Abstract
This is a visual representation of the abstract.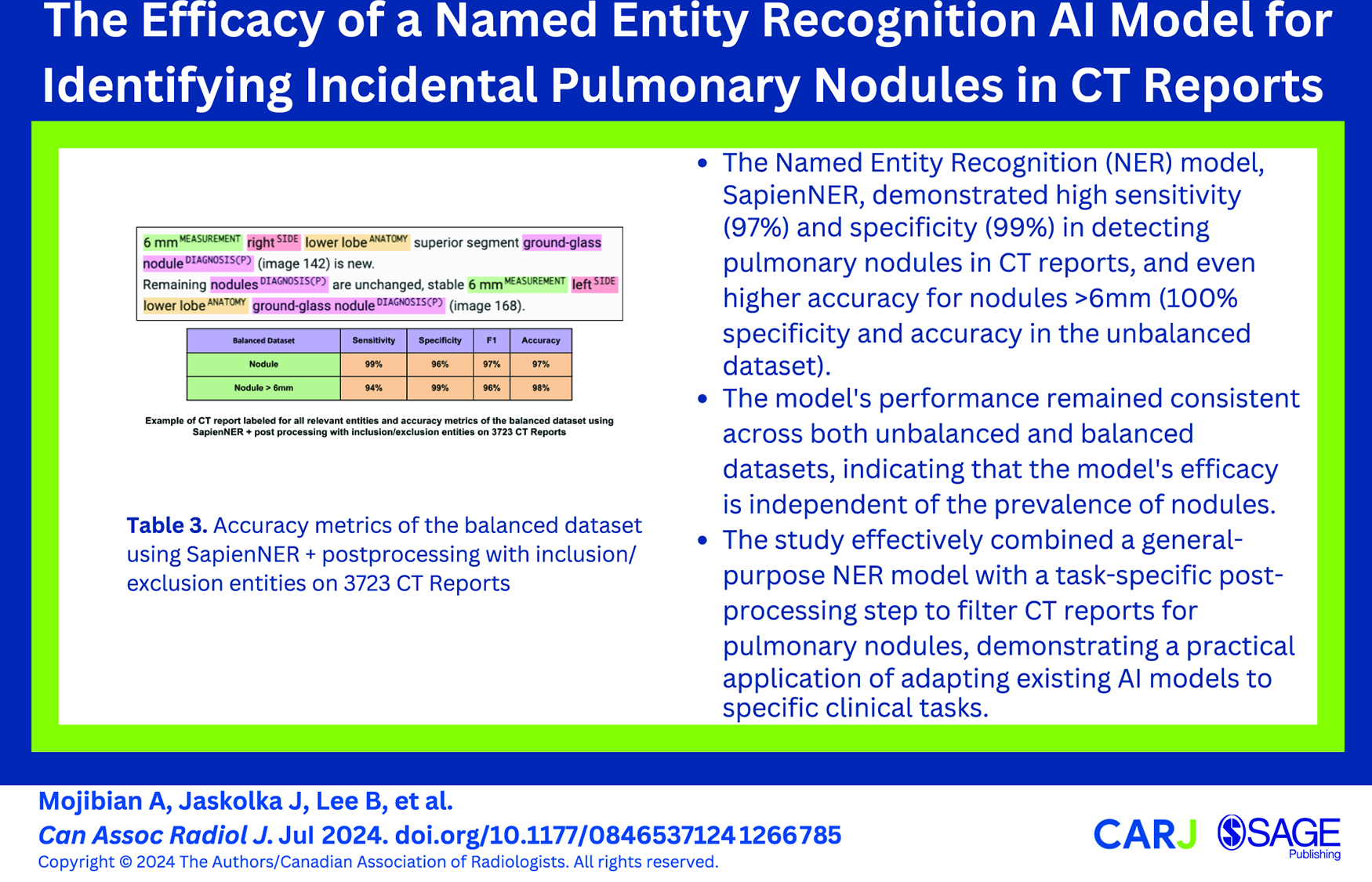
Introduction
As the leading cause of cancer mortality globally, lung cancer accounts for an estimated 1.8 million deaths annually. 1 Lung cancer screening programmes have demonstrated significant mortality reduction by detecting lung cancer at an earlier stage—often at stage 1 disease, defined as a pulmonary nodule less than 10 mm in diameter. Incidentally detected pulmonary nodules (IPNs), detected during CT scans that include some or all of the lungs, offer an important avenue for early lung cancer detection. Although the majority of IPNs are benign, an estimated 3% to 4% represent early-stage, potentially curable lung cancer. 2 The incidence of chest CT scans revealing lung nodules is notable, and these patients can be lost to follow-up, underscoring the need for streamlined, automated management. 2 The radiologist’s report of the imaging study is pivotal in determining subsequent patient management. 3
Despite the available literature advocating for standardized lexicons and structured reporting to improve clarity and consistency,4,5 the preference for narrative-free text persists in clinical practice, primarily due to the ease of its use in conveying nuanced clinical information. 3 The widespread adoption of Electronic Health Records (EHRs) has improved access to radiology reports, yet their unstructured format often impedes the precise identification or easy extraction of pertinent clinical information by referring physicians. In a study of Danish hospitals, 45.3% of pulmonary nodules mentioned in radiology reports were not followed up appropriately; reportedly, these unfollowed nodules later came to represent 2.5% of stage IV cases that could have been detected earlier with vigilant surveillance. 6
The ongoing growth in the volume of imaging studies will inevitably lead to increased detection of incidental findings requiring further evaluation and management. 7 This necessitates innovative methodologies that can offer consistent and expedited identification of important and actionable imaging findings from reports to reduce the variability inherent in human analysis of these reports. Automated extraction of important findings from unstructured reports can be used to automatically initiate appropriate notification and follow-up protocols, thereby decreasing the burden on clinicians while improving the quality of care that patients receive. 8
In the field of Natural Language Processing, Named Entity Recognition (NER) is the task performed by a machine learning model specifically trained to identify and classify parts of the text into “entities” such as names of people, organizations, and locations.8-10 In the case of NER models trained on medical information, the models identify and classify entities such as anatomy, positive/negative diagnosis, medication, and imaging modality. By themselves, medical NER models have established use cases for information retrieval and content categorization. However, when combined with a complementary relation extraction model, the semantic relationships between entities identified by the NER model can be established. The identification of entities and the following step of classification; the establishment of relations between those entities is referred to as Named Entity Recognition and Classification (NERC), yet, the terms NER and NERC are frequently used interchangeably. 10
Machine learning now often involves task and domain-adaption of pre-trained models like ULMFit and BERT11,12 through a fine-tuning step of these base models using additional supervised learning. This requires the use of specifically labelled datasets, which are costly and time-consuming to gather and validate. Also, costly GPU hardware is needed. As models grow, fine-tuning becomes more time-consuming and expensive. Hence, there is a need for a faster, cheaper approach to adapt models for new tasks.
The goal of this study was to test the efficacy of SapienNER, a commercially available general-purpose medical NER Model. The study tested SapienNER as a screening tool for pulmonary nodules identified in CT reports when used in combination with a site-specific post-processing step to filter CT reports to those with pulmonary nodules that potentially merit further clinical evaluation. 13
Methods
We conducted a retrospective study to determine the efficacy of combining a general-purpose NER model with a task-specific post-processing step to be used as a screening tool for pulmonary nodules identified in the reports of CT scans at our institution. The Institutional Research Ethics Board approved this study of our hospital with a waiver of informed consent.
Data Collection
Reports of all CT imaging studies that included a portion of the chest (lung-in-view) performed between August 2023 and November 2023 were identified. There were no exclusion criteria.
Extraction and De-Identification Process
A total of 9165 unstructured CT imaging reports were extracted from the electronic medical record (EMR) de-identified using custom commercial software (SapienSecure v2.3, Vancouver, Canada) and exported to a CSV file (Figure 1). The reports found in the EMR were formatted under headers such as Exam Type, History, Comparison, Technique, Findings, and Impression, with some offering additional headers specific to the anatomy undergoing diagnostic imaging. Each report in the dataset was manually reviewed by one of the authors (AM) and categorized into one of 3 categories: no nodules present, any nodule present, or nodule greater than 6 mm diameter present.

Workflow for de-identification of CT radiology reports.
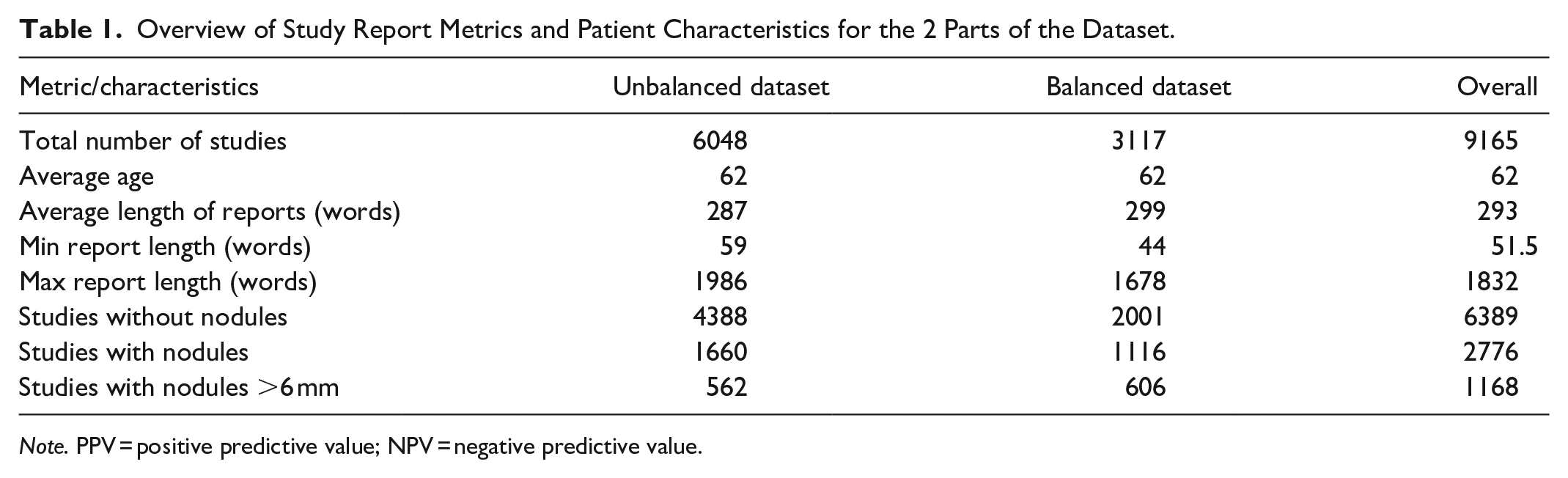
Due to the rarity of pulmonary nodules, the total number of studies without any nodules was expected to significantly exceed the number of studies with nodules present. This class imbalance can result in misleading metrics that are potentially skewed by the incidence of disease rather than the quality of the model. To address this, the total dataset was divided into an unbalanced dataset with an expected low incidence of nodules and a balanced dataset in which the number of reports with no lung nodule, any lung nodule, and any nodule >6 mm were equalized. 14 To prevent and data-leakage, the 2 datasets were selected from different time periods. The unbalanced dataset (6048 reports) consists of reports from August to September 2023, and the balanced dataset (3117 reports) consists of reports from October to November. Some patient and report characteristics are provided in Table 1.
Overview of Study Report Metrics and Patient Characteristics for the 2 Parts of the Dataset.
AI Model
SapienNER is a custom-developed commercially available general medical NER model which is based on the RoBERTa architecture. 12 It was adapted to named entity recognition by fine-tuning on an open-source dataset of over 7 million radiology reports, clinical reports, and discharge summaries across 3 geographical locations to deliver outputs across multiple entity categories, including anatomy and positive/negative diagnoses. 15 It was then further fine-tuned on a proprietary dataset of radiology reports and manually labelled for the named entity recognition and relation extraction task. The core model was not further fine-tuned or task-adapted for the task of lung nodule detection. The goal of this paper was not to develop a tailor-made model that is capable of doing one specific task very well but to evaluate the use of an existing model for usage in a retrospective cohorting system after filtering its outputs to provide task-specific results.
Named Entity Recognition and Classification (NERC)
NERC was performed on each of the de-identified reports. The reports were a mix of unstructured and structured reports, as dictated at the discretion of the reporting physician. As is typical in practice, the unstructured reports still had headers and sections to indicate the history, technique, report findings, and conclusion, even if the report section itself did not contain further structure. Inference was only done on the body and conclusion of the report and excluded the history, indication, and technique subsections. This was done to ensure that any nodule-related entities were reported as found on the images rather than mentioned in the meta-data of the report. For each document, the model identified all possible entities and provided the relationships between these entities (Figures 2 and 3).

Example of CT report labelled for all relevant entities—Medication, Imaging, Anatomy, Side, Diagnosis (positive), Diagnosis (negative), and measurement.
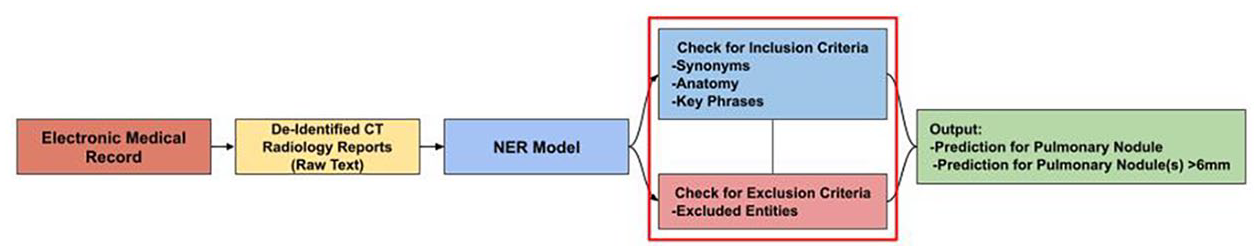
Summary of the automated data workflow for analyzing de-identified CT radiology reports to extract relevant entities and make predictions on pulmonary nodules using the Pulmonary Nodule Inclusion/Exclusion Criteria.
Pulmonary Nodule Inclusion/Exclusion Entities
To translate the general results of SapienNER to the specific use case of pulmonary nodule detection, a post-processing step was introduced. This step used a list of inclusion and exclusion entities created based on the specific lexicon of terms used to describe pulmonary nodules in CT reports (see Figure 4). The entities were chosen to maximize the sensitivity of nodule identification of reports, with the understanding that some of the identified nodules would potentially not be clinically significant. The size cutoff was chosen based on the Fleishner Society Criteria. When applied, the post-processing of NER results with these entities allowed for the categorization of studies into 3 categories: (1) those without nodules, (2) those with any pulmonary nodule, and (3) those with pulmonary nodules >6 mm. All detected lung nodules were included in the cohort, independent of reported stability relative to prior studies. The process of initial determination of the inclusion/exclusion criteria and the development of the code to process the NER model output was completed over several days.

Pulmonary nodule inclusion/exclusion entities.
Pulmonary nodule was a type of “diagnosis” entity. Various diagnosis entities considered equivalent to the pulmonary nodule class are presented in Figure 4. For any of these entities to be included for consideration, they were required to be related to an anatomy entity relevant to the lung and to not be related to or among any of the exclusion entities. For example, if a nodule entity was found to be linked to an anatomy entity such as “segment II of the liver,” it was excluded. However, if a nodule was found to be linked to the anatomy entity “Right Upper Lobe,” it was included. The same logic was applied for a size threshold of >6 mm. For example, if a pulmonary nodule entity was related to a measurement entity found to be less than 6 mm, it was categorized differently than a nodule related to a measurement entity of >6 mm. The 6 mm threshold was selected to be consistent with the widely accepted Fleischner Society Guidelines for evaluating the significance of incidentally detected pulmonary nodules. 13 All other entities identified by SapienNER but not included in Figure 4 and not related to a pulmonary nodule entity were ignored for this study.
Evaluation Metrics
To assess the performance of the model, the outputs were compared to the manually labelled ground truth. Binary performance of the system for determining nodule versus no nodule and nodule >6 mm versus no nodule was assessed. Sensitivity, specificity, precision, F1, and accuracy scores were calculated. Metrics were computed compared to the ground truth using Python and Scikit Learn. 16
Results
Raw, Unbalanced Dataset
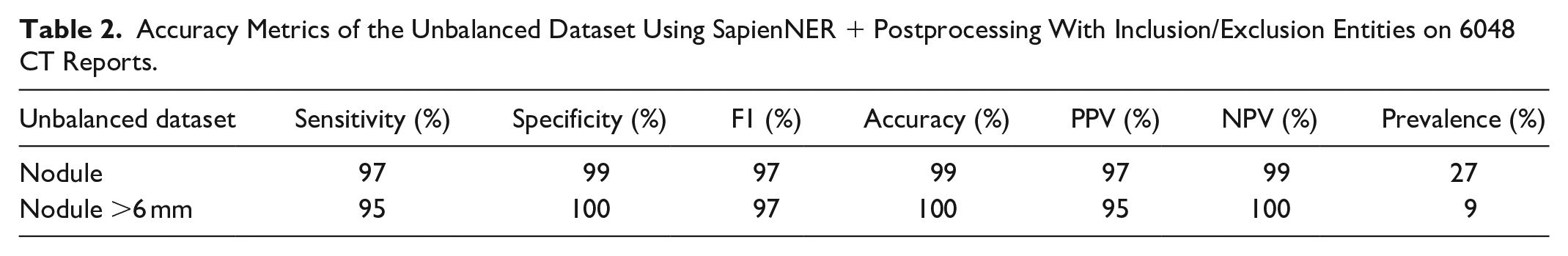
For the initial phase of the study, the dataset was unbalanced; in this form, the dataset reflected the real-world prevalence of pulmonary nodules in CT imaging reports, where the instances without nodules far outnumbered those with nodules. Unweighted metrics were used in the analysis to evaluate the model’s raw performance metrics without adjusting for the expected prevalence of nodules. The results of this analysis are presented in Table 2.
Accuracy Metrics of the Unbalanced Dataset Using SapienNER + Postprocessing With Inclusion/Exclusion Entities on 6048 CT Reports.
Balanced Dataset
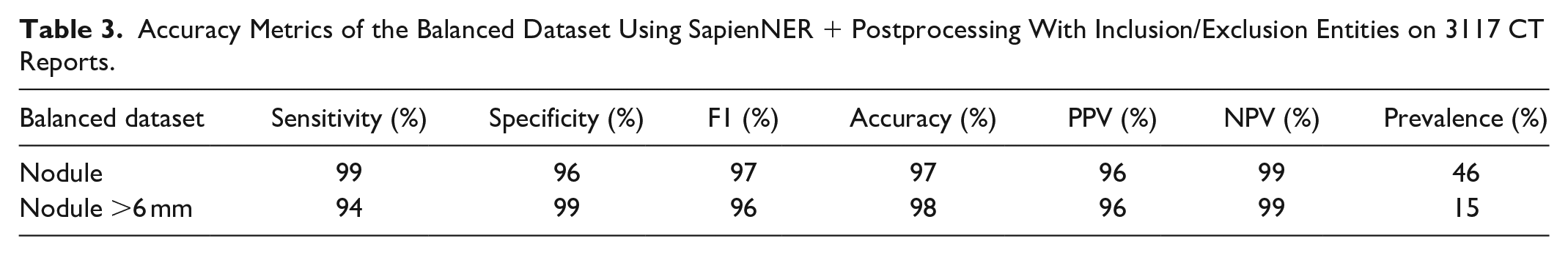
For the balanced dataset, unweighted metrics were calculated. The results of this analysis are presented in Table 3.
Accuracy Metrics of the Balanced Dataset Using SapienNER + Postprocessing With Inclusion/Exclusion Entities on 3117 CT Reports.
Discussion
This study demonstrates the efficacy of the medical NER model for the detection of reported pulmonary nodules in a real-world dataset. Using a generic medical NER model—SapienNER—post-processed with task-specific inclusion/exclusion criteria, we obtained excellent performance for the detection of reports describing lung nodules as well as classifying those reports into those that contained nodules >6 mm and those that did not. When applied to an unbalanced, real-world dataset of a combined dataset of 6048 CT radiology reports, SapienNER with simple post-processing was successful at identifying CT reports identifying any pulmonary nodule with a sensitivity of 97% and an accuracy of 99%. The system further identified reports of pulmonary nodules measuring >6 mm with a sensitivity of 95% and an accuracy of 100%.
Comparative literature on the use of established NER systems or the development of NER systems for the detection of pulmonary nodules is limited, with only 2 studies exploring the ground-up development of full NLP models.9,17 French et al 2019, developed 2 dedicated models, one “Nodule Model” and the other “Sizing Model” with a combined F1 score of 72.9%, recall of 60.3%, and precision of 90.2%. 18 These results are likely not adequate to warrant clinical implementation. One explanation for the improved results in our study would be the differences in model architecture and degree of development. French et al used an open-source, rule-based NLP tool named SimpleNLP as opposed to our study, which used a market-ready Medical NER model, SapienNER. Where SimpleNLP required a fine-tuning step, SapienNER was deployed without modifying the base model and achieving task specificity through the implementation of a post-processing step on the model outputs based on specific inclusion/exclusion criteria.
Potential Clinical Significance
The results of this study show this model’s potential for ensuring the appropriate follow-up process is initiated for patients with high-risk nodules as defined by the Fleischner Society Guidelines. 18 It is known that the majority of incidental nodules are benign, but the 3% to 5% that are malignant require recognition and appropriate follow-up. It is possible for NER to search thousands of reports in a few minutes to identify nodules requiring clinical follow-up. If a system like this could be implemented in a hospital RIS/PACS or outpatient EMR, it could automatically identify patients at risk of malignancy and trigger a manual notification or callback process. This, in turn, could decrease the number of patients lost to follow-up and result in a decreased number of preventable missed or late-stage cancer diagnoses. 6 This type of tool does not currently exist in a Canadian context and could be an important step forward to identify IPNs consistently, reducing clinician time, and ensuring no clinically relevant nodules are omitted.
This study demonstrated the ease with which an existing generic medical NER model can be adapted to assist with nodule detection and classification. Since the task-adaption was simply done by determining which entities would be included it was just a matter of applying this criteria to the generic output. This approach is significantly faster and cheaper than developing or fine-tuning custom, single-task models which can take weeks or months to accomplish when considering the time and money required for data collection, data curation, model development, and model training. Our study also highlighted the ease of creating systems for other NER-type problems including detection and flagging of other incidentally detected abnormalities on CT scans. Taking into consideration, the ever-expanding number of CT imaging studies being performed daily 19 and the high number of potentially significant incidental findings on these studies 20 an automated system for flagging needed follow-up studies is potentially very important. Although computer vision systems have been proposed for this type of task, using an NLP-based system on imaging reports as opposed to a computer vision system on the images has the advantage of easier development, easier deployment, faster inference time, and decreased burden on PACS databases. It also leverages the existing expertise and workflow of radiologists who are already providing an interpretation of these studies with high accuracy.21-24
Implications of Using Unweighted Metrics
The use of unweighted metrics in both unbalanced and balanced datasets offers a transparent view of SapienNER’s performance. In the unbalanced dataset, unweighted metrics highlighted the model’s capability to manage the skewed distribution of pulmonary nodules within CT reports, typical of daily practice. Meanwhile, the application of these same metrics to the balanced dataset allowed for a focused examination of the model’s precision and reliability when class prevalence was eliminated as a confounding factor in the assessment of the model’s performance. This dual approach underscores the model’s robustness and versatility in different analytical contexts. By including an analysis of the balanced dataset, we ensured that the evaluation of SapienNER was comprehensive and that it encompassed both the challenges of real-world data and the controlled conditions of balanced datasets.
One important limitation of this study is that it was performed at a single centre. Although the reporting language is generally quite similar across institutions and though the training data used for this study originated from multiple centres, further studies could incorporate multi-site validation to ensure the generalizability of the model. This study also did not incorporate other nodule risk assessment systems like “LUNG-RADs” or the Pan-Canadian Early Detection of Lung Cancer 25 which are frequently used for dedicated cancer screening studies. As such, the results of this study would be most applicable to detecting reports that have found incidental pulmonary nodules. However, future studies could explore the integration of multiple assessment systems and the use of models such as this in the lung cancer screening setting to help ensure appropriate risk stratification and follow-up of screen-detected pulmonary nodules.
Another limitation of the study was that the defined task was to identify any described pulmonary nodule, independent of its imaging characteristics (other than size) and the clinical context in which it was detected. This was chosen to maximize sensitivity for nodule detection and demonstrate the capability of an AI model to extract the presence of pulmonary nodules from reports while allowing clinical relevance to be determined by the referring physicians. Future studies could address the ability of altering the included and excluded entities of this system to specifically identify clinically relevant nodules based on parameters such as morphology, clinical context, and composition.
Finally, since the NLP approach is dependent on nodules being reported by the radiologist, it is possible that small nodules might not be included in the report. This could affect the rate of detection of nodules less than 6 mm. However, it would be very uncommon for a nodule >6 mm to be ignored by the reporting radiologist so the “clinically significant” nodule results would be unaffected by this concern. Future studies could also evaluate the combined use of NER models with Computer Vision to (a) allow for the extraction of valuable features from CT DICOMs and (b) further validate the findings of the NER model which is highly dependent on radiologist detection of reports.26-29
Conclusion
In summary, this study demonstrates the efficacy of a simple task-adaptation approach in conjunction with a commercially available medical NER model for detecting and size-cohorting of incidental pulmonary nodules from unstructured CT reports. Although further clinical studies are needed, this study shows the potential for this type of system for use in developing a robust follow-up strategy and preventing missed follow-up of potentially malignancy lung nodules. This study represents an important step in confirming the potential use of task adaptation of available NLP models for use in a healthcare setting with minimal effort in comparison to significant fine-tuning or model development from scratch.
Footnotes
Authors’ Note
Institution: Department of Radiology, University of British Columbia, UBC Radiology, Vancouver General Hospital, Gordon and Leslie Diamond Health Care Centre, 2775 Laurel Street, Vancouver, BC, Canada V5Z1M9.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Dr. William Parker owns the software IP used in this study, which is called SapienNER. No financial interest is associated with this study that would compromise its integrity. Dr. Savvas Nicolaou owns the software IP used in this study, which is called SapienNER. No financial interest is associated with this study that would compromise its integrity. Mr. Brian Lee owns the software IP used in this study which is SapienNER. No Financial Interest is associated with this study that would compromise its integrity. Mr. Alireza Mojibian is an employee at Sapien Machine Learning with no financial interest associated with the software IP that would compromise its integrity. Ms. Chloe Devine is an employee at Sapien Machine Learning with no financial interest associated with the software IP that would compromise its integrity.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
