Abstract
Animal models are essential tools for modern scientists to conduct biological experiments and investigate their hypotheses in vivo. However, for the past decade, raising the throughput of such animal experiments has been a great challenge. Conventionally, in vivo high-throughput assay was achieved through large-scale mutagen-driven forward genetic screening, which took years to find causal genes. In contrast, reverse genetics accelerated the causal gene identification process, but its throughput was also limited by 2 barriers, that is, the genome modification step and the time-consuming crossing step. Defined as genetics without crossing, next-generation genetics is able to produce gene-modified animals that can be analyzed at the founder generation (F0). This method is or can be accomplished through recent technological advances in gene editing and virus-based efficient gene modifications. Notably, next-generation genetics has accelerated the process of cross-species studies, and it will be a useful technique during animal experiments as it can provide genetic perturbation at an individual level without crossing. In this review, we begin by introducing the history of animal-based high-throughput analysis, with a specific focus on chronobiology. We then describe ways that gene modification efficiency during animal experiments was enhanced and why crossing remained a barrier to reaching higher efficiency. Moreover, we mention the Triple CRISPR as a critical technique for achieving next-generation genetics. Finally, we discuss the potential applications and limitations of next-generation mammalian genetics.
Animal models have contributed to the elucidation of mechanisms in various life systems. Specifically, studies using non-mammalian organisms, such as Escherichia coli, Caenorhabditis elegans, Saccharomyces cerevisiae, Schizosaccharomyces pombe, Neurospora, Cyanobacteria, and Drosophila melanogaster, have considerably contributed to the development of genetics and molecular biology. Moreover, studies on the KaiA-C in cyanobacteria and the clock genes in Neurospora crassa, such as FRQ, have made important contributions to the molecular mechanisms of the biological clock (Dunlap, 1999; Dunlap and Loros, 2004; Iwasaki and Kondo, 2004; Cohen and Golden, 2015; Dunlap and Loros, 2017; Loros, 2020). Epoch-making studies have utilized fruit flies in chronobiology, that is, isolation of per0, pers, and perl mutant flies, identifying causal gene period and proposing the circadian oscillation circuit based on the transcriptional-translational feedback loop (Konopka and Benzer, 1971; Reddy et al., 1984; Zehring et al., 1984; Hardin et al., 1990).
Mammalian studies have also contributed to life science, especially in the biomedical field. Rodents, such as hamsters, guinea pigs, rats, and mice, have been involved in animal experiments. Among these, mice are considered the king of model animals because (1) they are easy to handle (small storage space and easy to breed); (2) they have relatively short life cycles; (3) many inbred lines have been established; and (4) it is possible to generate gene-modified mice such as transgenic (TG), knockout (KO), and knock-in (KI) mice. Conventionally, the throughput of experiments involving genetically modified animals was relatively low, but the situation changed due to recent technological advances. In this review, we focused on high-throughput assays using mice supported by biotechnological advances. In addition, we summarized the history of high-throughput assays in chronobiology, discussed the barriers for raising throughputs, and introduced the “next-generation mammalian genetics,” which is defined as genetics without crossing (Susaki et al., 2017; Ukai et al., 2017, 2019). Moreover, applicable targets for next-generation genetics, such as genetically humanized animals, were also discussed.
ENU Mutant Mouse and Conventional KO Mouse
Forward Genetic Approach
One of the origins of the clock genes discovery is the studies using Drosophila. Although researchers back then employed fruit flies as a simple model, it took over a decade for scientists to identify the causal gene period from period mutant flies and 13 more years for them to find the human orthologue of drosophila period gene (Konopka and Benzer, 1971; Reddy et al., 1984; Zehring et al., 1984; Tei et al., 1997). Nowadays, mammalian molecular clock machinery is well described (Takahashi, 2017). Takahashi’s group opened the door for mammalian clock genetics through their success in forward genetic N-ethyl N-nitrosourea (ENU) screening, identifying a rhythm disorganized Clock mutant mouse and the causative Clock gene (Vitaterna et al., 1994; King et al., 1997). It is also noteworthy that mutagen-based screening continues to generate new findings, as represented by the discovery of Fbxl3 (Godinho et al., 2007; Siepka et al., 2007). In the case of natural circadian rhythm mutant Tau hamster, it took over a decade for the causative gene CKIε to be identified (Ralph and Menaker, 1988; Lowrey et al., 2000). As it is uncommon to discover natural-born clock mutants, mutagenesis screenings, due to their high efficiency, have become a precursor of high-throughput in vivo analysis.
Reverse Genetic Approach
Conventional KO Mouse
Through applying a gene-driven approach, several genes were recognized as clock genes, which are the primary components of the molecular clock mechanism. For instance, the specific functions of Per1-3 and Bmal1 genes in vivo were confirmed by generating conventional embryonic stem cell (ESC)-based KO mice (Zheng et al., 1999; Bunger et al., 2000; Shearman et al., 2000; Bae et al., 2001; Cermakian et al., 2001; Zheng et al., 2001). Similarly, Cry1 and Cry2 were identified as clock genes through analyzing ES-based KO mice (van der Horst et al., 1999; Vitaterna et al., 1999). It should be noted that a global collaboration to systematically knock out all genes in the mouse genome promoted the reverse genetics approach (Guan et al., 2010). International projects, such as the Mutant Mouse Resource & Research Center at UC Davis (https://mmrrc.ucdavis.edu/mmrrc-program/), aim to prepare KO-ESCs and KO mice libraries based on gene trapping, gene targeting, and/or gene-editing technologies. The International Mouse Phenotyping Consortium (IMPC) project (https://www.mousephenotype.org/) is creating a genome-wide annotation of gene functions, which is done through systematically developing and phenotyping a collection of standardized single-gene KO mice (Beckers et al., 2009; de Angelis et al., 2015; Dickinson et al., 2016). In 2020, Zhang et al. analyzed the indirect calorimetry (IC) data set from over 2000 wild-type mice collected by IMPC and established a machine learning program. They screened mice from 750 mutant lines and identified 5 novel genes involved in circadian misalignment (Zhang et al., 2020). Similarly, for-profit companies have KO mice libraries, and researchers can even order gene-modified animals online.
Conditional KO Mouse
Conditional KO mice are generated by various methods, including the Cre/LoxP system. They have been utilized to study the spatio-temporal function of clock genes. Bmal1 is a suitable target to study the spatio-temporal function of the molecular clock. Through a single-gene disruption, the normal circadian rhythmicity can be abolished (Bunger et al., 2000). As a pioneering study, Storch et al. (2007) developed mice with a conditional allele, Bmal1, and analyzed the effect of retina-specific loss of Bmal1 function by crossing mice with a CHX10-Cre transgene. Other groups also developed Bmal1 floxed mutant mice independently (Westgate et al., 2008; Shimba et al., 2011; Ratajczak et al., 2012). Here, floxed indicates a part of Bmal1 that is sandwiched by the LoxP sites. By crossing the floxed mice with the cell-type-specific Cre recombinase expressing mice, researchers performed various organ-specific loss of clock function studies on organs including muscle, bone, cartilage, adrenal cortex, kidney, and immune cells (Shimba et al., 2011; Gibbs et al., 2012; Takarada et al., 2012; Dyar et al., 2014; Dumbell et al., 2016; Nikolaeva et al., 2016; Takarada et al., 2017; Dan et al., 2020; Qian et al., 2020). In the brain, especially in the suprachiasmatic nucleus (SCN) of the clock center, the development of conditional Bmal1 KO mouse contributes to unveiling cell-type-specific molecular clock functions (Mieda, 2019; Hastings et al., 2020). Following are examples: GABA-neuron-specific Bmal1 KO mouse, AVP-neuron-specific Bmal1 KO mouse, and VIP-neuron-specific Bmal1 KO mouse (Mieda et al., 2015; Weaver et al., 2018; Todd et al., 2020). Along with the increasing demands on conditional KO tools, international repositories have offered such mice to the scientific community (McMurray et al., 2012; Murray et al., 2012; H. Lee, 2021). One representative depository is the JAX Cre Repository (https://www.jax.org). It offers more than 300 Cre tool strains, including Cre expressing strains, inducible Cre strains, Cre reporter strains, and loxP-flanked (floxed) strains (https://www.jax.org/research-and-faculty/resources/cre-repository).
High-Throughput Screening in Vitro
Mammalian Cell-based Screening
Compared with animal studies, cell-based analysis possesses its own notable advantages in terms of scale, space, and speed. Molecular clock studies in cultured mammalian cells began from the Rat-1 study by Schibler’s group (Balsalobre et al., 1998). Since the molecular oscillators in the cultured cells that the researchers used were identical to those in the SCN, they could investigate molecular clock function utilizing cultured cell lines (Yagita et al., 2001). Nowadays, numerous laboratories employ NIH3T3 fibroblast as a mouse clock model and U-2OS bone osteosarcoma as a human clock model (Akashi and Nishida, 2000; Vollmers et al., 2008; Isojima et al., 2009). Although the mammalian cell-based screening is not the primary focus of this review, we introduced it briefly here from a perspective of high-throughput assays.
Genetic Perturbations
Supported by the development of bioinformatics and laboratory automation, cell-based screening has become available. Particularly, it consists of genome-wide gene modification tools, including full-length cDNA, shRNA, and siRNA libraries. Using U-2OS cells, one group developed a siRNA-based circadian rhythm phenotype screening system based on a 384-well plate (Vollmers et al., 2008). Maier et al. (2009) tested 2660 RNAi constructs targeting kinases and phosphatases, and Zhang et al. (2009) knocked down approximately 1000 genes genome-wide by RNAi so as to investigate molecular clock components/modifiers. From the time when clustered regulatory interspaced short palindromic repeat (CRISPR)/CRISPR associated protein (Cas) 9 appeared, modifying the genomes of cell lines became easier. In terms of clock genes, lipofection-based Csnk1d and Csnk1e KO NIH3T3 cells, lentivirus-based Fbxl3 KO U-2OS cell, and adenovirus-based Bmal1 and Per2 KO U-2OS cell were reported (Korge et al., 2015; Guo et al., 2019; Jin et al., 2019). Notably, gene-editing based KO strategy will enable researchers to perform genome-wide KO analysis in vitro in the near future. Details of the CRISPR/Cas method will be discussed later.
Chemical Perturbations
Compared with genetic perturbations, chemical perturbations are more suitable for cell-based high-throughput analysis. In chronobiology, the concept of chemical biology appeared as early as in 2007, along with the many successful cases reported afterward (Liu et al., 2007). Two groups tested 1280 chemicals in a library of pharmacologically active compounds (LOPAC) for their effects on circadian period length (Hirota et al., 2008; Isojima et al., 2009). Through the study, our group succeeded to identify that temperature-insensitive phosphorylation of CKIδ defines period length. Shinohara et al. (2017) further identified aurintricarboxylic acid (ATA) as a dephosphorylation-specific inhibitor of CKIδ using LOPAC1280 and studied how temperature-independent phosphorylation reactions were achieved. Kay, Hirota, and their colleagues promoted a chemical-biological approach. They successfully identified small circadian rhythm modifying molecules utilizing U-2OS systems. These molecules include Longdaysisn, which targets CKIδ, CKIα, and ERK, LH846, which inhibits CKIδ specifically, and KL001, which prevents the ubiquitin-dependent degradation of CRYs (Hirota et al., 2010; Lee et al., 2011; Hirota et al., 2012). Moreover, Chen et al. (2012) performed a high-throughput screening of ~200,000 synthetic compounds using the fibroblasts from adult mouse’s ear, which harbored Per2::LucSV fusion gene, and found 11 circadian period and amplitude modifiers. He et al. (2016) screened an in-house collection of 5300 components using the heterozygous ClockΔ19/+ PER2::Luc reporter cells, which exhibited sustained reporter rhythms with low amplitudes, and discovered the circadian rhythm amplitude enhancer, Nobiletin. Reviews on various small circadian rhythm modifiers are available elsewhere (Hirota and Kay, 2015; Gloston et al., 2017; Chen et al., 2018; Miller and Hirota, 2020).
The Bottlenecks for High-Throughput Screening in Vivo
Two Bottlenecks of Genetically Modified Mice
Although forward genetic screenings were the pioneer of high-throughput analysis in vivo, it took years for such screenings to develop from the separation of mutants to the identification of causal genes. Reverse genetics altered the situation, but the generation and the preparation of gene-modified animals for testing phenotype remained laborious and time-consuming. The difficulties for increasing the experimental scales and experimental speeds exist in the genome manipulation and the crossing steps (Susaki et al., 2017) (Figure 1a).
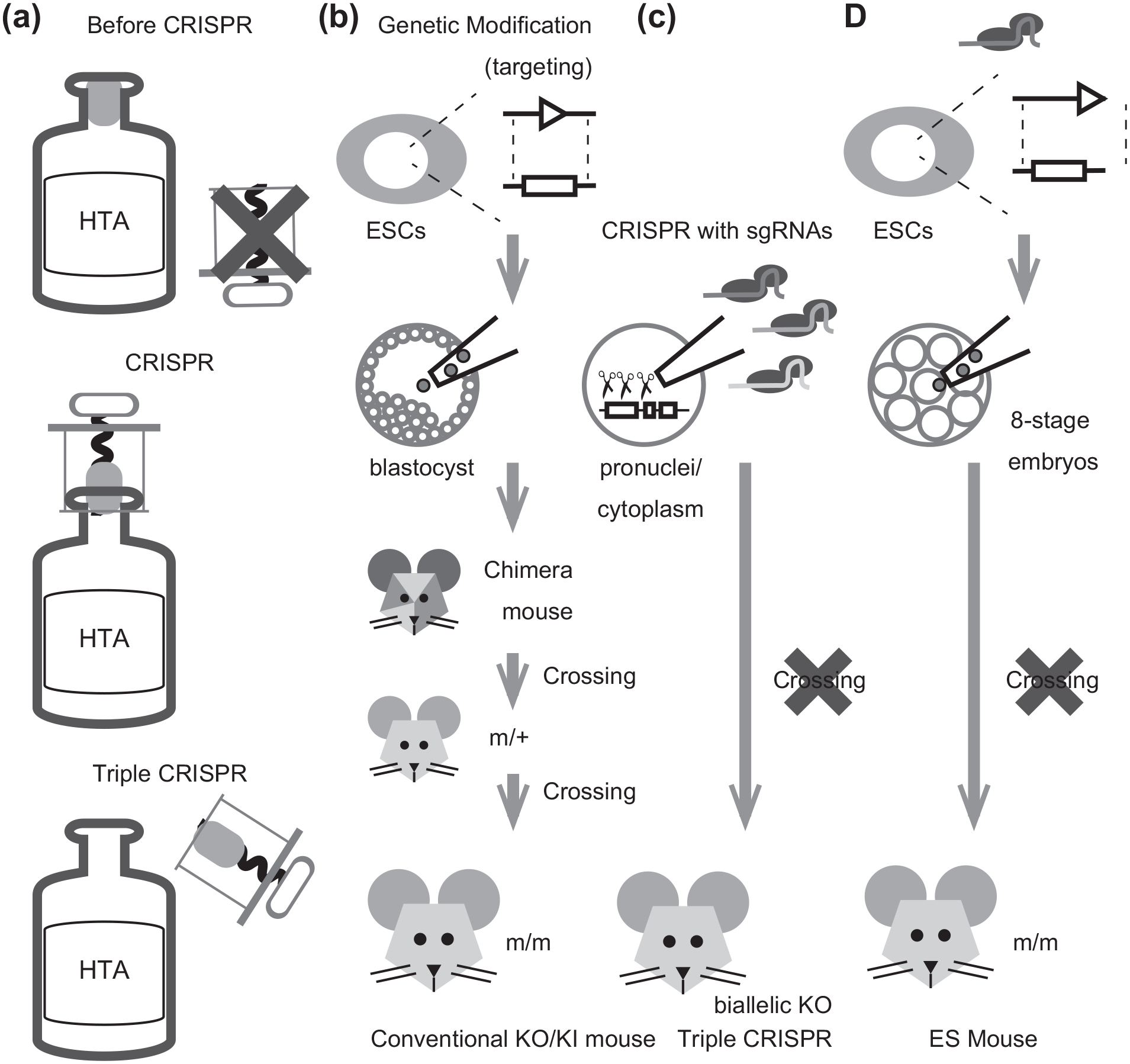
A conceptual illustration of next-generation mammalian genetics. (a) Three bottles analogy. Before the development of CRISPR technology, the bottle (high-throughput analysis in vivo; HTA) was sealed tightly (top), but the CRISPR technology had increased the efficiency of genetic modification in mice and thus loosened the stopper in the bottle. The Triple CRISPR method eliminated the need to cross, which is illustrated as pulling the stopper fully from the bottle (bottom). (b-d) Steps of gene modification animal production in conventional KO/KI mice (b), Triple CRISPR (c), and ES mice (d). Note that the crossing step is necessary for KO/KI mice. F0 chimera mice are crossed to wild-type mice and the F1 heterozygotes (m/+) are intercrossed to obtain the homozygotes (m/m). If ESCs are derived from the hybrids or the 129 strains, a further backcrossing step is needed (~10 generations) (b). In cases of the Triple CRISPR and ES mice, the F0 mice can be used for analysis and no further crossing needed (c-d). Details are described in the main texts. Square indicates the original gene and triangle indicates the mutated gene. Abbreviations: CRISPR = clustered regulatory interspaced short palindromic repeat; KO = knock out; KI = knock in; ESCs = embryonic stem cells; HTA = High-throughput analysis.
“Genomic Manipulations Are No Longer an Experimental Bottleneck”
Conventionally, a mutation needs to be introduced to the ESCs in order to produce a KO or KI mouse, which is a laborious process. A specific gene is either disrupted or inserted through gene targeting based on the spontaneous homologous recombination, which occurs at low frequencies (Austin et al., 2004; Capecchi, 2005). However, the genomic manipulation step has become easier due to the appearance of genome-editing technologies. Such technologies rely on DNA double-strand break repair mechanisms and have been applied in the forms of TALEN (Transcription activator-like effector nuclease), ZFN (Zinc finger nucleases), and CRISPR/Cas-based methods (Gaj et al., 2013; Chandrasegaran and Carroll, 2016). CRISPR/Cas9 system is composed of Cas9 DNA endonuclease, CRISPR RNA (crRNA), and trans-activating crRNA (tracrRNA). This tracrRNA typically binds to the crRNA and forms a ribonucleoprotein complex with Cas9. The crRNA then guides the Cas9 complex to the target sequence (Deltcheva et al., 2011). This 2-component RNA system was further simplified to synthetic single-guide RNA (sgRNA) consisting of a fusion of crRNA and tracrRNA. Today, the CRISPR/Cas9 system is widely applied in genome editing for knocking out genes (Adli, 2018). For example, introducing the Cas9 mRNA with 1 sgRNA into a zygote can efficiently generate mosaic mice containing KO cells. As stated in the Scientific Background part of the Nobel Prize in Chemistry 2020, “genomic manipulations are no longer an experimental bottleneck.”
Crossing Remains a Bottleneck
In terms of the conventional KO/KI mice, the KO/KI-ESCs are injected into the wild-type blastocysts. This process gives rise to the chimera mice developed from both the ESCs and the wild-type host’s embryo-derived cells. The obtained chimera mice (F0) are crossed with the wild-type mice, and their F1 generation carries a mutated genome if germline transmission occurs. According to Mendel’s Laws, a homozygous mutant is expected to appear in the F2 generation. This suggests that at least 9 months (3 months/generation x 3 generations) are needed to obtain the first homozygous mutant. Furthermore, the resulting mutants often need to backcross to some inbred strains, such as the C57BL/6, several times if the mutants are generated using the F1 hybrids or the 129 strain derived ESCs. These crossing steps cannot be abbreviated and remain a problem (Figure 1b). In contrast to the laborious methods for producing conventional KI/KO mice, the CRISPR/Cas9-based genome edited mice require no such backcrossing if they are generated based on a desired inbred genetic background. Because using multiple sgRNAs simultaneously can introduce mutations in various genes, KO mice with multiple dysfunctional genes can be obtained without crossing (Wang et al., 2013). However, in this case, the founder mice (F0) often contain a mosaic of wild-type and KO cells, and the rate of generating complete biallelic mutant mice is relatively low. Thus, though they could avoid the backcrossing issue, the mice generated using the CRISPR/Cas9 technology still consume time in experiments because in real practice, expanding colonies by crossing would still be necessary to obtain enough mice to analyze, and this takes time.
Next-Generation Mammalian Genetics
Genetics Without Crossing
To increase the throughput of analysis in vivo and propel mammalian genetics to the next generation, we must tackle the “crossing” issue. We define next-generation mammalian genetics as the “genetics without crossing” (Susaki et al., 2017; Ukai et al., 2017, 2019). This can be achieved through efficient KO/KI methods that generate analyzable KO/KI mice within one generation. There are 2 possibilities: generating whole-body biallelic mutants with almost 100% probability using the CRISPR/Cas9 method (or other gene-editing methods), or generating mice almost entirely from ESCs derived cells. From an ethical viewpoint, next-generation genetics contributes to reducing the number of experimental animals per study. Because the timing of the birth of F0 mutants depends on the experimental schedule, it is possible to systematically obtain animals that are matched in ages and genders. In addition, researchers do not obtain heterozygous mutants that are useful for mating but are normally useless in experiments (Figure 1c and 1d).
Triple-CRISPR Method
Triple-targeted CRISPR (Triple CRISPR) is a method that employs 3 kinds of sgRNA simultaneously targeting 1 gene. From a computational analysis, Sunagawa et al. found that a multiple-target CRISPR strategy is more efficient than a single-targeted CRISPR, which is consistent with the experimental results from a dual-target CRISPR case (Zhou et al., 2014). Experimentally, the biallelic KO efficiencies of a single gene (Tyr) using each of the 3 sgRNAs was 36.0%, 54.2%, and 64.7%, respectively. However, through Triple CRISPR, the biallelic KO efficiency of the Tyr gene increased to 97.5%, which was much higher than the minimum computational estimation, 82.6%. This increase was probably due to the irreversible mutations resulting from the Triple CRISPR. It is of note that the increased efficiency was reproducible by using other sets of 3 sgRNAs. Furthermore, the possibility of the off-target effects could be practically eliminated by comparing phenotypes of KO mice resulting from 2 different sets of 3 sgRNAs (Sunagawa et al., 2016).
Triple CRISPR has enabled us to perform gene-driven phenotype analysis at the F0 generation. This advancement allows us to implement high-throughput in vivo analyses, such as a comprehensive KO mice study of 1 gene family at a single laboratory level (Figure 2a). Following are some examples from previous sleep research, which employed the triple-CRISPR technology. Sunagawa et al. (2016) generated and analyzed the sleep phenotypes of KO mice with mutations in the Tyr, Bmal1, Hcrt, and NMDA receptors (5 genes). Tatsuki et al. (2016) found the importance of intracellular calcium concentration in sleep regulation by generating KOs of Ca2+-dependent K+ channels (8 genes), voltage gated Ca2+ channel α1 subunits (10 genes), plasma membrane Ca2+ ATPases (4 genes), and calcium/calmodulin-dependent protein kinase type II (4 genes). Niwa et al. (2018) identified muscarinic acetylcholine receptors, Chrm1 and Chrm3, as the essential genes for rapid eye movement (REM) sleep by generating and analyzing a total of 16 KO mice with dysfunctional nicotinic acetylcholine receptor (11 genes) and muscarinic acetylcholine receptors (5 genes). Yoshida et al. (2018) studied the importance of the leak potassium channel. They identified Kcnk9 as an essential regulator of sleep length by producing and examining KOs of leak potassium channel genes (14 genes).
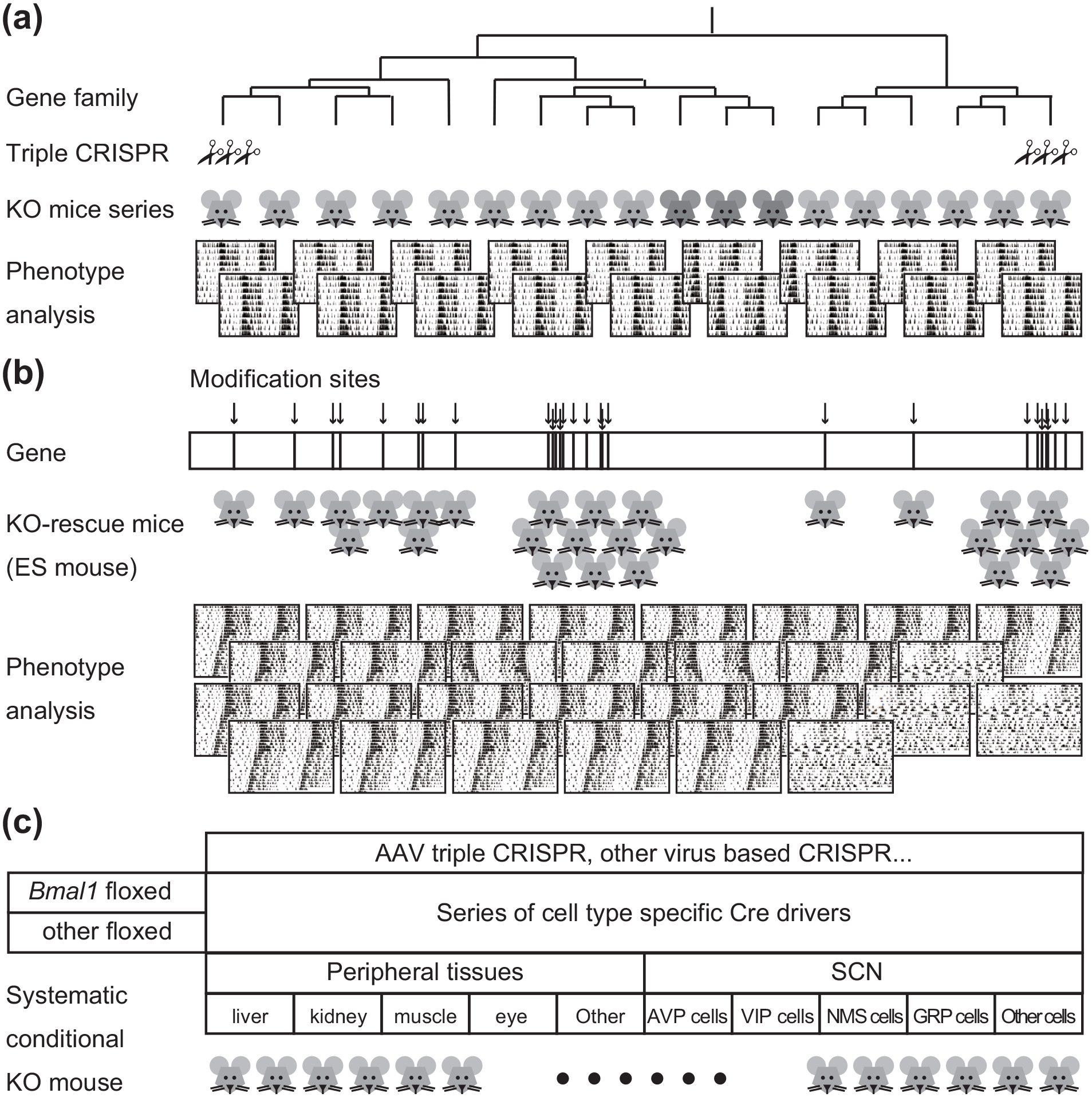
Comprehensive system-level analyses achieved by or will be achieved by next-generation mammalian genetics. (a) A comprehensive KO mice analysis covering the whole 1 gene family. In this review, we showed examples of KO mice covering almost all gene family, which includes Nr2s, Nr3s, Kcns, Cacna1s, Atp2bs, Camk2s, Chrnas, Chrnbs, Chrms, and Kcnks (Sunagawa et al., 2016; Tatsuki et al., 2016; Niwa et al., 2018; Yoshida et al., 2018). (b) A comprehensive gene structure–level analysis by KO-KI rescue using ES mouse. By changing the modification sites systematically (such as the phosphorylation sites) and introducing the gene-of-interest KO-ESCs, the corresponding series of KO-rescue ESCs/mice can be generated. In this review, the Cry1 phosphorylation sites were discussed as an example (Ode et al., 2017). (c) A systematic generation of conditional KO mice by the high-efficacy Cre/loxP system and/or the AAV-Triple-CRISPR technology. The expansion of Cre driver lines and/or the development of a virus-based CRISPR/Cas9 method will enable us to generate systematic conditional KO mice throughout all body/cell types. Abbreviations: KO = knock out; KI = knock in; ESCs = embryonic stem cells; AAV = adeno-associated virus; CRISPR = clustered regulatory interspaced short palindromic repeat; SCN = suprachiasmatic nucleus; ES cells = embryonic stem cells; AVP = arginine vasopressin; VIP = vasoactive intestinal peptide; NMS = neuromedin-S; GRP = gastrin-releasing peptide.
The ES Mouse
The ES KO Mouse
The ES mouse is almost entirely ESC-derived and carries mutations throughout the body in its F0 generation (Figure 1d). The tetraploid complementation method was first applied to generate the ES mouse (Duncan, 2005). The other approach was the microinjection of ESCs to 8-cell-stage embryos (Poueymirou et al., 2007; DeChiara et al., 2009, 2010). Huang et al. (2008) produced ES mice via ESCs injection to 4- or 8-cell embryos. ESCs are typically maintained with leukemia inhibitory factor (LIF) and serum (or BMP) to keep their undifferentiated states. LIF can activate STAT3, which additionally triggers Sox3 and Nanog. All these further activate Oct3/4, keeping the pluripotency of the ESCs (Niwa et al., 2009). Moreover, the ERK inhibitor, FGF receptor inhibitor, and GSK3 inhibitor (3i) enable efficient derivation and germline-competent propagation (Ying et al., 2008). Kiyonari et al. (2010) established and maintained C57BL/6 N derived ESC line with serum-free medium supplemented 3i, and they succeeded in producing the ES mouse. As an example, the feasibility of Kiyonari’s system in gene KO mice was shown in Chrm1/Chrm3 double KO mice production (Niwa et al., 2018). Although the ES KO mice are powerful tools, their importance pales in comparison to what can be done with the CRISPR editing. Only a limited number of groups still use this technique now.
The ES KI Mouse
The value of ES mice could be shown in the generation of KI mice. The reliability and robustness of the ES KI mouse method were witnessed in several studies (Susaki et al., 2014; Tainaka et al., 2014; Wang et al., 2016). Ukai et al. have improved the efficiency of the KI ESCs production, producing various types of KI ESCs in parallel (Ode et al., 2017; Ukai et al., 2017). Furthermore, combining their procedure with the ES mouse technology (Kiyonari et al., 2010), systematic production of diverse kinds of KI ES mice in parallel was achieved using a short period of time (2-3 months) (Ukai et al., 2019).
Loss-of-function and gain-of-function studies are necessary pairs to show causal relationships between gene(s) and phenotype. Antoch et al. illustrated a causal relationship between the behavioral phenotype of Clock mutant mouse and ClockΔ19 mutation via an in vivo complementation study. In this research, they generated TG mice harboring bacterial artificial chromosomes (BAC), including the wild-type Clock gene (Antoch et al., 1997). However, the inverse procedures called “KI and KO-rescue” remained arduous and expensive until recently. As the ES mouse technology enabled people to analyze various kinds of KI-KO rescue mice in parallel during a short duration, researchers were able to perform a high-throughput KI and KO-rescue study for analyzing phenotypes of various Cry1 mutants (Ode et al., 2017) (Figure 2b). First, Cry1-/-, Cry2-/- ES mice were generated and confirmed that they displayed no rhythmic behaviors as previously reported in conventional KO mice (van der Horst et al., 1999; Vitaterna et al., 1999). This arrhythmic phenotype was then rescued by inserting a wild-type Cry1 gene in the ROSA26 sites. Moreover, the authors selected 17 CRY1 mutations and added these mutated CRY1 genes into different ESCs, which lacked Cry genes. The mutant CRY1 rescue-ES mice were further analyzed. Compare with the wild-types, the Cry1(S243D)-rescued ES mouse displayed a shorter circadian rhythm behavior, whereas the Cry1(S243A)-rescued ES mouse showed a longer circadian rhythm behavior (Ode et al., 2017). Recent technological advances have enabled us to establish KI mice using the CRISPR/Cas technique (see below). The CRISPR/Cas-based technology, unlike ES mice, is more commonly employed by research groups when generating KI-KO rescue mice.
Generation of Region-Specific Gene Modification Animals in Short Duration
One-step Conditional KO Mouse
Although a significant amount of information has been obtained using the conventional KO/KI strategies, they generally lack spatial and temporal specificity, which can be improved by conditional KO approaches. In the Cre/LoxP system case, researchers are required to obtain both a Cre driver mouse and a floxed gene-of-interest mouse first and then mate them to obtain the desired conditional KO mouse. Extensive efforts have been made to accelerate the process of sandwiching part of a gene at the LoxP sites through the CRISPR/Cas-based KI strategy (Yang et al., 2013; Nakagawa et al., 2016; Nishizono et al., 2020). CRISPR/Cas9-based high-efficient KI technologies with electroporation have been reported and improved by many researchers (Horii et al., 2017; Miyasaka et al., 2018). A long single-stranded DNA (lss DNA) homology directed repair (HDR) approach sped up the generation of floxed animals (Yoshimi et al., 2016; Codner et al., 2018). Efficient addition with ssDNA inserts-CRISPR (Easi-CRISPR) is one of such technologies, which uses an lss DNA and Cas9 protein. Quadros et al. (2017) succeeded in generating 7 floxed mice and 6 KI mice with 8.5% to 100% and 25% to 67% success rates, respectively. Ohtsuka et al. showed that genome editing could be performed directly on fertilized eggs in oviducts and called this technique the improved-genome editing via oviductal nucleic acids delivery (iGONAD) (Ohtsuka et al., 2018; Gurumurthy et al., 2019). This method can also be employed for generating conditional alleles (Shang et al., 2021). Using these technologies with oocytes carrying tissue-specific Cre, 1-step generation of the conditional KO in F0 mouse could be accomplished (Horii et al., 2017; Miyasaka et al., 2018).
Cross-Species Studies Accelerated by Next-Generation Mammalian Genetics
Humanized Animal Models
Humanized animals are defined as experimental animal models xenotransplanted with human cells, tissues, organs, and/or expressing human gene products (Walsh et al., 2017; Brendel et al., 2020; Moriwaki et al., 2020; Stripecke et al., 2020). As a xenotransplantation model, humanized immune system mice, created from engrafting human stem cells to severely immunodeficient mice, are well-known examples (McCune et al., 1988; Mosier et al., 1988; Shultz et al., 1995; Ito et al., 2002; Allen et al., 2019). A genomically humanized mouse is a mouse model produced by introducing a human genomic region to a mouse, and it enables us to more comprehensively understand human health and diseases (Zhu et al., 2019; Moriwaki et al., 2020). At first, researchers employed transgenic mice with a piece of human genomic region. They then utilized a KI mouse and replaced the genome sequence with a human orthologous DNA (Devoy et al., 2011). In 2001, Fu’s group identified a S662G mutation in the human PER2 gene as a causal mutation for a kindred diagnosed with familial advanced sleep phase syndrome (FASPS) (Toh et al., 2001). Moreover, the group introduced a BAC harboring either the wild-type or the mutated PER2s, including the S662G mutation, to mice lacking the innate Per2 gene (Per2-/-). They discovered that the S662G-PER2/Per2-/- BAC-TG mouse exhibited a shorter behavioral rhythm in constant darkness (Y. Xu et al., 2007). Moreover, numerous coding and non-coding human PER3 polymorphisms are associated with diverse phenotypes and diseases (Archer et al., 2018). Particularly, Hasan et al. generated 2 lines of humanized PER3 mice, which contained a human PER3 variable-number tandem-repeat (VNTR) insertion at Per3 exon 17. It was concluded here that the PER3 VNTR polymorphism modified the homeostatic sleep response without significantly influencing circadian parameters (Hasan et al., 2014).
Improving the efficiency of humanized mouse production by KI and testing various types of KI mice in parallel are imperative steps in assessing the phenotypic diversity of diseases and/or evaluating the effects of multiple candidates of disease-related single nucleotide polymorphisms (SNPs). Notably, gene-editing technology will accelerate the development of KI-based-humanized mice. A KI method using single-stranded oligodeoxynucleotides (ssODNs) as a donor DNA in genome-editing tools, such as the CRISPR/Cas9, has become the most popular option in introducing an SNP (Bollen et al., 2018). In terms of middle-range insertion (a few hundred bp), KI methods that use lss DNA and/or double-strand DNA are employed by researchers (Miyasaka et al., 2018; Yao et al., 2018). For long-range insertion (approximately 200 kb), such as replacing an entire mouse gene with a human gene, a KI method utilizing 2 ssODNs is considered useful (Yoshimi et al., 2016). Currently these technologies are commercially provided, and researchers can even order humanized animals online. Ukai et al. (2019) further explained this humanized animals’ possibility in their previous review.
GWAS Studies for Considering Possible Candidates of Humanization
Many mental illnesses, including Alzheimer’s disease, bipolar disorder, dysthymia, major depression, Parkinson’s disease, psychosis, seasonal affective disorder, and schizophrenia, were found to be associated with clock genes, including BMAL1, CLOCK, CRY1, CRY2, NR1D1, PER1, PER2, PER3, RORA, and RORB (Jagannath et al., 2013, 2017). Thus, genetically modified mice harboring humanized clock genes with SNPs could contribute to unveiling the real associations between circadian clock machinery and psychiatric diseases. CLOCK-3111 T/C (rs1801260) is one of the well-described SNPs in the human CLOCK gene (Schuch et al., 2018). It was first identified as an SNP associated with diurnal preference and was further found to be associated with sleep disorders (Katzenberg et al., 1998; Serretti et al., 2005; Benedetti et al., 2007). However, some reports revealed no association between T3111C SNP and sleep phenotypes (Robilliard et al., 2002; Chang et al., 2011). To our knowledge, this mutation has not been studied in vivo using humanized mice, but it has been tested in vitro using a Clock-/- mouse embryonic fibroblast (Ozburn et al., 2016).
Chronotypes, or diurnal preference, are suggested as a polygenic basis of phenotypes. The first genome-wide association study (GWAS) on chronotypes was reported in 2007. The data obtained from 749 subjects were analyzed, and several genetic loci with suggestive linkages to habitual bedtime were identified, including CSNK2A2, PROK2, and CLOCK (Gottlieb et al., 2007). Kalmbach et al. reviewed the genetic basis of human chronotypes based on 3 large-scale GWASs; one of them comprised 89,283 samples of 23andMe and the other two included 100,420 and 128,266 samples respectively from the UK Biobank. They pointed out that RGS16, PER2, FBXL3, and AK5 were commonly recognized as the significant loci for chronotype through these 3 GWASs (Hu et al., 2016; Jones et al., 2016; Lane et al., 2016; Kalmbach et al., 2017). In 2019, using the accelerometer data from 85,670 UK Biobank participants, Jones et al. (2019) performed a GWAS of 8 derived sleep traits representing sleep quality, quantity, and timing and identified 47 genetic associations, and RGS16 was also included. Marukonen et al. (2020) investigated 8433 subjects from a Finnish general population. They discovered a novel genetic association between NR1D2 (RevErbβ) and chronotypes, but their GWAS analysis failed to show significant genome-wide associations. Although the studies above all illustrated a clear relationship between human genes and chronotypes, future researchers should keep in mind that significant loci varied in studies. As SNPs are also often correlated with nearby SNPs, discussing causality would be hard. Nevertheless, testing such candidate human genes and/or SNPs in mice in vivo would still provide valuable hints for understanding complicated phenotypes like the human chronotypes.
Future Perspectives
Virus-based CRISPR
Different types of viral vectors have their own advantages and disadvantages (Navabpour et al., 2020; Chuang et al., 2021). The adeno-associated virus (AAV)-based Triple CRISPR, which does not rely on the employment of ES cells or fertilized eggs, is under extensive investigation. Kim et al. developed a multiplexed CRISPR-Cas9 system in a single AAV to simultaneously knock out 3 clock genes, namely the CSAC (CRISPR/Cas9-based single AAV system targeting the circadian clock) (Kim et al., 2021). They constructed multiplexed U6 promoter-driven sgRNA expression cassettes for AAV packaging in which a maximum of 3 sgRNAs can be harbored. The researchers also developed CSAC-Pers targeting Per1 and Per2 simultaneously, CSAC-Crys targeting Cry1 and Cry2 at the same time, and CSAC-Bmal1 targeting Bmal1. Furthermore, they unveiled that an infection of the CSACs to the SCN reduced or abolished biological rhythms at both tissue and organismal levels (Kim et al., 2021). Since researchers can employ adult mice in these experiments, this virus-based approach is able to further shorten the duration of mutant animal experiments.
Spatio-temporal Analysis
Although next-generation genetics we mentioned above are powerful tools for generating systemic KOs and KIs, they still lack spatial and temporal specificities. Virus vectors enable us to introduce transgenes to various cell types efficiently, including neurons. Tso et al. injected AAV, 8-sgBmal, into the Aldh1 L1-Cre mice’s bilateral SCN. They found that the Aldh1 L1-Bmal1 KO mice, lacking the SCN astrocytes clock, showed longer period of behavioral rhythms and PER2::Luc activity rhythms from cultured SCN slices (Tso et al., 2017). In combination with appropriate surgical techniques (intracerebroventricular injections, the direct injection to the SCN, etc.), conditional gene modification techniques (e.g., Cre-loxP system), and high-throughput gene modification tools (e.g., Triple CRISPR), the virus-based transfection system will allow us to analyze cell-type-specific genetically modified animals in a very short duration (Figure 2c).
To evaluate cell-type specificity or time dependency of used conditional gene modification, whole organ–level expression analysis is desirable. Previously, it was laborious and unrealistic for researchers to make sections through the body and check the expression patterns of target genes at a whole organ level. Nowadays, however, scientists can use tissue-clearing technology to visualize and capture target gene expression seamlessly with light-sheet microscopy. Nonetheless, tissue-clearing technology itself is a powerful tool for circuit-level and organ-level understanding (Pan et al., 2016; Gradinaru et al., 2018; Ueda et al., 2020a, 2020b; Molbay et al., 2021).
Organism-level Systems Biology From Mice to Humans
Systems biology is a subfield of biology that understands life as integrated systems (Kitano, 2002). Life systems are composed of multi-layers such as genes, mRNA, protein, cells, tissue, and organs. From the beginning of the 21st century, systems biological approaches have succeeded at molecular and cellular levels. Organism-level systems biology might be achieved by next-generation genetics. At the same time, cross-sectional approaches between layers have been highlighted. Through combining gene and protein structure-based analyses with cellular-level functional assays, Xu et al. (2015) demonstrated that CRY1 competes for binding with coactivators to the intrinsically unstructured C-terminal transactivation domain of BMAL1 to establish a functional switch between activation and repression of CLOCK–BMAL1. Through a combination of structure, cellular, and mouse studies, our group succeeded in depicting a general picture of the mammalian circadian period control by Cry1 (Ode et al., 2017). Recently, the influence of phosphorylation on molecular timing was highlighted (Forger et al., 2007; Larrondo et al., 2015; Ode et al., 2017; Shinohara et al., 2017; Narasimamurthy et al., 2018; Ode and Ueda, 2018; Masuda et al., 2020; Philpott et al., 2020; Narasimamurthy and Virshup, 2021). Under such circumstances, these types of KI-KO mouse studies on various clock genes could assist in portraying a picture of a multidimensional real molecular clock.
Due to experimental and ethical limitations, researchers cannot perform the same experimental approaches on humans as they have implemented on mice. Humans have relatively long-life spans, diverse genetic backgrounds, and different environmental exposure histories, preventing them from becoming ideal experimental subjects. Consequently, systems biology in humans will be based on comparative knowledge from other species, especially mice. Plausible approaches could include performing a large-scale screening in the human population to elucidate correlations between diseases and genomic information, as discussed in the GWAS section. Furthermore, experimental mice studies are imperative for testing causalities. These tests will be completed quantitatively, in parallel, and done in a timely fashion. High-throughput analyses in vivo could undoubtedly contribute to such occasions.
Conclusion
In this article, we have introduced the history of animal-based high-throughput analyses in chronological order. Although ENU mutagenesis screenings showed their merits in their relatively high throughput, the identification step of these methods required a long time to detect the causal genes. Regarding this gene identification process, reverse genetics took less time, but it faced challenges in genome modification and crossing. Recently, next-generation genetics, which enabled researchers to evade the time-consuming crossing issue, emerged and shed light on high-throughput analyses. This method relied on the newly developed technologies, such as the Triple CRISPR. Some possible targets of this new method include humanized animals carrying human genes. Overall, studies from the past decades have offered precious experience and tools for developing ideal animal models. These efforts will ultimately contribute to unveiling the truth underlying human circadian rhythms.
Footnotes
Acknowledgements
We thank all the lab members at the University of Tokyo and RIKEN BDR. We apologize to researchers whose work is not cited due to space limitations. This work was supported by grants from the Brain/MINDS JP21dm0207049, Science and Technology Directorate Platform Program for Advanced Biological Medicine JP21am0401011, AMED CREST JP21gm0610006 (AMED/MEXT) (H.R.U.), Grant-in-aid for Scientific Research (S) JP25221004 (JSPS KAKENHI) (H.R.U.) and Scientific Research (C) JP20K07282 (JSPS KAKENHI) (Y.M.), HFSP Research Grant Program RGP0019/2018 (HFSP) (H.R.U.), ERATO JPMJER2001 (JST) (H.R.U.), MEXT Quantum Leap Flagship Program (MEXT QLEAP) Grant Number JPMXS0120330644, and an intramural Grant-in-aid from the RIKEN BDR (H.R.U.). H.R.U. is a co-inventor on a patent covering Triple CRISPR (PCT/JP2015/086259 [pending], patent applicant is RIKEN, other co-inventors are K. Sumiyama, G.A. Sunagawa, M. Ukai-Tadenuma, D. Perrin) and patent applications covering the CUBIC reagents (PCT/JP2014/070618 [pending], patent applicant is RIKEN, other co-inventors are E. A. Susaki and K. Tainaka; PCT/JP2017/016410 [pending], patent applicant is RIKEN, other co-inventors are K. Tainaka and T. Murakami) and a co-founder of CUBICStars Inc.
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
