Abstract
Cyanobacteria contain a circadian oscillator that can be reconstituted in vitro. In the reconstituted circadian oscillator, the phosphorylation state of KaiC oscillates with a circadian period, spending about 12 h in the phosphorylation phase and another 12 h in the dephosphorylation phase. Although some entrainment studies have been performed using the reconstituted oscillator, they were insufficient to fully explain entrainment mechanisms of the cyanobacterial circadian clock due to the lack of input pathway components in the in vitro oscillator reaction mixture. Here, we investigate how an input pathway component, CikA, affects the phosphorylation state of KaiC in vitro. In general, CikA affects the amplitude and period of the circadian oscillation of KaiC phosphorylation by competing with KaiA for the same binding site on KaiB. In the presence of CikA, KaiC switches from its dephosphorylation phase to its phosphorylation phase prematurely, due to an early release of KaiA from KaiB as a result of competitive binding between CikA and KaiA. This causes hyperphosphorylation of KaiC and lowers the amplitude of the circadian oscillation. The period of the KaiC phosphorylation oscillation is shortened by adding increased amounts of CikA. A constant period can be maintained as CikA is increased by proportionally decreasing the amount of KaiA. Our findings give insight into how to reconstitute the cyanobacterial circadian clock in vitro by the addition of an input pathway component, and explain how this affects circadian oscillations by directly interacting with the oscillator components.
A cyanobacterium is the simplest organism to have a circadian clock (Iwasaki and Kondo, 2004). Circadian clocks consist of 3 major components: a central oscillator, an input pathway, and an output pathway (Takahashi, 2004). The central oscillator in cyanobacteria is known as the minimal self-sustained circadian oscillator and is composed of 3 proteins, KaiA, KaiB, and KaiC. The oscillator can be reconstituted in vitro by mixing the Kai proteins with adenosine triphosphate (ATP) and magnesium (Kim et al., 2015; Nakajima et al., 2010). The phosphorylation/dephosphorylation state of KaiC exhibits a circadian rhythm regulated by alternating autokinase and autophosphatase activity (Kitayama et al., 2008). KaiC alone undergoes dephosphorylation in vitro due to its innate phosphatase activity (Kim et al., 2008). KaiA turns on the kinase activity of KaiC by binding to the A-loop, whereas KaiB turns the kinase activity off by sequestering KaiA from the A-loop (Chang et al., 2015; Kim et al., 2008).
Although the Kai proteins alone produce a robust circadian rhythm in vitro, there are several other proteins whose mutation results in abnormal rhythms in vivo (Boyd et al., 2013; Ivleva et al., 2006). In this study, we focus on the properties of CikA, a histidine kinase that has multiple effects on the in vivo clock (Schmitz et al., 2000). CikA-null mutants exhibit smaller magnitude phase shifts in response to dark pulses, and oscillate with a shortened period and reduced amplitude (Schmitz et al., 2000; Zhang et al., 2006). Based on these observations, CikA was originally classified as an input pathway component (circadian input kinase A). However, recent biochemical evidence that CikA acts to dephosphorylate the output pathway transcription factor RpaA suggests this may be a misnomer, and that the CikA-null phenotype arises from a disrupted output signal (Gutu and O’Shea, 2013). To distinguish between these possibilities, the physical properties of the oscillator must be monitored instead of the output. For this purpose, examining the reconstituted oscillator is advantageous because monitoring KaiC phosphorylation is easier in vitro than in vivo. Here we used the reconstituted in vitro oscillator to study the effect of CikA on KaiC phosphorylation.
As mentioned above, it has been suggested that CikA also functions as an output pathway component by acting as a phosphatase against RpaA, a transcription factor regulated by the circadian clock (Gutu and O’Shea, 2013; Markson et al., 2013). CikA’s phosphatase activity against RpaA is activated when CikA binds to KaiB (Chang et al., 2015; Tseng et al., 2017). Analysis of the 3-dimensional structure of the KaiBC protein complex has shown that CikA binds to KaiB at the same binding site that is used to sequester KaiA (Tseng et al., 2017). In this study, we explore the consequences of this competitive interaction between CikA and KaiA on the ability of KaiA to stimulate KaiC phosphorylation.
To examine whether CikA directly affects the phosphorylation and dephosphorylation of KaiC, the phosphorylation state of KaiC was monitored with CikA added to the reaction mixtures (3.4 µM KaiC, 150 mM NaCl, 20 mM Tris-HCl, 5 mM MgCl2, 0.5 mM EDTA, 1 mM ATP; pH 8.0). All assays were performed as described previously (Kim et al., 2015) without any modification. CikA was purified using the same method as used for KaiC purification. Various concentrations of CikA were added to the reaction mixtures at time zero, with KaiA (1.2 µM) or KaiB (3.4 µM) for phosphorylation (Fig. 1A) or dephosphorylation (Fig. 1B), respectively. The phosphorylation state of KaiC was monitored for 24 h by collecting 20 µL of the reaction mixture every 2 h to run sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Because the addition of CikA did not change the phosphorylation state of KaiC appreciably (Fig. 1A and 1B), we concluded that CikA does not directly affect the phosphorylation and dephosphorylation of KaiC.
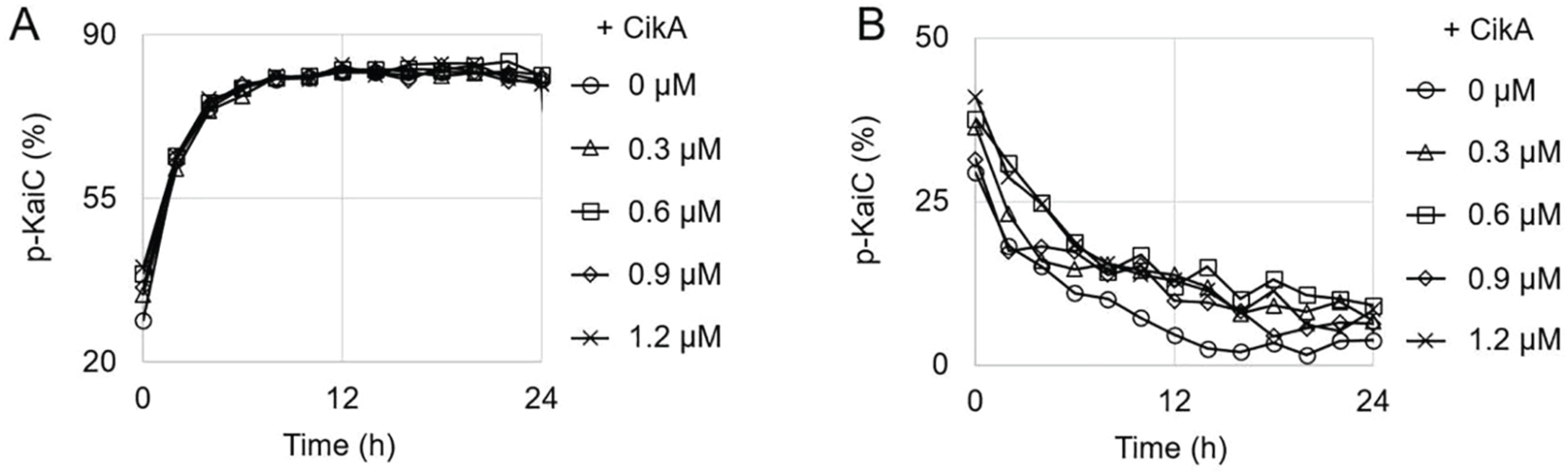
Direct effect of CikA on KaiC phosphorylation. (A) Phosphorylation traces of KaiC. KaiA (1.2 µM), KaiC (3.4 µM), and CikA (see the legend for concentrations) were mixed with adenosine triphosphate (ATP). The p-KaiC represents the percentage of total KaiC that is phosphorylated. The markers represent the experimental data points for KaiC phosphorylation (every 2 h). The graph is the best representative of 3 replicates. (B) Dephosphorylation traces of KaiC. KaiB (3.4 µM) was added instead of KaiA in the same reaction mixtures as in (A). Everything else is the same as in (A).
It has been suggested that CikA competes with KaiA for the binding site on KaiB to transfer the circadian signal generated by the circadian oscillator to the transcription factor (Chang et al., 2015). Structural analysis reveals that both CikA and KaiA bind competitively to the same site on KaiB bound to fully phosphorylated KaiC (Tseng et al., 2017). To observe the effect of this competitive binding on KaiC phosphorylation, we added various concentrations of CikA to the circadian oscillation reaction mixtures composed of KaiA, KaiB, and KaiC (Fig. 2A). The amplitudes of the oscillations decreased with each increase in CikA concentration (Fig. 2B). Compared with the oscillation in the absence of CikA, the amplitudes were decreased by 17% at 0.3 µM, 32% at 0.6 µM, 37% at 0.9 µM, and 54% at 1.2 µM of CikA. Phosphorylation levels at the peaks of the oscillation were not affected much by increasing the CikA concentration in the reaction mixtures (4% decreased with 1.2 µM of CikA); however, levels at the troughs were increased by 22% with 1.2 µM of CikA (Fig. 2A). Thus, overall, KaiC maintains a hyperphosphorylated state in the presence of CikA. CikA affected not only the amplitude but also the period of the circadian oscillation of KaiC phosphorylation. We computed period estimates from our data using 2 different methods: the Biodare2 (https://biodare2.ed.ac.uk/) implementation of the fast Fourier transform-nonlinear least squares (FFT-NLLS) procedure with linear detrending (Zielinski et al., 2014), and Lomb-Scargle periodograms with the MATLAB function plomb (Mathworks Inc, Natick, MA). These 2 approaches yielded very similar period estimates. We found that periods were shortened by increasing the CikA concentration in the reaction mixture (Fig. 2C). This result is compatible with a previous report performed with a controlled expression of CikA in cyanobacteria, in which increasing CikA expression results in a shorter period (Zhang et al., 2006). This hyperphosphorylation and the shortened period closely resemble the result of excess KaiA in the circadian oscillation reaction mixture (Kageyama et al., 2006; Nakajima et al., 2010). Because CikA competes with KaiA for binding to KaiB (Tseng et al., 2017), the addition of CikA in the circadian oscillation mixture leads to an increase in the concentration of free KaiA, which can bind to the A-loop of KaiC to activate the autokinase activity of KaiC (Kim et al., 2008). Since the addition of CikA produces a similar effect as the presence of excess KaiA in the reaction mixture (Kageyama et al., 2006; Nakajima et al., 2010; Rust et al., 2007), we decided to explore how varying CikA and KaiA concentrations in tandem affects the oscillation.
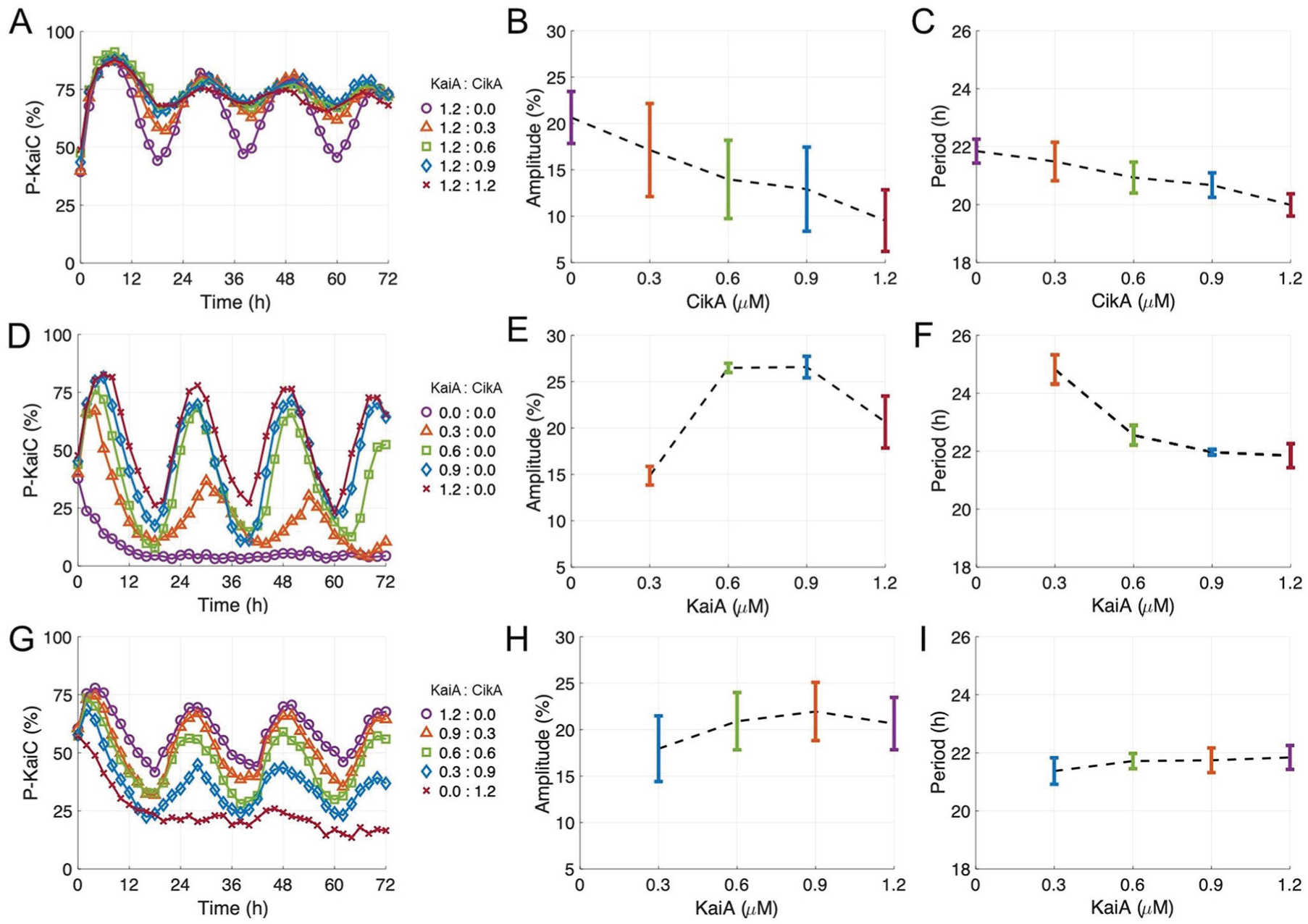
Effect of CikA on KaiC phosphorylation. Oscillation amplitude and period for various concentrations of KaiA and CikA (µM, listed as KaiA:CikA in legends) were calculated using FFT-NLLS. Error bars show standard error of the mean (SEM) from triplicates (except for the KaiA:CikA = 1.2:0.0 case, which had 6 replicates).
Earlier, it was reported that KaiA concentration in the reaction mixture is an important variable for the circadian period of KaiC phosphorylation. Previous reports (Kageyama et al., 2006; Nakajima et al., 2010) and our data demonstrate that decreasing KaiA concentration lengthens the period in the absence of CikA (Fig. 2D-F). We hypothesized that, in the presence of CikA, decreasing KaiA would not lead to lengthened periods, because more free KaiA would be released during the dephosphorylation phase of KaiC due to the competition with CikA for the binding site of KaiB. To test our hypothesis, we decreased the concentration of KaiA in the reaction mixture while simultaneously increasing the CikA concentration. The sum of the KaiA and CikA concentrations was maintained at 1.2 µM, which is the KaiA concentration specified in the original cyanobacterial circadian oscillation reaction mixture (Nakajima et al., 2005). The phosphorylation state of KaiC was monitored for 3 days to analyze the oscillatory properties of the reaction mixture. Differing from Figure 2D, here the phosphorylation state of KaiC maintained robust circadian rhythms with a stable circadian period as the CikA concentration was increased (Fig. 2G-I). In particular, the rhythms with small amplitudes at high concentrations of CikA in Figure 2B (up to a CikA concentration of 0.9 µM) can be recovered (at least 86%) into a robust amplitude circadian rhythm by decreasing the KaiA concentration (Fig. 2H). All reaction mixtures showed a similar circadian period (within an hour difference of each other), even though the KaiA and CikA concentration ratios were different in each reaction mixture (Fig. 2I). Thus, in the presence of CikA, a stable circadian oscillation can be generated with decreased KaiA concentrations despite KaiA concentration being an important variable for the period of KaiC phosphorylation (Kageyama et al., 2006; Nakajima et al., 2010). To explain this observation, we propose that CikA enhances the ability of KaiA to stimulate KaiC phosphorylation by competing KaiA out of its sequestered state. Specifically, because CikA competes with KaiA for the binding site on KaiB, it increases free KaiA, which stimulates the phosphorylation of KaiC by stabilizing the exposed conformation of the A-loop. Therefore, more CikA in the reaction mixture can release more free KaiA by occupying its binding site on KaiB, keeping the kinase activity of KaiC turned on with comparatively small amounts of total KaiA available.
In conclusion, we found that CikA affects the physical properties of the circadian oscillator, which implies input functionality. Although CikA does not have a large effect on the self-sustained period, it is still possible that day/night alternations in CikA abundance in cyanobacteria (Ivleva et al., 2005) play a role in entraining the KaiABC oscillator, since small changes in the self-sustained period can have a large effect on the entrained phase (Granada et al., 2013). We also propose a model for the mechanism of how CikA influences KaiC phosphorylation (Fig. 3): When KaiC is fully phosphorylated (Kim et al., 2008), KaiB binds to the B-loop of KaiC and sequesters KaiA from the A-loop of KaiC (Chang et al., 2015). If CikA is present at this point, it competes with KaiA for the binding site on KaiB. The displaced KaiA can bind to the A-loop to activate the kinase activity of KaiC (Fig. 3). As a result, CikA indirectly alters the phosphorylation state of KaiC.
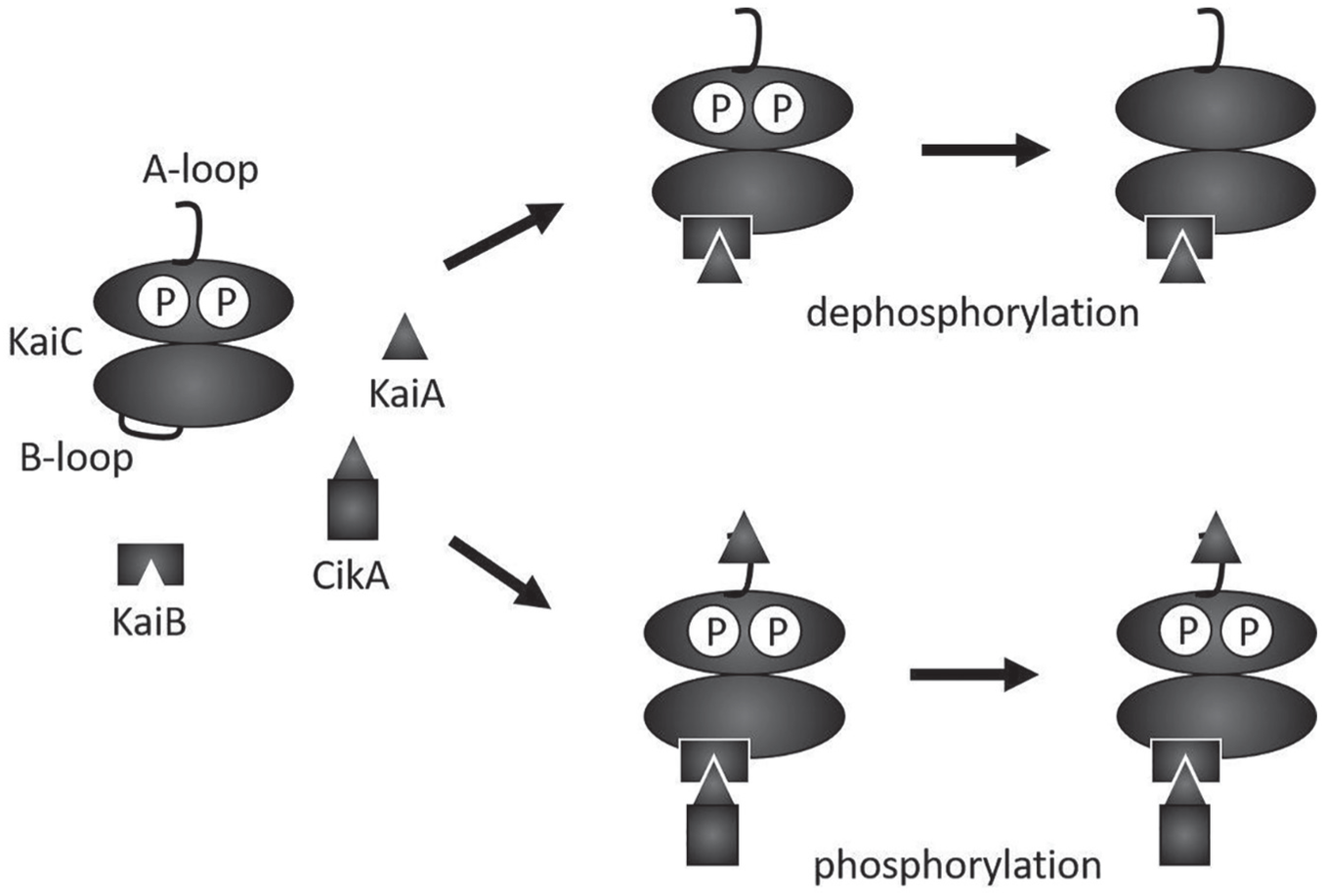
Model for the effect of CikA on KaiC phosphorylation. When KaiC is fully phosphorylated during the day, KaiB binds to the B-loop of KaiC and sequesters KaiA. CikA competes with KaiA for the binding site on KaiB. If CikA occupies the binding site, the KaiA released from KaiB binds to the A-loop of KaiC and keeps KaiC hyperphosphorylated.
Many current mathematical models of the cyanobacterial circadian oscillator have been made only considering interactions between the Kai proteins (Das et al., 2017; Leypunskiy et al., 2017; Miyoshi et al., 2007). By introducing CikA into the cyanobacterial circadian oscillator as an input component, it is possible to build a more accurate model of the cyanobacterial circadian clock. Furthermore, the functionality of CikA as an input component has not been examined extensively in vitro. Although a few in vitro entrainment studies have been performed without CikA, their results were partially inconsistent with the findings from in vivo experiments (Kim et al., 2012; Rust et al., 2011). Here, we have successfully generated a stable circadian period in the presence of CikA by varying the ratio between KaiA and CikA. This finding leads us one step closer to the in vitro reconstitution of the circadian clock, and not just the circadian oscillator, by adding the input and the output components to the reaction mixture. This will enable further studies on the entrainment of cyanobacterial circadian rhythms using phase response curves (Johnson, 1999) and entrainment maps (Diekman and Bose, 2016).
Footnotes
Acknowledgements
We thank Hana Nim for insightful discussions and Young Jeong, Helene Brochon, and Brianna Porr for technical assistance. This work was supported by a New Jersey Institute of Technology start-up grant to Y.I. Kim, and the National Science Foundation grant DMS-1555237 and the U.S. Army Research Office grant W911NF-16-1-0584 to C. Diekman.
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
