Abstract
This is a visual representation of the abstract.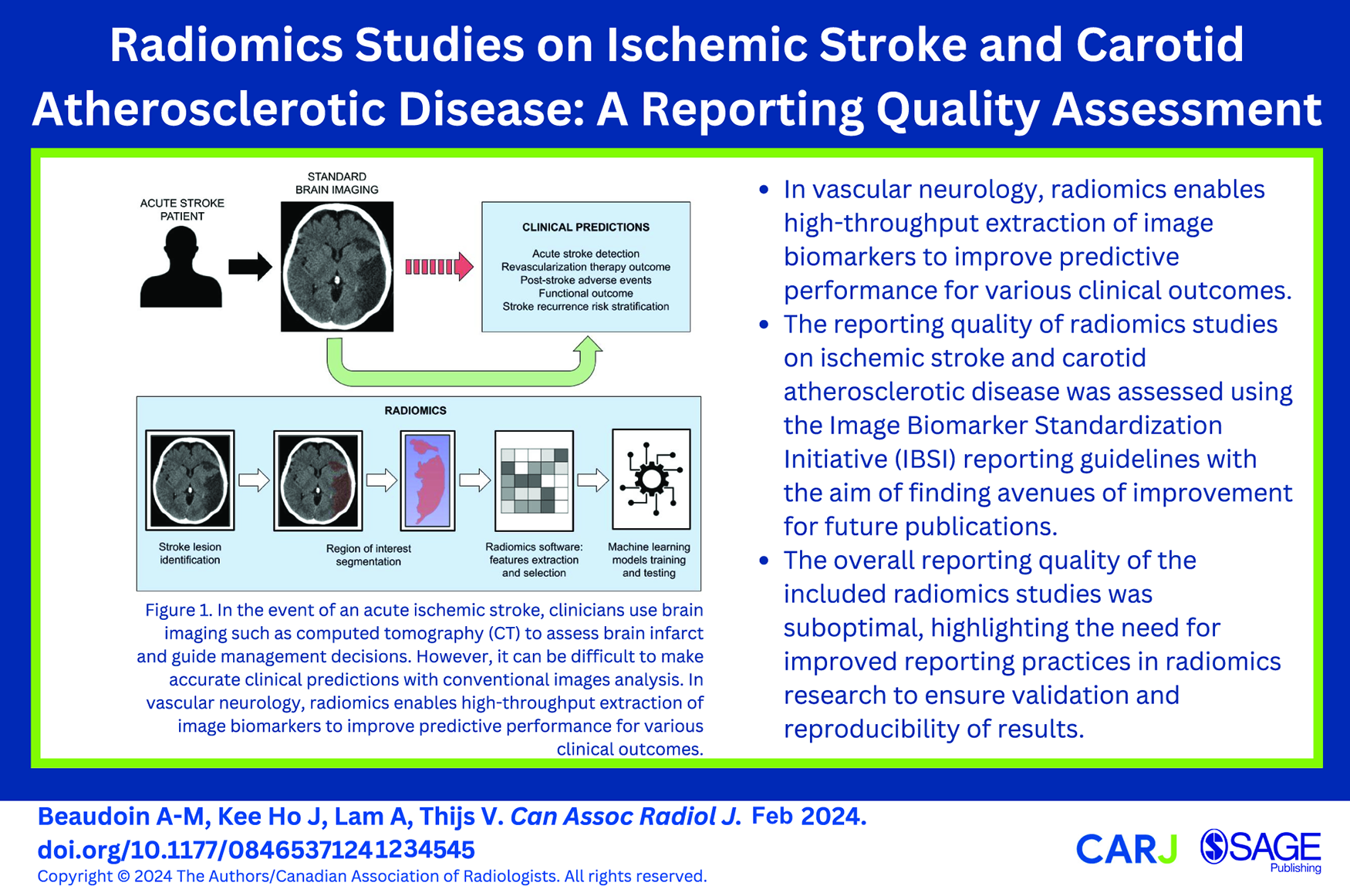
Introduction
Radiomics is an emerging field in medical imaging that involves the high-throughput extraction and analysis of quantitative features from medical images, such as computed tomography (CT), magnetic resonance imaging (MRI), or ultrasonography (US). 1 These features include shape, texture, intensity, and other characteristics that are often imperceptible to the human eye. The primary goal of radiomics is to convert medical images into mineable data, allowing for a more detailed and comprehensive analysis of various pathologies. By utilizing advanced computational techniques, it enables the identification of subtle patterns and biomarkers within images. These radiomics features can be correlated with clinical outcomes, response to treatment or prognosis, providing valuable insights into disease characterization, patient stratification, and personalized medicine.2,3
Initially developed in the field of oncology, radiomics has been increasingly applied in ischemic stroke research.4-7 The main applications of radiomics in ischemic stroke are acute lesion detection, outcome prediction after reperfusion therapies such as intravenous thrombolysis and mechanical thrombectomy, post-stroke adverse events risk identification, and long-term functional and cognitive outcome prediction. Moreover, radiomics holds promise in the evaluation and management of atherosclerotic disease. By identifying vulnerable plaques, radiomics could predict the risk of ischemic events beyond the degree of arterial stenosis. In the era of precision medicine, radiomics features could therefore facilitate personalized management strategies, including the use of new anti-inflammatory medications in non-surgical cases.
The key steps of the radiomics workflow are summarized in Figure 1. As radiomics is a relatively new tool for image analysis, lack of standardization, reproducibility, and validation is a challenge. To overcome the translational gap between radiomics advancements and clinical applications, researchers need to provide detailed manuscripts allowing replicability of methods and results. The Image Biomarker Standardization Initiative (IBSI) is an international collaborative effort aimed at standardizing the process of extracting image biomarkers from radiological images. 8 The IBSI provides definitions, image biomarker nomenclature, reporting guidelines as well as benchmark datasets and values to validate image processing and calculations (https://theibsi.github.io/). Since its publication in 2020, IBSI has gained recognition and acceptance within the field of radiomics, with leading experts suggesting that researchers should adhere to the information and framework provided through IBSI.9,10
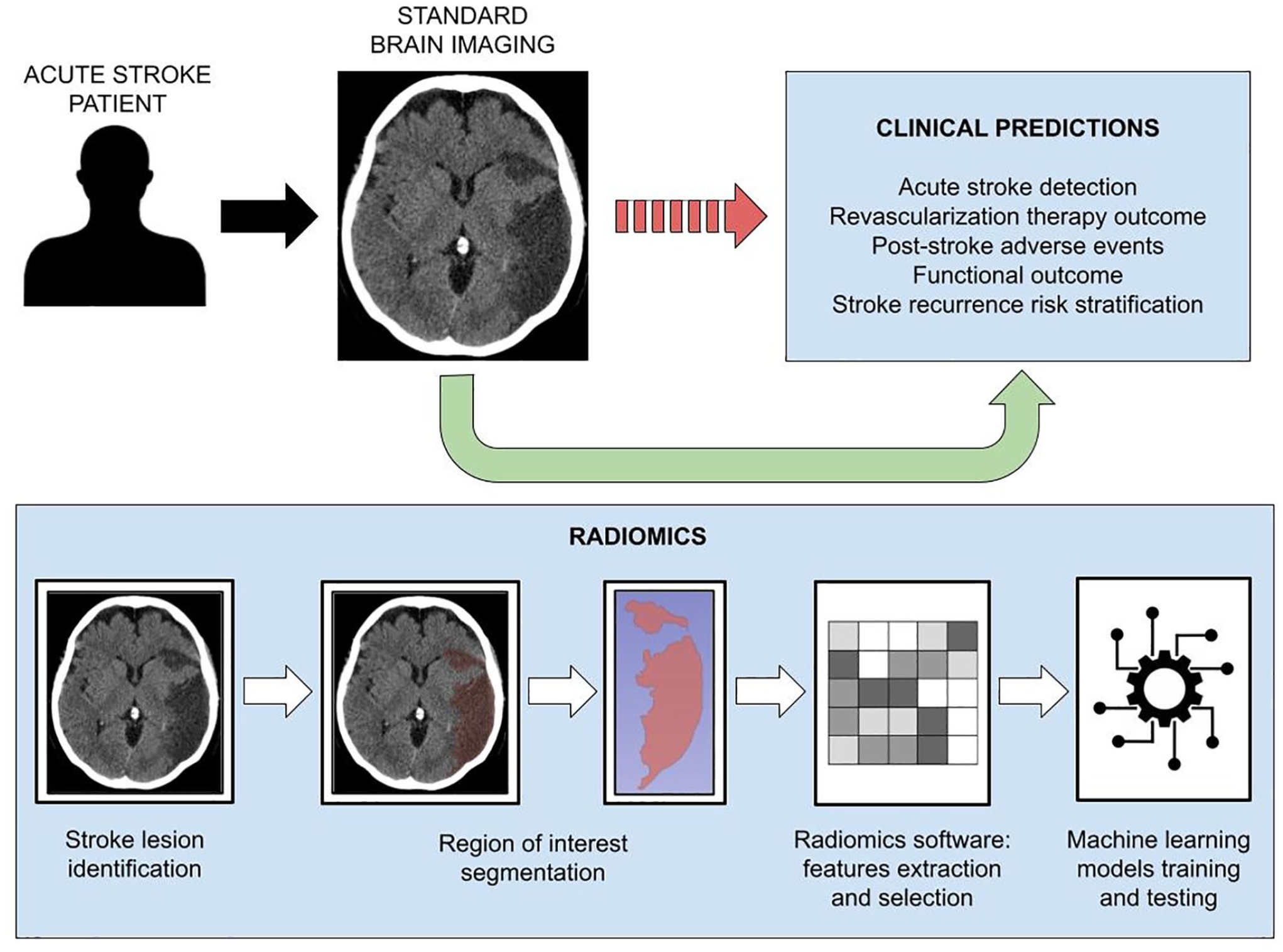
Key steps of the radiomics workflow.
Radiomics reporting guidelines are particularly helpful for authors to provide a complete report with reproducible results and can facilitate standardization and comparison of radiomics studies. Using the IBSI reference manual and reporting guidelines, we reviewed radiomics studies on ischemic stroke, intracranial artery disease, and carotid atherosclerotic disease. The aim of this work is to assess the reporting quality of these publications and provide recommendations to improve the quality of future reports.
Methods
Search Strategy and Study Selection
Using a systematic search strategy, the PubMed database was used to search all original research papers published in English before 1 September 2022. The terms utilized for the search are described in Supplemental Material 1. A total of 560 studies were imported into an online systematic review platform (Covidence, Veritas Health innovation Ltd, Melbourne Australia). After the removal of 73 duplicates, 487 studies were screened for eligibility according to the following inclusion criteria: (1) patients aged 18 years and older, (2) study design is either observational (cohort study, case-control study, cross-sectional study) or randomized controlled trial, and (3) study uses radiomics to investigate ischemic stroke-related issues (acute stroke detection, outcome prediction, adverse events identification) or atherosclerotic disease (carotid or intracranial arteries). Forty-one studies were included in the final analysis (Figure 2).11-51
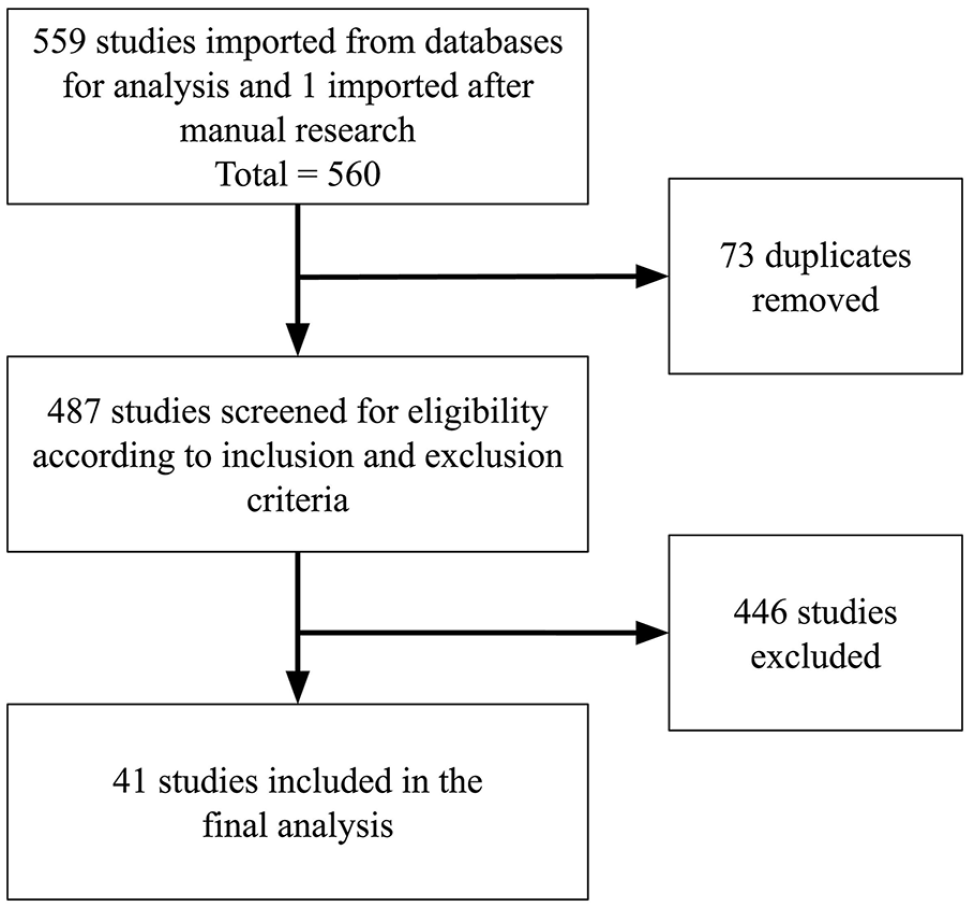
Flowchart of study selection.
Image Biomarker Classification
In radiomics studies, the aim is to identify a combination of these factors to provide a comprehensive understanding of a patient’s condition, guide treatment decisions, and estimate long-term outcomes. The selection of which factors to focus on depends on the specific research question of a study. Various types of factors can be considered to make clinical predictions, and these factors can broadly be categorized into diagnostic, predictive, and prognostic factors. However, these factors’ definitions may overlap, and the distinction between them can be subtle. Diagnostic factors help in distinguishing between different types of diseases or conditions. They are primarily used to aid in the initial diagnosis or classification of a patient’s condition. In this work, we included studies on ischemic stroke lesions detection, infarction onset time determination, high-risk atherosclerotic plaques identification, and studies differentiating intracranial contrast extravasation from intraparenchymal hemorrhage. Predictive factors provide information about how a patient is likely to respond to a particular treatment or intervention. Studies aimed at predicting successful recanalization after intravenous thrombolysis or mechanical thrombectomy were included in this category, as well as studies predicting post-stroke complications like malignant edema and hemorrhagic transformation. Finally, prognostic factors are used to estimate the patient’s prognosis. In this analysis, we included studies aimed at predicting functional outcome or cognitive impairment after stroke.
Reporting Quality Assessment
Using the IBSI reference manual and radiomics reporting guidelines, a reporting checklist for CT-based and MRI-based studies was created to verify reporting guidelines adherence (Tables S2 and S3). The objective was to investigate which items are most often missing or incomplete and which items are globally well reported. This analysis was performed individually by 3 investigators (AM.B., J.K.H., and A.L.) and results were then compared. Disagreements were resolved by discussion and consensus. The results are presented as raw scores, median and range or interquartile range (IQR), or percentages when appropriate.
Results
Characteristics of Radiomics Studies
The main characteristics of the 41 included studies are summarized in Table 1 and more details are provided in Table S1. The median number of cases per study was 141 (IQR 85-223), with a median of 106 (IQR 111-180) cases in the training dataset and 48 (IQR 42-79) cases in the validation dataset, for a ratio of 2.2:1. A total of 36 (88%) studies had a retrospective design and 32 (78%) used data from a single centre. Among the 29 studies with ischemic stroke as the main topic, 14 (48%) used CT images and 15 (52%) used MRI. When studying intracranial artery disease, MRI was utilized in all 3 studies. For the 9 studies on carotid artery disease, 6 (67%) publications used CT, 2 (22%) used MRI, and only 1 (11%) study used US images.
Characteristics of the 41 Included Radiomics Studies.
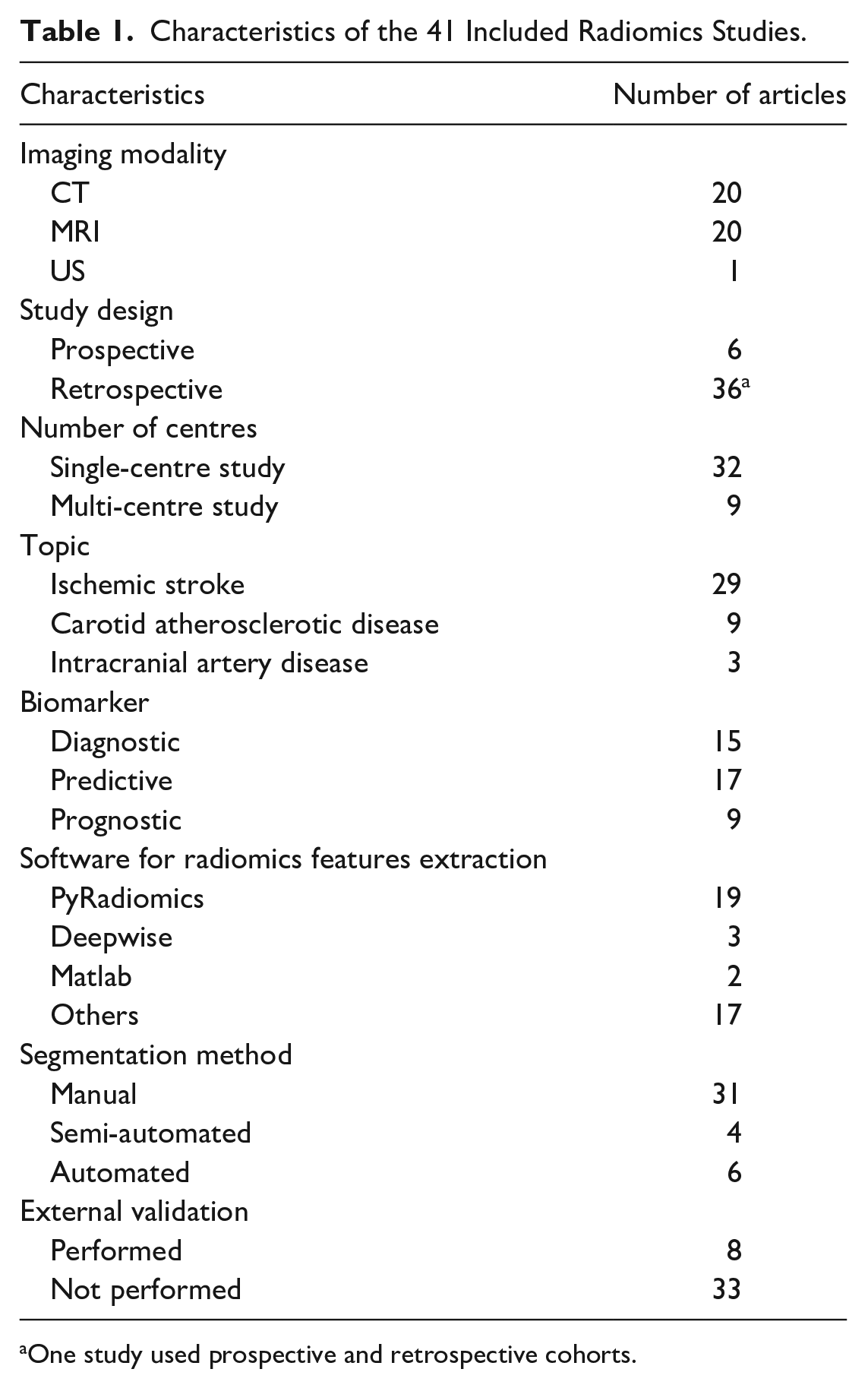
One study used prospective and retrospective cohorts.
The extraction of radiomics features was performed with PyRadiomics in nearly half of the included studies. Radiomics features were utilized as predictive biomarkers in 17 (41%) studies and prognostic biomarkers in 9 (22%) studies. They were used for diagnostic purposes in the remaining 15 (37%) studies. Regions of interest were manually segmented in 31 (75%) studies, while 4 (10%) used a semi-automated method and 6 (15%) performed a fully automated segmentation (Figure 3). Finally, 8 (20%) of the included publications validated their model with an external dataset.
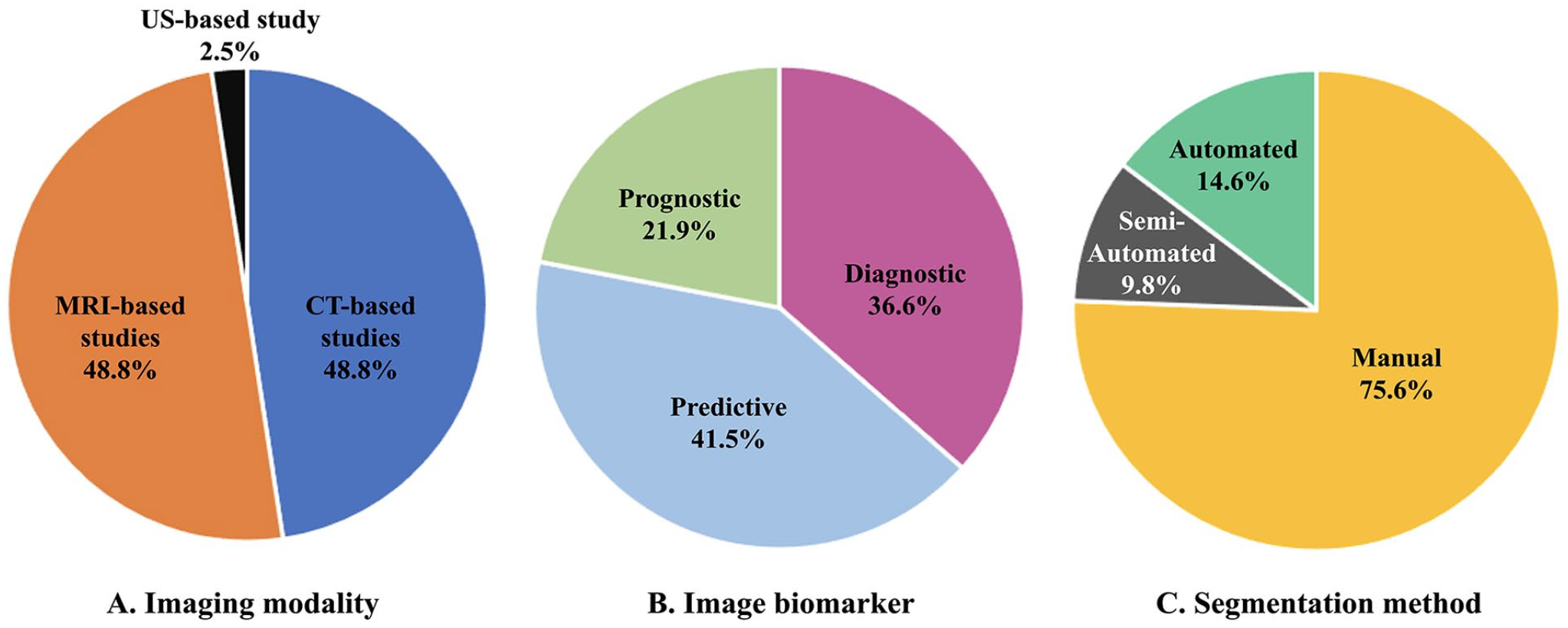
Characteristics of radiomics studies on ischemic stroke and carotid atherosclerotic disease.
Adherence to IBSI Reporting Guidelines
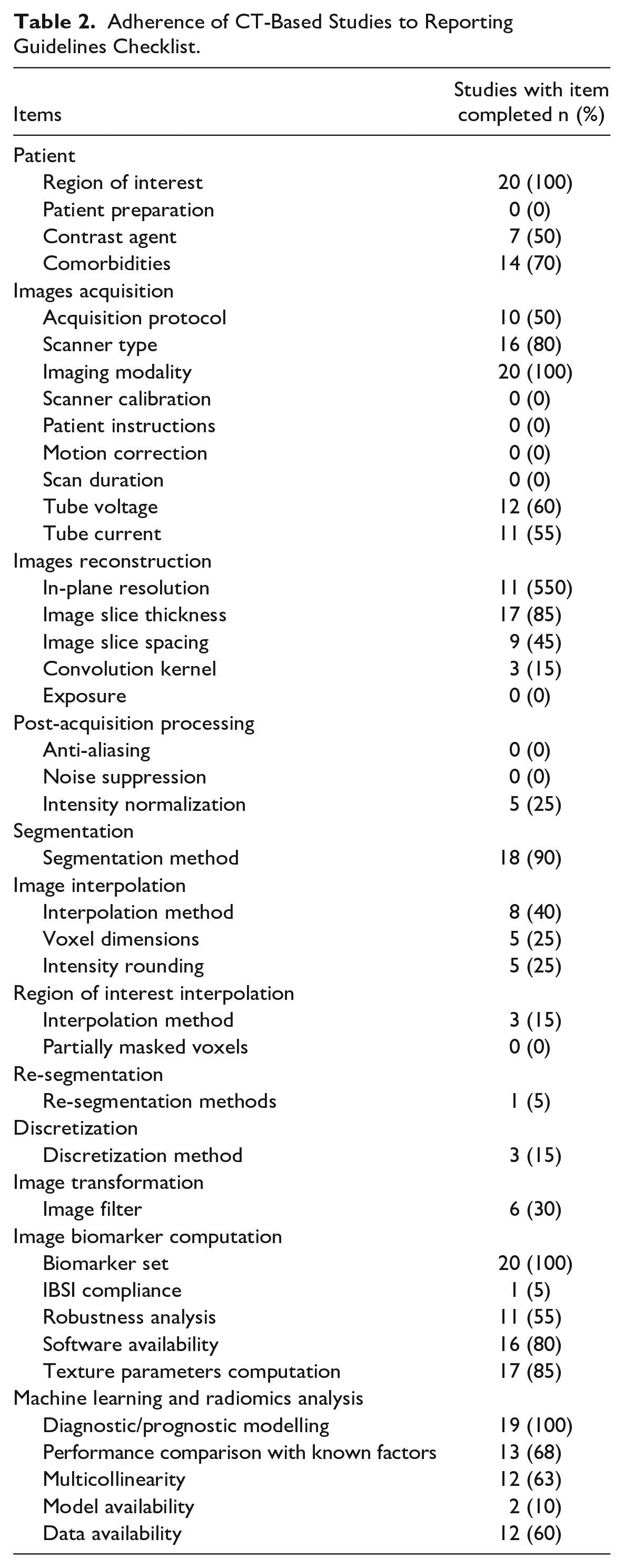
The included studies were reviewed and analyzed by the investigators, and the checklists were compared, with an inter-rater agreement of 92%. Disagreements were discussed, and a final decision was reached by consensus. In CT-based radiomics studies, the reporting checklist median score was 17/40 (range 12-25/40) (Table S2). As shown in Table 2, items such as the imaging modality used, the region of interest, and the image biomarker set utilized were well reported in all publications. On the other hand, some items were not reported in any of the reviewed studies, such as patient preparation and instructions before imaging, scanner’s last calibration, and scan duration among others. The details regarding image acquisition and reconstruction, post-acquisition image processing, and image biomarkers computation were inconsistently detailed across studies. Only one study 26 mentioned that the software used to extract the set of image biomarkers was compliant with IBSI feature reference values. All studies described how the machine learning models were built and most of them compared the performance of radiomics models with known clinical factors.
Adherence of CT-Based Studies to Reporting Guidelines Checklist.
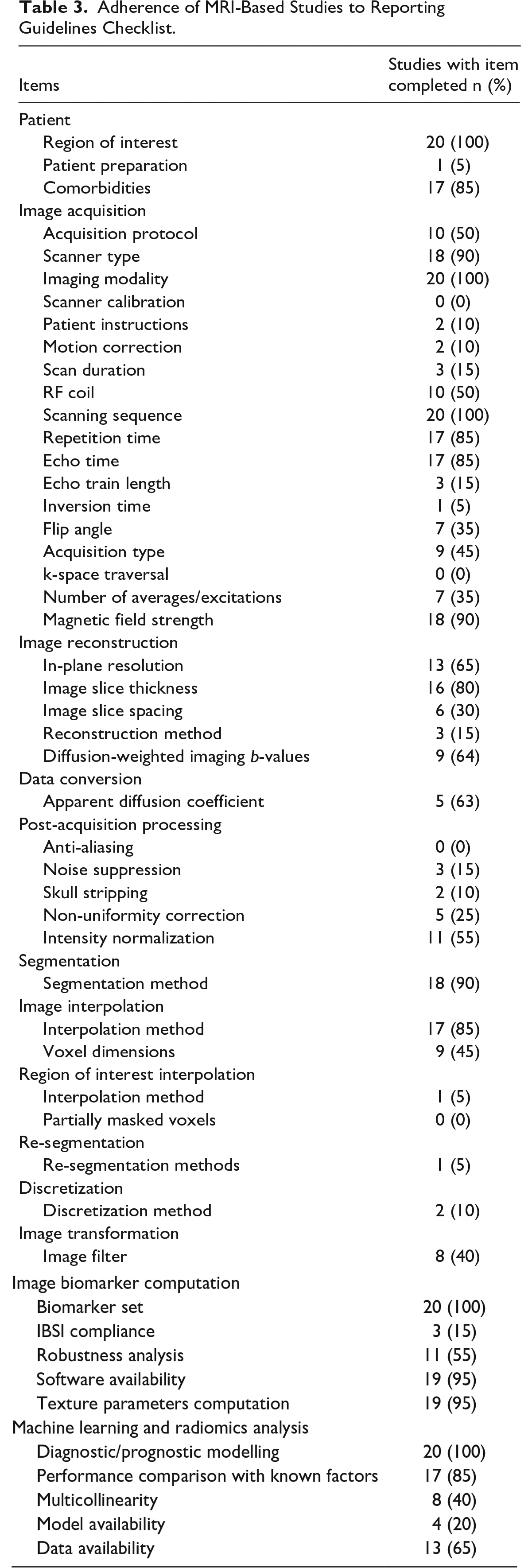
A separate reporting checklist was created for MRI-based studies and the median score was 22.5/50 (range 12-34/50) (Table S3). Items such as imaging modality, region of interest, scanning sequences, biomarker set, and machine learning modelling were reported in all these studies. Patients’ preparation and instructions were detailed in only 1 study, 36 and magnetic field strength was omitted in 2 publications. Post-acquisition image reconstruction and processing methods are incompletely reported in many publications. The detailed analysis is presented in Table 3.
Adherence of MRI-Based Studies to Reporting Guidelines Checklist.

Discussion
Radiomics, while offering significant potential in medical imaging and diagnosis, faces several challenges that impact its widespread adoption. One of the biggest challenges is reliable and complete reporting to ensure validation and reproducibility of results. Radiomics was first developed and primarily applied in the field of oncology. Some studies assessed the quality of radiomics studies in oncology with the aim of improving this field of research. A systematic review evaluated neuro-oncologic studies according to the radiomics quality score (RQS) and concluded that the quality of reporting was globally insufficient. 52 The RQS evaluates the quality of both the science and the reporting of radiomics studies with a list of 16 items. 53 Another study used the RQS and TRIPOD statement to assess the quality of science and reporting of radiomics in oncologic studies, also highlighting the need for improvements. 54 The TRIPOD statement is a checklist of 22 items, deemed essential for transparent reporting of a prediction model study. 55 This tool is not specific to radiomics, and some methodological aspects are not covered by the TRIPOD checklist. A combination with a second checklist like the RQS is therefore necessary.
A checklist specifically tailored to radiomics was lacking in the literature. Consequently, we created reporting quality checklists for CT and MRI-based studies to enable a comprehensive assessment of the included radiomics studies. IBSI is accepted by experts in the field and the reference manual provides an extensive list of items, including items specific to the chosen imaging modality. Thus, when the study was designed, IBSI was used as the primary source to elaborate the checklists. Interestingly, the CheckList for EvaluAtion of Radiomics research (CLEAR) reporting guideline was recently published. 56 This open-source checklist includes 58 items for the evaluation of radiomics research. It covers various aspects of a publication, from the title relevance to radiomics model availability. In the feature extraction description, the CLEAR guideline states that IBSI terminology for feature classes should be used. The authors should also indicate if the software adheres to the benchmarks and certification of IBSI. We encourage authors of future publications to refer to the CLEAR checklist to ensure the reporting quality of their manuscript and to facilitate the repeatability and reproducibility of studies.
Reviewing radiomics studies on ischemic stroke, intracranial and carotid atherosclerotic disease, we found that published studies on these topics do not provide sufficient detail, with global scores varying from 30% to 68% according to the IBSI-based reporting checklists. With the publication of reporting guidelines like the IBSI in 2020, one could expect an increase in reporting quality in more recent publications. The checklist scores of studies published until 2020 were compared to those published in 2021 and 2022. For CT-based studies, the mean score is 17/40 for both categories, with a range of 12 to 22/40 until 2020 and a range of 12 to 25/40 in 2021 to 2022. In the case of MRI-based studies, the mean score was 21/50 (range 19-22/50) until 2020 and 23/50 (range 12-34/50) in 2021 to 2022. Therefore, there was no significant improvement in reporting quality after the publication of IBSI in 2020. This finding highlights the lack of awareness of the importance of using established guidelines to enhance reporting quality and, consequently, the reproducibility of studies. We hope that the publication of the CLEAR checklist will have a more significant impact on the reporting quality of future radiomics studies.
Basic items such as imaging modality and region of interest were reported in all studies. These items are considered essential for the readers’ comprehension, and therefore cannot be overlooked. Conversely, certain items concerning image acquisition and processing were not included, probably for the sake of providing a concise report while limiting technical details. However, variability in imaging protocols across different scanners and institutions can affect the consistency and reproducibility of radiomic features. Standardization of imaging parameters (acquisition, reconstruction, and processing) is crucial to ensure that radiomics can be reliably used across different settings. It is therefore recommended that authors include the necessary details in their manuscript or in supplemental material, depending on their main audience and space availability.
Accurate segmentation of the region of interest in medical images is critical for radiomic analysis. Segmentation methodology was mentioned in 90% of the included studies. Manual segmentation is time-consuming and subject to inter-observer variability, while automated and semi-automated methods can sometimes lack accuracy and therefore need to be improve. Once the region of interest is delineated, image biomarker computation is performed. At this step, the report should include the set of image biomarkers and their definitions, the name and version of the utilized software, a statement about whether this software is able to reproduce the IBSI feature reference values or not, and how robustness of the biomarkers was assessed. Similarly, machine learning and radiomics analysis modelling should be detailed, including comparison with known clinical factors and assessment of multicollinearity.
To ensure reproducibility in radiomics research, availability of software and data is crucial. For example, PyRadiomics, used in many of the included studies, is an open-source Python package specifically designed for the extraction of radiomics features from medical images. In CT-based studies, the software used to extract radiomics features was reported in 80% of studies and data availability was specified in 60% of studies. The information regarding where radiomics models and necessary pre-processing information may be found was mentioned in only 10% of studies. In MRI-based studies, radiomics software was reported in most studies and data availability statement was present in 65% of publications. However, machine learning models were often not made available by the authors. It is therefore recommended that authors provide open-source code and data to allow replication of results by other investigators.
High-quality, large-scale imaging data are required for effective radiomic analysis. Acquiring such datasets can be challenging and retrospective studies are often easier to perform due to data availability, cost-effectiveness, and feasibility in rare conditions or outcomes. Nevertheless, retrospective studies are prone to several sources of bias such as selection and information bias. Large prospective multicentre studies with external validation are still needed for clinical validation and translation of radiomics models.
Conclusion
Radiomics harnesses the power of advanced image analysis to extract meaningful information from medical images. In cases of ischemic stroke, intracranial and carotid atherosclerotic disease, radiomics has the potential to enhance diagnosis accuracy and provide predictive insights into prognosis and clinical outcomes. However, the overall reporting quality of radiomics studies on these topics is suboptimal. Addressing this challenge and improving the quality of future publications is essential for the successful integration of radiomics into routine clinical practice.
Supplemental Material
sj-docx-1-caj-10.1177_08465371241234545 – Supplemental material for Radiomics Studies on Ischemic Stroke and Carotid Atherosclerotic Disease: A Reporting Quality Assessment
Supplemental material, sj-docx-1-caj-10.1177_08465371241234545 for Radiomics Studies on Ischemic Stroke and Carotid Atherosclerotic Disease: A Reporting Quality Assessment by Ann-Marie Beaudoin, Jan Kee Ho, Adrienne Lam and Vincent Thijs in Canadian Association of Radiologists Journal
Footnotes
Author Contributions
AMB researched literature, conceived the study, and analyzed the data. VT was involved in protocol development. AMB wrote the first draft of the manuscript. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors were supported by The Florey (Melbourne, Australia). AMB was also supported by the Société des Médecins de l’Université de Sherbrooke (Sherbrooke, Canada).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
