Abstract
Acutely after traumatic spinal cord injury (SCI), the immune system responds with an inflammatory cascade that promotes secondary damage to the spinal cord and systemic inflammation, which promotes persistent medical consequences. Here, we combined clinical and research data to evaluate cellular and molecular changes in the systemic immune system of individuals with SCI (SCI, N = 36) within 0–4 days after injury compared to uninjured individuals (CTL, N = 36). Analyzing blood samples by bulk-RNA Seq, 4752 differentially expressed (DE) gene transcripts were identified in SCI compared with CTLs, including increased expression of genes associated with inflammation and innate immunity (e.g., Neutrophil degranulation, Toll-Like Receptor signaling). Most participants with SCI had complete blood count data available, of whom 36% had elevated white blood cell and neutrophil counts, 24% had elevated monocytes, and 36% had lymphopenia. Significantly reduced expression of canonical natural killer (NK) cell, T cell and dendritic cell (DC) genes were identified, consistent with reduced frequencies of these cell types, determined by flow cytometry. Some molecular changes appeared to be influenced by motor completeness of injury. C-reactive protein, a validated clinical biomarker of inflammation, was significantly elevated after SCI, with levels higher in motor complete compared to motor incomplete injuries. This was also apparent for several other proinflammatory cytokines (e.g., High Mobility Group Box 1 protein, IL-6, IL-8). These data confirm and extend prior observations of cellular and molecular immunological changes, that may serve as potential biomarkers of injury severity, or as future therapeutic targets to improve health.
Introduction
There are an estimated 18,000 new traumatic spinal cord injuries (SCI) in the United Sates annually. 1 Individuals with acute traumatic SCI are at increased risk for infectious complications including respiratory infections and pneumonia. 2 There are several possible mechanisms contributing to increased infection risk, including persistent circulating inflammation that is present in most individuals with SCI, promoting an “inflammaging” phenotype. 3 Circulating inflammation may also reduce efficacy of neurorehabilitation and contribute to elevated risk of cardiometabolic syndrome that is observed in individuals with SCI.4–7 Due to their potential mechanistic relevance and clinical accessibility, immune mediators in the blood or spinal fluid (CSF) have also been proposed to serve as surrogate biomarkers of injury severity and of prognosis for neurological recovery.8–10
To characterize the systemic immune system changes after traumatic SCI, we conducted a prospective study to measure circulating immune phenotypes and neurological data from individuals recruited within the first three days after SCI and then at 3–6- and 12-months post injury. We previously reported immunological phenotypes from the first 12 participants who had blood samples collected at a minimum of 3 study visits. 11 Compared to uninjured controls (CTL), we demonstrated cellular and molecular changes that were associated with time from injury and neurological injury severity.
Here, we describe immunological phenotypes from all study participants with acute SCI (N = 36 total) using clinical data, bulk RNA-Seq, multicolor flow cytometry, and enzyme-linked immunosorbent assays (ELISAs). First, differences were elucidated in circulating gene expression after SCI compared with CTL. Second, changes in circulating immune cell subsets after SCI were characterized, using both clinical and research assays. Next, molecular changes were determined according to neurological injury severity (motor complete or incomplete). Lastly, levels of C-reactive protein (CRP) and other inflammatory proteins were compared between SCI and CTL and differences by neurological injury severity were determined.
Materials and Methods
Participants
This study was listed on ClinicalTrials.gov (#NCT02731027) as described. 11 Prior to enrollment, informed consent was obtained. Participants were recruited at Level One Trauma Centers affiliated with Northwell Health, The Kessler Institute for Rehabilitation, Thomas Jefferson University, University of Louisville, the Ohio State University Medical Center, and the University of British Columbia (2016–2020, prior to the COVID-19 pandemic). SCI was determined by clinical and radiographical presentation. Neurological level and severity of injury were determined by the International Standards for Neurologic Classification of Spinal Cord Injury (ISNCSCI). 12 Inclusion criteria for enrollment of individuals with SCI was: (1) ≥ 18 years of age, (2) traumatic SCI 0–3 days post-injury (dpi), (3) American Spinal Injury Association Impairment Scale (AIS) grade A–D, and (4) neurological injury level between cervical 4 and thoracic 10 (C4–T10). Potential participants were excluded if there was documented medical history of cancer, autoimmune disease, known previous SCI, pre-existing neurological disease (including any condition that would affect the ability to participate in the study or present an obvious confounder with the trajectory of functional recovery after SCI, including multiple sclerosis, Parkinson’s Disease, stroke, or other significant neurological condition). The Glasgow Coma Scale (GCS) ranges from 0 to 15. Scores were recorded for research for all (N = 36) participants and classified as severe (3–8), moderate (9–12), or mild (13–15). The trauma Injury Severity Scale (ISS) ranges from 0 to 75 and was recorded for research for most (N = 31/36) participants. The ISS scores were classified as very severe (≥26), severe (16–25), moderate (9–15), or mild (0–8). Inclusion criteria for the control group (CTL, N = 36) which was recruited at Northwell Health was: (1) ≥18 years of age, (2) without current or history of cancer, (3) without a history of SCI, autoimmune or neurodegenerative diseases.
This study aimed to collect blood samples and clinical data from patients with SCI during acute hospitalization, as well as at 3-, 6- and 12-months post SCI. 11 Previously, data from the 12 participants with SCI who had blood samples collected at a minimum of three follow up study visits, including the acute visit, was presented. 11 Here, we analyzed their acute data only with samples obtained during acute hospitalization (0–4 dpi) from an additional 24 participants with SCI (N = 36 total) who were enrolled in the study but who did not complete three follow up visits. Due to logistical challenges and medical considerations, not all blood samples from participants were available for all analyses. Data available from individuals with SCI were compared with data available from CTL who had similar demographic distributions (age and sex). The number of samples available for each immune analysis was: bulk RNA-sequencing (SCI N = 36, CTL N = 9), flow cytometry (SCI N = 14, CTL N = 14), and ELISAs (SCI N = 35, CTL N = 16). The number of samples collected at each dpi for RNA-Seq was: 0 dpi (N = 2), 1 dpi (N = 8), 2 dpi (N = 13), 3 dpi (N = 11), and 4 dpi (N = 2). For each type of analysis, there was no statistically significant difference in age and sex distribution between groups.
RNA-sequencing
Bulk RNA-Sequencing was performed as described previously. 11 Briefly, RNA was isolated from blood, libraries were prepared (Illumina TruSeq Stranded Total RNA with RiboZero Globin, Catalog #20020612) and 100 bp paired-end reads were collected on the Illumina NextSeq 500 as in 11 or NovaSeq 6000 platforms (Northwell Health). As described, trimmed reads were aligned using STAR to the human genome (hg38) assembly, filtered for expression >50, normalized using the Trimmed Mean of M-values method using the edgeR package embedded in Partek Genomics Flow software (St. Louis, MO, USA), and log2 transformed. 11 DE transcripts were determined by ANOVA with a fold change greater than 2.0, using the Benjamini–Hochberg false discovery rate (FDR) <0.05. For functional analysis of DE genes, if multiple transcripts for the same gene symbol were DE, then the transcript with the most significant p value for that gene was analyzed. Venn diagrams were created in Venny to determine genes that were unique to or shared across comparisons. 13 Functional analysis of DE genes was performed using the Reactome Pathway Database and the Human Gene Atlas within the Enricher platform. 14
CBC and flow cytometry
The earliest clinical complete blood count (CBC) data available for each participant was abstracted from medical records from participants with SCI (N = 33/36). CBC data was collected from participants at 0 dpi (N = 26), 1 dpi (N = 4), 2 dpi (N = 2), and 3 dpi (N = 1). Standard clinical reference range values used for differential white blood cells (WBC) were used (units: K/µL): WBC (4.8–10.8), neutrophils (1.8–8.0), lymphocytes (1.0–4.8), monocytes (0–0.8), and eosinophils (0–0.5). From fresh blood draws for research analyses, peripheral blood mononuclear cells (PBMCs) were purified by Ficoll-Hypaque, labeled with fluorescent multicolor antibody panels, and fixed (4% PFA) as described. 11 One to two million cells were stained per antibody cocktail, ∼150,000 total events were collected on a BD LSRII flow cytometer and analyzed using FlowJo software (FlowJo, LLC) with gating strategies described. 11 Briefly, cells were gated as leukocytes using forward scatter area (FSC-A) versus side scatter area (SSC-A). FSC-A and height (FSC-H) were then used to define singlets, which were further gated as major immune cell subtypes. 11
Enzyme-linked immunosorbent assays for immune factor profiling
Cytokines, chemokines, and growth factors were measured using commercially available single analyte ELISAs and multiplex bead-based assays, as described. 11 ELISAs were used according to standard protocols to measure CRP (IBL International #EU59131) and High Mobility Group Box 1 protein (HMGB1, IBL International # ST51011). Data were analyzed using GraphPad Prism 10. A multiplex assay was used to measure an additional 48 systemic immune mediators on a Bio-Plex 200 system and analyzed with Bio-Plex software (Bio-Plex Pro Human Cytokine Screening Panel, 48-plex, #12007283). Assay ranges and kit controls were within expected concentration ranges, average % CV across samples and analytes was 5.4%, with an average recovery of 100% and a range of 70–130%. For statistical analysis, measurements below detection limits were assigned a value of half the lowest limit of quantification (LLOQ). Analytes (N = 37/50) that were detectable in >50% of samples were compared between participants with SCI and CTLs, as well as between participants within the SCI group by motor completeness of injury (AIS A/B vs. AIS C/D).
Statistical methods
DE transcripts were defined as transcripts having a fold change ≥2.0, and an ANOVA Benjamini–Hochberg FDR of <0.05, using Partek Genomics Flow software. Among the DE genes identified by ANOVA, selected genes of interest that were statistically significant are presented as box plots for illustration and emphasis (Figs. 1C, 3E). Relative gene expression levels are presented (Figs. 1C, 3E), where boxes indicate the range from Q1 to Q3, lines represent the median, and “whiskers” extend from the 10th–90th percentile. For pairwise comparison of cell types between SCI and CTL, Mann–Whitney U tests were conducted to determine significance with a threshold set at p < 0.05, The results are presented in Figure 2 with significance levels indicated as follows: *p = 0.01–0.05, **p = 0.001–0.01, ***p = 0.0001–0.001. For comparison of CRP levels between SCI vs. CTL, a Kruskal Wallis test was performed with significance also defined as p < 0.05, as shown in Figure 4. Dunn’s multiple comparisons test was then carried out to identify which groups differ from one another, using similar p value definitions as shown in Figure 2. Prism GraphPad version 10 was used to perform statistical tests.
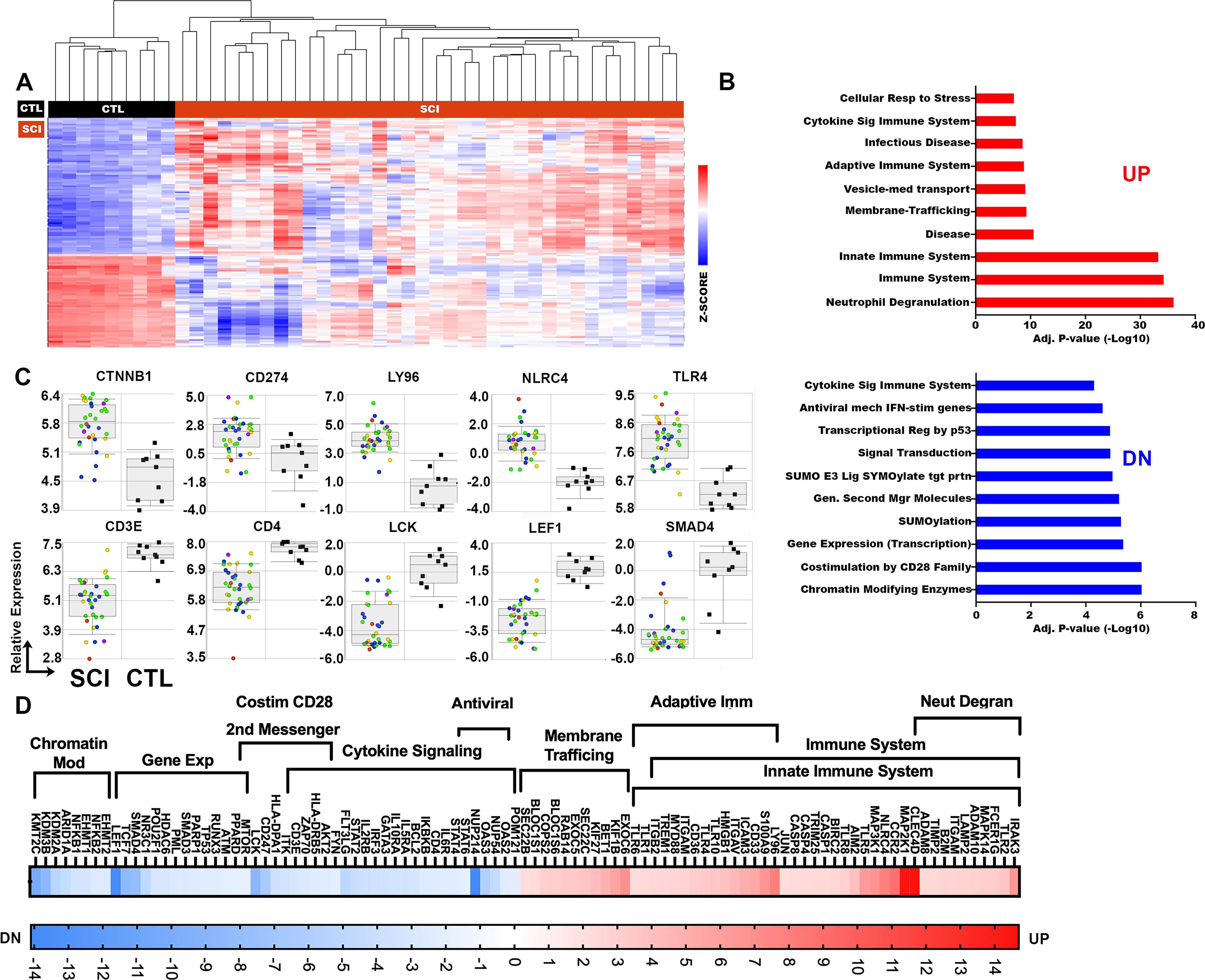
Profiling of whole blood gene expression acutely after spinal cord injury (SCI).
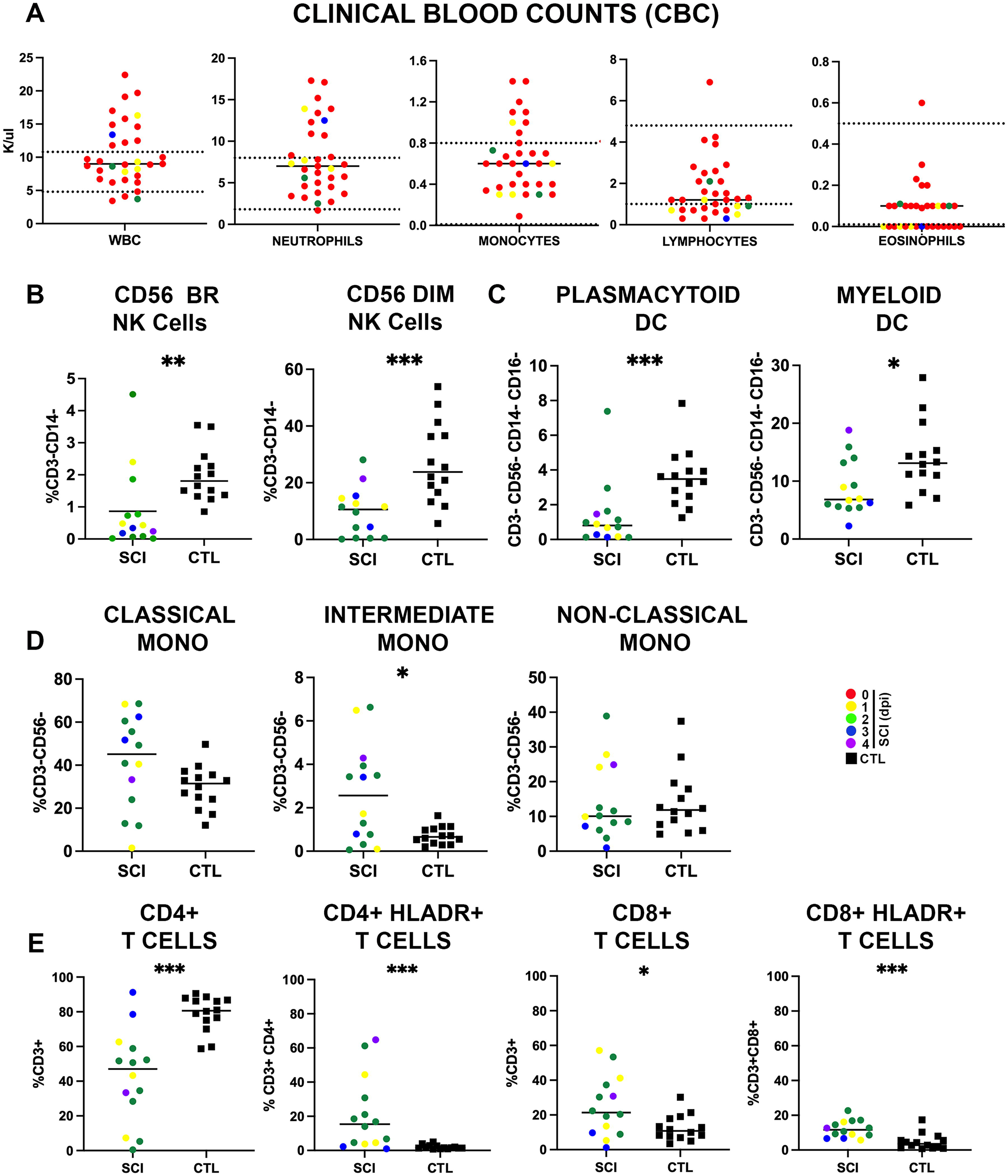
Phenotyping of immune cell subsets acutely after SCI.
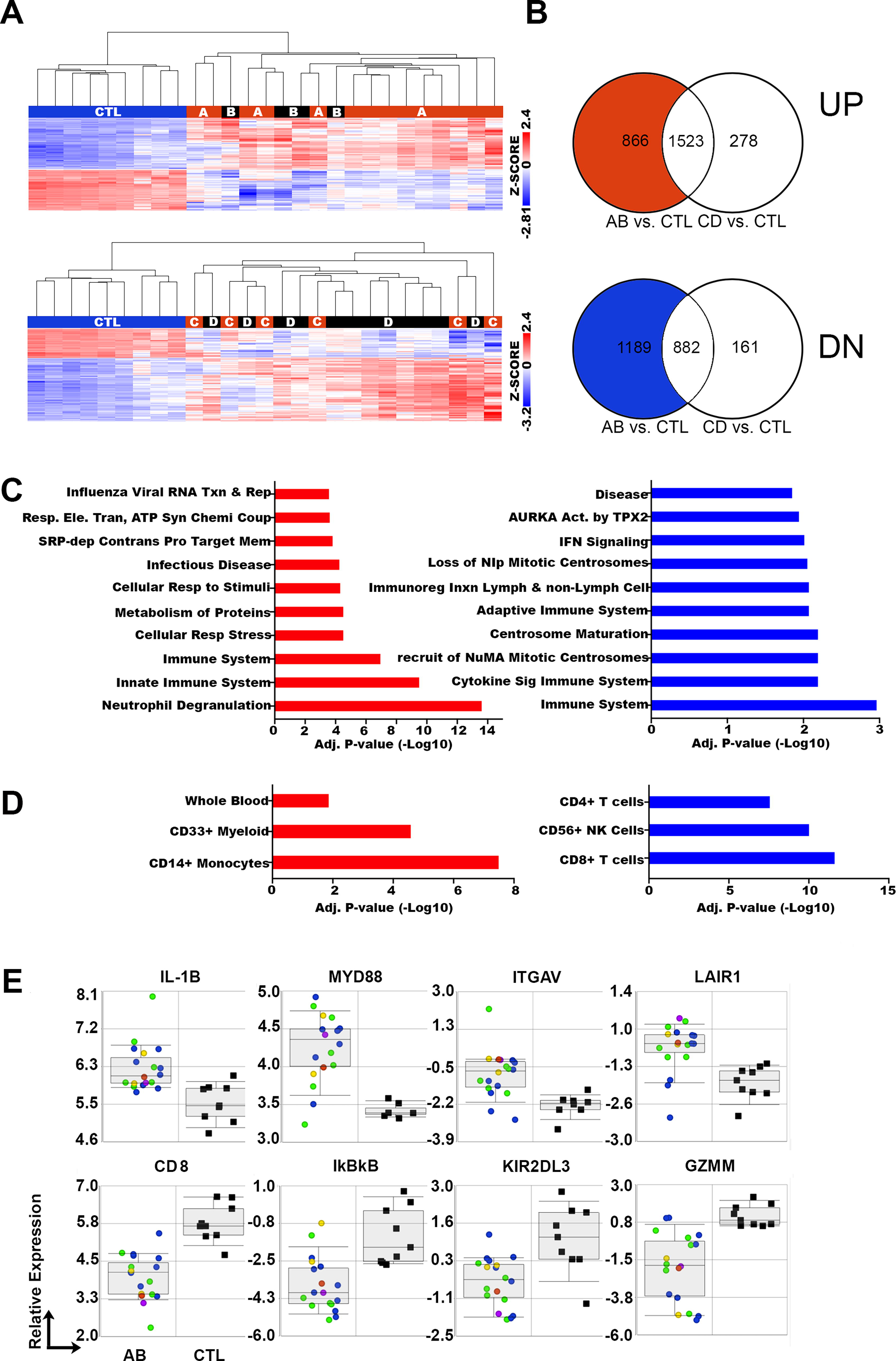
Pairwise comparison of gene expression profiles by injury severity.
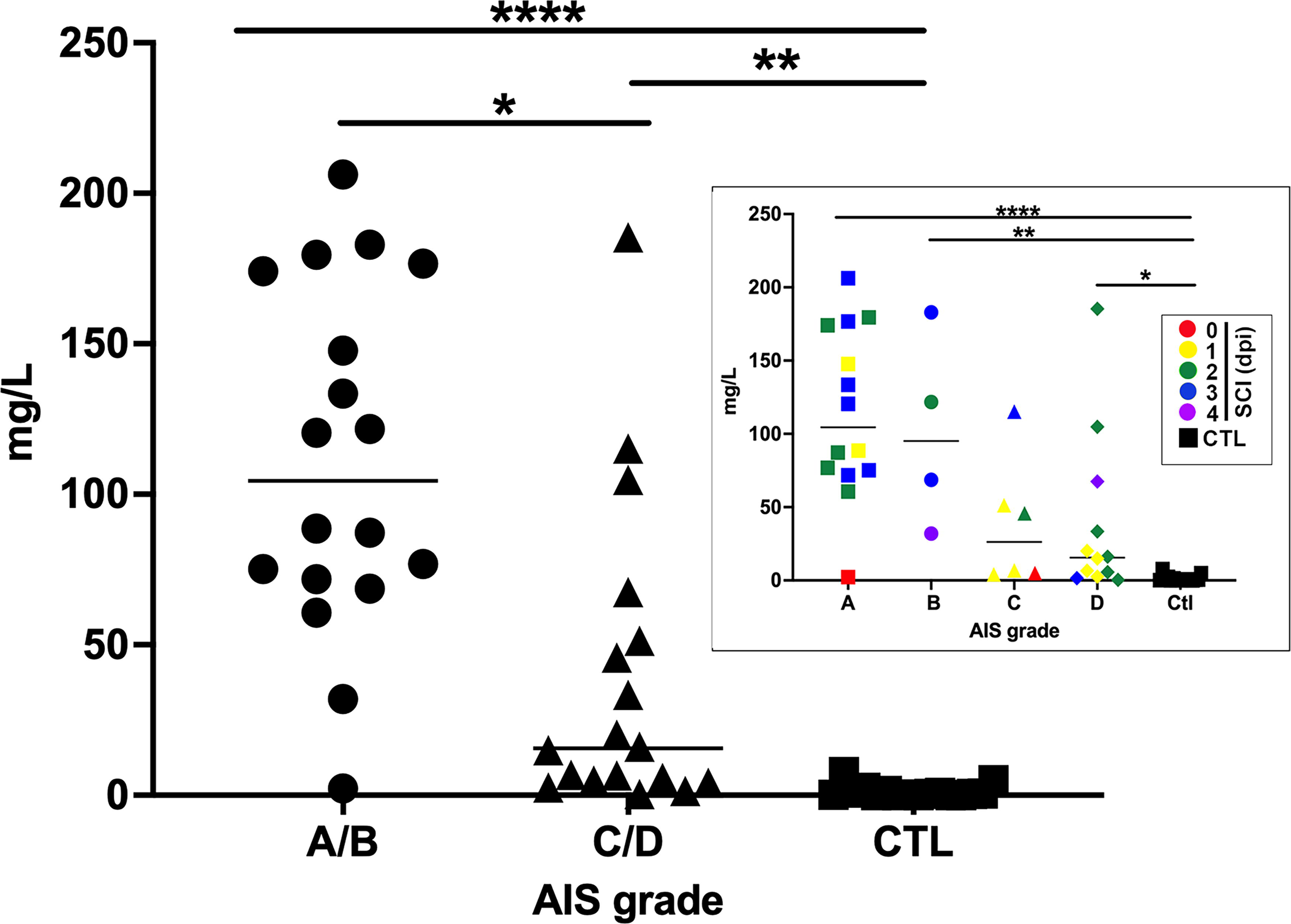
Injury status and severity correlate with C-reactive protein (CRP) levels. CRP is graphed according to injury severity (AIS A/B vs. C/D) compared to CTL. Line indicates median. Symbols represent individual participants. Circles = AIS A/B, Triangles = AIS C/D. Inset shows AIS individual grades: Symbols are color coded for days post SCI: zero (red), one (yellow), two (green), three (blue), four (purple). CTL samples are shown in black. Squares = AIS A Circles = AIS B, triangles = AIS C, diamonds = AIS D, black squares = CTL. Kruskal Wallis test was performed to compare SCI vs. CTL to determine significance defined as p < 0.05. Dunn’s multiple comparisons test done to compare each group. p Values for significant comparisons are shown: *p = 0.01–0.05, **p = 0.001–0.01, ***p = 0.0001–0.001, ****p < 0.0001.
Results
Participant characteristics
A summary of demographic and clinical characteristics of participants with SCI are shown in Table 1. SCI participants ranged in age from 21 to 85 years old and were mostly (75%) male. Uninjured controls were of similar age and sex distribution (see Methods). The most common neurological level of injury (NLI) was cervical (75%), followed by thoracic (25%), with most (92%) participants injured rostral to T6, the level of sympathetic outflow to immune organs. 15 Trauma injury severities were characterized according to the ISS, which ranges from 0 to 75, where scores were stratified as very severe (≥26), severe (16–25), moderate (9–15), or mild (0–8). As expected for individuals treated at Level One Trauma Centers, most (90%) participants had ISS scores in the severe or very severe range (Table 1). Neurological injury severities were characterized according to AIS grade: A (39%), B (11%), C (17%), and D (33%). Mechanisms of injury were: fall (61%), motor vehicle accident (19%), sports (11%), violence (3%), or other (6%). TBI was categorized by the GCS, where scores were stratified as severe (3–8), moderate (9–12), or mild (13–15, with 15 indicating neurologically intact). 16 Most participants in this study had GCS scores of mild (Table 1).
Summary of Demographics and Clinical Characteristics of Participants with


Changes in systemic immune gene expression acutely after SCI
There were 4752 DE genes between SCI and CTL groups defined as transcripts having a fold change ≥2.0, and an ANOVA Benjamini–Hochberg FDR of <0.05 (Supplementary Table S1). Hierarchical clustering of DE genes demonstrated distinct transcriptional profiles after SCI compared to CTL (Fig. 1A). Functional categories enriched among DE genes were determined using the Reactome Pathway Database (Fig. 1B). DE genes upregulated acutely after SCI were enriched significantly in categories related to neutrophil function, innate and adaptive immunity, membrane trafficking, and infection responses, among others. DE genes downregulated acutely after SCI were enriched significantly in categories related to T cell biology such as co-stimulation by CD28, signaling cascades like SUMOylation that regulate innate immunity, as well as antiviral and transcriptional responses. Relative expression of genes of interest is shown from the Reactome Database pathways that were up- or downregulated, such as TLR4 and its associated cell surface protein LY96, which are critical for innate immune responses (Fig. 1C). Relative expression of downregulated genes of interest is also shown, which are related to T cell signaling include CD3, CD4, LCK, LEF1, and SMAD4 (Fig. 1C). Relative expression patterns of genes from enriched Reactome Database pathways are shown, including chromatin modifications, cytokine signaling, membrane trafficking, and neutrophil degranulation (Fig. 1D).
Changes in systemic immune cell populations after SCI
Since CBC data are often the earliest immune outcome available after SCI, they were abstracted as available from participant medical records (N = 33) (Fig. 2A). All but one CBC record was obtained prior to the RNA-Seq sample from the same participant. Compared to standard reference values, WBCs were elevated above the reference range in 12 participants and reduced below the reference range in three participants. Neutrophils were elevated above the reference range in 12 participants and reduced below the reference range in one participant. Monocytes were elevated above the reference range in eight participants, seven of which also had elevated neutrophils. Lymphocytes were elevated above the reference range in one participant and reduced below the reference range in 12 participants, 6 of which also had elevated neutrophils. Eosinophils were elevated above the reference range in one participant. Levels for each cell type tended to be higher in AIS A/B vs. C/D injuries, but none were statistically significant (data not shown).
We next performed a deeper phenotypic analysis of circulating immune cell subsets using multicolor flow cytometry. A significant decrease in both the cytokine-producing CD56bright and the cytolytic CD56dim natural killer (NK) cell subsets after SCI compared with CTL was detected (Fig. 2B). Consistent with this observation from our pilot study, 11 several canonical NK cell genes were significantly downregulated after SCI compared to CTLs including granzyme M, perforin 1, CD7 and KIR genes (Supplementary Table S1).
Dendritic cells (DCs) provide a critical link between innate and adaptive immunity. Myeloid DCs (mDCs) are potent antigen presenting cells and plasmacytoid DCs (pDCs) are robust producers of IFN-alpha, particularly in response to viral infections. Both mDCs and pDCs were significantly reduced after SCI compared with CTL (Fig. 2C). Consistent with these observations, several DE genes that are typically associated with DCs were reduced significantly after SCI, including FLT3 and FLT3LG, which are critical for DC development and function (Supplementary Table S1).17,18 In contrast, the transcription factor IRF8, which regulates mDC and promotes DC development, 19 was significantly increased (Supplementary Table S1). We also measured relative proportions of monocyte subsets, which are crucial for the innate immune response. We observed a trend of increased classical monocytes and a significant increase in intermediate monocytes after SCI compared to CTL (Fig. 2D). As in the initial cohort of 12 patients, 11 median fluorescence intensity of surface HLA-DR, a marker of activation, was significantly decreased on all monocyte subsets after SCI compared with CTL (classical monocytes: 937 ± 64 vs. 1689 ± 150 mean±SEM, p < 0.0001; intermediate monocytes: 4340 ± 531 vs. 9437 ± 790 mean±SEM p < 0.0001; and non-classical monocytes: 1697 ± 170 vs. 2925 ± 267 mean±SEM, p < 0.0005 acute SCI vs. CTL respectively, Mann–Whitney U Test). Compared with CTL, there was an increased expression of many DE genes associated with (but not exclusive to) monocyte activation 20 acutely after SCI. These included Arachidonate 5-Lipoxygenase (ALOX5), an enzyme found in monocytes that catalyzes the production of pro-inflammatory leukotrienes, 21 C-C Motif Chemokine Receptor 2 (CCR2), a chemokine receptor that plays a role in monocyte recruitment to sites of inflammation 22 and Myeloid Cell Nuclear Differentiation Antigen (MNDA), a myeloid cell nuclear protein which is associated with chronic inflammatory diseases.23–25
CD4+ and CD8+ T cell subsets were changed after SCI
Lymphopenia has been observed in individuals with acute SCI,26,27 and CBC counts indicated that lymphocytes were reduced in a third of participants with SCI (Fig. 2A). Flow cytometry also showed a reduced percentage of CD4+ T cells after SCI, of which a greater proportion had elevated surface levels of the activation marker HLA-DR (Fig. 2E). In contrast, CD8+ T cells were increased after SCI, and displayed elevated surface levels of HLA-DR (Fig. 2E). We observed decreased expression of DE genes related to CD4+ T cells after SCI compared to CTLs, including CD3, CD4, the transcription factor Lymphoid Enhancer Binding Factor 1 (LEF1), and the tyrosine kinase Lymphocyte Cell-Specific Protein-Tyrosine Kinase (LCK) (Fig. 1C). Conversely, we observed increased expression of other DE genes including proinflammatory IL-4, a cytokine produced by activated T cells, as well as the G-protein coupled receptor that promotes T cell apoptosis, GPR65, and the cell surface adhesion molecule L-selectin (SELL). 28
Correlation of injury severity and immunological phenotypes
To identify potential changes in gene expression acutely after SCI that correlated with injury severity, samples from individuals with motor complete (AIS A/B: N = 18) or incomplete (AIS C/D, N = 18) injuries were compared in a pairwise manner to CTL (ANOVA, FDR <0.05, ≥2.0-fold change, Fig. 3, Supplementary Table S1). Compared to CTLs, there were 2389 up- and 2071 downregulated DE genes in the AIS A/B samples (Fig. 3A–B). Of these, there were 1523 up- and 882 downregulated DE genes shared in samples from motor incomplete injuries (Fig. 3B), while 866 up- and 1189 downregulated genes were unique to samples from motor complete. Reactome Database pathway analysis of DE genes upregulated uniquely in motor complete injuries were enriched in pathways related to neutrophil function, innate immune system function, stress responses, infection responses, and other pathways (Fig. 3C). Reactome Database pathway analysis of DE genes downregulated uniquely in motor complete were enriched in pathways related to immune system function, cytokine signaling, adaptive immunity, interferon signaling, and others (Fig. 3C). Human Gene Atlas analysis of DE genes upregulated uniquely in motor complete injuries were enriched in genes related to monocyte and myeloid cell types (Fig. 3D). In contrast, DE genes downregulated uniquely in motor complete injuries were enriched in genes related to CD4+ and CD8+ T cells, as well as NK cells (Fig. 3D). Selected up- and downregulated genes from Reactome Pathways “Neutrophil Degranulation,” “Innate Immune system,” “Immune System,” and “Cytokine signaling” are shown in Figure 3E.
Individuals with AIS A injuries have worse physical and psychosocial health outcomes than individuals with AIS B injuries. 29 Although there were few participants with acute AIS B injuries enrolled in this study, we performed an exploratory pairwise comparison of differences in gene expression in individuals with AIS A (N = 14) or AIS B (N = 4) injuries (Supplementary Fig. S1). Compared to CTL, most DE genes were common between AIS A and B injuries (Supplementary Fig. S1B). However, Reactome Database pathway analysis of DE genes upregulated uniquely in AIS A injuries included categories related to protein catabolism, adaptive immunity, neutrophil function and others (Supplementary Fig. S1C). The only significantly decreased category in AIS A injuries was interferon signaling (Supplementary Fig. S1C).
In addition to changes in gene expression, we also analyzed the presence and levels of inflammatory proteins in samples from participants with SCI compared to CTL. CRP, a validated clinical biomarker of inflammation, was elevated in participants with SCI compared to CTL, in a manner reflecting neurological injury severity (Kruskall–Wallis was performed to determine significance, defined as p < 0.05, followed by Dunn’s multiple comparisons test to compare groups with one another, with significance shown in Fig. 4, Table 2). We identified an additional 15 inflammatory mediators that were significantly different after SCI from CTL (Table 2), of which the majority were elevated after SCI. For example, both HGF and HMGB1 were elevated after SCI compared to CTL, which was also observed at the transcriptional level. IL-6 and IL-8 were also elevated after SCI, in accordance with both our own and other independent previous studies.9,11,30 We also identified nine factors that were significantly different between samples from motor complete (AIS A/B) and incomplete injuries (AIS C/D), of which eight were elevated in participants with motor complete injuries. Among these, only the HGF and IL-10 were also upregulated at the transcriptional level in participants with motor complete injuries.
Circulating Inflammatory Proteins That Significantly Differ by Injury Status (A) or Severity (B)
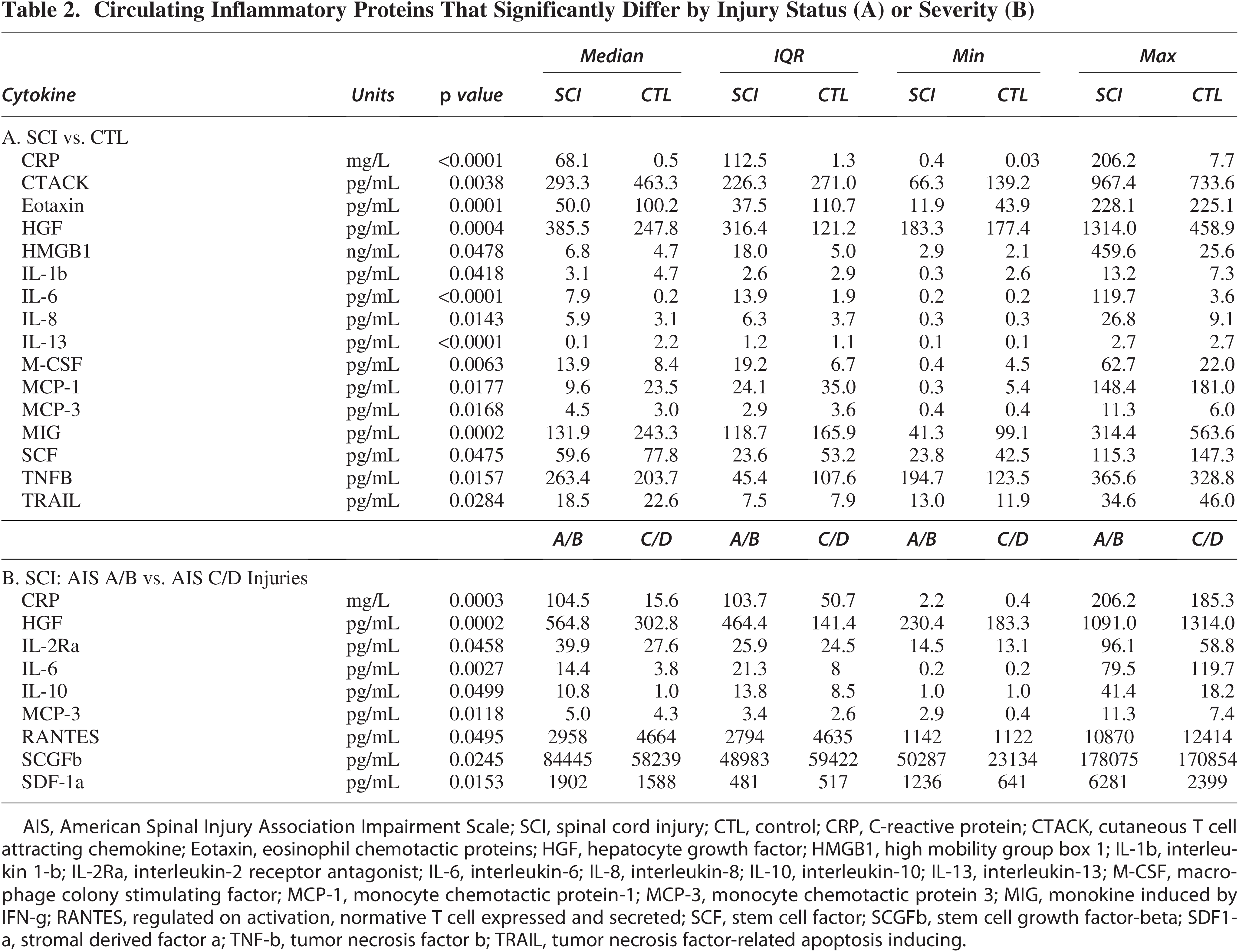
AIS, American Spinal Injury Association Impairment Scale; SCI, spinal cord injury; CTL, control; CRP, C-reactive protein; CTACK, cutaneous T cell attracting chemokine; Eotaxin, eosinophil chemotactic proteins; HGF, hepatocyte growth factor; HMGB1, high mobility group box 1; IL-1b, interleukin 1-b; IL-2Ra, interleukin-2 receptor antagonist; IL-6, interleukin-6; IL-8, interleukin-8; IL-10, interleukin-10; IL-13, interleukin-13; M-CSF, macrophage colony stimulating factor; MCP-1, monocyte chemotactic protein-1; MCP-3, monocyte chemotactic protein 3; MIG, monokine induced by IFN-g; RANTES, regulated on activation, normative T cell expressed and secreted; SCF, stem cell factor; SCGFb, stem cell growth factor-beta; SDF1-a, stromal derived factor a; TNF-b, tumor necrosis factor b; TRAIL, tumor necrosis factor-related apoptosis inducing.
Discussion
Pneumonia and septicemia are leading causes of death acutely after SCI, and people with SCI are more than 80 times more likely to die of sepsis than uninjured persons.31,32 Infections are leading causes of rehospitalization for individuals within the first year after SCI.1,33 In a database study of individuals with acute motor complete cervical SCI, the presence of pneumonia or post-operative wound infections was identified as an independent risk factor for poor neurological recovery in the first year after SCI. 34 Increased risk for infections after SCI is partially due to respiratory muscle paralysis reducing the ability to clear secretions, resulting in accumulation of fluid and microbial pathogens in the lung. 2 However, additional mechanisms have been proposed to contribute to increased infection risk after SCI, including depression of the immune system itself that may result from reduced descending supraspinal control of the autonomic nervous system.27,35
To investigate immune responses in individuals with acute SCI, this study analyzed a comprehensive array of clinical and research assays to elucidate cellular and molecular changes. We first examined the effect of injury status on systemic immune gene expression, which confirmed and extended previous studies showing distinct gene expression profiles in acute SCI.9,11,36–41 As in our pilot report and in independent studies, genes related to neutrophil function, inflammation, and innate immunity were upregulated acutely after SCI8,11,37,40,41 This included genes related to TLR signaling, which are pattern recognition receptors (PRR) that recognize intracellular and extracellular pathogens that were upregulated in our pilot study and in other studies of individuals with acute or chronic SCI.11,42,43 In contrast, genes related to adaptive immunity, such as those involved in T cell function, antiviral immunity, and other cell signaling pathways such as chromatin modifications and gene regulation, were downregulated acutely after SCI. We also detected elevated levels of inflammatory proteins after SCI. These included CRP, which is a marker of risk for cardiovascular disease, as well as IL-6, IL-8, and the endogenous TLR4 ligand HMGB1, which is neurotoxic in animal models of SCI. 44
Changes in circulating white blood cells have been shown to reflect neurological injury severity and proposed to be prognostic of neurological recovery after SCI. They are also relevant to infection susceptibility and systemic inflammation.9,26,45,46 Acute neutrophilia and lymphopenia have been associated with acute neurological injury severity and functional recovery, and higher neutrophil to lymphocyte ratios are an independent prognostic factor for 6-month outcomes.9,26,45,47 Using clinical data, more than a third of study participants had elevated white blood cells, neutrophilia, and lymphopenia, while approximately a third of participants had elevated monocytes. Perhaps due to differences in number or characteristics of participants, we did not observe significant correlations of CBC measures with neurological injury severity in this study. Using flow cytometry, we did observe decreased NK cells, CD4+T cells, pDCs and mDCs, confirming and extending data from our pilot study and independent studies of acute SCI.11,26,27,35,45,48 We also identified elevated monocyte subsets by flow cytometry, which are not included in standard CBC measurements.11,27,35,49
As we showed previously, 11 neurological injury severity was reflected in systemic gene expression in this larger acute SCI cohort. Pairwise analysis of AIS A/B vs. CTL or AIS C/D vs. CTL showed more than 800 genes distinctly upregulated and more than 1000 genes uniquely downregulated in samples from AIS A/B vs. CTL. DE genes uniquely upregulated after AIS A/B injuries included pathways related to neutrophil function, responses to infectious diseases and others. As we showed previously, there was an enrichment of genes upregulated after AIS A/B injuries related to TLR signaling such as the TLR adaptor MYD88 and the inflammatory cytokine IL-1b.11,42,43,50,51 DE genes that were uniquely downregulated after AIS A/B injuries were related to immune system function, cytokine signaling, centrosome modifications, and interferon signaling. These included genes related to T cell function such as CD8, and NK cell genes KIR2DL3 and granzyme.
Some elevated inflammatory proteins also reflected neurological injury severity, including CRP, which was more than sixfold higher in samples from AIS A/B vs. C/D injuries. As CRP is routinely measured in clinical labs, it may thus be of particular interest to explore this as a potential biomarker of acute injury severity, as has also been suggested for sepsis. 52 In addition, we confirmed elevated IL-6 in AIS A/B vs. C/D injuries,9,11,53 which may justify future exploration of FDA-approved biologics such as Tocilizumab, Sarilumab, Siltuximab, and Satralizumab in preclinical models and eventually, in the acute SCI population. However, recent data demonstrate a beneficial role for IL-6 in the brain in preclinical models of TBI and SCI, suggesting that additional animal studies may provide deeper insights in the context of different CNS injury paradigms.54,55
There are broad estimates of concomitant acute SCI and TBI. 56 Studies of patients with acute TBI have also demonstrated the presence of specific inflammatory biomarkers, and blood-based biomarkers are being proposed in the newest NIH framework for TBI classification.16,57 The majority of participants in this study included individuals with SCI who were recruited within the first three days of traumatic injury and who were diagnosed with SCI by the ISNCSCI exam, which requires a patient to be awake and alert in order to follow the instructions to complete the exam. Thus, patients with severe TBI may have been excluded if they would have been unable to complete a reliable ISNCSCI exam within the enrollment timeframe of 0–3 dpi. This is consistent with the high GCS scores for most participants (Table 1), indicating that they were neurologically intact. Early SCI diagnosis is a major challenge for the field and this challenge underlies the search for objective blood-based biomarkers of injury status and severity.
There were several limitations of this study. Since half of the injuries were incomplete tetraplegia, and most participants (N = 33/36) sustained injuries at or rostral to T6, we were unable to examine correlations between immune outcomes and all clinical characteristics of interest, such as injury level, which has been suggested to contribute to immune system changes after SCI via damage to the SNS innervation of immune organs. This hypothesis predicts that individuals with injuries at or rostral to T6, where immune organs are innervated, would have the greatest immune changes. 15 In 2023, the SCI Model System Annual Report described that most injuries were incomplete tetraplegia and 68.7% of injuries occurred at or rostral to T6. Injury level may also reflect other aspects of SCI, such as mechanism or velocity of injury, which could impact inflammation regardless of the sympathetic system. In our pilot report, we also highlighted limitations such as limits to interpreting results of bulk-RNA seq, which are derived from a mixture of circulating white blood cells. This contrasts with single-cell RNA-sequencing, which is more technically challenging to pursue in a multi-center acute trauma study, but facilitates an understanding of gene expression changes within specific cell types. While we can hypothesize that a reduction of expression of specific genes such as NK cell genes derives from the reduced percentage of circulating NK cells observed after SCI, we cannot draw direct conclusions, as some genes typically expressed by NK cells may also be expressed by other immune cell types and bulk RNA-Seq cannot distinguish between these possibilities. However, in agreement with this study, a recent study of single-cell RNA-Seq in humanized mice also showed decreased T cell and antigen processing and presentation. 58 Also, we did not compare data here to that obtained from individuals with non-neurological trauma, where additional immune outcomes may be informative.48,49
Conclusions
Despite the above limitations, this study provides novel data that confirms and extends our knowledge of immunological changes in individuals with acute SCI. Using multiple immunological outcomes, including clinical lab data, transcriptional analysis, flow cytometry, and proteomic measures, we define distinct aspects of the immune system that are impacted acutely after SCI. This study again supports a neurogenic molecular signature of acute SCI, including some changes that are clinically feasible to measure in the acute clinical setting, such as CRP. Other changes observed support a rationale for testing cytokine-directed therapies in acute SCI that are approved for use in other clinical populations.
As hope for clinical trials in acute SCI expands, different types of biomarkers are being evaluated for their potential to reflect injury status and neurological injury severity, help identify individuals most appropriate for specific therapeutic strategies, stratify enrollment in clinical trials, evaluate success of treatments such as surgical decompression, and predict long-term neurological recovery. These include neurochemical factors such as gene expression and protein levels in blood or CSF,8,11,41 clinical imaging parameters such as MRI,59–61 and white blood cell responses.26,45 Inflammatory (IL-6, IL-8, MCP-1) and neuronal injury factors (tau, S100B, GFAP, UCH-L1) have also been identified in CSF that may predict recovery.8,10,62 As an increasing amount of data is collected from individuals with acute SCI, we may envision development of a composite score reflecting several biomarkers, including immune outcomes, providing a deeper physiological phenotype of an individual with acute SCI.
Transparency, Rigor, and Reproducibility
This observational study was registered on Clinicaltrials.gov (NCT02731027) and conducted as described. 11 The study did not reach the desired sample size of 100 to yield 80% power to detect a correlation of 0.28 or greater (2-tailed alpha = 0.05, with no adjustment for multiple testing) using a simplified power calculation for estimating Spearman correlations between AIS grade and a given inflammatory biomarker level. There were 36 participants with blood samples from the acute visit that passed quality control for RNA-sequencing. Participants were not informed of or able to guess fluid biomarker levels. Participant IDs were de-identified for analyses and personnel who were blinded to participant characteristics handled samples and made quality control decisions. Equipment and analytical resources were from commercial suppliers (see Methods). Statistical analyses were performed in Prism GraphPad, SAS, and Partek Genomics Flow, using algorithms and methods pre-specified by the commercial software. The inclusion and exclusion criteria are based on established standards in the field. Biological samples obtained by the investigators are being used for additional analyses; there are limited remaining quantities of some samples. We are not conducting replication studies. Requests for flow cytometry files will be considered in accordance with regulatory guidelines on an ad hoc basis. De-identified RNA-Seq data from this study are available in the GEO repository, accession #GSE293559. This article will be published under a Creative Commons Open Access license, enabling it to be freely available upon publication at https://TBD.
For ELISAs, samples were assayed in duplicate and performed with standards according to manufactures’ protocols. For singleplex ELISAs, R2 values and average %CVs are as follows: CRP: R2=1.0, %CV = 7.5; HMGB1: R2=0.998, %CV= 4.1. On the multiplex panel, 37/48 analytes were detectable in >50% of the samples and further analyzed. The average %CV across all samples and the lower limits of quantification (LLOQ, pg/mL) for each of these analytes is as follows: CTACK (4.32, 2.27), Eotaxin (4.0, 0.19), FGF basic (4.53, 3.72), G-CSF (6.68, 9.19), GROa (4.21, 103.61), HGF (5.52, 13.86), IFN-g (5.63, 2.16), IL-10 (5.71, 2.06), IL-13 (5.2, 0.25), IL-16 (3.98, 2.19), IL-18 (3.91, 0.716), IL-1a (6.07, 4.47), IL-1b (6.32, 0.56), IL-1ra (6.84, 8.23), IL-2Ra (6.34, 0.93), IL-4 (5.52, 0.22), IL-6 (6.6, 0.42), IL-7 (5.37, 3.73), IL-8 (7.73, 0.66), IL-9 (6.04, 5.08), IP-10 (4.87, 3.12), LIF (5.11, 7.09), M-CSF (5.05, 0.78), MCP-1 (6.35, 0.64), MCP-3 (5.51,0.71), MIF (3.69,2.96), MIG (4.59,7.55), MIP-1a (5.78,0.06), MIP-1b (4.39,0.49), PDGF-bb (5.87, 12.7), RANTES (3.06, 1.91), SCF (5.55, 4.26), SCGF-b (3.19, 203.8), SDF-1a (3.42, 23.39), TNF-a (5.27, 9.6), TNF-b (6.36, 2.26), and TRAIL (5.39, 0.84).
Authors’ Contributions
Conceptualization and study design: O.B., C.S., A.L., P.G., G.F., S.K., S.H., M.A.B., B.K., A.S., and M.B. Methodology: D.M., C.P., A.L., C.S., P.G., O.B., and J.S. Formal analyses: D.M., C.P., A.L., C.S., and O.B. Investigation: D.M., C.P., A.L., C.S., A.C., G.F., M.A.B., M.B., J.H., T.B., B.K., A.S., and O.B. Data curation: O.B., D.M., and A.C. Writing original draft: D.M. and O.B. Editing: S.K., J.H., T.B., J.S., and B.K. Funding acquisition: O.B. All the authors reviewed and approved article.
Footnotes
Acknowledgment
The authors thank Dr. Martin Lesser for consultations on the initial study design and biostatistical plan.
Author Disclosure Statement
No competing financial interests exist.
Funding Information
This work was supported by the Office of the Assistant Secretary of Defense for Health Affairs under Award No. W81XWH-15-1-0614 (grant to O.B.). Opinions, interpretations, conclusions and recommendations are those of the author and are not necessarily endorsed by the Department of Defense. This work was also supported by grants to O.B. from the New York State Spinal Cord Injury Research Board.
Supplemental Material
Supplemental Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
