Abstract
To detect the blood genomic DNA methylation in coke oven workers and find a possible early screening index for occupational lung cancer, 74 coke oven workers as the exposed group and 47 water pump workers as the controls were surveyed, and urine samples and peripheral blood mononuclear cells (PBMCs) were collected. Airborne benzo[a]pyrene (B[a]P) levels in workplace and urinary 1-hydroxypyrene (1-OH-Py) levels were determined by high-performance liquid chromatography. DNA damage of PBMCs and the p14(ARK), p15(INK4b) and p16(INK4a) gene CpG island methylation in the promoter region were detected by comet assay and methylation-specific polymerase chain reaction techniques, respectively. Results show that compared with the controls, concentration of airborne B[a]Ps was elevated in the coke plant, and urinary 1-OH-Py’s level and DNA olive tail moment in comet assay were significantly increased in the coke oven workers, and p14(ARK), p15(INK4b) and p16(INK4a) gene methylation rates were also significantly increased. With the working years and urinary 1-OH-Py’s level, the rates of p14(ARK) and p16(INK4a) gene methylation were significantly increased while that of p15(INK4b) gene methylation displayed no statistical change. We conclude that PBMCs’ p14(ARK) and p16(INK4a) gene methylation may be used for screening and warning lung cancer in coke oven workers.
Keywords
Introduction
Polycyclic aromatic hydrocarbons (PAHs), emitting from the coke oven operation and production, are the primary cause of occupational lung cancer and other occupation-related respiratory system disorders. PAHs, formed mainly from the pyrolysis and incomplete combustion of various kinds of organic matter such as coal tar, coking coal and woods, include benzo[a]pyrene (B[a]P), naphthalene, pyrene, benzo[a]anthracene, chrysene, fluoranthene, anthracene and so on. B[a]P, one of the main members of PAH mixtures, accounts for over 60% of the total carcinogenic activity of PAHs and exhibits carcinogenicity and mutagenicity. The mortality of lung cancer increases by 5% when the concentration of airborne B[a]P increases by 0.001 μg/m3. 1 So B[a]P is usually regarded as a representative of PAHs due to its widespread existence, stability in the environment and correlation with PAHs. 1
B[a]P-induced DNA damage (mainly DNA double-strand breaks) is recognized as a critical biological event in the process of B[a]P’s carcinogenesis and mutagenesis. 2 Abnormal methylation of genomic DNA (mainly DNA CpG island hypermethylation) is regarded as one of the most important epigenetic carcinogenic mechanisms. DNA methylation is the methyl transfer from the S-adenosylmethionine methyl donor to a specific base of the DNA sequence under the catalysis of DNA methyl transferase, which usually happens in the cytosine site of the CpG island in the promoter region. The cytosine is methylated and forms the 5-methylated cytosine in mammalian cells. 3 Genomic DNA CpG island methylation often causes gene mutation and the replacement of cytosine by thymine, gene silencing, chromosome instability, activation of oncogenes and inactivation of anti-oncogenes. 4 Anti-oncogenes such as p14(ARK), p15(INK4b) and p16(INK4a) encoding cell cycle regulatory proteins negatively regulate the cell cycle and inhibit cell proliferation. 5 In cancer tissue, DNA hypermethylations of p14(ARK), p15(INK4b) and p16(INK4a) genes were found in the CpG island of the promoter regions and were correlated with the inactivations of p14(ARK), p15(INK4b) and p16(INK4a) genes. 6 That is to say, DNA methylation of p14(ARK), p15(INK4b) and p16(INK4a) genes is involved in the tumourgenesis and is an early biologic event. Therefore, we aim to explore the blood DNA CpG island methylation of p14(ARK), p15(INK4b) and p16(INK4a) genes in coke oven workers and evaluated whether DNA methylation can be a potential screening and warning index for occupational lung cancer in coke oven workers.
Methods
Subjects
An occupationally B[a]P-exposed group was composed of 74 male Han nationality coke oven workers employed more than 3 years in the same coke plant of a large state-owned company. A working shift lasted 8 h and each worker’s task was relatively constant over 1 year. Controls were 47 male water pump workers in the water supply plant of the same company, matched with the exposed group by age, gender, working years, nationality, socioeconomic status, life style and health status except for occupational exposure to PAHs. Controls had neither visited the coke plant in the previous 3 months. The workplace of controls was about 5 km upwind from the coke plant. All subjects lived in the same residential area and over 2.5 km upwind from the coke plant. The research protocol was approved by the Ethics and Human Subject Committees of Shanxi Medical University, and written informed consents were signed by all subjects themselves.
Reagents
Blood genome DNA extraction kit (Takara Bio Inc., China) and EZ DNA methylation-Gold kit™ (ZYMO Research, Irvine, California, USA) were purchased through the local agents. And the primer sequences were referenced 7 and synthesized by the TAKARA Biotechnology Company Ltd (Dalian, China) and thehe primer sequences are listed in Table 1.
Primer sequences information of genes.

M: methylation primer; U: non-methylation primer; F: forward primer; R: reverse primer.
Questionnaire survey
A structured questionnaire designed by our research group and pre-surveyed prior to the final investigation was used to survey the subjects’ age, gender, nationality, occupational history, working years, smoking and drinking habits, frequency of eating fried and coal-baked foods, histories of personal and family diseases, medication used in the previous 2 weeks and so on. Smoking was defined as currently smoking no less than 10 cigarettes per day over the last year; while drinking was defined as currently drinking wine, beer or spirits no less than 3 times a week for the last 6 months. Subjects who had histories of personal and family tumour diseases were excluded. Urine and blood samples of the subjects were collected in fast in the morning after a night shift. As a final result, 74 exposed workers and 47 controls completed the questions and were included in the study.
Workplace monitoring
Air samplers were located in the bottom, side and top of ovens in the workplace of coke plant and controls, respectively, and two paralleled automatic air sampling pumps were also fixed at personal breathing zone heights and worked at a flow rate of 2 L/min continuously for 6 h during the working day over three consecutive days. Air pressure and temperature data were recorded and used to calculate the standard sampling volume. After sampling, the filters were removed from the sampling pumps, sealed in a clean container, stored at 4°C and transported to the laboratory. B[a]P in the filters was extracted 4 times by hexamethylene and alkaline aluminium oxide and then analyzed using a high-performance liquid chromatography (HPLC) equipped with a fluorescence detector according to the methods for determination of PAHs in the air of workplace (GBZT 160 44-2004).
Internal dose detection
Post-shift urine samples were collected in 50 mL polyethylene plastic tubes in the morning and stored at −80°C until 1-OH-Py levels were analyzed by HPLC according to the standard method.
8
Identification and quantification of B[a]P were accomplished by comparing retention times and peak area ratios of samples with those of standards. Carbazole was used as an internal standard substance. The lowest limit of detection of urinary 1-OH-Py was 1.0 ng/mL urine. Measurements below a concentration of 1.0 ng/mL were given a value of 1.0 ng/mL ratio to
The urinary creatinine (Cr) reacted with alkaline picrate, and the Cr-picrate complex was measured using a UV-V is spectrophotometer (TU-1800S, Bejing Purkinje General Instrument Co., Ltd China) at a wavelength of 520 nm. The concentration of 1-OH-Py was expressed as micromoles per mole Cr (μmol/mol Cr).
Detection for DNA damage
Elbow venous blood samples of 4 mL were drawn from each subject while fasting in the morning. The blood was anticoagulated using ethylenediaminetetraacetic acid and equally divided into two tubes. One tube of blood was frozen at −80°C for DNA extraction. And another tube of blood was used to isolate the peripheral blood mononuclear cells (PBMCs) using Ficoll liquids for DNA damage detection with comet assay. Five random fields of vision cells were counted and photographed with a magnification of 200× under a BX51 fluorescence microscope (Olympus, Japan). The olive tail moments (OTMs) of tailed cells were measured by comet assay software project and expressed in micrometer.
Genomic DNA extraction and methylation detection
The frozen whole blood was melted under room temperature and genomic DNA was extracted with Takara blood genome DNA extraction kit. The purity and concentration of genomic DNA were determined by UV-Vis spectrophotometry. The DNA sample was purified and modified with EZ DNA methylation-Gold kit™ , and then 2 µL DNA (200 ng/µL) was used to detect the methylation by methylation-specific polymerase chain reaction (MS-PCR, MSP). The MSP reaction system consisted of two parallel PCR tubes; one tube was used for detecting DNA methylation and another for DNA non-methylation, each tube with 25 µL volumes containing 10× buffer 2.5 µL, 0.2 mmol/L deoxyribonucleoside triphosphate, 0.05 U Taq Plus DNases, 0.2 µmol/L forward primer and reverse primer and 400 ng DNA template. The methylation primers were added into the DNA methylation tube and non-methylation primers were added to the non-methylation tube. In the meantime, a blank control without DNA template was used for co-amplification. And the amplification was carried out as follows: initial denaturation at 95°C for 3 min, denaturation at 95°C for 30 s, annealing at 62°C (p14(ARK)-M), 60°C (p14(ARK)-U, p15(INK4b)-M, p15(INK4b)-U), 65°C (p16(INK4a)-M), 61°C (p16(INK4a)-U) for 40 s, extension at 72°C for 30 s, 35 cycles, then extension for 5 min at 72°C. The PCR product was identified the methylation using 2% agarose gel electrophoresis. When DNA is methylated, one band appears in the methylation lane; and when DNA is non-methylated, one band appears in the non-methylation lane and no band in the methylation lane.
Data processing and statistical analysis
Data were input into a personal computer with software EpiData 3.0 and then were analyzed by Statistival Package for Social Sciences version 11.5 for Windows (SPSS Inc., Chicago, Illinois, USA). Urinary 1-OH-Py levels were log 10 transformed to normalize the distribution. Student’s t test was used to compare the difference of 1-OH-Py levels, ages, working years and OTM values in comet assay between the two groups, and χ 2 test was used to analyze the habits of smoking, drinking and DNA methylation rates among groups. All p values were 0.05 with two-sided.
Results
Workplace monitoring
The mean concentrations of airborne B[a]P were 16.8 ± 11.2, 177.5 ± 40.6 and 1286.5 ± 466.3 ng/m3 at the bottom, side and top of the coke oven, respectively, and were higher than the mean value at the workplace of control (8.7 ± 6.7 ng/m3). The mean B[a]P concentration at the top of the coke oven (1286.5 ng/m3) was over 147 times of that at the controls’ workplaces (8.6 ng/m3), far exceeded the maximum allowable concentration 150 ng/m3 in the workplace of airborne B[a]P set by the ex-Soviet Union.
Subjects’ general information, urinary 1-OH-Py levels and DNA damage
Table 2 shows that age, working years, smoking and drinking habits were matched between the exposed group and the control group (p > 0.05). Urinary 1-OH-Py levels and OTM values in comet assay were significantly increased in the exposed groups compared with the controls (p < 0.05 or p < 0.01). Furthermore, the OTM values in comet assay were correlated with urinary 1-OH-Py levels, and the Pearson correlation coefficient was 0.361.
General information, urinary 1-OH-Py levels and OTM values (mean ± SD) of subjects.a

1-OH-Py: 1-hydroxypyrene; OTM: olive tail moment.
a t means the statistical value for t test of the quantitative data between groups, and χ 2 represents the statistical value for χ 2 test of the qualitative data between groups.
b p < 0.05 or p < 0.01: significant difference compared with the control group.
DNA methylation of p14(ARK), p15(INK4b) and p16(INK4a) genes
Table 3 indicates that the detection rates of p14(ARK), p15(INK4b) and p16(INK4a) gene methylation were 10.7, 10.7 and 22.3% in the exposed group, respectively, and were significantly increased compared with the control group (p < 0.05 or p < 0.01).
Comparison results of DNA methylation between the two groups N (%).a
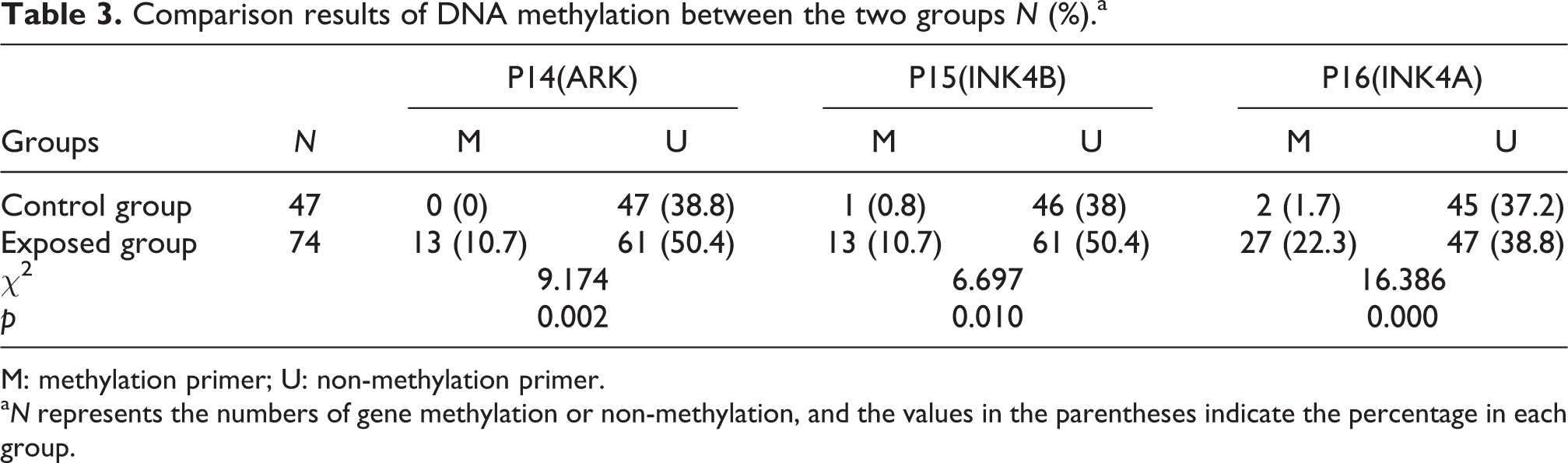
M: methylation primer; U: non-methylation primer.
a N represents the numbers of gene methylation or non-methylation, and the values in the parentheses indicate the percentage in each group.
The exposed workers were categorized into three groups according to the 30th and 70th percentiles of working years, <13 year group (n = 24), 13–24 year group (n = 32) and 25–29 year group (n = 18). Table 4 indicates that the detection rates were gradually increased with the working years in p14(ARK) and p16(INK4a) gene methylation, however, which did not show an increasing or decreasing trend in p15(INK4b) gene. P14(ARK) gene methylation was significantly increased with the working years (p < 0.05). There was no significant difference in p16(INK4a) gene methylation among the groups (p > 0.05).
DNA methylation changes with the working years in the exposed group N (%).a

M: methylation primer; U: non-methylation primer.
a N represents the numbers of gene methylation or non-methylation, and the values in the parentheses indicate the percentage in each group.
All subjects were categorized into three groups according to the 25th and 75th percentiles of their urinary 1-OH-Py levels, <0.18 μmol/mol Cr group (n = 33), 0.18–0.50 μmol/mol Cr group (n = 59) and 0.51–0.60 μmol/mol Cr (n = 29). Table 5 indicates that the detection rates of p14(ARK), p15(INK4b) and p16(INK4a) gene methylation were gradually increased with the levels of urinary 1-OH-Py. And there were significant differences in p14(ARK) and p16(INK4a) gene methylation among the three groups (p < 0.01 or p < 0.05), yet the difference of p15(INK4b) gene methylation was not significant among the groups (p > 0.05).
DNA methylation changes with the 1-OH-Py’s levels in subjects N (%).a

1-OH-Py: 1-hydroxypyrene; M: methylation primer; U: non-methylation primer.
a N represents the numbers of gene methylation or non-methylation, and the values in the parentheses indicate the percentage in each group.
Discussion
PAHs, a kind of important occupational and environmental pollutant, are by-products mainly from coke oven production, which exhibit carcinogenicity and mutagenicity. The morbidity and mortality of lung cancer were extremely high in the occupational exposure population such as the coke oven workers. B[a]P is regarded as the major pathogen of lung cancer in coke oven workers. DNA damage (mainly DNA double-strand breaks) induced by B[a]P is the main mechanism of carcinogenicity. In this research, the concentration of airborne B[a]P was significantly higher in the coke oven plant than that in the workplace of controls and increased with the altitude of coke oven. Urinary 1-OH-Py levels and DNA OTM values in comet assay were significantly increased in the coke oven workers in comparison with the control group. Moreover, DNA values in comet assay were correlated with the 1-OH-Py levels and B[a]P contents in the workplace.
DNA methylation is one of the most important epigenetic carcinogenic mechanisms. DNA methylation of the CpG island in the tumour suppressor gene promoter make the tumour suppressor gene silence, induce immoderate cell growth and proliferation and ultimately result in the occurrence of tumours. 9 Tumour suppressor genes P14(ARK), p15(INK4b) and p16(INK4a) are all located in 9p21 where the mutation occurred frequently in tumour tissues. 10 P14, p15 and p16 proteins suppress cell cycle by arresting cell cycle progression in the S phase and inhibit cell proliferation. The DNA promoter hypermethylation in CpG island of p14(ARK), p15(INK4b) and p16(INK4a) genes, widespread in tumour tissues, can cause the inactivation of these genes and their encoded proteins. DNA hypermethylation in the promoter regions of p14(ARK), p15(INK4b) and p16(INK4a) genes was detected in pulmonary squamous cell carcinoma. 7 Patients showed the worst prognosis with p14(ARK), p15(INK4b), and p16(INK4a) genes’ inactivation. 11
In this research, the PBMCs’ p14(ARK), p15(INK4b) and p16(INK4a) gene hypermethylation in CpG island were significantly increased in the coke oven workers compared with the control group. DNA hypermethylation of p14(ARK) and p16(INK4a) genes gradually increased with the working years and urinary 1-OH-Py levels. And p15(INK4b) gene hypermethylation showed an increasing trend with urinary 1-OH-Py levels though the difference was not significant statistical.
P14(ARK), p15(INK4b) and p16(INK4a) genes are all located in the same region of chromosome 9p21 and exhibit inhibition of cell cycle G 1 phase progression. But no correlation was found between p15(INK4b) gene methylation and p14(ARK) or p16(INK4a) gene methylation. 12 P15(INK4B) promoter methylation seldom seems to occur in solid tumours but is a major gene silencing mechanism in haematological malignancies. P14(ARK) and p16(INK4a) gene methylation in the promoter region often occurs in solid tumours, leukaemias and lymphomas. 13 P14(ARK) gene hypermethylation was regarded as an early event in the process of lung carcinoma 14 and can be used as a marker or predictor for tumour diagnosis or recurrence. 15
Usually, P16(INK4a), is inactivated by gene mutation and gene hypermethylation. And p16(INK4a) gene methylation in the promoter region is the primary reason for p16(INK4a) gene inactivation or silencing. Inactivation of p16(INK4a) gene showed loss of p16(INK4a) protein expression 16 and was significantly correlated with tumour size, tumour grade, 10 and pathological classification, and was reversed by the methylation-specific inhibitors. 17 Moreover, p16(INK4a) gene methylation was not significantly correlated with p15(INK4b) and p14(ARK) gene methylations. 10 In addition, PBMCs’ p16(INK4a) gene methylation in patients with lung cancers may reflect the tumour-bearing status in body. 18
This research indicates that the detection rates of PBMCs’ p14(ARK) and p16(INK4a) gene methylation were significantly higher in the B[a]P-exposed workers and were dependent on the working years and the internal exposure in the coke oven workers. Yang et al. reported that p16(INK4a) gene methylation was associated with DNA damage and PAHs’ internal exposure in the PAH-exposed workers and might be an essential biomarker for PAHs’ exposure and for early diagnosis of cancer. 19 In conclusion, p14(ARK) and p16(INK4a) gene hypermethylation in PBMCs might be a good biomarker for the lung cancer early screening and warning in coke oven workers.
Footnotes
Acknowledgements
The authors acknowledge the medical staff of the Center of Disease Control and Prevention, Taiyuan Iron and Steel Company (CDCTISCO), Taiyuan, People’s republic of China, for assistance with collecting air, urine and blood samples. We also gratefully and sincerely thank the administrative personnel of the CDCTISCO for their great help in organizing and facilitating the investigation and for their thoughtful comments. We would also like to thank graduates and postgraduates of Shanxi Medical University for their selfless contribution to this study. More importantly, we sincerely thank the workers who consented to participate in this study.
Conflict of interest
The authors declared no conflicts of interest.
Ethics approval
This study was conducted with the approval of the Ethics and Human Subject Committees of Shanxi Medical University.
Funding
This study was supported by the grant from the National Natural Science Foundation of China (81001258); the grant from Key Science and Technology Project of Shanxi Province (201003111107-2); the grants from Shanxi Provincial Foundation for Leaders of Disciplines in Science (2011); and Shanxi Province Science Foundation for Youths (2013021033-3).
