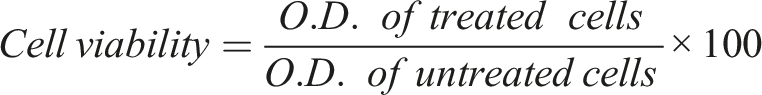Abstract
The rise of antimicrobial drug resistance among microorganisms presents a global challenge to clinicians. Therefore, it is essential to investigate drug delivery systems to combat resistant bacteria and fungi. This study examined the potential and mode of action of vancomycin-conjugated gold nanoparticles (PEI-AuNP@Van) to enhance vancomycin’s biocidal activity against C. tropicalis, C. albicans, E. coli, and P. aeruginosa. Drug conjugation and nanoparticle characterization were assessed using UV-Vis spectroscopy, X-ray diffraction, TEM, ATR-FTIR, and fluorescence spectroscopy. Effective vancomycin conjugation on polyethyleneimine-stabilized gold nanoparticles was achieved via electrostatic interactions or hydrogen bonding between the COO-/OH groups of vancomycin and the NH- groups of polyethyleneimine, yielding nanoparticles with a narrow size distribution and high zeta potential. The high luminescence of the nanoparticles facilitated their detection in microbial cells. PEI-AuNP@Van was internalized in C. albicans and C. tropicalis but showed surface adsorption in E. coli and P. aeruginosa. The in vitro results indicated that the nanodelivery system exhibited superior biocidal activity against the tested strains compared to free vancomycin and unconjugated AuNPs. The mode of action of PEI-AuNP@Van was cell-type-dependent, involving intracellular reactive oxygen species accumulation, cell membrane integrity loss, and apoptosis. The development of antimicrobial nanoformulations using AuNPs and efficient conjugation systems offers a promising approach to address antimicrobial drug resistance.
Introduction
Human diseases associated with microorganisms (e.g., bacteria, fungi, parasites, and viruses) typically respond to the administration of antimicrobial agents. However, emerging drug resistance among microorganisms, particularly Candida spp. And Gram-negative bacteria, poses a major global health challenge. Drug-resistant microbial infections are associated with increased mortality and substantial medical costs among hospitalized patients.1,2 For example, the high mortality rates among those with Candida species infections are attributed to invasive systemic infections and septicemia, particularly in immunocompromised individuals.1–3 Candida species (e.g., C. albicans, C. parapsilosis, C. tropicalis, C. glabrata, and C. krusei) inhabit healthy hosts but cause invasive infections when host immunity is compromised. 4 Despite the availability of antifungal treatments, Candida infection mortality rates remain high, at 45%. 5 The long-term use of azole drugs has led to increased antifungal resistance in Candida spp. 6 In 2019, the WHO reported approximately 4.95 million deaths related to antimicrobial-resistant (AMR) bacteria, primarily from the ESKAPE pathogens (S. aureus, S. pneumoniae, E. coli, K. pneumoniae, A. baumannii, and P. aeruginosa). 7 The rapid emergence of multidrug resistance (MDR) in Gram-negative bacteria severely threatens public health8,9; bacteria known as “superbugs” resist nearly all antibiotics and are prevalent in medical settings; this phenomenon severely limits treatment options for severe cases.
Nanotechnology may be used for addressing communicable and non-communicable diseases; for example, nanoparticles may be used as drug carriers. The physicochemical characteristics of nanoparticles (e.g., shape, surface chemistry, and size) can be modulated; as such, nanoparticles are versatile materials for combating bacterial and fungal infections.10,11 Due to their high drug-loading capacity and capability for crossing biological membranes, surface-modified nanoparticles are excellent drug carriers.12,13 Various metal nanoparticles and their oxides (e.g., silver, gold, aluminum oxide, zinc oxide, titanium dioxide, and copper oxide) have been extensively assessed, either alone or in combination with antibiotics or other functional molecules, with regard to their potential in treating infections.14–17 Gold nanoparticles (AuNPs) have garnered significant attention because of their biocompatibility, optical properties, and ability to be surface-modified with specific drugs. 18 Recent studies have successfully demonstrated AuNPs as carriers for the delivery of antimicrobial agents.19–24 Additionally, the high volume-surface ratio observed in AuNPs allows for substantial drug loading and efficient delivery to infection sites.
Mu et al. demonstrated that Ag@Au bimetallic core cell NPs that were loaded with ciprofloxacin exhibited enhanced antibacterial activity against Escherichia coli. 25 Similarly, Shaker et al. evaluated carbapenem-conjugated gold nanoparticles (meropenem and imipenem) against K. pneumoniae, P. mirabilis, and A. baumannii. Nanoparticles showed loading efficiencies of 72% for imipenem and 74% for meropenem. Imipenem-loaded nanoparticles reduced the minimum inhibitory concentration (MIC) fourfold; meropenem-loaded nanoparticles reduced the MIC threefold. 26 Other studies have also enhanced antibacterial drug efficacy using AuNPs as a drug carrier. Hagbani et al. used vancomycin-functionalized gold nanoparticles (V-GNPs) for treating bacterial infections. 10 The loading efficiency that was noted for vancomycin onto V-GNPs was 86.2%; the antibacterial activity of vancomycin-loaded V-GNPs was 1.4- to 1.8-fold higher against Gram-negative E. coli, K. oxytoca, and P. aeruginosa, and Gram-positive S. aureus in comparison to pure vancomycin. 10 Various strategies have been employed to improve antifungal efficacy via the conjugation of drugs or small molecules. For instance, the anti-Candida effects of indolicidin were compared with those of a host defense peptide that was conjugated with gold nanoparticles against fluconazole-resistant C. albicans isolates. 27 Numerous studies support the use of antibacterial and antifungal agents. Hsu et al. described the antifungal properties of polymyxin B, a cationic lipid oligopeptide typically utilized for the treatment of Gram-negative bacterial infections against Fusarium spp. 28 Similarly, Zhai et al. considered the anti-cryptococcal activity of polymyxin B. 29 Other antifungal agents have also been examined for efficacy against various fungal species. In addition, antifungal agents (e.g., caspofungin) have been evaluated for their bactericidal activity against vancomycin-resistant E. faecium. Thus, exploration of antibacterial agents against drug-resistant fungal isolates is warranted. This study considers the synthesis and enhancement of the in vitro antimicrobial activity of highly stable vancomycin-conjugated gold nanoparticles against C. albicans, C. tropicalis, and the Gram-negative bacteria E. coli and P. aeruginosa. Vancomycin, a known antibiotic for treating Gram-positive bacterial infections among terminally ill ICU patients, is a glycopeptide that exhibits a unique mode of action that inhibits the second stage of cell wall synthesis within bacteria. 30 However, Gram-negative bacteria resist vancomycin interaction with specific C-terminal peptides because of their outer lipopolysaccharide (LPS) membrane, which prevents drug access. 31 Therefore, this study explored the stabilization of gold nanoparticles using polyethyleneimine (PEI), a cationic polymer used for delivering nucleic acids and genes to mammalian cells, with vancomycin as a payload. This study suggests a method to combat antimicrobial drug resistance through rational conjugation and site-specific delivery of classical antibacterial and broad-spectrum biocidal agents.
Experimental section
Material and methods
All of the reagents and the materials were of analytical quality. Vancomycin, chloroauric acid trihydrate, polyethyleneimine, amphotericin B, and meropenem were purchased from Sigma Aldrich (Mumbai, India). Microbial cultures and SDA (Sabouraud dextrose agar) were obtained from RPMI (Roswell Park Memorial Institute, Buffalo, NY, USA); Mueller Hinton agar (MHA) and Mueller Hinton broth (MHB) were acquired from Hi-Media Laboratories Limited (Mumbai, India). The solvents were obtained from Merck Life Science Private Limited (Bangalore, Karnataka, India). Other required glassware and plastic wares were purchased from Tarson Products Private Limited (Kolkata, West Bengal, India). All of the experiments were conducted in ultra-purified HPLC-grade water.
Microbial strains and growth condition
In this study, we used the ATCC strains of C. tropicalis (ATCC 750), E. coli (ATCC 25,922), C. albicans (ATCC 90,028), and P. aeruginosa (ATCC 27,853). The microbial strains were identified via conventional microbiological procedures, including morphological and cultural characteristics; pure strains were subcultured on MHA for bacterial stains and SD agar slants for fungus and stored at −20°C for further experiments.
Synthesis of colloidal vancomycin-conjugated gold nanoparticles
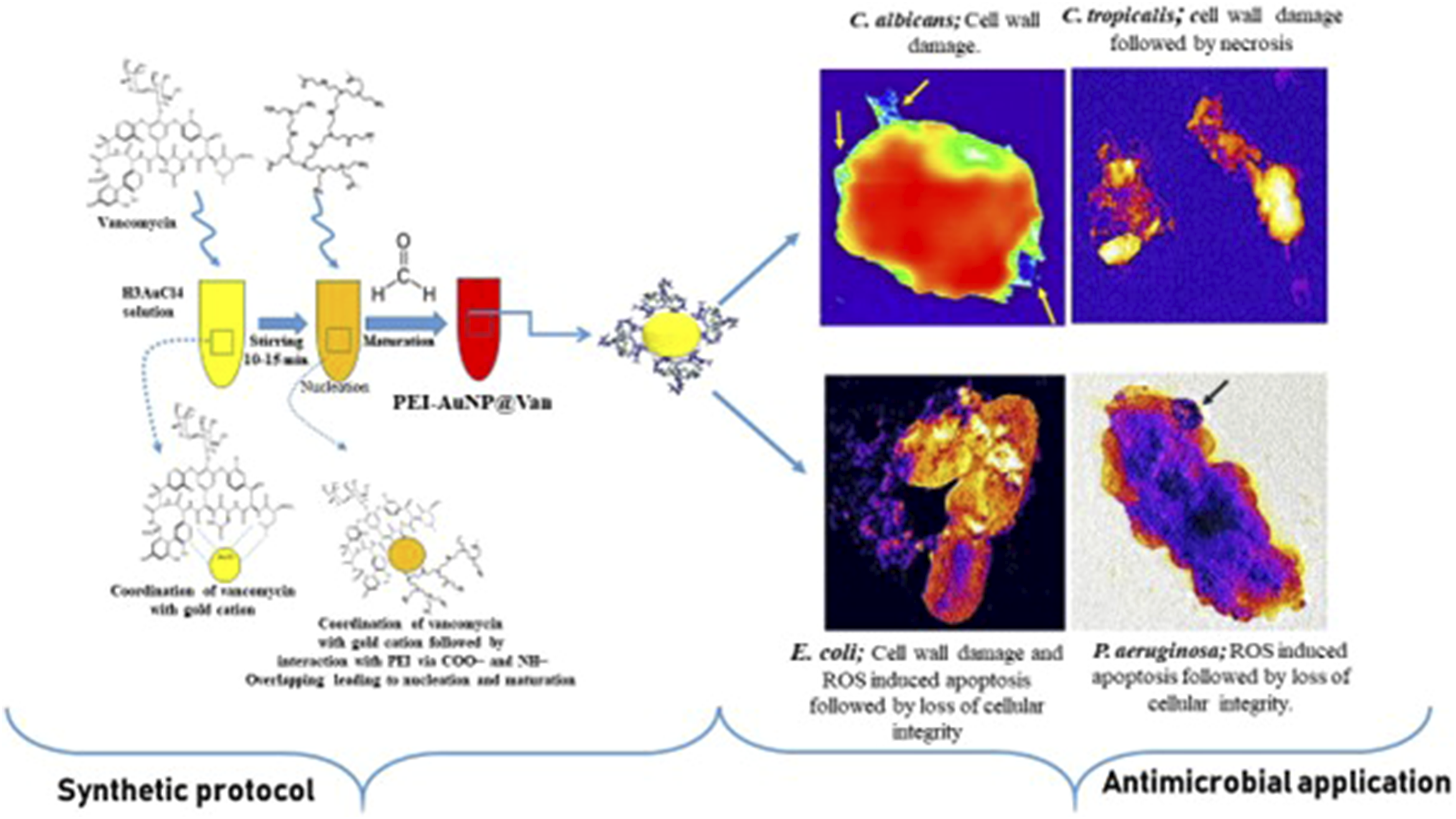
The vancomycin-conjugated colloidal gold nanoparticles were synthesized and described recently by Tiwari et al.32,33 Briefly, 200 μL (10 mM, HAuCl4.3H2O) was combined with an aqueous solution (40 μL, 2 mg/mL) of vancomycin within a glass vial; this step was followed via continuous stirring for 5–10 min using a magnetic stirrer. Immediately, an aqueous solution (30 μL of 4 mg/mL, mw. 60,000 Da.) of PEI (polyethyleneimine) was added; this step was followed by the addition of 30 μL of formaldehyde (35% w/v). The reaction mixture was stirred for 40–60 min at room temperature, which resulted in the dark red-pink color that was observed in PEI-AuNP@Van.
Physical characterization of prepared nanoparticles
The synthesized PEI-AuNP@Van was characterized using a U-2900 UV-Vis spectroscopy instrument (Hitachi, Tokyo, Japan) and transmission electron microscopy (TEM; FEI Tecnai G2 20 S Twin, Hillsboro, Oregon, USA). Nanoparticle size was analyzed using ImageJ software (NIH, Bethesda, MD, USA), and data were plotted with Origin 8.5 software (Northampton, MA, USA). Dynamic light scattering (DLS) and zeta potential measurements were undertaken using a Malvern Nano Zeta Sizer (Malvern, UK). X-ray diffraction (XRD) was performed using a Rigaku Mini-Flex 600 (Tokyo, Japan) on a nanoparticle film on a silicate glass slide covering 10–80° 2θ. The fluorescence emission spectra in water were recorded using a Hitachi F7000 spectrophotometer (Tokyo, Japan). Atomic force microscopy (AFM) measurements were conducted on nanostructured films of colloidal gold nanoparticles on silanized glass plates using a Bruker Dimension Edge AFM in tapping mode at ambient conditions. Attenuated total reflectance-Fourier transform infrared (ATR-FTIR) spectroscopy was utilized to analyze conformational changes upon vancomycin loading onto PEI-AuNPs with a Nicolet iS5 spectrometer (Thermo Electron Scientific Instruments, Waltham, MA, USA), and spectra were obtained with 4 cm−1 resolution from 4000 to 400 cm−1.
Minimal inhibitory concentration (MIC) determination
In vitro minimum inhibitory concentration (MIC) of PEI-AuNP (unconjugated) and PEI-AuNP@Van against strains of C. tropicalis (ATCC 750), E. coli (ATCC 25,922), C. albicans (ATCC 90,028), and P. aeruginosa (ATCC 27,853) were determined using a serial dilution (in purified water) technique in a sterile 96-well microtiter plate as previously described.34,35 A 50 µg/mL PEI-AuNP@Van aqueous suspension was prepared; 100 µL was dispensed into the first well and then serially diluted to final concentrations ranging from 0.38 to 50 µg/mL. The positive controls (amphotericin B and meropenem at 32 µg/mL) and a negative control (distilled water) were similarly diluted. Each well received 100 µL of freshly prepared microbial culture; the plate was then incubated statically at 37°C for 24 h. Further, a 10-µL aliquot from each well was then sub-cultured on MHA (bacteria) and SD agar plates (fungal strains) for 24 h, and microbial growth was observed. MIC values corresponded to the PEI-AuNP@Van concentration that inhibited or slowed growth when compared to the control.
Microbial cell viability assay
The viability of PEI-AuNP (unconjugated) and PEI-AuNP@Van-exposed C. albicans, C. tropicalis, E. coli, and P. aeruginosa cells was assessed using propidium iodide (PI), a membrane-impermeable dye as previously described by Yadav et.al.. 36 Briefly, freshly grown bacterial and fungal cultures were washed in PBS (100 mM, pH 7.2) followed by resuspension in MHB (for bacteria) and RPMI-1640 (for fungal strains); this step was followed by treatment with PEI-AuNP and PEI-AuNP@Van at their respective MIC, along with positive and growth controls, and incubation for 4 h at 37°C. Aliquots were withdrawn from each tube and then centrifuged at 3500 r/min for 5 min, followed by repeated washing and resuspension in PBS. The cells were stained with 10 µL of PI (1 mg/mL stock; λex/em = 535/617 nm), incubated at room temperature in the dark for 30 min, and then washed with PBS to remove excess dye. The treated cells were evaluated under an SP5 AOBS Super-resolution laser-scanning confocal microscope (Leica, Wetzlar, Germany) with a 40X oil immersion objective.
Determination of post-treatment dead population
To understand the percentage of dead cells after exposure to PEI-AuNP (unconjugated) and PEI-AuNP@Van, freshly grown cultures of C. albicans, C. tropicalis, E. coli, and P. aeruginosa were prepared. A 5 mL culture (107 CFU/mL for bacteria and 105 CFU/mL for fungi) in MHB and RPMI-1640 medium, respectively, received PEI-AuNP and PEI-AuNP@Van at their respective MICs. Each tube was subsequently incubated at 37°C for 4 h. Post-incubation, 2 mL aliquots were pelleted at 3500 r/min for 5 min and washed with PBS (100 mM, pH 7.2). PEI-AuNP@Van-exposed cells were stained with PI (10 µL; 1 mg/mL) and incubated in the dark for 30 min at room temperature. The stained cells were washed with PBS to remove the excess stain. The data acquisition was undertaken using BD FACSAria Fusion Flow cytometry (BD Biosciences, Franklin Lakes, NJ, USA) and BD Accuri C6 software, with measurements obtained using 20 mW laser illumination (λ = 488 nm). The assays were performed at a low sample rate (14 μL min−1), with 104 events for fungi and 106 events for bacteria, and were measured logarithmically.
Endogenous ROS accumulation study
PEI-AuNP@Van-treated C. albicans, C. tropicalis, E. coli, and P. aeruginosa cells were assessed for endogenous reactive oxygen species (ROS) production after 4 h. Flow cytometry with 2′, 7′-dichlorofluorescein-diacetate (DCFH-DA) as a ROS marker was employed for the measurements. 37 The cultures were exposed to MIC values, washed with PBS (100 mM, pH 7.2), and adjusted to 108 CFU/mL for bacteria and 106 CFU/mL for fungi. Cell suspensions were incubated with 5 μM DCFH-DA for 30 min, followed by BD FACSAria Fusion Flow cytometry analysis (BD Biosciences, Franklin Lakes, NJ, USA). Data analysis was performed using BD Accuri C6 software based on light scatter and fluorescence signals from 20 mW laser illumination (λ = 488 nm). Measurements were made logarithmically; measurements were obtained at a low sample rate (14 μL/min−1), with 104 events for fungal cells and 106 events for bacteria.
Phosphatidylserine externalization (apoptosis) assay
Freshly harvested (log phase) cells of the aforementioned strains were treated with PEI-AuNP@Van at their respective MICs at 37°C for 4 h; this step was followed by washing with PBS (100 mM, pH 7.2) and incubation for the next 20 min in an annexin V binding buffer that contained 5 μL/mL of Annexin V-FITC and 2 μL/mL of PI (FITC-annexin V apoptosis detection kit (BD PharmingenTM, Franklin Lakes, NJ, USA)) along with the untreated control. Further, recovered cells were visualized using an SP5 AOBS super-resolution laser-scanning confocal microscope (Leica, Wetzlar, Germany) with a 40X oil immersion objective lenses.
Raman spectroscopic studies of treated cells
Raman spectroscopy was undertaken to understand any bimolecular changes in microbial cells after exposure with PEI-AuNP@Van along with drug and growth control cells of C. albicans, C. tropicalis, E. coli, and P. aeruginosa cells. The exposed cells were pelleted and then washed twice with PBS to remove any debris. Then, the sample was placed on a microscopic glass slide and then dried under ambient conditions for 30 min. Raman measurements were obtained using a WI-Tec alpha 300 RA/S instrument (WI-Tec GmbH, Berlin, Germany). The excitation source was a Nd: YAG laser (λ = 532 nm), which provided a maximum power output of 40 mW. The laser power attenuated to ∼16 mW at the sampling point. The dispersed intensity of light associated with the signal from the grating was studied with a Peltier-cooled charge-coupled device (CCD); the spectra were recorded over the range of 400-2400 cm−1.
Multivariate data analysis
Principal Component Analysis (PCA) is an unsupervised technique that was used to examine spectral datasets and identify the primary patterns of variability within the spectral matrix of various samples. This method transformed correlated variables into uncorrelated ones, termed PC scores, which preserved the variability in the spectral data and eliminated random noise. The initial PC accounted for the majority of the variation that was observed in the spectral data; subsequent PCs elucidated the remaining variability. 38 While PCA is effective in revealing clusters within datasets, other biological factors with less significance, such as spectral characteristics unrelated to pathological conditions, may influence the data distribution in scatter plots. Consequently, to achieve enhanced discrimination, a supervised constraint was introduced to PCA, which placed greater emphasis on the underlying spectral information in the classification process. 39 The PCA analysis was performed using Origin 2018 software (Origin Lab. Corp, Northampton, MA, USA).
Ultra-structural visualization (Transmission electron microscopy) studies
For the transmission Electron Microscopy (TEM) (G2 20 S Twin Technai, FEI instrument, Hillsboro, OR, USA) studies, freshly harvested cells of C. albicans, C. tropicalis, E. coli, and P. aeruginosa were prepared and treated with PEI-AuNP@Van at their respective MICs and then incubated for 4 h at 37°C. After that, the culture was pelleted down, washed with PBS, and fixed with Karnowasky solution (2% paraformaldehyde, 2.5% glutaraldehyde) in 0.1 M sodium phosphate buffer (pH 7.4) for 1 h. Further, the treated samples and control samples were drop cast onto a carbon-coated copper grid with a mess size of 300, dried for 1 h under ambient conditions, and visualized.
Cytotoxicity evaluation of PEI-AuNP@Van on HEK-293 cell line
In order to understand the toxic effects of PEI-AuNP@Van on cells, human embryonic kidney (HEK-293) cells were cultured in DMEM supplemented with 10% FBS and 1% antibiotics (Penicillin/Streptomycin). Cells were incubated at 37°C in a humidified environment with 5% CO2 (Thermo Scientific Heracellvios 160i CO2 incubator). A total of 10,000 cells were seeded into each well of 96-well tissue culture plates and then left overnight. On the following day, the medium was replaced with 200 µL of the fresh culture medium that contained varying concentrations of PEI-AuNP@Van (5–100 µg/mL); the cytotoxicity was examined after 24 h. Following exposure, a 5 mg mL−1 solution of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) in PBS pH 7.2) was added and incubated for 3 h at 37°C. The medium was then removed, and the resulting formazan crystals were solubilized in 100 µL of DMSO. After 20 min, the absorbance of the solution was evaluated at λ:570 nm with a microplate reader. The cell viability value was described as a percentage (%) relative to untreated cells using the formula as follows:
Statistical analysis
All of the experiments were conducted three independent times. The data were represented in mean ± standard deviation (SD) format. The statistical significance was evaluated with ordinary one-way ANOVA tests. In this study, the p values <0.05, <0.01, and <0.001 were noted to indicate the statistical significance.
Results
Synthesis and characterization of vancomycin-conjugated gold nanoparticles
Tiwari et al. (2023) optimized a one-pot procedure for synthesizing vancomycin-conjugated AuNPs.32,33 The resulting PEI-AuNP@Van nanoparticles exhibited an absorption maximum at 520 nm, with vancomycin interactions evaluated using UV-Vis spectrometry (Figure 1(a)). These spherical nanoparticles had a uniform size distribution, with an average diameter of 7.5 nm and a standard deviation of 0.5 nm. The smallest particle measured 3 nm, and the largest measured 12 nm (Figure 1(b) and (d)). Zeta potential analysis (Figure 1(c)) showed a sharp bimodal distribution around 48 mV, indicating a positive charge and stable dispersibility over several months. X-ray diffractometry data (2θ = 10°-80°) revealed peaks at 38.4°, 44.6°, 64.6°, and 77.7°, corresponding to hkl values of (111), (200), (220), and (311), indicating that the particles were face-centered cubic (FCC) crystalline (Figure 1(e)). AFM results (Figure 1(f)) agree with the TEM observations (Figure 1(a) and (b)). Physical characteristics of as-prepared vancomycin-conjugated nanoparticles (PEI-AuNP@Van). (a) UV–visible spectrum, (b) TEM micrograph, (c) zeta potential distribution, (d) particle size distribution plot, (e) X-ray diffractogram, and (f) AFM micrograph of nanoparticles. (Fig 1 (a, d and e) Adopted from ref. 33).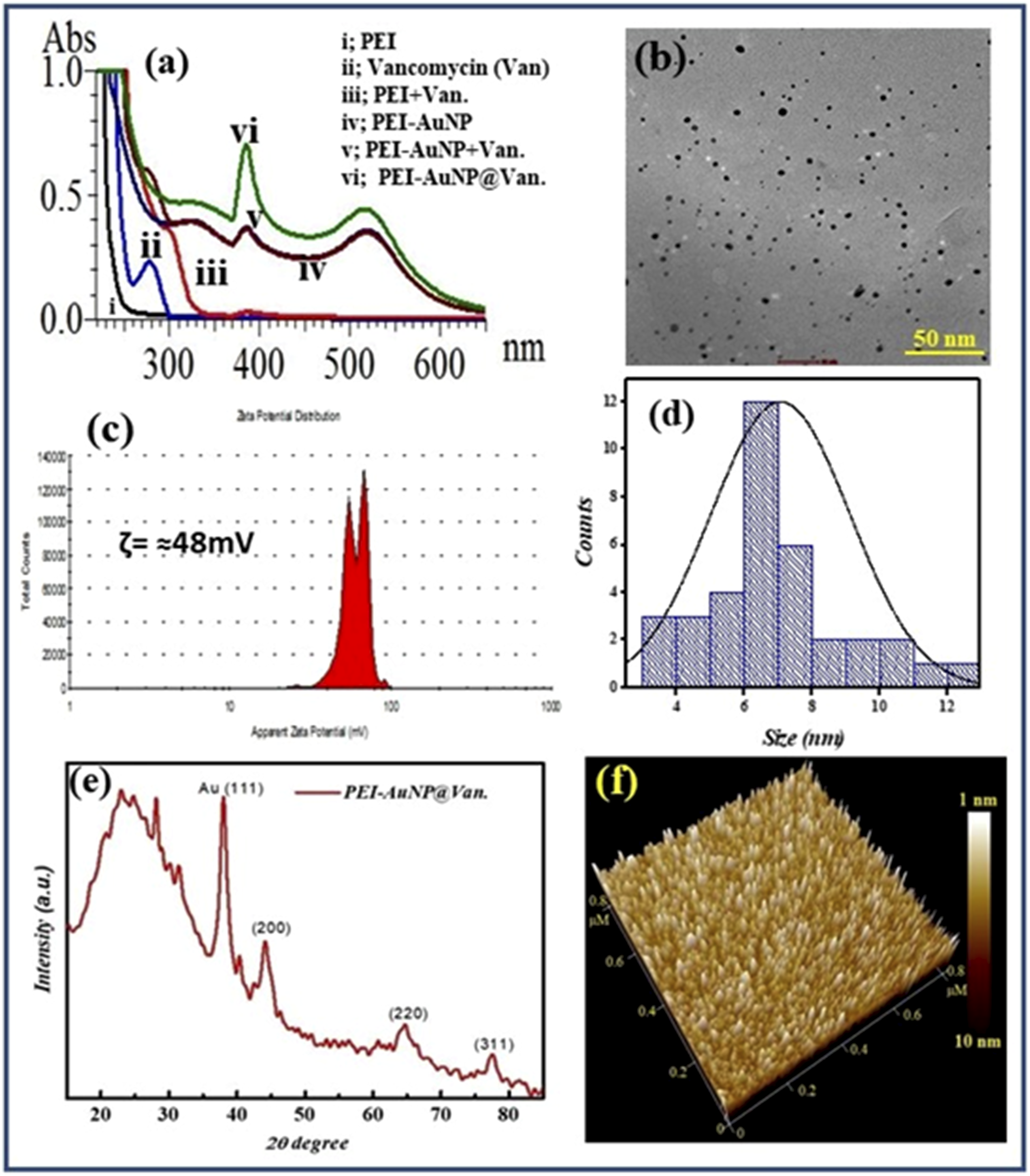
ATR-FTIR spectroscopy was used to analyze vancomycin conjugation on PEI-AuNPs, providing semi-quantitative data that are challenging to obtain via UV-visible spectrometry. Figure 2 shows the spectra of pure vancomycin, PEI-AuNPs, and PEI-AuNP@Van. Pure vancomycin exhibited peaks at 3313 cm−1 (O–H stretching), 2918 cm−1 (C–H stretching), 2110 cm−1 (C≡C stretching), 1635 cm−1 (C = O stretching), 1435 cm−1 (O–H bending), 1383 cm−1 (C = C stretching), 1244 cm−1 (C–N stretching), and 1194 cm−1 (C–O stretching), with a strong peak at 1024 cm−1 (C–N/O–H stretching) (Figure 2, black line). Unconjugated AuNPs (PEI-AuNPs) exhibited peaks at 3313 cm−1 (O–H stretching), 1635 cm−1 (C = O stretching), and 1231 cm−1 (C–N stretching) (Figure 2, red line). Vancomycin-conjugated AuNPs (PEI-AuNP@Van) exhibited shifts and new peaks, such as 2843 cm−1 (C-H stretching), weakening at 1435 cm−1 (O–H bending), and 1115 cm−1 (C–O stretching of the secondary alcohol), with a shift from 1024 to 1015 cm−1 (C–OH stretching) (Figure 2, blue line), confirming the presence of vancomycin on the PEI-AuNP surface. (a) ATR-FTIR spectra of free vancomycin (black line), unconjugated PEI-AuNPs (red line), and vancomycin-conjugated PEI-AuNPs (PEI-AuNP@Van; blue line).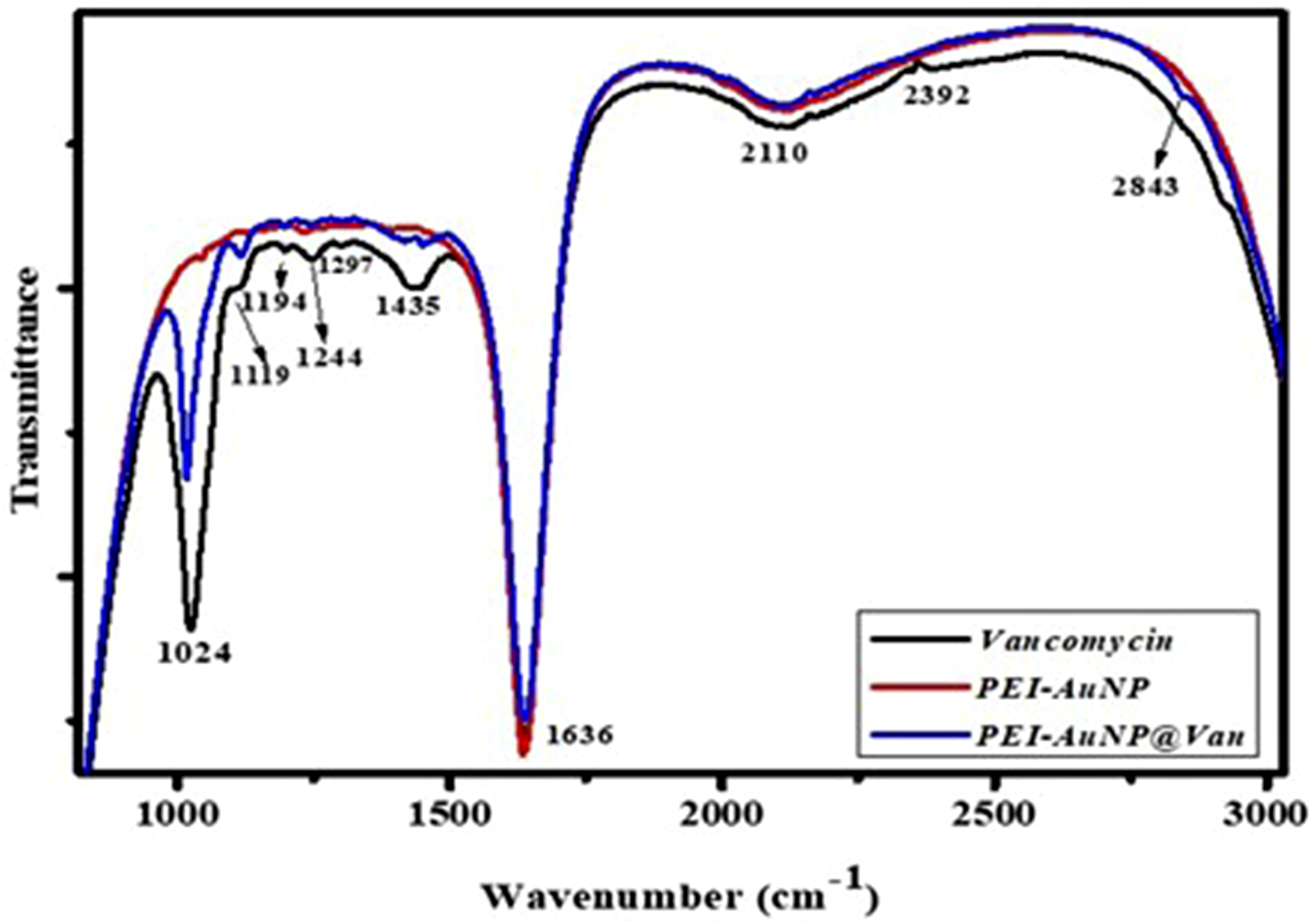
PEI-AuNP@Van nanoparticles have potent antimicrobial activity
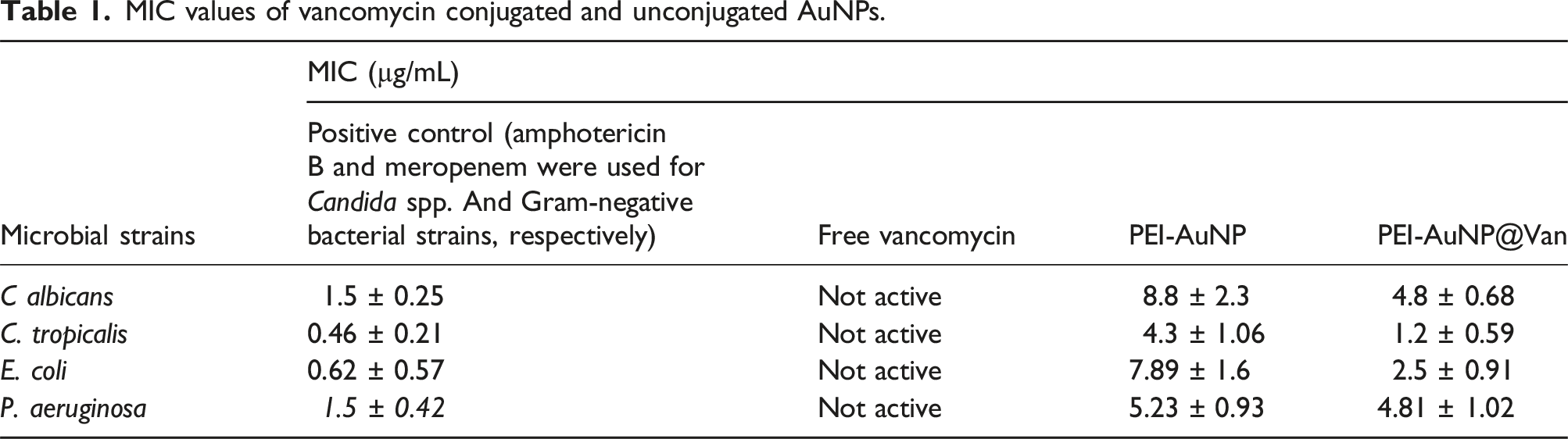
MIC values of vancomycin conjugated and unconjugated AuNPs.
Cell membrane integrity was compromised by PEI-AuNP@Van
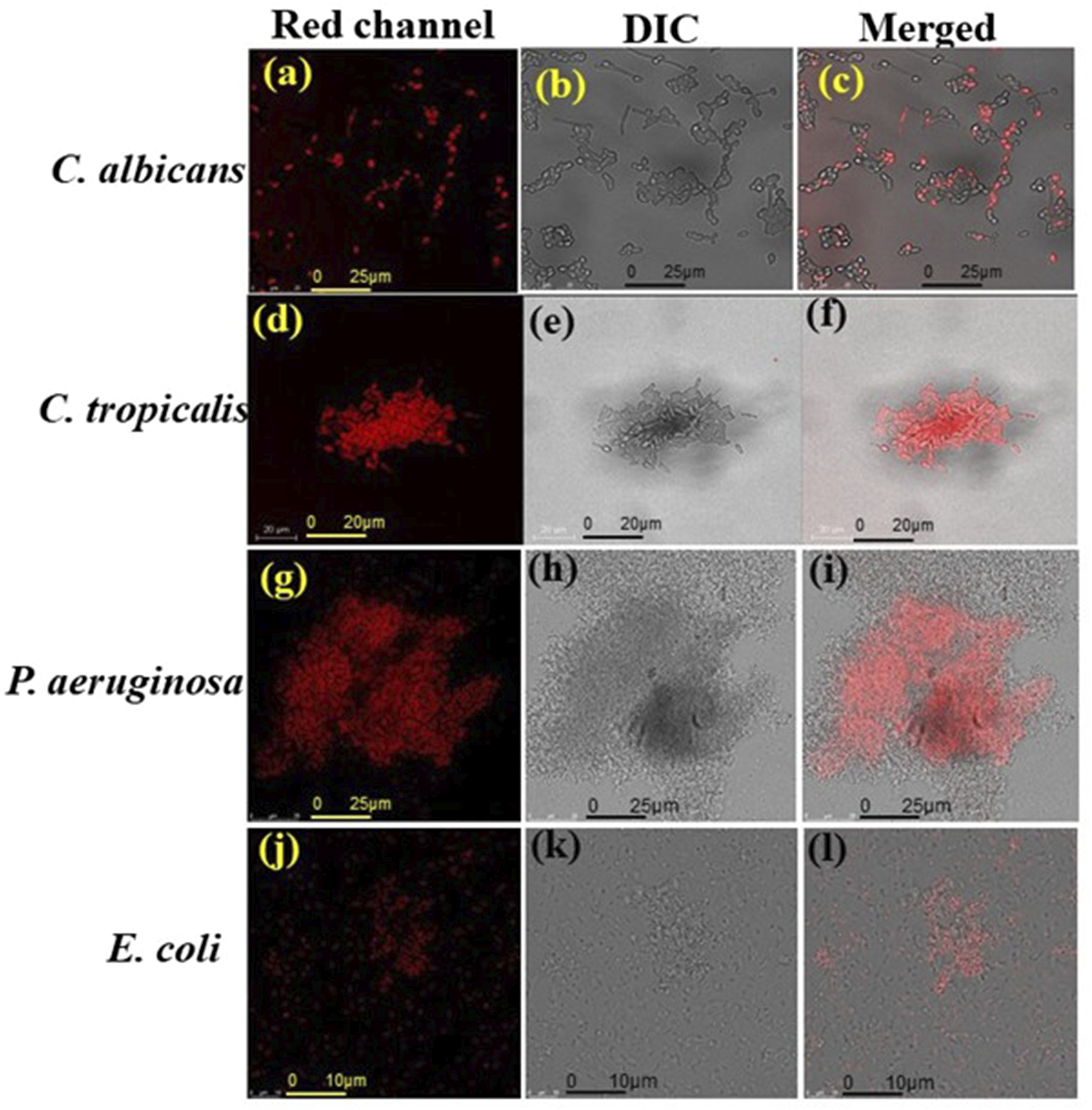
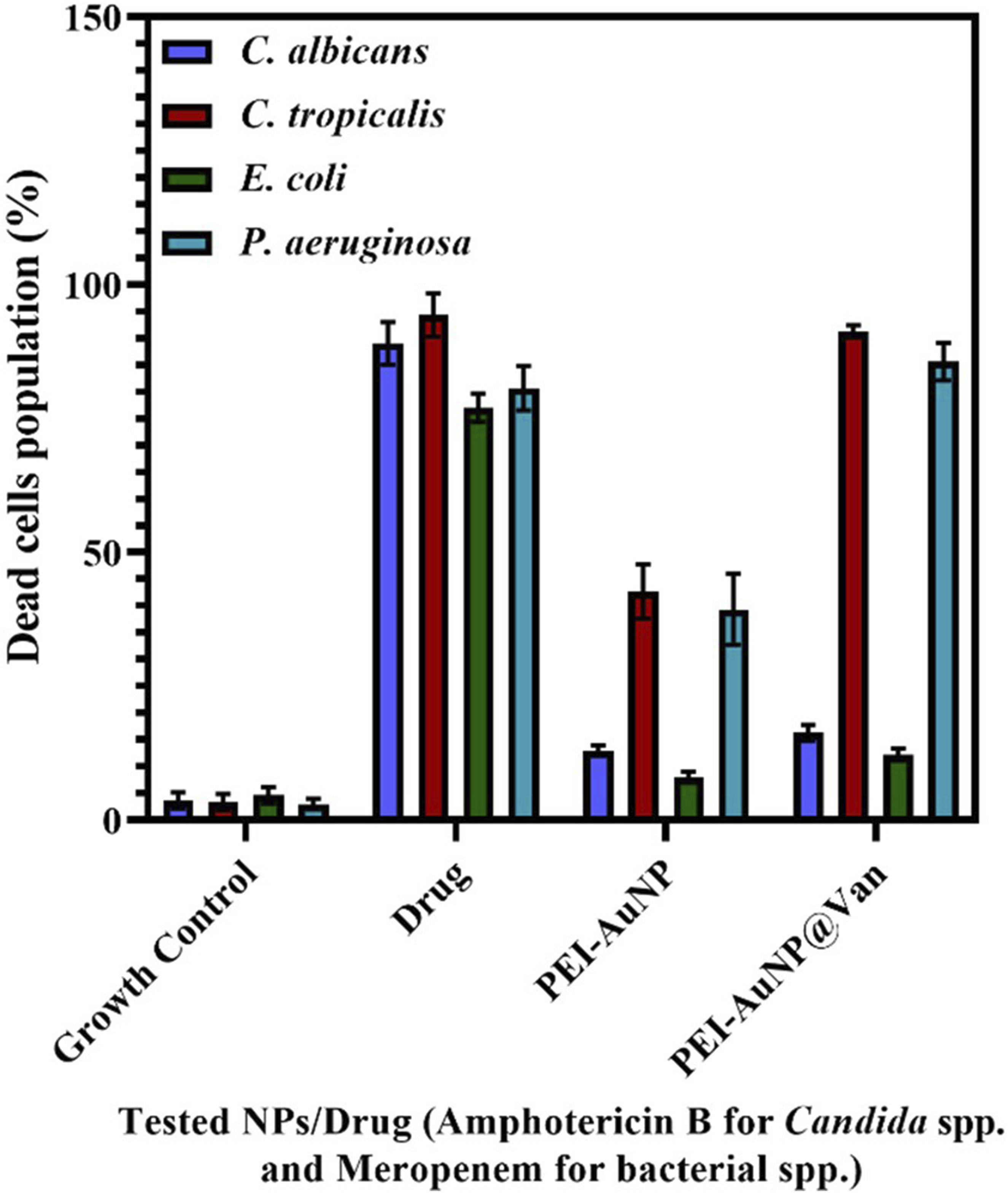
When PEI-AuNP@Van encountered microbial cells, it bound to a specific site on the cell surface, resulting in antimicrobial activity. As a fluorescent nanoparticle, PEI-AuNP@Van could be detected using specific excitation/emission ranges under confocal microscopy. As shown in Figure 3(a)–(h), PEI-AuNP@Van nanoparticles localized intracellularly in C. albicans and C. tropicalis cells, but adsorbed onto the cell surface of E. coli and P. aeruginosa after 4 h of exposure (Supplement Figure S1). A viability assay with propidium iodide dye (Figure 4) confirmed that PEI-AuNP@Van compromised the membranes of the treated cells. The percentage of dead cells was assessed via fluorescence-activated cell sorting with propidium iodide (PI) dye (Figure 5 and Supplement Figure S3), showing significant membrane compromise in C. albicans (16.3%), C. tropicalis (91.8%), E. coli (12%), and P. aeruginosa (85.8%). Unconjugated PEI-AuNPs showed a relatively low killing activity (Figure 5), whereas the standard antimicrobial drug exhibited excellent cell membrane-damaging behavior. Confocal micrographs of the post-exposure localization of PEI-AuNP@Van. C. albicans: (a) DIC panel (b) red panel (propidium iodide), (c) blue emission (PEI-AuNP@Van), and (d) all merged panel. C. tropicalis; (e) DIC panel, (f) red emission panel (propidium iodide), (g) blue emission (PEI-AuNP@Van) panel, and (h) merged panel. Confocal micrographs representing dead cells after the exposure to PEI-AuNP@Van for 4 h. C. albicans (a: PI emission panel; b: DIC panel and c: red, DIC merged panel); C. tropicalis, (d: PI emission panel; e: DIC panel and f: red, DIC merged panel); E. coli (g: PI emission panel; h: DIC panel and i: red, DIC merged panel) and P. aeruginosa (j: PI emission panel; k: DIC panel and l: red, DIC merged panel). Graph representing dead cell populations after 4 h of exposure to PEI-AuNP@Van along with amphotericin B and meropenem (for Candida spp. And Gram-negative bacteria, respectively) and unconjugated PEI-AuNPs. Data are presented as mean values of three independent experimental replicates.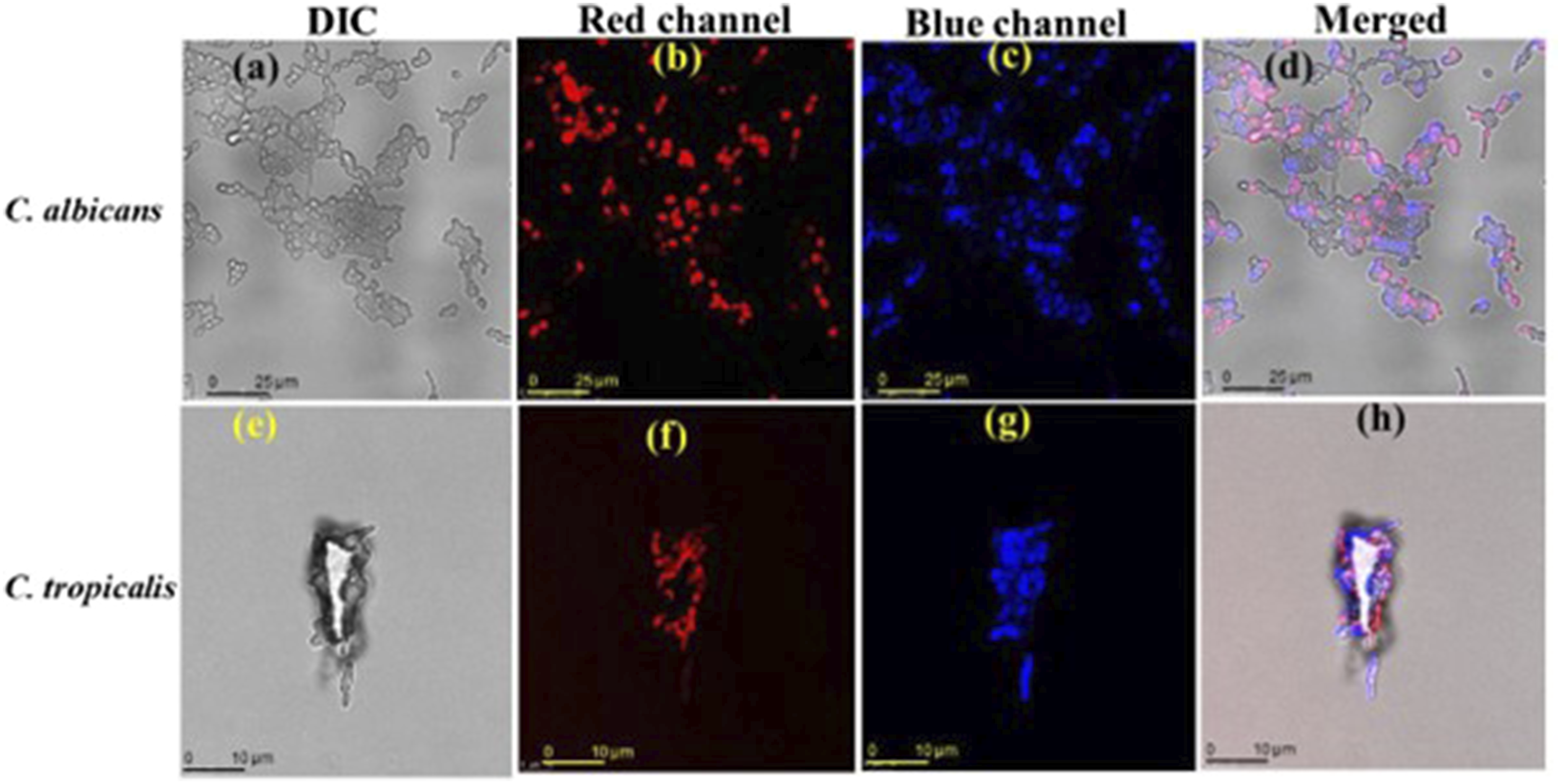
PEI-AuNP@Van induces endogenous ROS accumulation and apoptotic features in a cell type-dependent manner
Endogenous ROS generation was monitored via flow cytometry using an active dye, 2, 7 dichlorofluorescein diacetate (DCFH-DA). Interestingly, the results indicated differences in the intracellular ROS accumulation profile in the PEI-AuNP@Van exposed strains (Figure 6 and Supplement Figure S3). Compared with the untreated growth control, the PEI-AuNP- and PEI-AuNP@Van-treated cells demonstrated a substantial increase in DCF fluorescence. After treatment with PEI-AuNP@Van C. albicans (11.38 %; Figure 6(a)), C. tropicalis (25.02 %; Figure 6(b)), E. coli (30.4%; Figure 6(c)), and P. aeruginosa (36.0 %; Figure 6(d)) populations demonstrated elevated endogenous reactive oxygen species as compared to PEI-AuNP and the standard drug control. In contrast, untreated control cells showed negligible ROS generation. These results justify the ROS induction activity of PEI-AuNP@Van in treated microbial cultures. Graph representing endogenous ROS-accumulated populations of 4 h exposed cultures of tested microbial species with PEI-AuNP@Van along with drug and unconjugated PEI-AuNPs. (a) C. albicans; (b) C. tropicalis; (c) E. coli, and (d) P. aeruginosa. The data are described as the mean value of three independent experimental replicates, and an ordinary one-way ANOVA was completed to compare the tested variables (p value *** = <0.001; ** = 0.007; * = 0.04, and ns = > 0.05).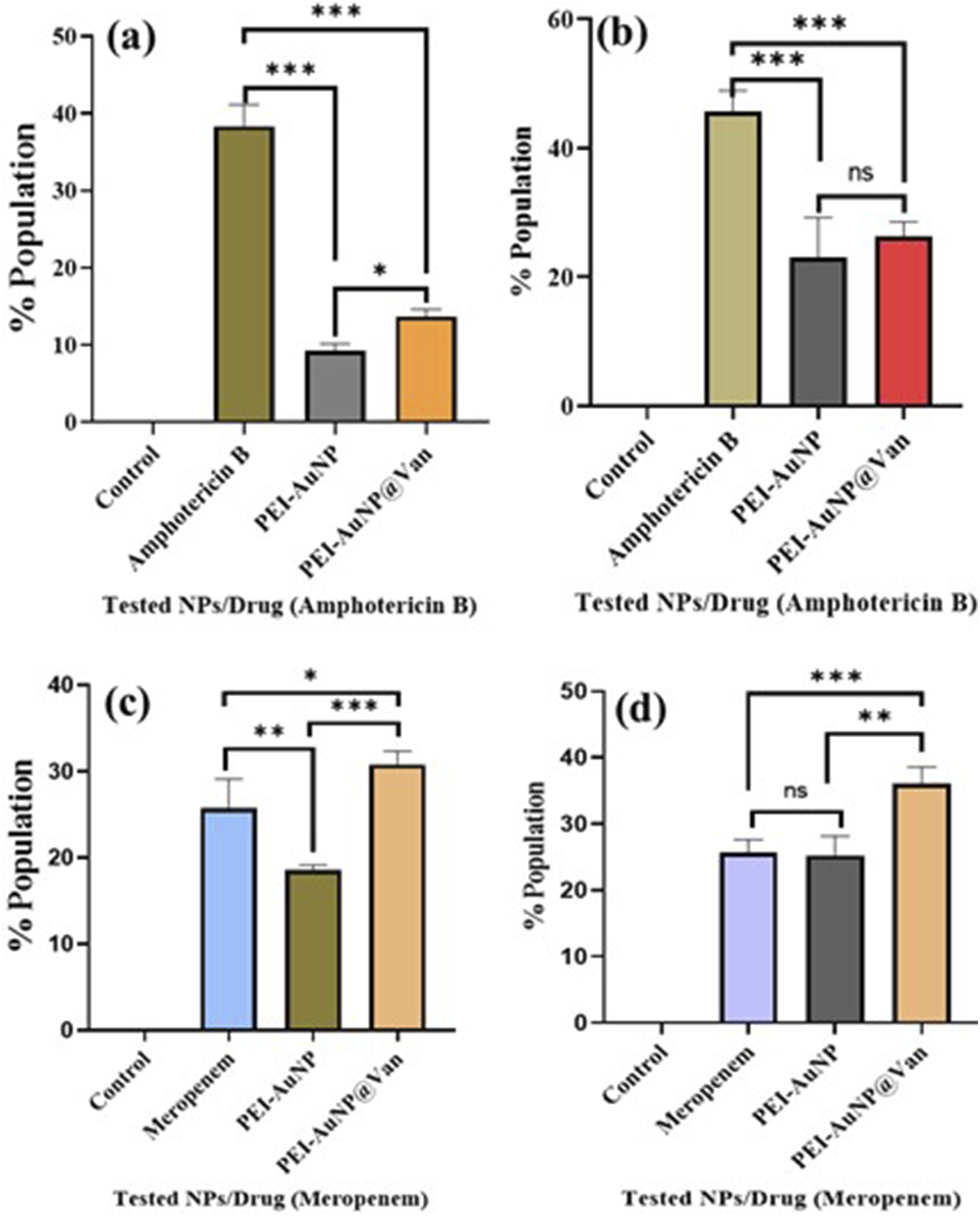
To assess whether PEI-AuNP@Van-treated C. albicans, C. tropicalis, E. coli, and P. aeruginosa cells exhibit apoptotic phenomena, annexin V-FITC/PI and confocal microscopy studies were performed. The results indicate that all treated microbial strains, except C. albicans (Figure 7(a)–(d))), displayed early and late apoptotic properties (as shown in Figure 7). Notably, P. aeruginosa (Figure 7(m)–(p)) showed significant apoptotic behavior, followed by E. coli (Figure 7(i)–(l)). Confocal micrographs representing phosphatidylcholine externalization assay using the Annexin-V/PI staining system after 4 h exposure to PEI-AuNP@Van against microbial species. C. albicans: (a) DIC panel, (b) green panel (Annexin-V stained cells), (c) red panel (PI stained cells), and (d) all merged panel; C. tropicalis: (e) DIC panel, (f) green panel (Annexin-V stained cells), (g) red panel (PI stained cells), and (h) all merged panel); E. coli: (i) DIC panel, (j) green panel (Annexin-V stained cells), (k) red panel (PI stained cells), and (l) all merged panel); and P. aeruginosa: (m) DIC panel, (n) green panel (Annexin-V stained cells), (o) red panel (PI stained cells), and (p) all merged panel).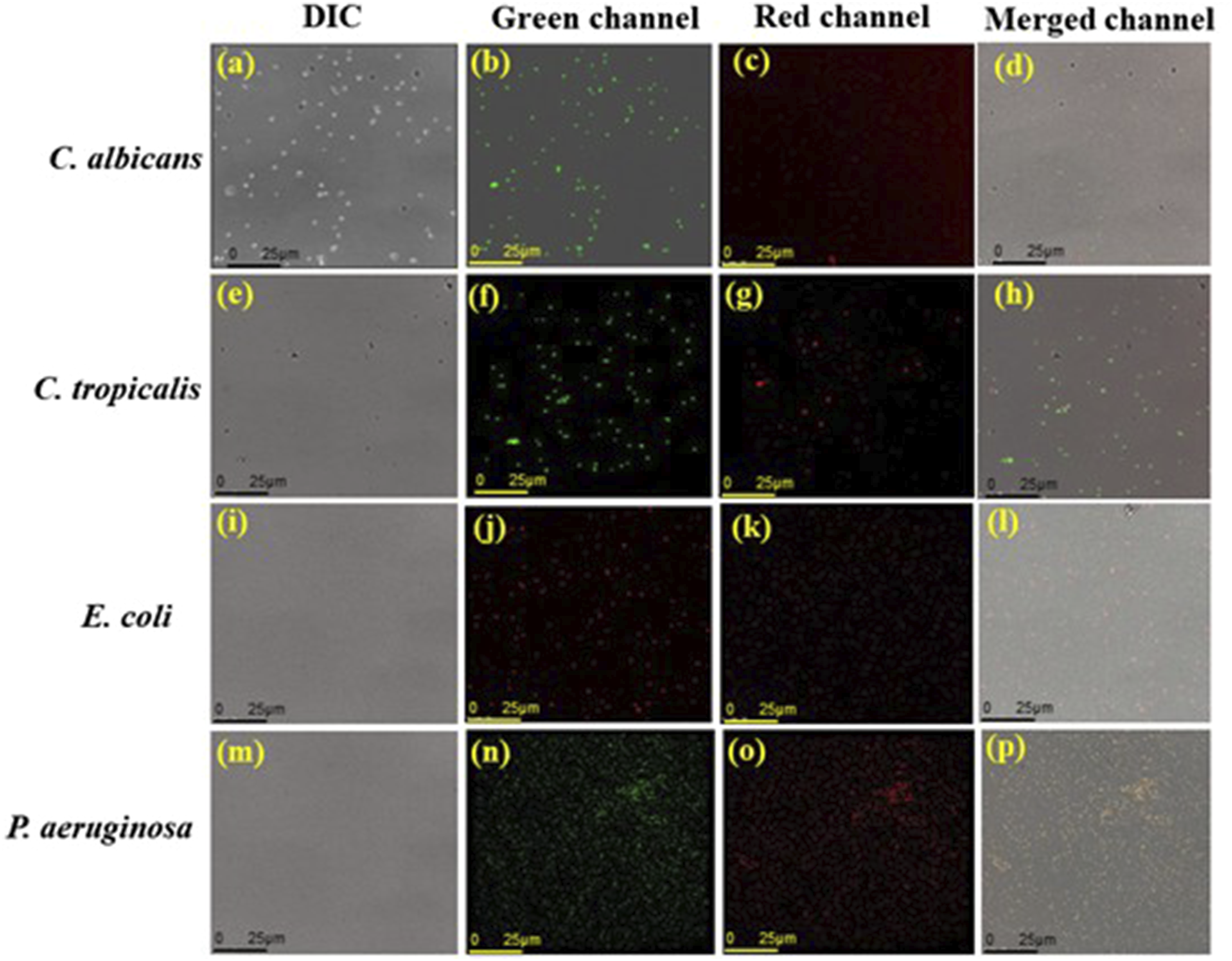
Raman spectroscopy analysis
Raman spectrometry of C. albicans, C. tropicalis, E. coli, and P. aeruginosa was performed after treatment with PEI-AuNP@Van; the recorded spectra matched and validated with previously reported Raman band fingerprints and molecular assignments; multivariate data analysis and mean spectral analysis were performed.38–50
Raman spectroscopy of C. albicans and C. tropicalis
Figure 8(a) and (d) show the Raman spectra that were obtained from untreated cells and PEI-AuNP@Van- and amphotericin B-treated C. albicans and C. tropicalis cells, respectively. In the untreated cells, Raman bands appeared at 487 cm−1 (CCO bending and COHO torsion in chitin), 531 cm−1 (CCC, β(CCO), OCO bending in chitin), and 539 cm−1. Features at 726-735 cm−1 (CO stretching, ring deformation of glucosyl unit/adenine, and glycosides) were observed in both species. Bands at 808-810 cm−1 correspond to asymmetric OPO phosphodiester stretching; the 1040 cm−1 band is linked to C–C stretching (phospholipids, chitin) and C–N stretching. The 1135 cm−1 band relates to = C–O–C= (unsaturated fatty acids in lipids), which is common to both species. The 1248 cm−1 band in C. tropicalis was linked to amide III (random) bonding and was absent in C. albicans. Both species showed bands at 1314 and 1323 cm−1 (amide III bonding, protein, or CH deformation), and CH2 deformation bands at 1449 and 1452 cm−1, respectively. The 1582 cm−1 band in C. tropicalis (absent in C. albicans) was attributed to NH2 deformations/amide III of proteins; the 1663 cm−1 band, common to both, was linked to chitin. These peaks corresponded to the Candida cell wall components: mannoproteins, glucan, and chitin. The spectra of amphotericin B- and PEI-AuNP@Van-treated cells were compared with those of untreated cells to study the mode of action and structural changes of PEI-AuNP@Van. Differences were noted at 587 cm−1 (absent in C. albicans), 590 cm−1, 676 cm−1, 685 cm−1, 852 cm−1 (absent in C. albicans), 914 cm−1, 1125 cm−1, 1162 cm−1, 1214 cm−1 (absent in C. albicans), 1314 cm−1, 1582 cm−1 (absent in C. albicans), 1602 cm−1, and 1653 cm−1. These features (Figure 8(a) and (d) and Supplement Table S1) are attributed to in-plane bending (CCC), β(CCO), and β(OCO) in chitin glycosidic linkage (587, 590 cm−1); guanine and tyrosine (676 cm−1); in-plane CCO bending in glucosyl unit (685 cm−1, absent in C. tropicalis); CH2 twisting, CC stretching in glucosyl unit (852 cm−1), CC bond stretching (914 cm−1, absent in C. tropicalis); C–O ring, aromatic amino acids in proteins (1162 cm−1, absent in C. tropicalis), amide III (random), and thymine (1214 cm−1, absent in C. albicans). Raman spectra and PCA analysis of PEI-AuNP@Van treated microbial cells. (a) Raman spectrum of C. albicans: (i) untreated cell control, (ii) amphotericin B treated, and (iii) PEI-AuNP@Van treated. (b) Loading plot and (c) k-cluster plot. (d) Raman spectrum of C. tropicalis: (i) untreated cell control, (ii) amphotericin B treated, and (iii) PEI-AuNP@Van treated. (e) Loading plot and (f) k-cluster plot. (g) Raman spectrum of E. coli: (i) untreated cell control, (ii) meropenem-treated, and (iii) PEI-AuNP@Van-treated. (h) Loading plot and (i) k-cluster plot. (j) Raman spectrum of P. aeruginosa: (i) untreated growth control, (ii) meropenem treated, and (iii) PEI-AuNP@Van treated. (k) Loading plot and (l) k-cluster plot.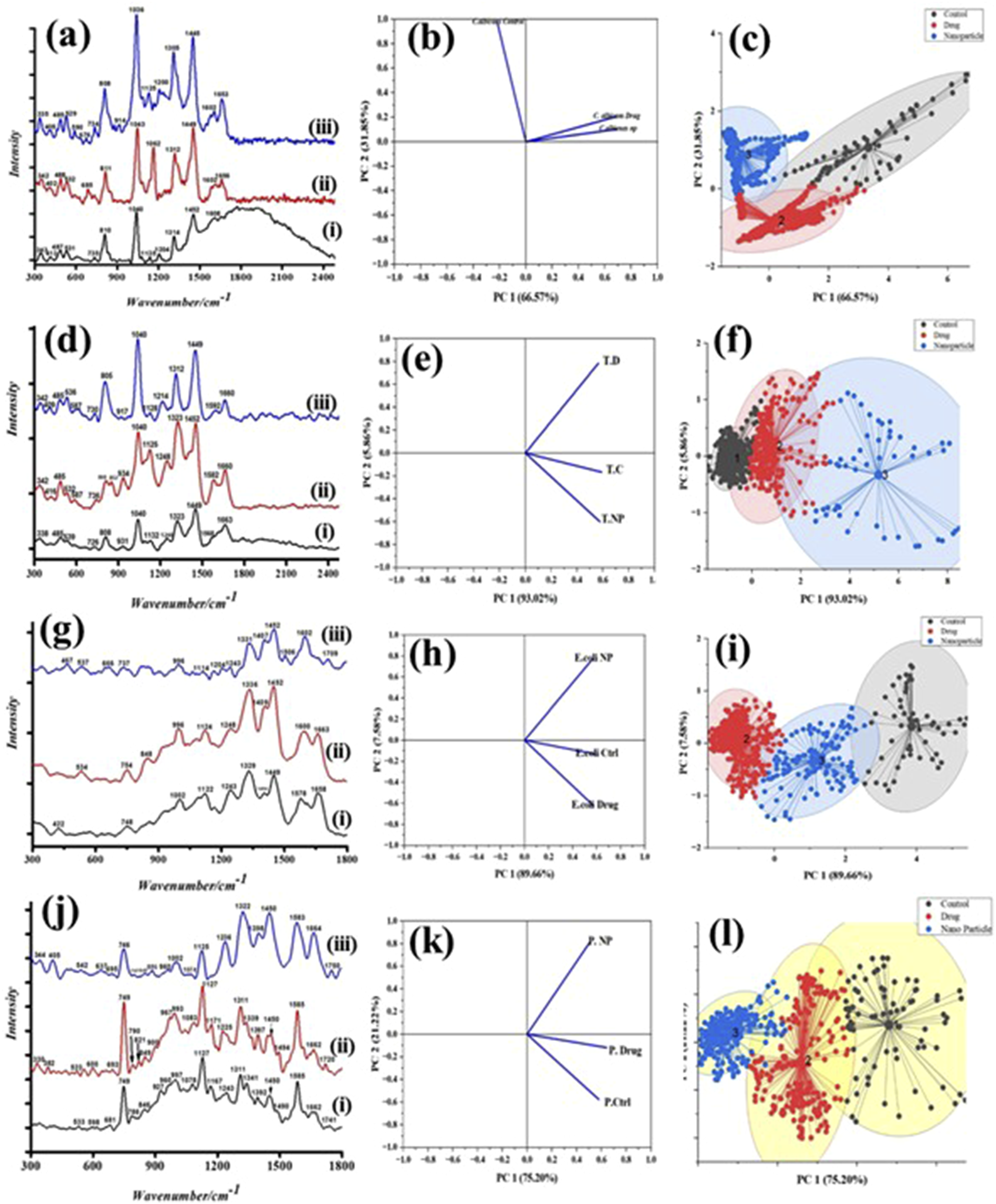
Raman spectroscopy analysis of E. coli and P. aeruginosa
Unlike fungal cells, Gram-negative bacteria exhibit a thin peptidoglycan cell wall that is enclosed by a lipopolysaccharide outer membrane. Raman spectroscopy of untreated, PEI-AuNP@Van, and meropenem-treated E. coli and P. aeruginosa cells (Figure 8(c) and (d)) revealed weak and strong peaks between 400 and 1660 cm−1. Peaks included 748 cm−1 (C–S (protein)/CH2 rocking/adenine, peptidoglycan), 997 or 1002 cm−1 (in-plane (CH), Phe, (CC) aromatic ring, ring-breathing (CC) polysaccharides), 1122 or 1127 cm−1 (stretching (C–C) skeletal of acyl backbone in lipid), 1167 cm−1 (C–H in-plane bending in tyrosine, absent in E. coli), 1243 cm−1 (asymmetric phosphate stretching, amide III), 1311 cm−1 (CH2 wagging and twisting), 1329 cm−1 (COH (mono, oligosaccharide) amide III, bend (CH), (OH) (polysaccharide) d (OH), in-plane (CH2), 1341 cm−1 (amide III, Trp; bend (CH), d (OH) (polysaccharide), 1392-1404 cm−1 (symm (CO2) alanine), 1450 cm−1 (CH2) (fatty acid chains, phospholipids, amino side-chains of proteins); CH3, CH2, COH (mono or oligosaccharides), 1578-1585 cm−1 (νCOO−; C-C, N–H deformation; C–N stretching (amide II)), and 1658 cm−1 (amide I), indicating fingerprint bonds of untreated cells (Figure 8(c) and (d)). Comparison of treated E. coli and P. aeruginosa cells with untreated controls showed spectral band differences (Figure 8(c) and (d)). Peaks at 534 cm−1, 848 cm−1, 1506 cm−1, and 1600 cm−1 were absent in untreated E. coli, while 637 cm−1 and 1662 cm−1 were absent in P. aeruginosa. (Figure 8(c) and (d)).
Principle component analysis
The principal component analysis (PCA) of Raman spectral datasets for C. albicans, C. tropicalis, E. coli, and P. aeruginosa (Figure 8(b), (c), (e), (f), (h), (i), (k) and (l)) show three distinct clusters, each indicated by a different color, formed by the separation of Raman spectra. Each dot within a cluster represents a single spectrum, with clusters aligned along the positive axis of the first principal component (PC-1). These clusters represent spectral datasets for candida strains, E. coli, and P. aeruginosa exposed to PEI-AuNP@Van, amphotericin B, and meropenem, respectively, as well as untreated cells. PC-1 explained the largest data variability: 66.57% (Figure 8(b) and (c))), 93.02% (Figure 8(e) and (f))), 89.66% (Figure 8(h) and (i))), and 75.20% (Figure 8(k) and (l))) for C. albicans, C. tropicalis, E. coli, and P. aeruginosa, respectively, with PC-2 accounting for the remaining variability. PCA identified and verified distinctive Raman spectral features, showing that Raman spectroscopy could detect spectral changes owing to biochemical alterations induced by PEI-AuNP@Van in the exposed strains.
Ultrastructural (TEM) analysis
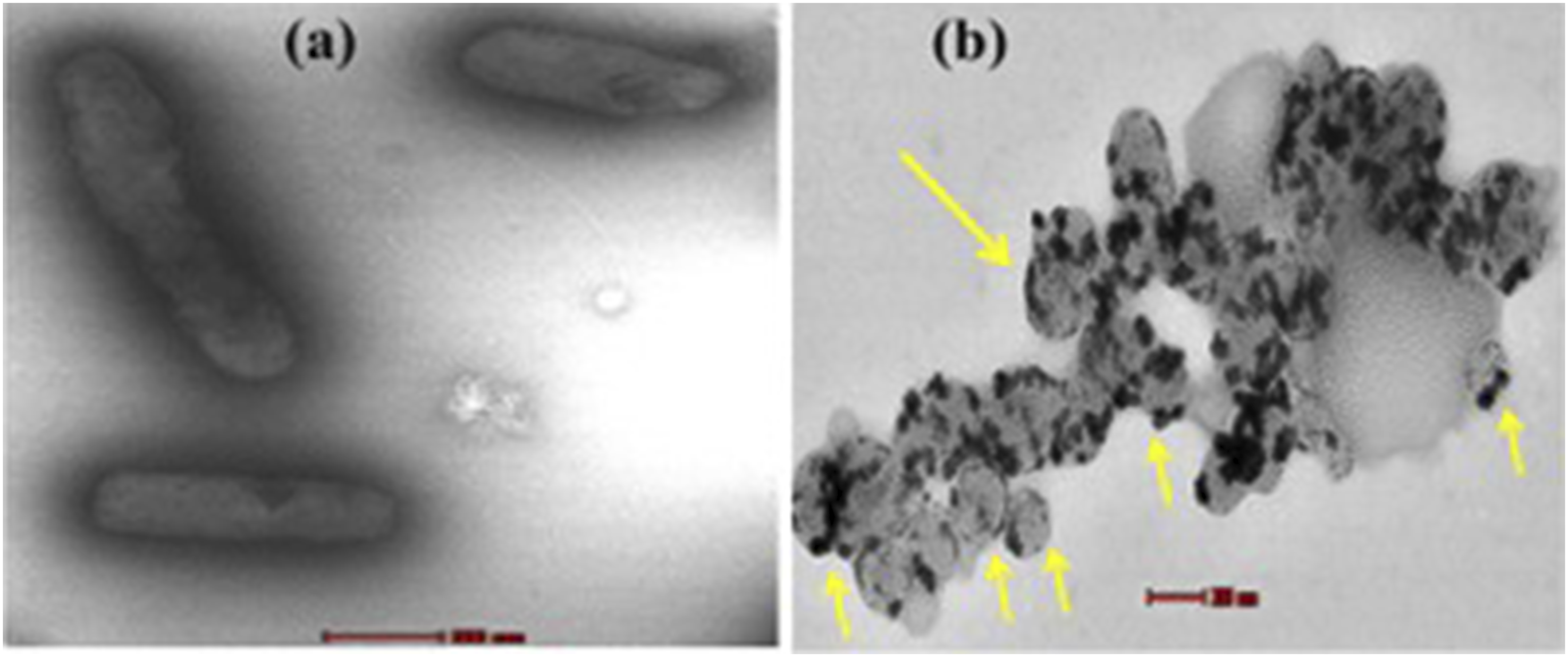
The TEM analysis revealed the cell complexity-dependent antimicrobial activity of PEI-AuNP@Van. Figure 9 demonstrates the cell wall-damaging action in C. albicans and C. tropicalis (Figure 9(a)–(d)). However, the selective surface binding was noted in E. coli and P. aeruginosa cells (Figure 9(e)–(h) yellow arrow). The results confirmed that the PEI-AuNP@Van surface interaction dynamics and mechanochemical impact were different for fungi and Gram-negative bacteria (as shown in Figure 9). Additionally, an intriguing observation was made in a few E. coli populations treated with PEI-AuNP@Van; numerous spherical structures with surface-attached nanoparticles were visible (Figure 10), suggesting an PEI-AuNP@Van-induced apoptotic phenomenon in bacterial strains. Transmission electron micrographs of PEI-AuNP@Van-exposed microbial strains and the respective untreated control cells. (a) untreated control cells of C. albicans, (b) treated with PEI-AuNP@Van, (c) untreated control cells of C. tropicalis, (d) treated with PEI-AuNP@Van, (e) untreated control cells of E. coli, (f) treated with PEI-AuNP@Van, (g) untreated control cells of P. aeruginosa, and (h) treated with PEI-AuNP@Van. TEM micrographs of PEI-AuNP@Van treated E. coli cells showing apoptotic behavior; (a) cell control and (b) treated cell.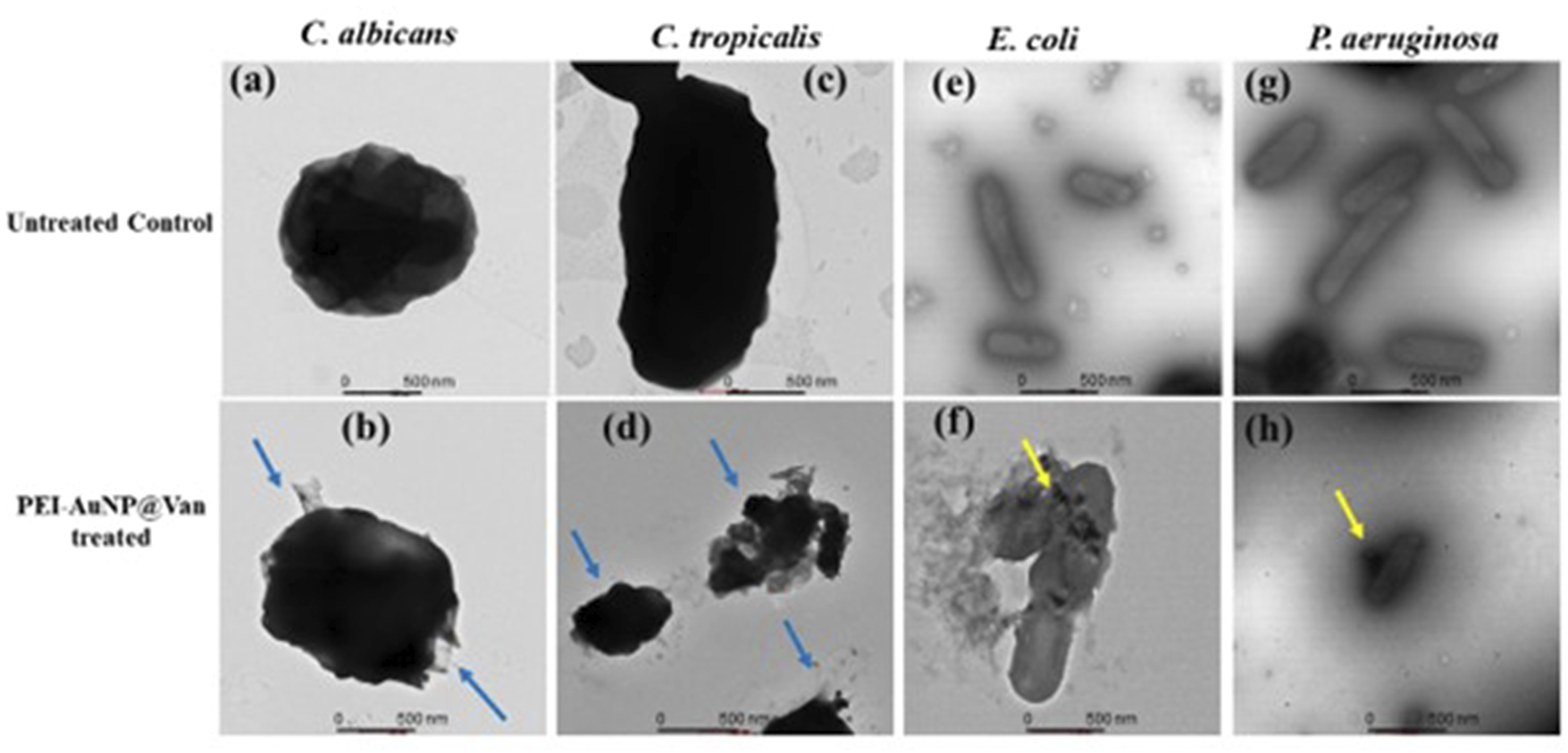

Cytotoxicity assay involving the HEK-293 cell line
The conventional MTT assay was utilized to investigate the mammalian cell cytotoxicity of the PEI-AuNP@Van nanosystem in cells from the Human Embryonic Kidney (HEK) 293 cell line. Specifically, HEK-293 cells were incubated with PEI-AuNP@Van at different concentrations; the results showed that PEI-AuNP@Van exhibited moderate concentration-dependent cytotoxicity (as demonstrated in Figure 11). PEI-AuNP@Van exhibited approximately 84% viability at a concentration of 10 µg/mL, whereas it had the highest cell toxicity (47%) at 100 µg/mL. These results indicate that PEI-AuNP@Van caused cell toxicity at higher concentrations. Biocompatibility of PEI-AuNP@Van particles against HEK cells.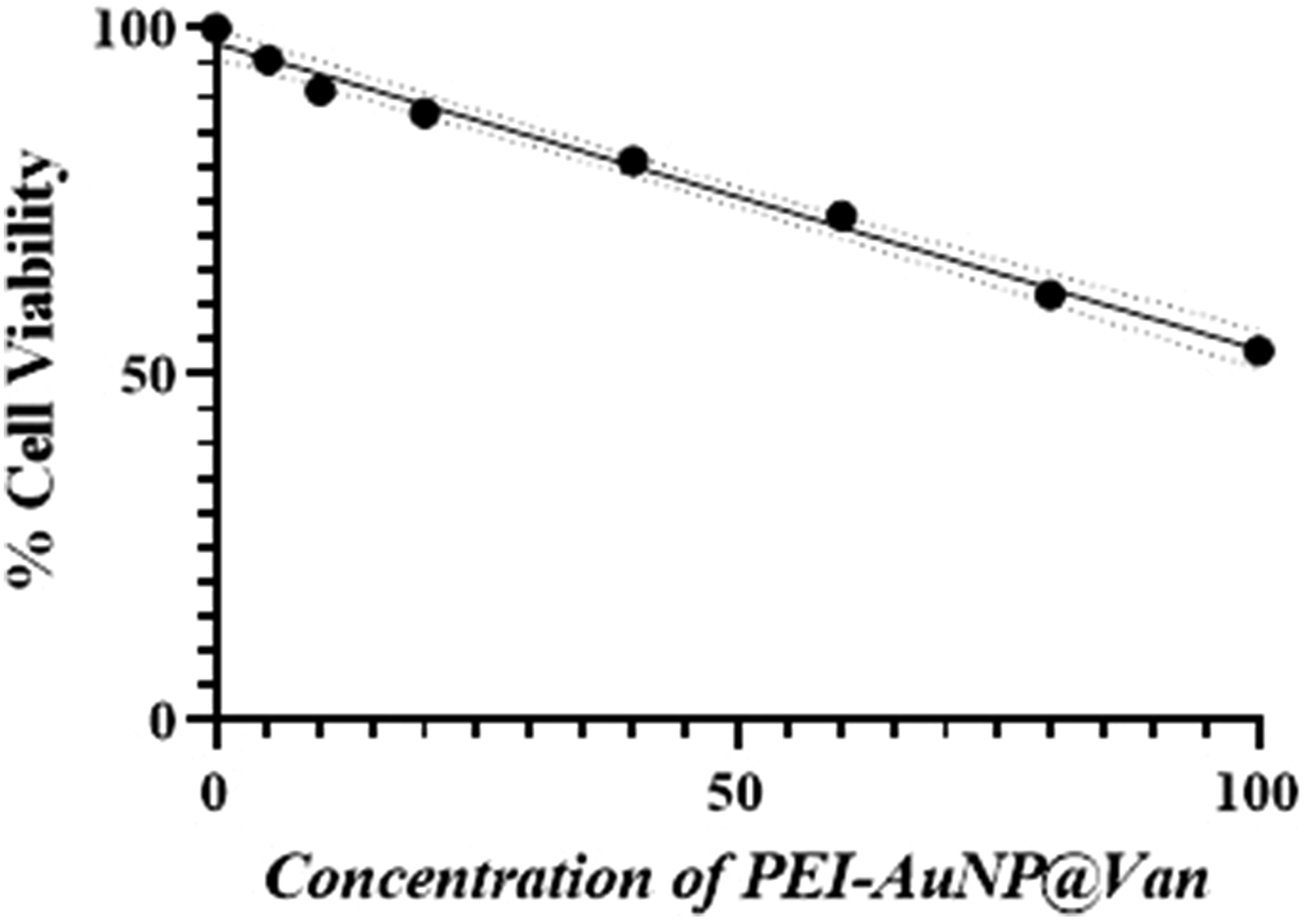
Discussion
AuNPs are useful in medical applications such as imaging and drug delivery. They can be linked to large molecules, including antimicrobial drugs, via surface interactions. The growing issue of antimicrobial resistance and declining efficacy of current drugs pose a global health threat, necessitating new antimicrobial alternatives for infections caused by Candida spp. And Gram-negative bacteria. Our team optimized a rapid protocol for developing fluorescent vancomycin-conjugated gold nanoparticles (PEI-AuNP@Van); the prepared nano-system was explored for its antimicrobial applications.32,33 In this system, vancomycin coordinates with gold cations and the NH- group of PEI interacts with the COO/OH groups of vancomycin via electrostatic interactions or hydrogen bonding, as confirmed by ATR-FTIR spectroscopy (Figure 2) and supported by Hagbani et al. 2022. 10 Further studies are required to fully understand these mechanisms. Vancomycin significantly affected the nanogeometry of PEI-AuNP@Van, resulting in the formation of spherical particles (<10 nm; Figure 1(d)). Electrostatic stability measurements using zeta potential studies revealed a high value (48 mV) at ambient temperature (Figure 1(c)) due to non-equivalent vancomycin conjugation with PEI-AuNPs, which leads to slight polydispersity (4-12 nm; Figure 1(d)) and results in a bimodal distribution of the zeta potential. To validate
These findings, AFM was conducted to understand the morphology and polydispersion of PEI-AuNP@Van. The Z-axis data from this analysis clearly indicated that the nanoparticles were spherical, with an average diameter that ranged from 4 to 10 nm (Figure 1(f)). The size distribution observed between these two characterization techniques can be attributed to the increased likelihood of nanoparticle conjugation with the silanized surface during the AFM measurements. The intrinsic fluorescence of vancomycin made PEI-AuNP@Van highly luminescent and excitation-dependent (λex = 270–360 nm; λem = 420 nm) with a quantum yield of 15.40%, making the materials traceable under UV exposure.32,33
Vancomycin inhibits RNA synthesis by forming hydrogen bonds with NAM/NAG residues, preventing their incorporation into the peptidoglycan matrix. 51 The outer lipopolysaccharide (LPS) layer of Gram-negative bacteria obstructs vancomycin access to the peptidoglycan layer, making it ineffective. Yeast cell walls, composed of chitin, proteins, and mannoproteins, differ structurally from bacterial cell walls. 52 PEI-stabilized metal nanoparticles interact with bacterial and fungal cell surfaces via electrostatic forces, permeabilizing the LPS layer and permitting vancomycin to reach the peptidoglycan layer, leading to cell death.53–56 PEI-AuNP@Van exhibited stronger antimicrobial effects against Candida spp. And Gram-negative bacteria than unconjugated PEI-AuNPs, while free vancomycin remained inactive (Table 1). C. tropicalis and P. aeruginosa were more sensitive to PEI-AuNP@Van compared to C. albicans and E. coli strains. Notably, C. tropicalis exhibited a significantly higher level of chitin than C. albicans. In contrast, the majority of Candida species showed similar levels of glucan in their cell walls, with C. tropicalis being the exception, demonstrating a markedly lower glucan content than C. albicans. 57 Hence, this cell wall structural variation probably led to the species-dependent mode of action of PEI-AuNP@Van. Cellular localization studies showed PEI-AuNP@Van inside C. albicans and C. tropicalis (Figure 3 and Supplement Figure S1), and on the surfaces of E. coli and P. aeruginosa, again indicating selective binding and antimicrobial action based on structural differences. High cell membrane integrity loss was observed in C. tropicalis (91.8%) and P. aeruginosa (85.8%) using propidium iodide staining and dead cell enumeration (Figures 4 and 5), while C. albicans and E. coli exhibited slower death rates after 4 h of exposure to PEI-AuNP@Van at MIC values. Unconjugated PEI-AuNPs also significantly killed C. albicans and E. coli strains, indicating a minimal role of vancomycin. Additional investigations are necessary to fully understand the species-specific mode of action of PEI-AuNP@Van.
The action of PEI-AuNP@Van on various strains was evaluated using assays and imaging techniques including ROS accumulation, apoptosis assays, Raman spectroscopy, and TEM. While essential for cellular functions, excessive ROS causes oxidative stress, leading to apoptosis or necrosis.58,59 Flow cytometry with DCFH-DA staining revealed cell type-dependent ROS induction profiles in treated strains (Figure 6 and Supplement Figure S3). PEI-AuNPs and PEI-AuNP@Van induced similar ROS levels in C. albicans and C. tropicalis, suggesting that vancomycin reduced ROS production (Figure 6(b)). However, ROS accumulation alone did not fully explain the PEI-AuNP@Van activity relative to the total dead population (Figure 5). 59 To explore alternative mechanisms, an apoptosis assay using annexin V-FITC/PI staining was performed. AuNPs are known to penetrate cell membranes and induce apoptosis-like death (including DNA damage, membrane depolarization, and caspase-like protein activation) in fungi and Gram-negative bacteria without increasing ROS levels. 59 Thus, multiple programmed cell death mechanisms may vary with microbial diversity. 60 During early apoptosis, phosphatidylserine translocates from the inner leaflet to the outer leaflet of the plasma membrane.61,62 Annexin V then binds to this externalized phosphatidylserine when in the presence of Ca2+ ions. In this double-staining approach, the early apoptotic cells were stained only by annexin V-FITC; necrotic and late apoptotic cells were stained by both PI and annexin V-FITC. An apoptosis assay using annexin V-FITC/PI staining investigated other mechanisms, as AuNPs can induce apoptosis-like death in fungi and Gram-negative bacteria without increasing ROS levels. 59 PEI-AuNP@Van-treated C. albicans cells exhibited early apoptosis properties (Figure 7(a)–(d)), while C. tropicalis, E. coli, and P. aeruginosa showed both early and late apoptotic/necrotic behavior (Figure 7(e)–(p)). This suggests ROS-independent apoptosis and cell death in C. albicans, 63 whereas C. tropicalis, E. coli (Figure 10), and P. aeruginosa undergo ROS-induced apoptosis, indicating multiple modes of action for PEI-AuNP@Van depending on the cell type.
Raman spectroscopy can be utilized to understand chemical bonding in biological macromolecules after interaction with metal nanoparticles, revealing significant structural and compositional changes in cells, including modifications of the nucleic acid phosphate backbone, disrupted protein hydrogen bonds, and reduced lipid molecule stacking forces. 64 In treated C. albicans, Raman bands shifting at 590, 676, 914, 1125, 1314, 1602, and 1653 cm−1 indicated alterations in chitin and unsaturated lipid chains (Figure 8(a)–(c)); this result was confirmed by TEM analysis showing damaged cell walls (Figure 9(a) and (b)). C. tropicalis exhibited similar Raman shifts at 526, 587, 852, 917, 1312, and 1592 cm−1, suggesting changes in chitin, lipid, and protein structures (Figure 8(d)–(f)), with complete cellular rupture observed under TEM (Figure 9(c) and (d)); this finding may be potentially due to excessive ROS accumulation, leading to cellular collapse. 65 E. coli and P. aeruginosa, despite similar cellular compositions, exhibit molecular differences in their Raman spectra (Figure 8(g) and (j)). Untreated E. coli (i), meropenem-treated (ii), and PEI-AuNP@Van-treated cells (iii) display peaks at 530-550 cm−1 and 848 cm−1, which correspond to peptidoglycan deformation and various molecular vibrations. Glycosidic linkage deformation supports the action of vancomycin, and spectral changes (e.g., a peak shift from 1122 cm−1 to 1114 cm−1) indicate membrane lipid and carbohydrate stretching 66 ; these findings are aligned with TEM images of peptidoglycan sheet fractures (Figure 9(e) and (f)). P. aeruginosa showed no significant Raman band shifts between treatments (Figure 8(j)), with minor changes at 533 cm−1 to 542 cm−1 and 786 cm−1 to 797 cm−1, which were related to peptidoglycan and lipid stretching. TEM revealed no cellular fracture in P. aeruginosa; however, PEI-AuNP@Van nanoparticles were adsorbed on the cell surface (Figure 9(g) and (h)). Additionally, an intriguing observation was made in a few E. coli populations treated with PEI-AuNP@Van; numerous spherical structures with surface-attached nanoparticles were visible (Figure 10), suggesting an PEI-AuNP@Van-induced apoptotic phenomenon in bacterial strains.
Conclusions
Vancomycin-conjugated PEI-stabilized gold nanoparticles (PEI-AuNP@Van) were synthesized and evaluated for antimicrobial activity against C. albicans, C. tropicalis, E. coli, and P. aeruginosa, along with their modes of action. PEI-AuNP@Van showed significant antimicrobial effects within 4 h, with MIC values of 4.8 µg/mL for C. albicans, 1.2 µg/mL for C. tropicalis, 2.5 µg/mL for E. coli, and 4.81 µg/mL for P. aeruginosa. These nanoparticles emitted blue fluorescence (418-430 nm), facilitating tracking of intracellular localization in C. albicans and C. tropicalis, and adsorption on E. coli and P. aeruginosa surfaces. Viability assays revealed that 16.3% of C. albicans, 91.8% of C. tropicalis, 12% of E. coli, and 85.8% of P. aeruginosa populations were killed within 4 h of treatment. PEI-AuNP@Van-treated cells accumulated endogenous ROS in 11.38% of C. albicans, 25.02% of C. tropicalis, 30.4% of E. coli, and 36.0% of P. aeruginosa populations. Phosphatidylcholine externalization indicated apoptosis-like behavior in all strains, with fungal strains showing early apoptosis. Raman spectroscopy and TEM analysis revealed multiple modes of action: ROS-dependent and independent apoptosis and cell wall damage in C. albicans and C. tropicalis, and ROS-induced apoptosis and peptidoglycan sheet deterioration in E. coli and P. aeruginosa. PEI-AuNP@Van nanoparticles effectively targeted a broad range of pathogenic microorganisms. However, comprehensive biocompatibility evaluations (both in vitro and in vivo) are essential. The efficacy of the PEI-AuNP@Van nanoparticles against planktonic cells has been documented, but their effectiveness against biofilms of mono- or mixed-species remains to be studied.
Supplemental Material
Supplemental Material - Making vancomycin a potent broad-spectrum antimicrobial agent using polyaziridine-stabilized gold nanoparticles as a delivery vehicle
Supplemental Material for Making vancomycin a potent broad-spectrum antimicrobial agent using polyaziridine-stabilized gold nanoparticles as a delivery vehicle by Atul Kumar Tiwari, Aishwarya Nikhil, Avinash Chaurasia, Prem C. Pandey, Roger J. Narayan and Munesh Kumar Gupta in Journal of Biomaterials Applications.
Footnotes
Acknowledgments
The authors would like to thank the Central Instrument Facility, Indian Institute of Technology (BHU), for characterization of the nanoparticles, the Central Discovery Centre (CDC), Banaras Hindu University, for providing access to flow cytometry and Raman spectroscopy, and SATHI BHU for laser scanning super-resolution microscopy. The authors also thank SAIF, AIIMS, New Delhi, India, for TEM imaging of biological samples.
Authors Contribution
AN and AKT contributed equally. Conceptualization: AKT and MKG; Data curation: AKT; Formal analysis: AKT; Funding acquisition: MKG and RJN; Investigation: AN and AKT; Methodology: AN, AKT and MKG; Project administration: MKG and RJN; Resources: MKG, AC & PCP; Software: AKT; Supervision: PCP, and RJN; Validation: AKT and MKG; Visualization: AN, AKT and MKG; Roles/Writing - original draft: AKT; Writing - review and editing: AKT and RJN. All of the authors reviewed and agreed to this submission.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is partially supported by an Incentive Grant under IoE to Dr. MK Gupta (R/Dev/D/IoE/seed & Incentive grant_III/2022-23/49235).
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.