Abstract
The neurotoxin paraquat (PQ) causes apoptosis of dopaminergic neurons in mammalian cell culture and animal models, mimicking an important pathological feature of Parkinson’s disease (PD). The phosphoinositide 3-kinase (PI3K)/Akt pathway is critical for several major survival signals in central nervous system neurons. Phosphatidylinositol 3-kinase 55 kDa gamma (p55PIK) is a regulatory subunit of PI3Ks with important roles in cell proliferation, antiapoptosis, and cell cycle progression. However, p55PIK involvement in mechanisms regarding progression and maintenance of neurodegenerative diseases is largely undetermined. We used PQ-induced apoptosis in human dopaminergic SH-SY5Y cells to investigate the association between p55PIK expression levels, subcellular location, and apoptosis. p55PIK expression was reduced in SH-SY5Y cells and p55PIK messenger RNA and protein expression levels were decreased after PQ treatment. Apoptosis induced by PQ was associated with caspase activation and decreased p55PIK expression. Restoration of p55PIK expression was observed after coincubation with a caspase inhibitor. Overexpressed full-length p55PIK in SH-SY5Y and human embryonic kidney 293 cells showed specific distribution in the nucleus and was cleaved in vitro by recombinant caspase 6 (C6), but not C3 and C7. A p55PIK construct lacking 24 N-terminal amino acids (N24) was tested for the presence of a potential C6-recognizable sequence and was found to express its proteins outside the nucleus. The results suggest that p55PIK may be involved in PQ-induced apoptosis signal transduction and that N24 is crucial for p55PIK subcellular localization. Thus, p55PIK could be a substrate of activated C6 during apoptosis, leading to loss of original biological functions and redistribution to disturb cell cycle progression.
Introduction
Phosphatidylinositol 3-kinases (PI3Ks) are intracellular transducers with lipid and serine/threonine kinase activities that are composed of different heterodimers containing a catalytic subunit from a large family of regulatory subunits. 1 PI3Ks are recruited and activated by multiple receptor tyrosine kinases. 2 After activation, PI3Ks catalyze phosphorylation at position 3 in the inositol ring of phosphoinositides, resulting in 3-phosphoinositides, primarily phosphatidylinositol-3-trisphosphate (PIP3). These lipids act as docking sites for recruitment at the plasma membrane of protein-bearing pleckstrin homology domains, such as Akt/PKB, PDK1, BTK, and PLCγ. Binding to PIP3 lipids activates these proteins to signal a wide array of downstream effectors that ultimately lead to multiple cellular responses. 3 –7
Phosphoinositide 3-kinase regulatory subunit 3, also known as phosphatidylinositol 3-kinase 55 kDa gamma (p55PIK; p55γ ), is a PI3K regulatory subunit. Two common Src-homology-2 (SH2) domains at the C-terminus are capable of binding phosphorylated tyrosines that link this pathway to receptor tyrosine kinase signal transduction cascades, including an inter-SH2 domain that binds to the PI3K catalytic subunit 8 and a unique 24 amino acid N-terminal end (N24). p55PIK was originally isolated by expression library screening for proteins that interact with phosphorylated insulin receptor substrate 1 and was found to interact with insulin-like growth factor 1 receptor (IGF1R). 3,8,9 The retinoblastoma protein (Rb), a key regulator of cell cycle progression, associates with p55 kDa (p55α and p55γ ) regulatory subunits of PI3K in vivo and in vitro. The Rb-p55γ interaction observed in quiescent cells appears to be necessary for initiation of cell cycle progression. 10
p55PIK expression levels are low in adipocyte tissues and skeletal muscle compared to brain and testis tissues, suggesting a restricted role of p55PIK in neuronal signaling. In rat PC12 cells, PI3K is necessary at multiple stages of neurite outgrowth during nerve growth factor-stimulated differentiation. Other studies have reported that p55PIK messenger RNA (mRNA) expression levels are significantly upregulated in ovarian cancer compared with normal ovaries, while knockdown of p55PIK expression by small interfering RNA significantly increased apoptosis in cultured ovarian cancer cell lines. 11 These data strongly suggest that the PI3K pathway may gain function via p55PIK amplification and overexpression in ovarian cancer. Apoptosis plays a critical role in the development of the central nervous system and may be involved in many neurodegenerative diseases. PI3K/Akt activation is a promising strategy to achieve neuronal protection or prevent cell death. 12 In contrast, inhibition of this survival pathway might induce apoptosis. Data have indicated that the PI3K catalytic subunit can selectively interact with regulatory subunits that mediate signaling of specific PI3K pathways. 13 However, the signaling mechanisms for specific pathways and functions mediated by specific regulatory subunits of PI3K are not well understood. Specific contributions of each subunit in the progression and maintenance of neurodegenerative diseases is largely undetermined.
The agricultural pesticide paraquat (PQ) is a potent inducer of apoptosis in neuronal cells and may increase the risk for Parkinson’s disease (PD) in human populations. 14 In this study, PQ-induced apoptosis was evaluated in dopaminergic SH-SY5Y cells as a PD apoptotic model to investigate changes in and regulation of p55PIK expression and subcellular localization.
Materials and methods
Materials
Dulbecco’s modified Eagle’s medium/Ham’s Nutrient Mixture F-12 (DMEM/F12), DMEM high glucose medium, penicillin, streptomycin, and fetal bovine serum (FBS) were obtained from HyClone Laboratories (Logan, Utah, USA). PQ, 4′,6-diamidino-2-phenylindole (DAPI), and N-benzyloxycarbonyl-Val-Ala-Asp fluorome-thylketone (z-VAD-fmk) were purchased from Sigma-Aldrich (St Louis, Missouri, USA). Trizol® reagent, lipofectamine™ 2000, and the Neon transfection device were obtained from Invitrogen (Carlsbad, California, USA). PrimeScript® reverse transcription (RT) reagent kit and SYBR® Prime EX Taq™ and the bicinchoninic acid (BCA) protein assay kit and enhanced chemiluminescent (ECL) Plus reagents were obtained from TaKaRa Biotechnology (Shiga, Japan) and Beyotime Biotechnology (Beijing, China), respectively. Antibodies to β-actin (SC-47778), caspase 3(C3; SC-7148), Poly (adenosine diphosphate-ribose) polymerase (PARP; SC-8007), and p55PIK (SC-50289) were obtained from from Santa Cruz Biotechnology (Santa Cruz, California, USA), and recombinant human C3, C6, and C7 were obtained from BioVision (Milpitas, California, USA).
Cell culture and drug treatment
Human neuroblastoma SH-SY5Y and human embryonic kidney 293 (HEK293) cells were grown in DMEM/F12 and DMEM high glucose medium, respectively, containing 100 U/ml penicillin, 100 μg/ml streptomycin, and 10% FBS at 37°C with 5% carbon dioxide in a humidified incubator. Based on a previous study, logarithmic growth phase SH-SY5Y cells were incubated with 250 μM PQ for 36 h to induce apoptosis. 15 For the caspase activity inhibition assay, SH-SY5Y cells were preincubated with 50 μM z-VAD-fmk for 30 min before PQ treatment.
RT-PCR and real-time PCR
RNA was extracted from cells cultured under experimental conditions (control cells or 250 μM PQ for 36 h) using Trizol reagent according to the manufacturer’s instructions. RT was performed using a PrimeScript RT reagent kit according to the manufacturer’s instructions. Complementary DNA (cDNA) primers were designed as follows: β-actin, forward (F): 5′-AGA TGA CCC AGA TCA TGT TTG-3′; reverse R: 5′-ATC ACG ATG CCA GTG GTA-3′ and p55PIK, F: 5′-AAG GAG TGA GAC AGA AAC GC-3′; R: 5′-TGC TTG TAC TCG ATT GAT ATC CTC-3′. Thermal cycling conditions were 95°C for 30 s, 40 cycles at 95°C for 5s and 60°C for 34 s.
Real-time polymerase chain reaction (PCR) was performed with an ABI 7500 with SYBR Prime EX Taq according to the manufacturer’s instructions. The relative transcript amount of the target gene was normalized to β-actin using the quantitatiive comparative cycle of threshold fluorescence (ΔΔC T) method. Threshold fluorescence values were 0.2929 for p55PIK and 0.2691 for β-actin. The final results of real-time PCR were expressed as the ratio of mRNA relative to the control. Each sample was run in triplicate, and each PCR experiment included three nontemplate control wells.
Western blot analysis
Cells under experimental conditions were harvested and lysed in an appropriate volume of enhanced radioimmunoprecipitation assay buffer containing 1 mM phenylmethylsulfonyl fluoride for 30 min. Protein concentrations were measured using the BCA protein assay kit. Equal amounts of protein were analyzed by 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred to polyvinylidene difluoride at 70 V for 60 min using a mini-trans blot cell apparatus (Bio-Rad, Hercules, C alifornia, USA). Membranes were incubated overnight at 4°C with primary antibodies anti-p55PIK, anti-C3, anti-PARP (all at a 1:100 dilution) and anti-β-actin (1:1000 dilution). After washing with phosphate-buffered saline with Tween 20 (PBST) for three 10-min periods, membranes were incubated for 60 min at 25°C with horseradish peroxidase-conjugated secondary antibodies (1:2000 dilution). After washing with PBST for three 10-min periods, detection of bound antibodies was visualized by chemiluminescence using ECL Plus reagent. β-actin content was analyzed as a control. Densitometry values were normalized to β-actin levels and the control lane in each blot.
Construction of eukaryotic expression plasmids with the p55PIK gene and its mutant
The full-length p55PIK coding sequence was amplified by PCR from the cDNA library of human peripheral blood cells (primer F:5′-GGA ATT CCT GAC AGC GCG ATG TAC AAT-3′ and primer R: 5′-GGG GTA CCC CTT CCT CTT TTC TGC A-3′). PCR products were subcloned into a pEGFP-N1 plasmid using EcoR I and Kpn I restriction enzyme sites. To compare the functional differences between p55PIK proteins with and without N24, we constructed a recombinant pEGFP-N1-Δ1–50 p55PIK that only encodes 51–461 amino acids of p55PIK. A set of primers (F: 5′-CGG AAT TCC GAT GGG AAT GAA GGA CA-3′ and reverse primer with the same sequence as full-length P55PIK) was used for PCR amplification. The resulting products were subcloned as previously described. DNA sequences of all constructs were verified using standard sequencing protocols.
Transient transfection
SH-SY5Y cells were transiently transfected using a Neon transfection device with an electroporation protocol of conditioning at 1600 V pulse voltage, 30 ms pulse width, and one pulse. HEK293 cells were transfected using lipofectamine 2000, with 4 μg of plasmid DNA per 10-cm2 tissue culture dish according to the manufacturer’s protocol.
Cell fluorescence microscopy analysis
SH-SY5Y or HEK293 cells were seeded onto glass coverslips. After transfecting for 48 h with pEGFP full-length p55PIK or Δ1–50 p55PIK, cells were directly observed under inverted fluorescence microscopy. Transfected cells were placed into 3% paraformaldehyde for 15 min and the coverslips were incubated with DAPI solution (5 μg/ml) at 37°C for 15 min. Cells were observed via fluorescence microscopy (TCS SPE; Leica, Heidelberg, Germany).
Caspase substrate assay
Full-length p55PIK or Δ1–50 p55PIK were overexpressed in SH-SY5Y cells and HEK293 cells. The cells were rinsed twice with cold PBS, removed by scraping, and centrifuged at 900g for 5 min at 4°C. Cells were resuspended in caspase assay buffer (50 mM hydroxyethyl piperazine ethanesulfonic acid (HEPES), pH 7.2, 50 mM sodium chloride, 0.1% 3-[(3-cholamidopropyl)-dimethylammonio]-1-propane sulfonate (CHAPS), 10 mM ethylenediaminetetraacetic acid, 5% glycerol, and 10 mM dithiothreitol), sonicated, and centrifuged at 13,000g for 5 min at 4°C. Supernatants were incubated in the presence or absence of recombinant human C3, C6, and C7 (protein/caspase = 50:1 wt/unit) for 4 h at 37°C. Western blot analysis was performed with an anti-p55PIK antibody.
Statistical analysis
For statistical analysis, the mean and standard deviation from at least three experiments were calculated. In all the experiments, data were analyzed using the t test, and the values of p < 0.05 were considered statistically significant.
Results
p55PIK mRNA and protein expression levels are altered during PQ-induced SH-SY5Y apoptosis
SH-SY5Y cells were exposed to 250 μM PQ for 36 h and observed for the formation of apoptotic dopaminergic cells. Pro-casp 3, a prototypical caspase often used as an apoptotic marker, was activated as an effector C3 (Figure 1(a) and (b)). PARP, a nuclear DNA-binding zinc finger protein and a C3 substrate, was proteolytically cleaved to yield 85 kDa fragment (Figure 1(a) and (b)). Morphological changes in the cell nucleus were observed by fluorescence microscopy, and apoptotic nuclei in SH-SY5Y cells were visible following DAPI staining after PQ treatment (Figure 1(c)). PQ induces SH-SY5Y apoptosis, which may be caspase dependent.

Neurotoxin PQ-induced SH-SY5Y cell apoptosis. (a,b): PARP and pro-C3 in SH-SY5Y cells were analyzed by Western blot. SH-SY5Y cells were treated with 250 μM PQ for 36 h, and specific antibodies were used to detect cleaved PARP and pro-C3. β-actin was used to confirm equal loading of the extracts. Protein levels were quantified using densitometric analysis and normalized to β-actin (error bars are SE; n = 3). *p < 0.05. (c) Cells cultured on coverslips were fixed and stained with DAPI for detection of nucleic pathology. Arrows indicate apoptotic bodies. Bar: 10 μm. These findings were consistent over three separate experiments. C3: caspase 3; PQ: paraquat; PARP: poly (adenosine diphosphate-ribose) polymerase; DAPI: 4′,6-diamidino-2-phenylindole.
To explore whether the p55PIK subunit of PI3Ks associated with cell survival signaling is involved in PQ-induced apoptosis, we measured p55PIK mRNA expression by real-time quantitative PCR in SH-SY5Y cells exposed to PQ. p55PIK mRNA levels were reduced in PQ-treated SH-SY5Y cells compared with the control cells (Figure 2(a)). p55PIK protein expression in SH-SY5Y cells and SH-SY5Y cells with overexpressed recombinant full-length p55PIK (exposed to 250 μM PQ for 36 h) were investigated using an antibody specific to the N-terminal of p55PIK. As shown in Figure 2(b) to (e), p55PIK protein expression was downregulated in the PQ treatment groups compared with the controls. We further examined whether the downregulation of p55PIK protein expression due to PQ exposure could be counteracted by coincubation with z-VAD-fmk, a cell-permeable pan-caspase inhibitor that irreversibly binds to the catalytic site of caspase proteases. As expected, coincubation of PQ with z-VAD-fmk resulted in a significant increase in p55PIK expression (Figure 2(d) and (e)). These findings suggest that downregulation of p55PIK expression is involved in PQ-induced SH-SY5Y cell apoptosis.

Effects of PQ on p55PIK mRNA and protein levels. (a) p55PIK mRNA expression levels in SH-SY5Y cells treated with or without PQ. The quantitative histogram represents statistical data from real-time PCR. (b,c) p55PIK protein expression levels in SH-SY5Y cells exposed to PQ determined by Western blotting. Protein levels were quantified by densitometric analysis and normalized to β-actin (error bars are SE; n = 3). (d,e) Overexpressed p55PIK protein in SH-SY5Y cells exposed and not exposed to PQ and incubated with z-VAD-fmk before exposure to PQ. Western blotting was performed and protein levels were quantified. *p < 0.05; **p < 0.01. These findings were consistent over three separate experiments. PQ: paraquat; p55PIK: phosphatidylinositol 3-kinase 55 kDa gamma; mRNA: messenger RNA; z-VAD-fmk: N-benzyloxycarbonyl-Val-Ala-Asp fluorome-thylketone.
Identification of a C6 recognizable sequence in the N-terminal of p55PIK
The caspase subgroup for apoptosis includes C3, C6, and C7. Interaction between activated caspases and specific substrates completes the apoptotic process. To determine whether caspases directly utilize p55PIK as a substrate and participate in PQ-induced cell apoptosis, we overexpressed recombinant full-length and Δ1–50 p55PIK in SH-SY5Y and HEK293 cells. The two recombinant proteins were detected as 78 kDa and 72 kDa proteins using Western blotting (Figure 3(a) and (b)). Cell lysates containing the two recombinant proteins were incubated with human recombinant C3, C6, and C7. Expressed full-length p55PIK in SH-SY5Y cells was reduced by recombinant C6, but changes were not observed with exposure to C3 or C7 (Figure 3(a) and (c)). Δ1–50 p55PIK expression in SH-SY5Y cells did not change after exposure to any of the three caspases (Figure 3(a) and (c)). Full-length p55PIK expressed in HEK293 cells was cleaved by recombinant C6 into a novel fragment between the full-length and Δ1–50 p55PIK in addition to the reduced full-length p55PIK (Figure 3(b) and (c)), although cleavage was not observed in Δ1–50 p55PIK (Figure 3(b) and (c)) as was observed in SH-SY5Y cells. The results indicate that there may be a C6-recognizable sequence in the p55PIK protein, implying that p55PIK might be a downstream substrate of the apoptotic signaling pathway and that a PQ-induced reduction in p55PIK levels could be due to effector caspase proteolysis. Lack of Δ1–50 p55PIK cleavage by recombinant C6 suggests that the C6 cleavage site in p55PIK must be located in the N-terminal region.

Caspase-mediated cleaved p55PIK. (a) SH-SY5Y cell lysates containing overexpressed full-length p55PIK and Δ1–50 p55PIK were incubated with human recombinant C3, C6, and C7, respectively. Western blots were performed and samples were probed with a primary antibody to p55PIK. (b) HEK293 cell lysates containing overexpressed full-length p55PIK and Δ1–50 p55PIK were incubated with human recombinant C3, C6, and C7, respectively. Western blots were performed and samples were probed with a primary antibody to p55PIK. (c) Protein levels were quantified by densitometric analysis and normalized to controls. *p < 0.05 (error bars are SE; n = 3). p55PIK: phosphatidylinositol 3-kinase 55 kDa gamma; C3: caspase 3; C6: caspase 6: C7: caspase 7; HEK293: human embryonic kidney 293.
N-terminal p55PIK domain determines subcellular localization
p55PIK was expressed at lower levels in SH-SY5Y cells. Previous studies have shown that N24 in p55PIK has an important function. To understand p55PIK distribution, dopaminergic SH-SY5Y cells were transfected with full-length or Δ1–50 p55PIK expression plasmids using a Neon transfection device for 48 h before direct observation under fluorescence microscopy. The ratio of the cells detected using a fluorescent signal was higher in the full-length p55PIK group, but most of the positive cells appeared to have morphological changes and were floating in the culture medium, implying that p55PIK overexpression may lead to abnormal cell population. After SH-SY5Y cells were fixed with paraformaldehyde, full-length p55PIK was primarily expressed in the nuclei, and some cells were condensed with a pearl-like appearance (Figure 4(a) to (c)). Compared with the full-length p55PIK, Δ1–50 p55PIK had increased expression and distribution, even in the cytoplasm of SH-SY5Y cells (Figure 4(d) to (f)).
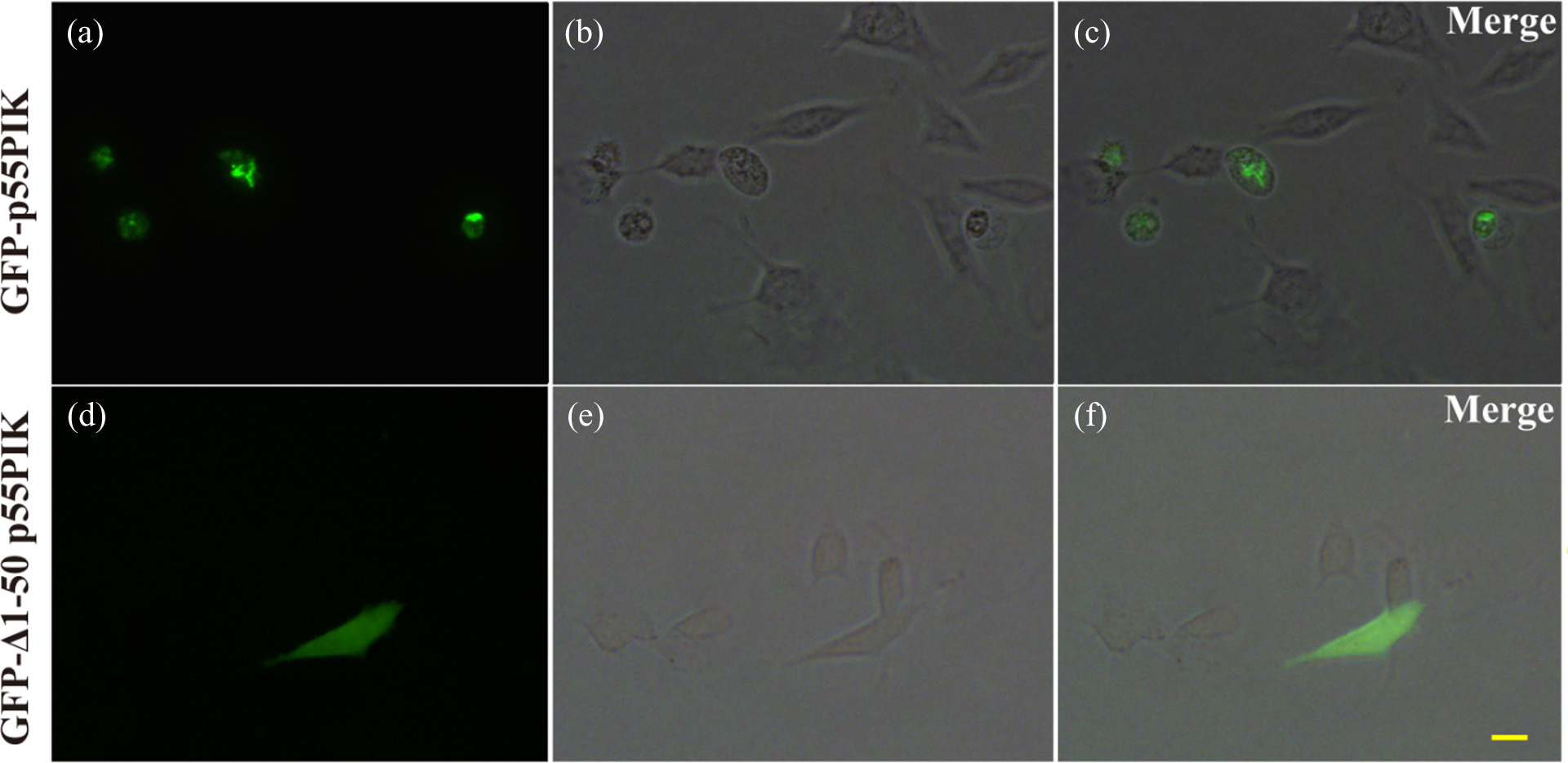
Subcellular location of full-length p55PIK and Δ1–50 p55PIK recombinants in SH-SY5Y cells. pEGFP-p55PIK (a to c) or pEGFP-Δ1–50 p55PIK (d to f) were transfected into SH-SY5Y cells via the Neon transfection device. (a,d) Green fluorescent cells represent transfected GFP-p55PIK recombinants; b,e, and c,f are phase and merge images, respectively. Scale bars: 10 μm. GFP: green fluorescent protein; p55PIK: phosphatidylinositol 3-kinase 55 kDa gamma.
To rule out possible impacts of electroporation transfection on SH-SY5Y cells, pEGFP-N1 vectors or pEGFP-N1-HIPPI expressing a protein interactor associated with Huntington’s disease were also transfected. Under the same electroporation conditions, there were no obvious morphological changes or floating cells as observed in SH-SY5Y cells (data not shown). In addition, we transfected full-length p55PIK and Δ1–50 p55PIK plasmids into HEK293 cells. As in SH-SY5Y cells, similar subcellular localization was observed (Figure 5(a) to (f)). These data suggest that the N-terminal domain of p55PIK may determine subcellular localization and imply that this region has an important function.

Expression and distribution of full-length and Δ1–50 p55PIK recombinants in HEK293 cells. pEGFP-N1-full length p55PIK (a to c) and pEGFP-N1-Δ1–50 p55PIK (d to f) were transfected into HEK293 cells using the Fugene-6 reagent. a,d Green fluorescent cells represent transfected GFP-p55PIK recombinants; b,e and c,f are phase and merge images, respectively. Scale bars: 10 μm. GFP: green fluorescent protein; p55PIK: phosphatidylinositol 3-kinase 55 kDa gamma; HEK293: human embryonic kidney.
Discussion
Although the importance of genetic inheritance and environmental toxins has been debated, the cause of PD remains unknown. Exogenous environmental toxins, such as PQ and rotenone, have been linked to PD. 16,17 Similarly, although exposure to PQ triggers the death of neurons in vitro and in vivo, the exact cause of selective dopaminergic neuronal loss is unknown. PQ-induced apoptotic cell death has been studied in several cell lines in which it causes widespread oxidative and nitrosative stress. 18 PQ generates reactive oxygen species and induces selective neurodegeneration in dopaminergic neurons in the substantia nigra pars compacta. 19 In a previous study, associations between PD and disturbances in neurotransmitter transmission in dopaminergic neurons were observed by investigating interactions between secretogranin III (SCG3), which is closely associated with secretory granules in neuroendocrine cells, and PQ or dopamine (DA) under PQ- and/or DA-induced apoptosis. 15 The results suggested that SCG3 might be a new candidate target in the neuroendocrine pathway. However, involvement of other signal transducers in PQ-induced dopaminergic neuron apoptosis is not known. PI3K activation is a common signaling element that plays an important role in regulating biological responses. These cellular events are known to regulate cell growth, survival, proliferation, and movement. 20 Moreover, the PI3K/Akt pathway may prevent SH-EP1 cells from apoptosis caused by 1-methyl-4-phenylpyridinium, a positively charged molecule, which has a similar chemical structure as PQ. 6 p55PIK, an important regulatory subunit of PI3Ks, has been shown to be exclusively expressed in a few tissues and highly expressed in many tumors. 11,21 IGF-2 may activate PI3K by inducing interactions between p55PIK and the tyrosine-phosphorylated intracellular complex, resulting in growth-promoting abilities through the PI3K/Akt pathway. p55PIK knockdown inhibits these abilities and increases the possibility of cellular apoptosis. 11 These results indicate that p55PIK is involved in the PI3K/Akt signaling pathway and decreased p55PIK may be related to cell apoptosis. In this study, PQ induced apoptosis in dopaminergic SH-SY5Y cells. Characteristic apoptotic features included caspase activation and PARP cleavage as well as nuclear condensation/fragmentation. These results are consistent with the previous reports. Interestingly, p55PIK mRNA and protein expression levels were decreased in the context of PQ-induced dopaminergic cell apoptosis, and downregulation of p55PIK protein expression was partially restored by z-VAD-fmk. To the best of our knowledge, this is the first study to provide evidence that changes in expression of the regulatory subunit p55PIK may be associated with PQ-induced dopaminergic cell apoptosis. p55PIK expression levels directly affect PI3K activity and interactive functions with phosphorylated partners, potentially leading to inhibition of cell growth or apoptosis. Thus, p55PIK may be a new candidate target associated with cell growth, survival, proliferation, and apoptosis through PI3K pathways.
In general, the most salient feature of the caspase family is its specificity for substrate cleavage after Asp residues. 22 The optimal target sequence for C3 and C7 is DXXD↑G, where D is Asp, G is Gly, and X is any amino acid. The optimal target sequence for C6 is VXXD↑G, where D is Asp, G is Gly, V is Val, and X is any amino acid. 23 According to this theory, the target sites for caspase cleavage in p55PIK are shown in Figure 6. We found a DXXD recognition sequence for C3 or C7, but not the common DEHD or DEID sequences, where D is Asp, E is Glu, H is His, and I is Ile. Amino acids 30 to 33 at the end of the N24 domain of p55PIK are IEMD, where M is Met, which is generally in accordance with C6-recognizable sequences, except for Met given that the IETD and IEYD motifs, where T is Thr and Y is Tyr, were previously reported as target sequences of C6 by Klaiman et al. 24 To validate the potential cleavage sites recognized by caspases in p55PIK and target sequences, we coincubated human recombinant C3, C6 and C7 with overexpressed full-length p55PIK and Δ1–50 p55PIK in SH-SY5Y and HEK293 cells, respectively. We observed a reduction in full-length p55PIK protein in SH-SY5Y cells after treatment with C6 and a reduction in full-length p55PIK protein and production of a degradation fragment between full-length p55PIK and Δ1–50 p55PIK in HEK293 cells. There are clearly two different degradation mechanisms in the two cell lines. Because the degraded fragment cleaved by C6 was not detected in overexpressed SH-SY5Y cells, we speculate that the specific degradation fragment may be further degraded by proteases in SH-SY5Y cells that are not present in HEK293 cells. Further studies involving protease inhibition will assist in determining p55PIK cleavage by C6 in SH-SY5Y cells. Interestingly, overexpressed Δ1–50 p55PIK in both SH-SY5Y and HEK293 cells was not cleaved by recombinant C6. We suggest that the C6 cleavage site in p55PIK is located within 50 amino acids of the N-terminal, likely in the IEMD sequence. Thus, there is at least one cleavage site for C6 in the p55PIK protein, and decreased p55PIK may relate to C6 activity because p55PIK is a C6 substrate in vitro.

Putative target sites for caspase cleavage in p55PIK. C3 and C7 recognize a consensus DXXD motif, where D is Asp and X is any amino acid. C6 recognizes a consensus VXXD motif, where V is Val. IEMD sequence, where E is Glu, H is His, M is Met, and I is Ile, located on N24 of p55PIK may be a new C6-recognizable motif. C6: caspase 6; C3: caspase 3; C7: caspase 7; p55PIK: phosphatidylinositol 3-kinase 55 kDa gamma; N24: 24 N-terminal amino acids.
The p55PIK regulatory subunit is distributed in a tissue-specific manner, suggesting cell-specific functions. Functional studies have indicated its role with respect to subcellular distribution, PI3K activity, and affinity toward intracellular docking molecules. 25 Previous data revealed that p55PIK is almost entirely localized in the nucleus. As a result, nuclear compartmentalization likely enables p55PIK to modulate nuclear processes such as cell cycle progression. 26 Nuclear compartmentalization also may limit the role of p55PIK in Akt phosphorylation because only a small fraction of total Akt is in the nucleus and only this pool of Akt is potentially activated by p55PIK. 10 Recent research demonstrated that p55PIK colocalizes with IGF1R in bodies and dendrites of Purkinje neurons, but that p55PIK immunoreactivity in astrocytes shows a nuclear pattern, 4 implying subcellular-specific functions. More notably, the 24 N-terminal residues of p55PIK form a new domain that provides interaction sites for other proteins or results in localization of p55PIK to specific cell compartments. Hu et al. reported that p55PIK specifically interacts with Rb protein through the unique NH2 terminus of p55PIK. This interaction is critical for cell proliferation and cell cycle progression. 10 Confocal microscopy and biochemical analysis have demonstrated the presence of p55PIK binding with the Rb protein in the nucleus. 26 Overexpressed N24 blocked cell proliferation and induced cell cycle arrest in cancer cell lines, which specifically inhibited PI3K signaling pathways mediated by p55PIK. In the present study, we overexpressed full-length p55PIK and Δ1–50 p55PIK in both SH-SY5Y and HEK293 cell lines to observe subcellular distribution and location. Although morphological observations revealed that full-length p55PIK was expressed in the nucleus, the majority of positive cells was floating or showed an abnormal morphology with condensed nuclei 48 h after transfection. However, Δ1–50 p55PIK without N24 was also distributed in the cytoplasm after transfection without alterations in the shape of positive cells. These results further confirm that the p55PIK N24 peptide is specifically nucleophilic. Moreover, p55PIK may not be involved in cell cycle progression modulated by signaling pathways in the cell nucleus in the absence of an N24 motif. Regarding unexpected morphological changes and cell floatation after transfection with full-length p55PIK, we speculate that p55PIK overexpression may disrupt the balance dominated by PI3K signaling pathways or directly interact with phosphorylated substrates to trigger other signals because the p55PIK subunit is expressed at low levels in SH-SY5Y dopaminergic neurons. Further studies will focus on overproduction of the p55PIK protein and regulation of cell cycle progression based on Rb-dependent or Rb-independent mechanisms as well as determining direct interactions with other phosphorylated partners.
Excessive activation of C6 is an early marker of neuronal dysfunction and has been implicated in the pathogenesis of Huntington’s disease 27 and Alzheimer’s disease. 28,29 Endogenous caspase-cleaved huntingtin fragments have a markedly different subcellular localization depending on the amino acid position of proteolysis. Active C6 is nuclear. Pro-C6 can be found in the nucleus and does not require translocation for activation. Many C6 substrates, such as lamin A and PARP, are nuclear or predominantly cleaved in the nucleus. 27 Interestingly, C6 is the main caspase effector involved in apoptosis induced by PI3K inhibition, while C3 is not significantly activated. 7 In this study, p55PIK was reduced in PQ-induced SH-SY5Y cell apoptosis, overexpressed p55PIK was primarily localized in the nuclei, and the N-terminal of p55PIK may be cleaved by C6. Thus, the cell nucleus might be an important subcellular site of p55PIK proteolysis by C6 in PQ-induced SH-SY5Y apoptosis. The specific and different subcellular localization of endogenously generated N24 and Δ1–50 p55PIK fragments suggest that these components are trafficked to precise locations after cleavage or that p55PIK cleavage only occurs in specific locations in the cell.
Footnotes
Conflicts of interest
The authors declared no conflicts of interest.
Funding
This work was supported by a grant from the National Natural Science Foundation of China (81172713 to Dr Hao Pang).
