Abstract
In the cyanobacterial circadian clock, a core oscillator comprising the proteins KaiA, KaiB, and KaiC keeps time based on a rhythmic phosphorylation of KaiC, and histidine protein kinases relay temporal information from the KaiABC complex to regulate gene expression. The kinases SasA and CikA engage directly with the oscillator and are responsible for modulating the phosphorylation and dephosphorylation throughout the circadian day of the response-regulator transcription factor RpaA; the phosphorylation state of RpaA in turn determines circadian gene expression. We recently showed that either CikA or SasA can drive rhythmic phosphorylation and DNA binding of RpaA in an in vitro system. However, when SasA is absent in vivo, a bioluminescence reporter gene shows a very low expression and amplitude rhythm, indicating CikA kinase activity is not sufficient to activate gene expression. We questioned why CikA cannot serve as a robust kinase for RpaA in the absence of SasA in the cell. Here, we investigated post-translational modifications of CikA and found KaiC-dependent phosphorylation sites of CikA that dramatically affect its activity. Phosphomimetic mutants of these sites showed that the phosphorylated version of CikA is not functional. Our data show that inverse correlation of KaiC levels and these inhibitory phosphorylation sites can explain the lower CikA activity in a SasA knockout background. We conclude that these phosphorylation sites act as a rheostat for CikA activity and are regulated by KaiC levels.
Keywords
Circadian clocks are self-sustained oscillators that provide an internal sense of time for individual cells and/or whole organisms. Post-translational modifications are known to regulate the activity of clock components. A striking example is the phosphorylation of the animal clock protein Period by casein kinase 1 epsilon, which affects clock function from Drosophila to humans. In addition, expressing different phosphomimetic mutant variants of the core mammalian circadian oscillator protein CRY1 produces changes in period length or arrhythmic gene expression (Kloss et al., 1998; Liu and Zhang, 2016; Maywood et al., 2014; Toh et al., 2001). In the
Missing from this model is the evidence that CikA, when it is not associated with the Kai complex, is a better kinase/phosphatase for RpaA than is SasA (Chavan et al., 2021). Moreover, either CikA or SasA can alone provide rhythmic output from the Kai complex in vitro, with CikA catalyzing RpaA phosphorylation in the day portion of the cycle and actively dephosphorylating RpaA in the night, when CikA is associated with KaiBC. SasA appears to provide only daytime kinase activity, with a slower RpaA autodephosphorylation. These data support the role of CikA as a kinase for the circadian clock in vivo without the requirement of SasA.
The current study aimed to reconcile in vitro and in vivo data. Consistent with the classic model, deletion of CikA results in elevated
Materials and Methods
Immunoprecipitation and Mass Spectrometry of FLAG-CikA
Cultures used for IP were grown under a 2-day 12:12 light/dark cycle to entrain and then released to constant light. In initial experiments, cells were collected at CT12 (12 h after lights on) for immunoprecipitation (IP) and tandem mass spectrometry (LC-MS/MS) as described below and reported in Figure 1a and Table S1. Based on these results, for further experiments, cells were collected at CT4 (4 h after lights on) and CT22 (22 h after lights on) from the strains indicated in the figure legends and frozen in liquid nitrogen (reported in Figure 4). Cells were lysed by vortexing in TAP buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1.5 mM MgCl2, 0.1% Triton-X, 0.5 mM dithiothreitol (DTT), 1 mM phenylmethylsulfonyl fluoride (PMSF), protease inhibitor cocktail tablet Roche, REF11836170001) and glass beads (Sigma, G9143). The clearing centrifugation step was done for 10 min at 15,000 rpm at 4 °C. Supernatants were collected. Meanwhile, FLAG-M2 magnetic beads (30 µl per sample) were washed with 5 column volumes (CV) of Tris-buffered saline (TBS, 50 mM Tris pH 8.0, 100 mM NaCl) 3 times and then equilibrated with 5 CV of TAP buffer 3 times. The cleared supernatant was incubated with FLAG-M2 magnetic beads at 4 °C on a rotating wheel for 4 h. The FLAG-M2 magnetic beads were then washed with 5 CV of TAP buffer 3 times and 5 CV of TBS until the TBS absorbance at 280 nm was less than 0.05. The FLAG-M2 magnetic beads were stored at −80 °C. Afterwards, protein samples were diluted in TNE (50 mM Tris pH 8.0, 100 mM NaCl, 1 mM ethylenediaminetetraacetic acid [EDTA]) buffer. RapiGest SF reagent (Waters Corp.) was added to the mix to a final concentration of 0.1%, and samples were boiled for 5 min. Tris(2-carboxyethyl)phosphine (TCEP) was added to 1 mM (final concentration), and the samples were incubated at 37 °C for 30 min. Subsequently, the samples were carboxymethylated with 0.5 mg/ml of iodoacetamide for 30 min at 37 °C, followed by neutralization with 2 mM TCEP (final concentration). Protein samples prepared as above were digested with trypsin (trypsin:protein ratio = 1:50) overnight at 37 °C. RapiGest was degraded and removed by treating the samples with 250 mM HCl at 37 °C for 1 h, followed by centrifugation at 14,000 rpm for 30 min at 4 °C. The soluble fraction was then transferred to a new tube, and the peptides were extracted and desalted using C18 desalting columns (Thermo Scientific, PI-87782). Peptides were quantified using a Bicinchoninic Acid protein assay, and a total of 1 µg of peptides were injected for LC-MS analysis.
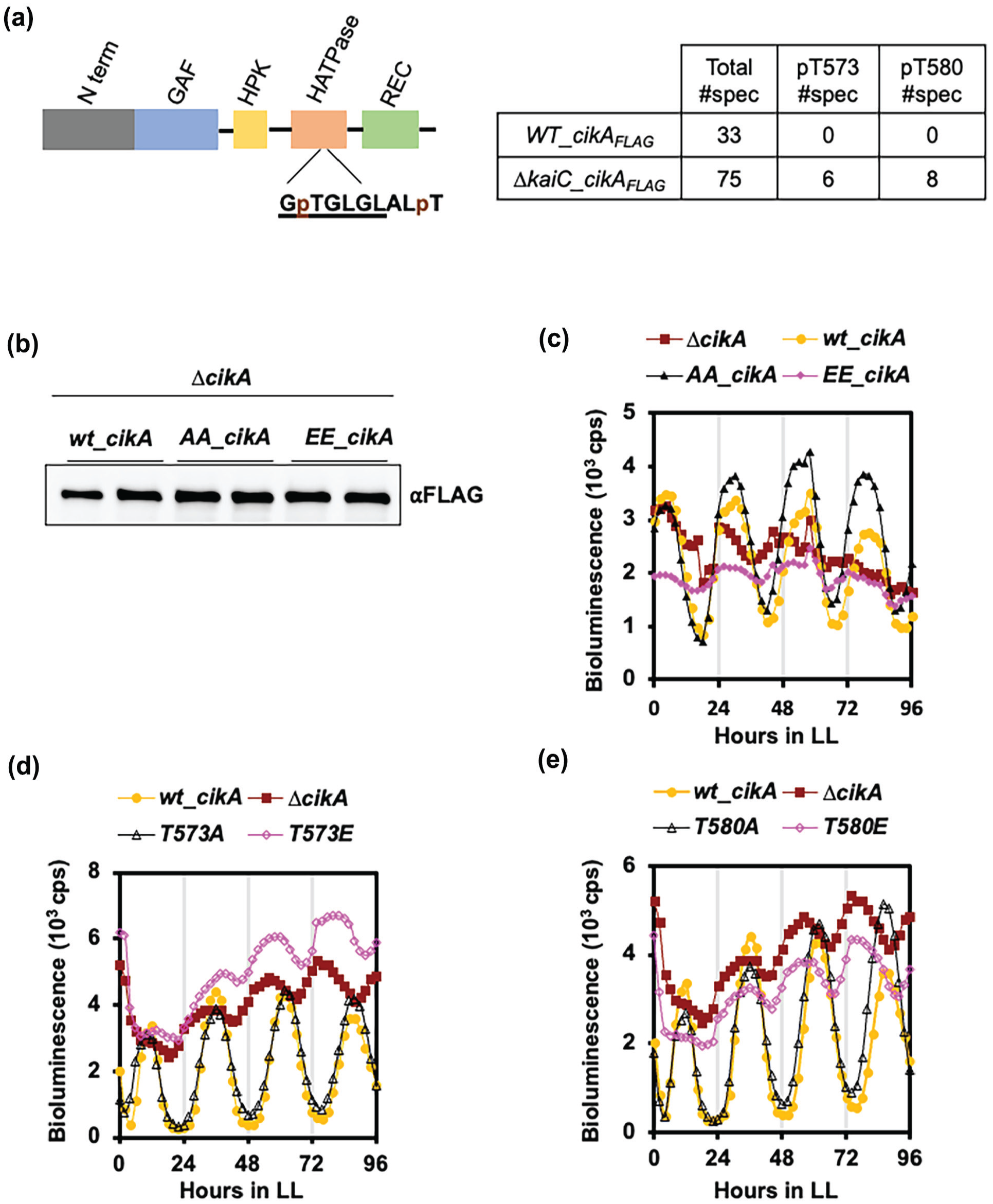
A phosphomimetic mutant of CikA cannot substitute for wt_CikA. (a) Scheme showing the conserved domains of CikA (left panel). Amino acids in the conserved G-2 box are underlined. Phosphorylated threonines (T573 and T580), identified by LC-MS/MS from samples collected at CT12, are labeled with “p.” Peptides that include phosphorylated threonines are shown in the table for the
Tandem Mass Spectrometry
Trypsin-digested peptides were analyzed by ultra-high-pressure liquid chromatography (UPLC) coupled with LC-MS/MS using nanospray ionization. The nanospray ionization experiments were performed using an Orbitrap Fusion Lumos hybrid mass spectrometer (Thermo) interfaced with a nano-scale reversed-phase UPLC (Thermo Dionex UltiMate 3000 RSLC nano System) using a 25-cm, 75-micron ID glass capillary packed with 1.7-µm C18 (130) BEH beads (Waters Corporation). Peptides were eluted from the C18 column into the mass spectrometer using a linear gradient (5%-80%) of acetonitrile (ACN) at a flow rate of 375 µl/min for 1.5 h. The buffers used to create the ACN gradient were Buffer A (98% H2O, 2% ACN, 0.1% formic acid) and Buffer B (100% ACN, 0.1% formic acid). Mass spectrometer parameters were as follows: an MS1 survey scan using the Orbitrap detector (mass range [m/z]: 400-1500 [using quadrupole isolation], 120,000 resolution setting, spray voltage of 2200 V, ion transfer tube temperature of 275 °C, Automatic Gain Control target of 400,000, and maximum injection time of 50 ms) was followed by data-dependent scans (top speed for most intense ions, with charge state set to only include +2-5 ions, and 5-second exclusion time, while selecting ions with minimal intensities of 50,000, during which the collision event was carried out in the high-energy collision cell (HCD Collision Energy of 30%), and the fragment masses were analyzed in the ion trap mass analyzer (with an ion trap scan rate of turbo, first mass m/z was 100, AGC target 5000, and maximum injection time of 35 ms). Protein identification was carried out using Peaks Studio 8.5 (Bioinformatics Solutions Inc.). MS analysis was performed at the “Molecular Mass Spectrometry Facility, UC San Diego.”
Protein Extraction
Western Blot Analysis
Total protein extract from
Purification of GST-Tagged CikA and Mutants of CikA
The gene encoding CikA from
In Vitro Phosphorylation Assay of RpaA
Purified 0.65-µM wt-CikA and 2.5-µM RpaA, 0.65-µM AA-CikA and 2.5-µM RpaA, 0.65-µM EE_CikA and 2.5-µM RpaA were incubated in in vitro reaction buffer (20-mM Tris-HCl pH 8.0, 150-mM NaCl, 5-mM MgCl2, 1-mM ATP, 0.5-mM EDTA) at 30 °C for 1 h. Only 2.5-µM RpaA in in vitro reaction buffer was used as control. After 1 h, 7 µL from each reaction was immediately mixed with 2x Laemmli buffer and loaded on a pre-prepared Phos-tag gel for further western blot analysis.
Partial Trypsin Digestion
Recombinant WT_CikA and double-mutant variants (AA_CikA, EE_CikA) were purified from
Microscopy
Indicated
Bioluminescence Reporter Assay
Flask-grown cultures were diluted to a density of OD750 = 0.05 and grown under constant light until reaching a density of OD750 = 0.2. Then, 20 µL of each culture was mixed with 10 µL of 100-mM firefly luciferin and placed on a Grenier Bio-One 96-well microplate (Cat #655073) filled with BG-11 agar medium. Plates were covered with clear tape to prevent drying, and holes were poked using a sterile needle to allow air transfer. Cultures were synchronized by incubating the plate under 12 h:12 h light (~30 µE m−² s−¹ light)/dark conditions for 2 days and then returned to constant light conditions for bioluminescence sampling. Bioluminescence of
Results
CikA Is Phosphorylated at Thr573 and Thr580 When KaiC Is Absent
While deletion of CikA regulates circadian phase resetting, period length, and cell size in
Phosphomimetic Variants of CikA Cannot Substitute for WT CikA
To investigate the importance of CikA Thr573 and Thr580 phosphorylation sites on CikA function, we exchanged both sites to alanine (AA_CikA) to prevent phosphorylation of CikA, and to glutamate (EE_CikA) to mimic phosphorylated CikA. Alleles that expressed FLAG-tagged WT_CikA, AA_CikA, or EE_CikA from the native
Kinase Activity of EE_CikA on RpaA Is Diminished Due to a Possible Conformation Change
CikA is both a kinase and phosphatase for RpaA (Gutu and O’Shea, 2013). Our in vivo data suggest that the EE_CikA variant produces a
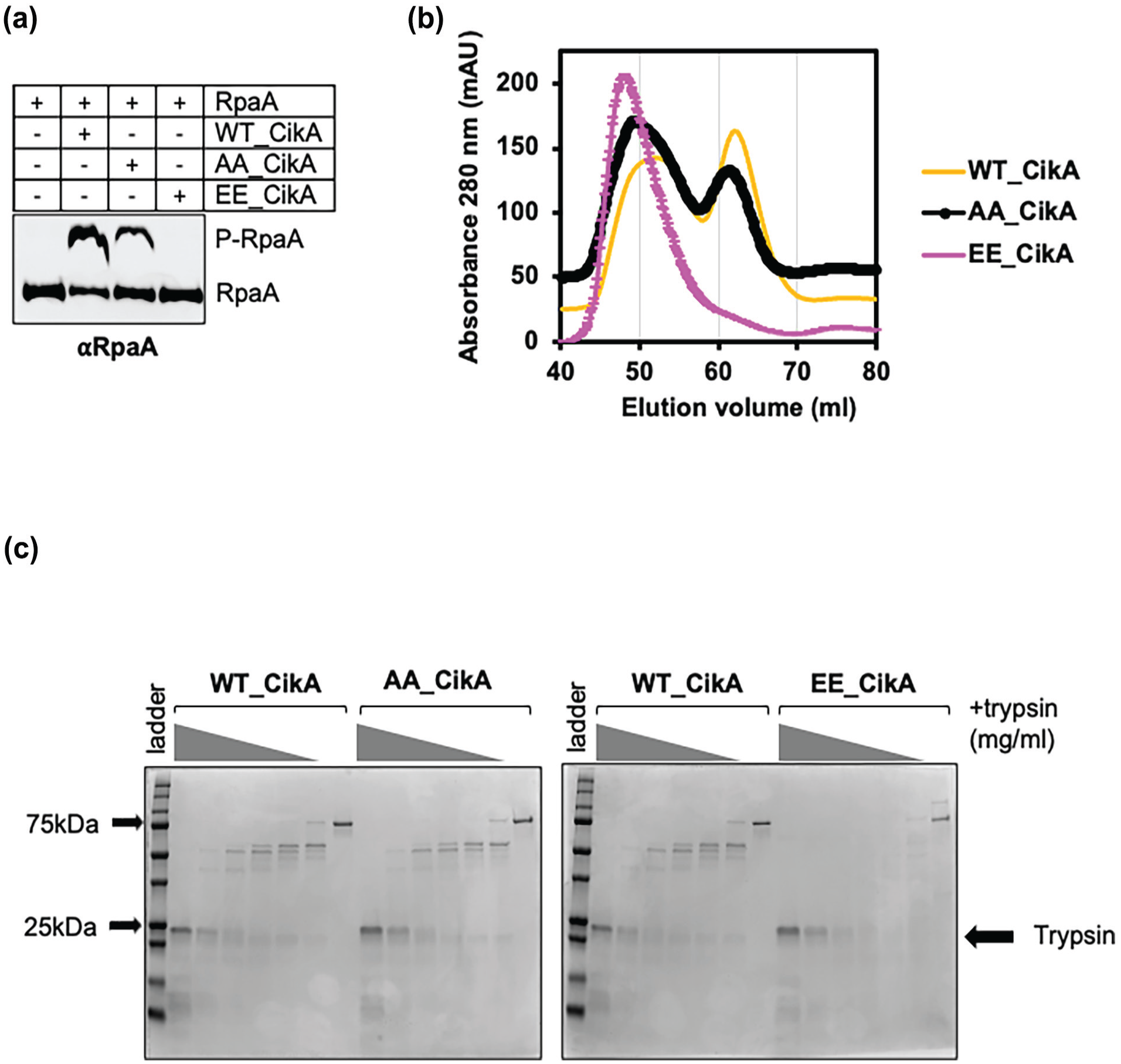
Kinase activity of EE-CikA on RpaA is diminished due to a possible conformation change. (a) Western blot showing the phosphorylated and unphosphorylated form of purified RpaA from Phos-tag SDS-PAGE after incubation with WT_CikA, AA_CikA, or EE_CikA, respectively. (b) Combined size-exclusion column elution graphs showing the WT_CikA, AA_CikA, and EE_CikA elutions. (c) Coomassie-stained SDS-gels showing partial trypsin digestion of CikA variants. Purified CikA variants incubated with trypsin at 1 mg/mL, 0.33 mg/mL, 0.11 mg/mL, 0.04 mg/mL, 0.012 mg/mL, 0.004 mg/mL, and 0 mg/mL.
These phosphorylation sites are around the G2-box, which forms a flexible loop in histidine kinases (Bilwes et al., 1999). To test whether phosphorylation at this region induces a conformational change, we performed size-exclusion chromatography of purified CikA variants. WT_CikA and AA_CikA eluted as two peaks, whereas EE_CikA eluted as a single high-molecular-weight peak, which suggests that EE_CikA has a different conformation compared to WT_or AA_CikA (Figure 2b). Due to elution in the high-molecular-weight range, we hypothesized that EE_CikA has a more open structure. To test this hypothesis, we utilized partial trypsin digestion to compare sensitivity of WT_CikA, AA_CikA, and EE_CikA to trypsin. EE_CikA was more sensitive to trypsin, consistent with a more open conformation than WT_CikA and AA_CikA (Figure 2c). In addition, purified EE_CikA ran slightly slower than WT_CikA or AA_CikA when analyzed by Phos-tag SDS-PAGE (Supplemental Figure 2D). Together, the data suggest that phosphorylation at Thr573 and Thr580 induces a conformational change on CikA that inhibits its kinase activity against RpaA.
AA_CikA Compensates for the Low-Amplitude Rhythm Observed in ∆sasA
A recent study showed that CikA can be a more robust kinase as well as phosphatase for RpaA than SasA in the presence of KaiABC in vitro (Chavan et al., 2021). However, a
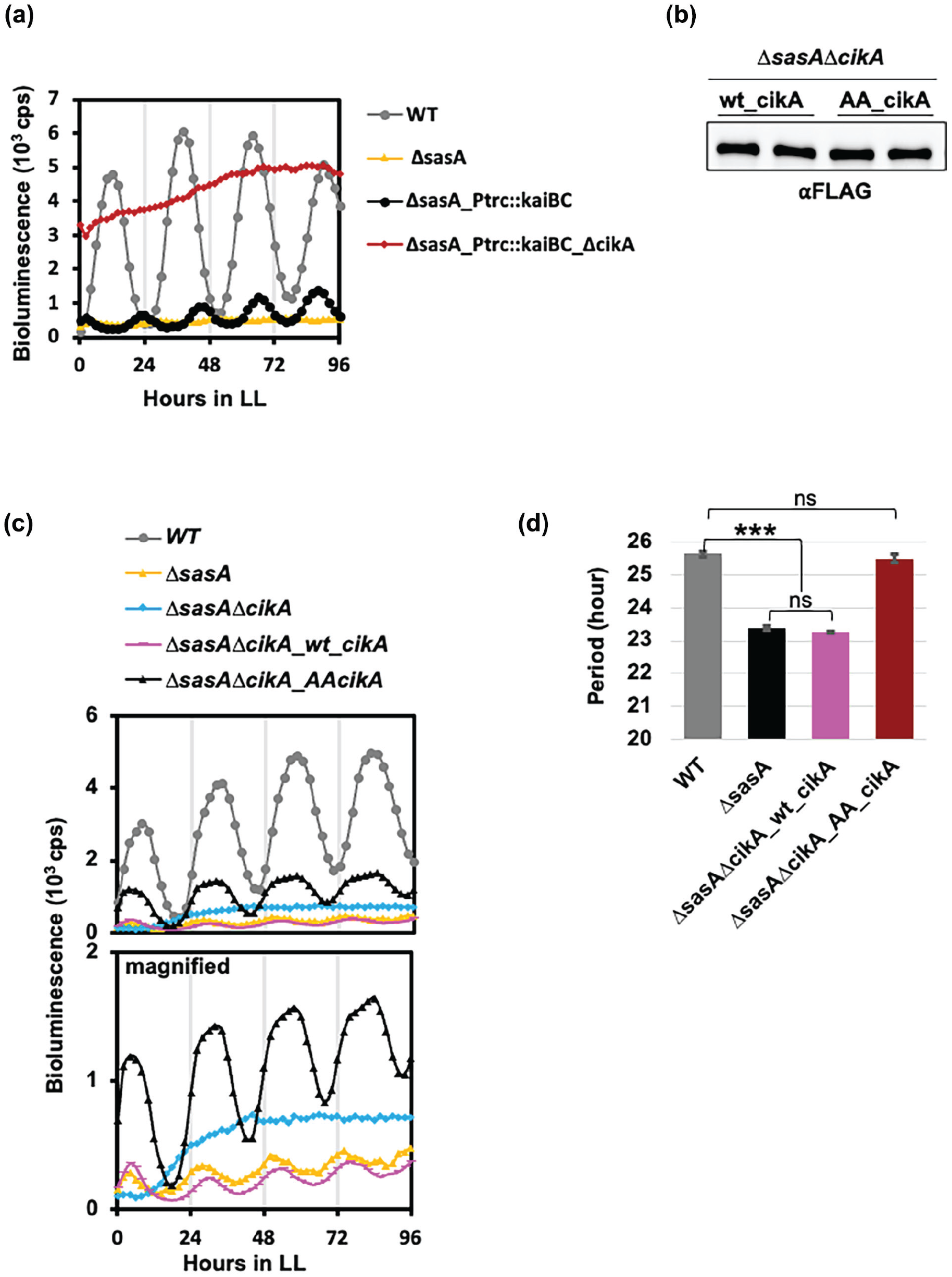
AA_CikA compensates for low-amplitude rhythm and short period observed in ∆
To test whether the low-amplitude gene expression rhythm in the ∆
Time-Dependent Interaction Partners of CikA Require KaiABC Interaction
Our data suggest that CikA could form at least two different complexes; one on the KaiABC complex (hereafter, Kai complex) and the other not interacting with the Kai complex. In order to identify potential time-dependent CikA interaction partners, we analyzed the CikA complexes at CT4 (off the Kai complex) and CT22 (on the Kai complex). We identified 125 proteins that were retrieved with WT_CikA compared to a mock control (Figure 4a, upper panel) (Supplemental Table 2). Among these proteins, 43 interacted regardless of the time of the day, whereas 25 were enriched at CT4, and 57 were enriched at CT22 (Supplemental Table 2). We also used the C644R mutant, which binds poorly to the Kai complex, as a control to find out the contribution of CikA-KaiABC interaction on the CikA interactome. The total number of interaction partners was decreased for C644R, and especially for CT22-specific ones (Figure 4a, lower panel, Supplemental Figure 4B). CikA peptides detected by MS in different samples were comparable, minimizing the contribution of IP efficiencies on the detection of interaction partners (Supplemental Table 2). We analyzed the fold enrichment of 125 WT_CikA-interacting candidates in CT4 vs CT22 (-10lgP fold > 1.5) in WT_CikA and C644R experiments to see whether the C644R substitution changes time-specific interaction. Among the CT4-enriched group, the C644R substitution resulted in decreased interaction specifically at CT4 (Figure 4b, left panel, Supplemental Table 3). Interestingly, half of the CT22-enriched interaction partners still associated with C644R, albeit with loss of the time specificity (Figure 4b, right panel, Group 1, Supplemental Table 3). Nonetheless, half of CT22-specific interaction partners were lost with C644R (Figure 4b, right panel, Group 2). Only a few interaction partners of WT_CikA behaved similarly when C644R was used (Figure 4b, right panel, Group 3). Together, the data suggest that CikA-KaiABC interaction determines both the time and intensity of the majority of CikA interactions.
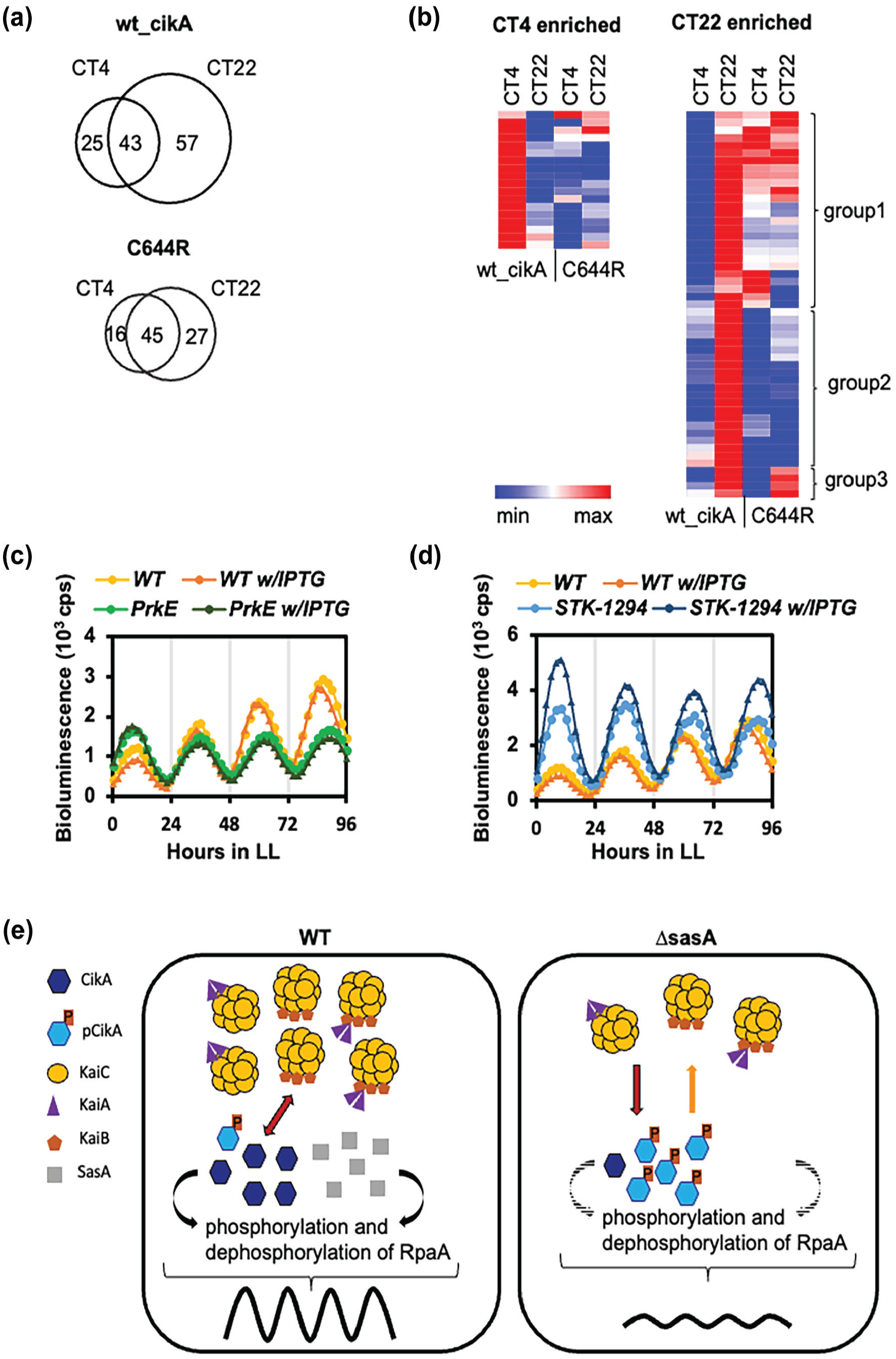
Time-dependent interaction partners of CikA are also dependent on clock interaction. (a) Venn diagrams showing the number of putative interaction partners of CikA with wt_CikA (upper panel) and the C644R mutant (lower panel) at CT4 and CT22. For stringent analysis, the potential interaction partners with the 50 > -10lgP value were excluded. (b) Heat map showing the significance (-10lgP) of the time-specific interaction partners of CikA. Interaction partners enriched in CT4 (>1.5 fold) and CT22 (>1.5 fold) are included in WT and C644R backgrounds. The color scale ranges from red to blue, where red indicates higher significance, and blue indicates lower significance of interaction. (c) Bioluminescence reporter assay showing the expression of PkaiBC::luc under constant light for WT and PrkE (WT_Ptrc::prkE) with and without IPTG. Each trace represents the average of two different clones (
Among the putative interaction partners of CikA and C644R, we found two serine/threonine kinases (STK) as potential interactors of CikA:
Discussion
Although CikA was shown to have a significant role in the cyanobacterial circadian clock almost 25 years ago, the biochemical regulation of its kinase and phosphatase activities is not well understood. Recent work has shown that the in vitro clock can guide in vivo experiments to yield deeper insights from mutant phenotypes than were gleaned during initial characterization. For example, a combination of in vitro and in vivo experiments recently explained why the circadian clock in a
Similarly, in vitro clock experiments challenged the interpretation of CikA’s role in the clock as a phosphatase, with SasA dominating the role as kinase. CikA in vitro is a potent kinase of RpaA, and it supports very robust rhythmic phosphorylation of RpaA in a clock reaction. We sought to answer why RpaA phosphorylation levels are so low, and rhythms are of such low amplitude, in the absence of SasA in vivo. One possible answer to this question was that CikA activity is reduced by cellular factors in a way that does not occur during the in vitro clock cycle. Here, we showed that CikA is inhibited by phosphorylation when KaiC levels are low. This mechanism unravels, at least partially, why CikA does not support strong rhythms of gene expression in vivo when SasA is absent. Low levels of KaiC in a
Exchange of these phosphorylation sites to glutamate, either individually or in combination, rendered CikA inactive. Thr573 site resides inside, and Thr580 just after, the G2-box that acts as a nucleotide-binding motif in histidine kinases. Previous studies showed by mutation of the glycine residues that the G2-box plays an important role in modulating phosphatase and/or kinase activities (Tanaka et al., 1998; Zhu and Inouye, 2002). Our data indicate that a phosphomimetic (EE) version of Thr573 and Thr580 does not have any kinase activity against RpaA. Because the G2-box is where ATP binds, we hypothesize that phosphorylation at this site could interfere with ATP binding due to the ionic interactions with phosphorylated threonine and ATP. Another possible regulation could be a conformation change of CikA upon phosphorylation at these sites that affects partner interaction. This idea is supported by the higher sensitivity of EE_CikA to limited proteolysis. Previous studies focusing on the role of G2-box in histidine kinases focused on exchange of conserved glycine residues. S/T residues are found in the second position of many G2-boxes of histidine kinases in bacteria (Kim and Forst, 2001). Hence, it would be of interest whether histidine kinases in other organisms are regulated by inhibitory phosphorylation of G2-box sites.
CikA is proposed to interact with the Kai clock complex primarily during the night, and this interaction is decreased when CikA is mutated on the C644 site to Arg (C644R) (Tseng et al., 2017). When we compared the putative interaction partners of CikA during day (CT4) and night (CT22), we observed more interaction partners at night when CikA is on the Kai complex. The C644R mutant lost most of this discrimination between day and night putative interaction partners. In addition, when CikA affinity to the oscillator complex is decreased as in C644R, both the time and intensity of the interactions are affected, suggesting that the clock orchestrates the CikA interactome. Moreover, phosphorylation of T573 and/or T580 in the C644R_CikA mutant was greater in the day than in night, when CikA interaction with the core clock is more limited due to both circadian time and the C644R mutation. Together, these data support our hypothesis that CikA is selectively inactivated and, when phosphorylated, has a potential conformation change that prevents its interaction with partners, including RpaA. While the KaiABC/CikA/RpaA clock can function in vitro without the requirement of other feedback loops, this inhibitory phosphorylation of CikA could enhance the robustness of the clock by limiting CikA activity to the subpopulation that is interacting with the clock. In future, it would be of interest to use strains that carry the unphosphoryatable AA_CikA variant to determine whether selective CikA inactivation contributes to the robustness of the circadian clock.
Supplemental Material
sj-docx-1-jbr-10.1177_07487304251338156 – Supplemental material for Clock-Dependent Phosphorylation of CikA Regulates Its Activity
Supplemental material, sj-docx-1-jbr-10.1177_07487304251338156 for Clock-Dependent Phosphorylation of CikA Regulates Its Activity by Cigdem Sancar and Susan S. Golden in Journal of Biological Rhythms
Supplemental Material
sj-xlsx-2-jbr-10.1177_07487304251338156 – Supplemental material for Clock-Dependent Phosphorylation of CikA Regulates Its Activity
Supplemental material, sj-xlsx-2-jbr-10.1177_07487304251338156 for Clock-Dependent Phosphorylation of CikA Regulates Its Activity by Cigdem Sancar and Susan S. Golden in Journal of Biological Rhythms
Supplemental Material
sj-xlsx-3-jbr-10.1177_07487304251338156 – Supplemental material for Clock-Dependent Phosphorylation of CikA Regulates Its Activity
Supplemental material, sj-xlsx-3-jbr-10.1177_07487304251338156 for Clock-Dependent Phosphorylation of CikA Regulates Its Activity by Cigdem Sancar and Susan S. Golden in Journal of Biological Rhythms
Supplemental Material
sj-xlsx-4-jbr-10.1177_07487304251338156 – Supplemental material for Clock-Dependent Phosphorylation of CikA Regulates Its Activity
Supplemental material, sj-xlsx-4-jbr-10.1177_07487304251338156 for Clock-Dependent Phosphorylation of CikA Regulates Its Activity by Cigdem Sancar and Susan S. Golden in Journal of Biological Rhythms
Footnotes
Acknowledgements
Research reported in this publication was supported by the National Institute of General Medical Sciences of the National Institutes of Health under Award Number R35GM118290. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Conflict of Interest Statement
The authors have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Considerations
This article does not contain any studies with human or animal participants.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
