed to contribute to the remodeling process present in asthma. 28 It is unknown whether the production or activity of ADAM33 in asthma is increased or reduced. Overproduction or enhanced activity of ADAM33 may lead to excessive shedding of inflammatory mediators, compatible with the enhanced airway wall inflammation present in asthma. Shedding and thereby overproduction of growth factors may furthermore induce proliferation of smooth muscle cells and fibroblasts. These features may lead to the remodeling process present in the airways of asthmatic patients. However, the present results have suggested that this may also reflect a general phenomenon because polymorphisms in ADAM33 could be associated with acceleration in the decline in the ventilatory functions in subjects exposed to wood dust containing allergic agents.
S1 SNP is located in exon S, which encodes the transmembrane region. This polymorphism causes an amino acid change (valine to isoleucine), but it is unknown whether this also modifies the structure of the ADAM33 protein. In the present results, even there was no significant difference in the ventilatory functions of the wood workers according to the SNP S1, duration of exposure played a significant role in its decline, especially among S1 GA and S1 AA for FEV1 and FVC; the indices of chronic obstructive pulmonary disease and among S1 GG and S1GA for PEF; the index of bronchial asthma according to the American Thoracic Society 22 and the Egyptian Society of Chest and Tuberculosis. 29 However, further studies will be needed.
SNP Q1 locus is located in the intron just before exons Q, P, and R, which comprise the epidermal growth factor domain. 30 The intronic Q1 SNP may influence the splicing of the β-variant 31 and disturb the maturation of ADAM33 protein. Therefore, ADAM33 protein may not be anchored correctly in the membrane and may not be able to exert its function. 30 This may lead to progressive destruction of alveolar tissue and thereby enhance accelerated decline in lung functions. 32
In the present study, the ventilatory functions in the workers exposed to wood dust were declined in all the Q1 SNP than normal, but without a significant difference. But the ventilatory indices FEV1 and FVC were significantly declining with the duration of exposure in the workers with Q1 GA. Therefore, subsequent effects of the maturation of ADAM33 protein in the workers with SNP Q1 may result in a defect in tissue repair after inflammation-induced damage due to their exposure to wood dust, and this phenomenon increases with the prolongation of the duration of exposure.
In human subjects, it is important to consider what surrogate markers of remodeling have been used in genetic studies. Baseline FEV1 and airway hyperresponsiveness are determined by a complex interplay of factors, including nonremodeling mechanisms. 33 Currently, this impairment was increased with the duration of exposure to wood industries. It is concluded that ventilatory functions in wood workers were impaired and stratification of results showed a dose–response effect for years of wood dust exposure on the ventilatory functions.
In conclusion, polymorphisms in the ADAM33 gene were associated with an obvious decline in ventilatory functions, mainly FEV1, and this may be helpful in the prediction of susceptible individuals among workers. SNPs in ADAM33 constitute important risk factors for the development of respiratory diseases in individuals exposed to dusty environmental factors such as wood dust. Moreover, arginase activity may play an associated important role in increasing the decline of ventilatory functions in those workers.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
