Abstract
The circadian clock is the broadly conserved, protein-based, timekeeping mechanism that synchronizes biology to the Earth’s 24-h light-dark cycle. Studies of the mechanisms of circadian timekeeping have placed great focus on the role that individual protein-protein interactions play in the creation of the timekeeping loop. However, research has shown that clock proteins most commonly act as part of large macromolecular protein complexes to facilitate circadian control over physiology. The formation of these complexes has led to the large-scale study of the proteins that comprise these complexes, termed here “circadian interactomics.” Circadian interactomic studies of the macromolecular protein complexes that comprise the circadian clock have uncovered many basic principles of circadian timekeeping as well as mechanisms of circadian control over cellular physiology. In this review, we examine the wealth of knowledge accumulated using circadian interactomics approaches to investigate the macromolecular complexes of the core circadian clock, including insights into the core mechanisms that impart circadian timing and the clock’s regulation of many physiological processes. We examine data acquired from the investigation of the macromolecular complexes centered on both the activating and repressing arm of the circadian clock and from many circadian model organisms.
With their description of proteins in the 1830s, Gerardu Mulder and Jons Berzelius identified a basic biochemical building block that proved essential to the fabric of life (Hartley, 1951). Since this initial discovery, researchers have gone to great lengths to elucidate the functionality of individual proteins in a biomolecular process. However, this research has ubiquitously shown that proteins do not act alone. Instead, they often perform as one of many components in a macromolecular protein complex to fulfill the needs of a cell (Frieden, 1971; Chautard et al., 2009; Miryala et al., 2018). It is essential to learn how all the components of these protein complexes work together to appreciate how a single protein contributes to cellular function. Thus, the study of a protein’s interactome, termed protein-protein “interactomics,” has become a common research focus to study interactivity in macromolecular protein complexes and how this interactivity produces functional changes within living organisms.
An exemplar of a vital biological process that is enabled by macromolecular protein complexes are circadian clocks. Circadian clocks are the molecular machinery that time 24-h biological rhythms inherent across nearly all branches of life, allowing an organism to anticipate daily changes with the Earth’s light-dark cycle (Dunlap, 1999; Panda et al., 2002; Montenegro-Montero and Larrondo, 2012; Hurley et al., 2016). These intrinsic clocks allow organisms to anticipate environmental changes by organizing physiology and behavior such that appropriate activities occur at biologically advantageous times. For example, longevity, predator avoidance, and reproductive fitness are increased in strains that maintain a clock versus strains with mutations that cause an organisms’ clock to malfunction (DeCoursey et al., 1997; Klarsfeld and Rouyer, 1998; Ouyang et al., 1998). Conversely, because the circadian clock is so tightly intertwined with many organismal systems, a disrupted circadian clock can be detrimental to an organism and chronic circadian disruption has been implicated at many levels of human disease, with higher rates of mortality, cancer, diabetes, stroke, neurodegeneration, and heart disease all associated with circadian disruption (Fu et al., 2002; Davidson et al., 2006; Sahar and Sassone-Corsi, 2007; Straif et al., 2007; Brown et al., 2009; Scheer et al., 2009; Pan et al., 2011; Cutolo, 2012; Greene, 2012; Narasimamurthy et al., 2012; Evans and Davidson, 2013; Mattis and Sehgal, 2016).
The circadian clock comprises an evolutionarily conserved oscillator commonly involving a transcriptional-translational negative-feedback loop (TTFL) that cycles with a period roughly equal to the 24-h day (Bell-Pedersen et al., 2005; Y. Liu and Bell-Pedersen, 2006; Diernfellner and Schafmeier, 2011). In animals and fungi, this protein-based clock is a two-component system consisting of a positive arm and a negative arm. The positive arm, a heterodimeric transcription factor complex, transcriptionally activates the production of the negative arm. Once translated, the components of the negative arm form large, temporally regulated, macromolecular complexes, including the proteins of the positive arm, leading to the repression of the positive arm and closing the loop. The negative arm then goes through a precisely timed half-life, after which it is degraded, allowing the positive arm to reactivate and restart the cycle (Dunlap, 1999; Bell-Pedersen et al., 2005; Albrecht, 2012; Beckwith and Yanovsky, 2014).
Circadian regulation of cellular physiology, termed the clock’s “output,” is believed to stem from positive-arm transcriptional activation beyond the promoter of the negative arm, leading to an oscillation in a group of genes beyond those involved in the core clock call “clock-controlled genes” or “ccgs” (Aronson et al., 1994; Crosthwaite et al., 1997; Hurley et al., 2015). Up to 80% of mRNAs are reported to oscillate in simian models, demonstrating that the circadian cycle is one of the most extensive transcriptional regulatory networks in biology (Menet et al., 2012; Beckwith and Yanovsky, 2014; Menet et al., 2014; Hurley et al., 2015; Mure et al., 2018). While bacterial clocks, and to a certain extent the plant clock, differ somewhat from this basic mechanism, many of the same principles that are used to keep time in animals and fungi apply to clocks in bacteria and plants (Bell-Pedersen et al., 2005; Brunner and Schafmeier, 2006; McClung, 2006; Rosbash, 2009; Mori et al., 2015; H. Huang and Nusinow, 2016).
Research focused on the investigation of individual interactions between the negative arm and the positive arm of the clock has defined many of the mechanisms that time circadian rhythms (e.g., Crosthwaite et al., 1997; Grima et al., 2002; Dunlap and Loros, 2004; Triqueneaux et al., 2004; Akashi and Takumi, 2005; Boothroyd et al., 2007; Yakir et al., 2007; Dong et al., 2008; Kadener et al., 2008; Smith et al., 2010; Hardin, 2011; Gendron et al., 2012; Beckwith and Yanovsky, 2014; Zhang et al., 2014; Gustafson and Partch, 2015; Sancar et al., 2015; Mure et al., 2018). However, the mechanisms that initiate or terminate the circadian cycle have not been completely elucidated using a one-by-one protein analysis approach. Moreover, although transcriptional activation via the positive arm accounts for some of the circadian coordination within cells, a plethora of proteomics data gathered over circadian time has demonstrated that there is post-transcriptional regulation of circadian output and the source of this regulation is uncertain (Reddy et al., 2006; Kojima et al., 2011; Mauvoisin et al., 2014; Robles et al., 2014; J. Wang et al., 2017; Hurley et al., 2018; Collins et al., 2021). In addition, recent investigations have demonstrated that the core proteins of the circadian clock are either intrinsically disordered proteins or proteins with intrinsically disordered regions, a class of proteins that exists in a heterogeneous ensemble of conformations and has a large, transient, interactome (Partch et al., 2005; Tompa, 2012; Hurley et al., 2013; Oldfield and Dunker, 2014; Gustafson and Partch, 2015; Pelham et al., 2018, 2020). All of the above have led many circadian researchers to shift their studies to the macromolecular protein complexes of the core clock proteins. In this review, we evaluate the state of circadian interactomics by focusing on the research that has been carried out in several of the major circadian model organisms. We divide our discussion into two categories, the macromolecular complexes that coalesce around the negative arm and the macromolecular complexes that coalesce around the positive arm. Although this is by no means an exhaustive review, we endeavor to highlight major contributions from the study of circadian macromolecular complexes as well as the novel questions that circadian interactomic studies have generated.
Macromolecular Complexes that Coalesce Around the Negative Arm of the Circadian Clock
It is well-established that the negative arm of the circadian TTFL acts as the repressor of the core clock mechanism by inhibiting the activity of the positive arm, ostensibly via temporal phosphorylation events, although this mechanism has yet to be cemented (Aronson et al., 1994; Hardin, 2005; Albrecht, 2012; Gustafson and Partch, 2015; B. Wang et al., 2019). The negative arm also serves to set the period of the clock through temporal phosphorylation events that affect either the repressive activity or the degradation of negative-arm proteins (Hardin, 2005; He et al., 2006; Mehra et al., 2010; Larrondo et al., 2015; Takahashi, 2015; Hurley et al., 2016; Ode and Ueda, 2018). To perform both of these functions, the core negative-arm proteins recruit many kinases and phosphatases that facilitate the required phosphorylation (Ko et al., 2002; Eide et al., 2005; He et al., 2006; Baker et al., 2009; Lamia et al., 2009; Brody et al., 2010; Hayasaka et al., 2017; Ode and Ueda, 2018). However, recently published studies of the negative-arm interactome have suggested potential mechanisms of repression beyond phosphorylation. Moreover, these interactome studies have highlighted novel roles for the negative arm in the regulation of circadian output. We address each of these findings below.
Core Clock Cyanobacterial Macromolecular Complexes Interact With Proteins Required for Photosynthesis
A unicellular cyanobacterium, Synechococcus, has been used as a model for prokaryotic cellular clocks since the discovery of a circadian oscillation in its nitrogen fixation and amino acid uptake (Cohen and Golden, 2015; Swan et al., 2018). Unlike the TTFL in eukaryotic cells, the circadian timekeeping mechanism in cyanobacteria is a post-translational oscillator comprised of three core proteins: KaiA, KaiB, and KaiC (Figure 1). In brief, this post-transcriptional oscillator initiates during the subjective morning, when KaiA binds to KaiC to promote the latter’s autokinase activity (Y. I. Kim et al., 2008). As the day progresses, the protein SasA binds to KiaC to activate the protein RpaA and regulate the clock’s output. In the subjective evening, KaiB switches folds upon binding cooperatively to KaiC. Once bound to KaiC, separate KaiB proteins on KaiC can bind to the autoinhibited state of KaiA and CikA. KaiB-CikA interactions allow CikA to dephosphorylate and thereby inactivate RpaA, closing the cycle (Y. I. Kim et al., 2008; Chang et al., 2012; Swan et al., 2018). It is this cycle of KaiC phosphorylation and dephosphorylation that sets the length of the day in the cyanobacterial clock and times rhythms in protein oscillations (Guerreiro et al., 2014; Cohen and Golden, 2015).
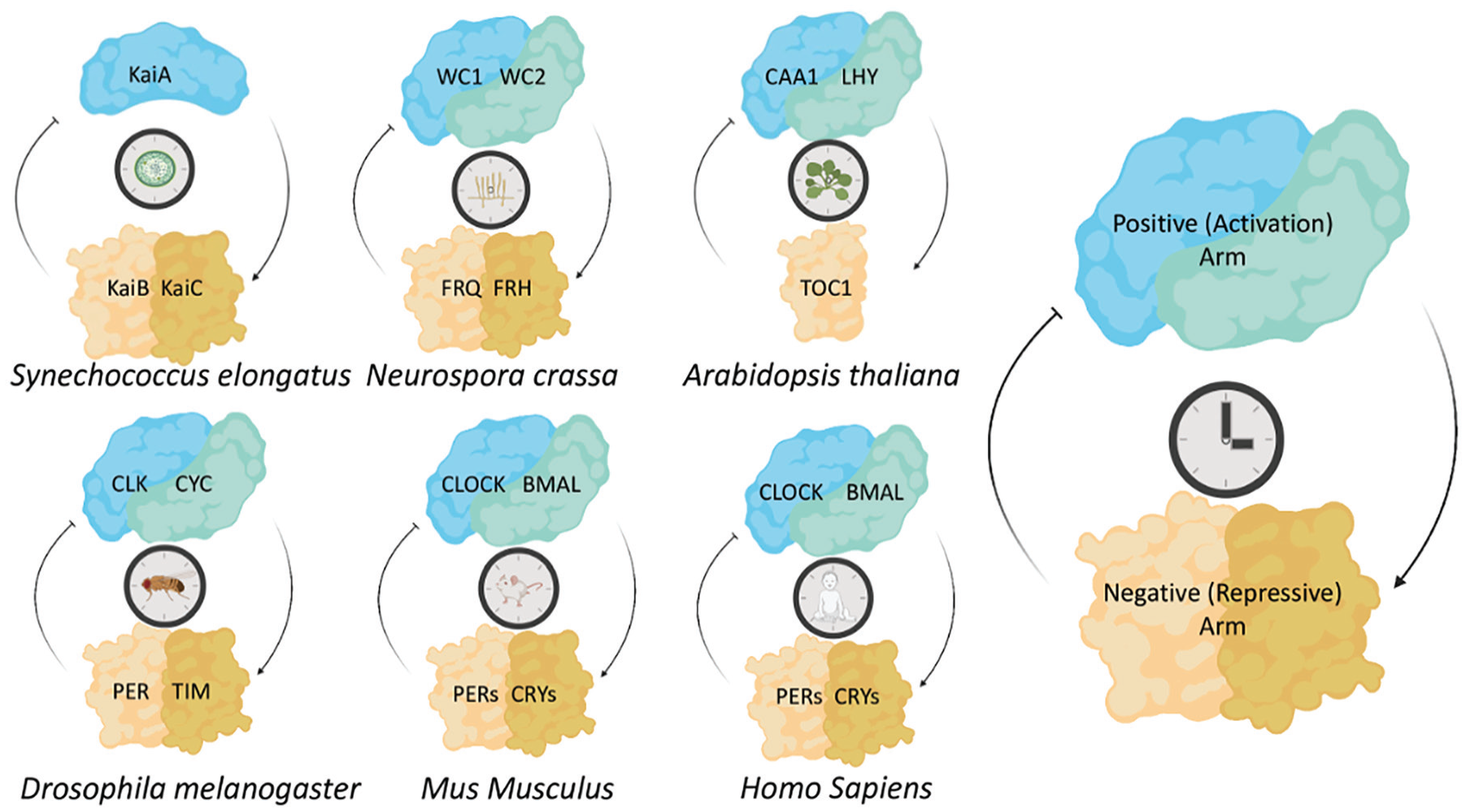
The protein-based timing mechanisms of the circadian clock are evolutionarily conserved. A simplified diagrammatic representation of the core circadian clock proteins across many circadian model organisms. The central mechanism consists of positive elements that activate the negative elements, which in turn act to repress the activation by the positive elements. Core clock components from each model system are represented in their associated feedback loops. Abbreviations: WC1 = White Collar-1; WC2 = White Collar-2; FRQ = FREQUENCY; FRH = FRQ-interacting RNA Helicase; LHY = LATE ELONGATED HYPOCOTYL; TOC1 = TIMING OF CAB EXPRESSION; CLK = CLOCK; CYC = CYCLE; CLOCK = Circadian locomotor output cycles kaput; BMAL = Brain and Muscle ARNT-Like ; PER = PERIOD; TIM = TIMELESS; CRY = CRYPTOCHROME; CCA1 = CIRCADIAN CLOCK-ASSOCIATED-1.
Beyond the interaction with KaiA and other clock output proteins like SasA, CikA, and RpaA, circadian interactomics studies have shown KaiB and KaiC complex with CO2 processing and concentrating proteins as well as light-harvesting protein complexes (U. K. Aryal et al., 2018; Swan et al., 2018). In a global study of protein complexes in the cyanobacterium Cyanothece, both KaiB and KaiC were consistently found in complex with many subunits of phycocyanin and allophycocyanin, both of which are important complexes for light harvesting and photosynthesis in cyanobacteria (U. K. Aryal et al., 2018). KaiB and KaiC were also found in complex with many subunits of the CO2 concentrating protein (Ccm), which prepares CO2 for use by the carbon dioxide–fixing enzyme Rubisco complexes, signifying that the post-translational arm of the bacterial clock may play a role in CO2 fixation and control output beyond its interaction with RpaA (U. K. Aryal et al., 2018; Figure 2). Independently of KiaC, KaiB also forms a complex with the Rubisco large (RbcL) and small (RbcS) subunits, suggesting that KaiB is involved in preparation of CO2 outside of its core clock function. In total, circadian interactomics approaches demonstrated that core proteins of the cyanobacterial clock are connected with many physiological processes in the cell, which may regulate physiology through mechanisms beyond SasA and RpaA or provide a source of environmental input into the clock.
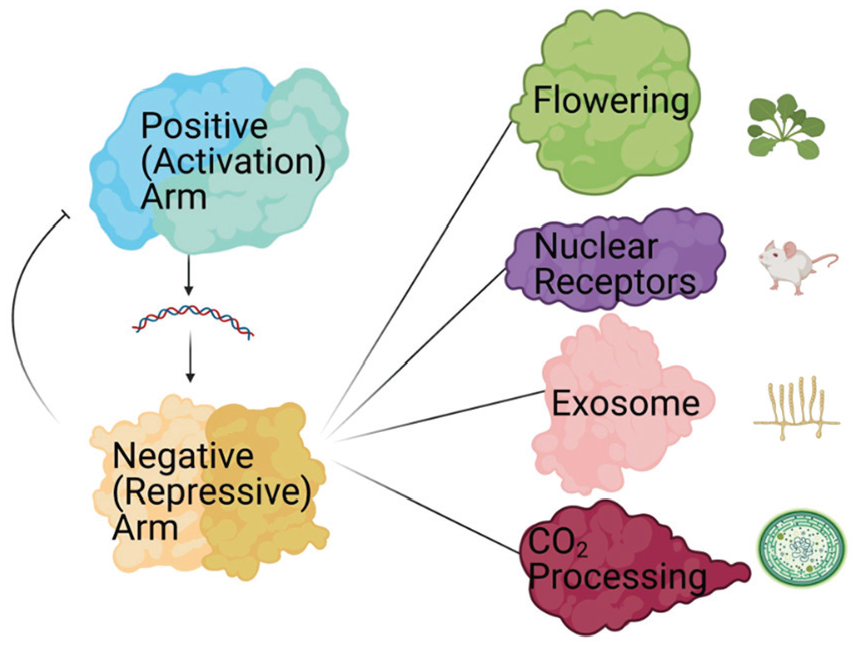
Circadian negative-arm complexes aid in the regulation of many cellular processes. A pictorial illustration of cellular processes that the negative arm has been linked to based on circadian interactomics data. The organisms in which these discoveries were made are represented next to each cellular process.
Constitutive Interactions in the Clocks of Fungi Regulate Negative-arm and Other Cellular mRNA Half-lives
A model for non-hierarchical eukaryotic clocks, Neurospora crassa first gained notoriety as a genetic model for the one-gene/one-enzyme theory proposed by Beadle and Tatum in 1941. This primed Neurospora’s readiness as a model for circadian research when Pittendrigh described the visual phenotypic clock output of conidial formation of Neurospora growing on race tubes (Beadle and Tatum, 1941; Pittendrigh et al., 1959). Since then, Neurospora has contributed greatly to the elucidation of the genetic and biochemical mechanisms of eukaryotic clocks, as well as to the fields of recombination, gene action, and cell growth in eukaryotic cells (Loros, 1998; Fuller et al., 2014; Roche et al., 2014; Jankowski et al., 2020). In Neurospora, the core negative-arm protein is FREQUENCY (FRQ; Dunlap and Loros, 2004; Cha et al., 2015; Pelham et al., 2018). FRQ is produced in the early subjective morning, immediately dimerizing with itself as well as binding to its chaperon protein FRQ-interacting RNA Helicase (FRH; Guo et al., 2010; Shi et al., 2010; Hurley et al., 2013). Together, these proteins form the FRQ-FRH complex (FFC), which is transported from the cytoplasm to the nucleus in a time-specific manner (Luo et al., 1998). In the nucleus, the FFC and recruited kinases and phosphatases, for example, Casein Kinase-1 (CK-1), interact with the Neurospora positive-arm complex, the White Collar Complex (WCC), inactivating the transcriptional activity of the WCC and inhibiting the production of frq, closing the loop (Figure 1; He et al., 2006). In the subjective evening, through a series of phosphorylation events, FRQ is inactivated and then targeted for degradation, thus setting the period and restarting the clock (Aronson et al., 1994; Dunlap, 1999; He and Liu, 2005; Larrondo et al., 2015).
Phosphorylation of FRQ also correlates with oscillations in the components of FRQ macromolecular complexes (Baker et al., 2009). Moreover, while some core clock interactors, for example, FRH, were found consistently in FRQ macromolecular complexes throughout the circadian day, others, for example, the WCC and CK-1, were found at greater levels in the early subjective morning, coinciding with lower amounts, and lower phosphorylation levels, of FRQ (Baker et al., 2009). These interactomics analyses of FRQ identified a large number of proteins in complex with the FFC that are not related to the core clock, suggesting that the FFC may play a physiological role beyond the repression of the WCC (Baker et al., 2009). For example, proteins that regulate the cell cycle were found among FRQ interactors, relevant as the cell cycle has been shown to be under clock control (Hong et al., 2014).
Beyond its interaction with FRQ, FRH forms a complex with RRP44 and RRP6, two key components of the exosome that enable the degradation of mis-polyadenylated mRNA (Figure 2; Guo et al., 2009). The interaction between FRQ, FRH, and the exosome facilitates the rhythmic degradation of frq mRNA and other mRNAs (Guo et al., 2009). In concordance with this, the disordered region of FRH that is key to the interaction between FRQ and FRH contacts the helicase active site on FRH, suggesting the FRQ-FRH interaction may regulate the helicase activity of FRH, representing a mechanism of cell-wide circadian post-transcriptional regulation (Jia et al., 2012; Hurley et al., 2013; Conrad et al., 2016; Morales et al., 2018). In total, interactomics research in Neurospora has highlighted avenues of potential core clock regulation and control of physiology (Guo et al., 2009; Hurley et al., 2013; Conrad et al., 2016).
Clock Interactors Regulate Phosphorylation and Localization of the Negative-arm proteins in Drosophila
In Drosophila melanogaster, a foundational model system of the molecular circadian clock since the discovery of a circadian rhythm in eclosion, the core TTFL, is formed by the transcriptional activation of period (per) and timeless (tim) by the heterodimer formed by CLOCK (CLK) and CYCLE (CYC; Pittendrigh, 1954; Blau, 2001). Newly produced PER and TIM are phosphorylated by the kinase DOUBLETIME (DBT), heterodimerize, and upon interaction with CLK-CYC repress their own transcription. Phosphorylation promotes PER and TIM degradation, which causes dephosphorylation of CLK and allows the cycle to reinitiate (Figure 1; Hardin, 2011; Patke et al., 2020). An intermediate between bacterial and fungal clocks and clocks in mammals, the Drosophila circadian system is partly hierarchical and partly autonomous, with roughly 150 pace-making neurons in the brain coordinating among themselves to set circadian timing while CRY in peripheral cells can coordinate the light response in isolation (Allada and Chung, 2010).
In Drosophila, interactomics studies have identified novel core clock proteins. Through an interactomics approach, DBT was found to interact with BRIDE OF DOUBLETIME (BDBT), inhibiting DBT autophosphorylation and allowing DBT to properly phosphorylate, localize, and target PER for degradation. Without BDBT, DBT is hyperphosphorylated, leading to reduced DBT levels, hypophosphorylated PER, and a change in clock period and PER localization (Fan et al., 2013). Another interactomics study in Drosophila demonstrated a direct interaction between Casein Kinase 1 alpha (CK1a) and DBT. CK1a collaborates with DBT to phosphorylate, temporally regulate the degradation, and control the subcellular localization of PER (Lam et al., 2018). Combined, the interaction of CK1a and DBT regulate PERs return to the nucleus and, by extension, PER repression of positive-arm proteins (Lam et al., 2018). The necessity of the interaction between DBT, BDBT, and CK1a for proper PER homeostasis demonstrates the importance of diverse macromolecular complexes for core circadian functions.
Mammalian Clock Negative-arm Macromolecular Complexes Regulate Promoter Activation and Transcription
Similar to insects, Mus Musculus and Homo sapiens clocks are complex, hierarchical, systems (Bass and Takahashi, 2010). The master clock, located in the suprachiasmatic nucleus (SCN), communicates environmental cues and temporal regulation to peripheral clocks (Ueyama et al., 1999; Balsalobre et al., 2000; Lee et al., 2001; Reppert and Weaver, 2001). These peripheral tissue clocks are essential to organismal health, regulating many complex physiological systems and chronic disruptions to circadian rhythms have been connected to a number of pathologies (Stevens et al., 2007; Bass and Takahashi, 2010; Evans and Davidson, 2013). The mammalian negative arm is composed of the Period (per) and Cryptochrome (cry) genes. PERs and CRYs are transcribed via positive-arm activation from the bHLH-PAS domain containing proteins Circadian locomotor output cycles kaput (CLOCK) and Brain and Muscle ARNT-Like 1 (BMAL1). The PERs and CRYs are translated in the cytoplasm, where they recruit various kinases, after which they are imported back into the nucleus and repress the transcriptional activity of BMAL1/CLOCK (Figure 1; Takahashi, 2015). An auxiliary negative-feedback loop is formed by BMAL1/CLOCK transcriptional activation of the rev-erb genes, which when translated repress the expression of BMAL1 (Takahashi, 2015).
Like FRQ in fungi, human PERs have been shown to have an extensive interactome, with interactor functions ranging from the immune response to metabolic regulation. In a study using human microvascular endothelial cells (HMEC1s), environmentally dependent complexes centered on human PER2 (hPER2) contained as many as 1284 proteins. Proteins identified in this interactomics study were involved in cholesterol biosynthesis, phagosome maturation, TriCarboxylic Acid (TCA)-cycle, RNA signaling, and tRNA charging (Oyama et al., 2019). As each interactor is likely a component of a broadly efficacious pathway (e.g., the TCA-cycle is required for countless cellular processes), these interactions portend a cascading circadian regulation beyond the immediate interactors that could play a role in the regulation of cellular physiology.
Circadian interactomics studies have shown that mouse PER2 (mPER2) is also a hub protein with many interactors. One global interactomics study showed that mPER2 is part of macromolecular complexes that regulate transcription. mPER2 was found in complexes with both polypyrimidine tract binding protein-associated splicing factor (PSF) and SIN3A. PSF and SIN3A recruit histone deacetylases to the per promotor regions to repress transcription when mPER2 interacts with the BMAL1/CLOCK complex (Duong et al., 2011). In addition, mPER2 interacts with Suv39h1, Suv39h2, HP1γ, and Kap1, all of which are histone methyltransferase subunits involved in the methylation of H3K9 (Duong and Weitz, 2014). When mPER2 interacts with the BMAL1/CLOCK complex, these methyltransferases facilitate di- and tri-methylation at H3K9 and deacetylation that represses transcriptional activity at the mper2 promoter (Duong and Weitz, 2014). mPER2 also interacts with RNA helicases DDX5, DHX9, and SETX, as well as RNA Polymerase II, suggesting that mPER2 also plays a role in the regulation of transcription (Padmanabhan et al., 2012). These murine interactomics studies have demonstrated the mechanistic role that the negative arm plays in closing the feedback loop of the circadian clock.
Interactomics studies in mice have also established novel functions for the negative arm in the regulation of clock output. In a targeted analysis of mPER2 macromolecular complexes, several nuclear receptors were identified, including PPAR alpha and REV-ERB alpha. Nuclear receptor-mPER2 interaction allows mPER2 to act as a direct coregulator of nuclear receptor-mediated transcriptional regulation via its binding to the promoter of Bmal1 and other nuclear-receptor target genes (Figure 2; Schmutz et al., 2010). Moreover, mPER complexes outside the nucleus that do not involve the positive arm interact with a wide range of proteins, including a guanine nucleotide exchange factor for Rab GTPases involved in receptor-mediated endocytosis, GAPVD1. The GAPVD1-mPER interaction proved to be essential for proper circadian timing (R. P. Aryal et al., 2017).
In total, mammalian clock interactome analyses demonstrate negative-arm clock proteins have a direct role in the regulation of promoter activation and transcription of the positive arm and other clock-controlled genes. Moreover, the extent of the proteins involved in negative-arm macromolecular complexes suggest there are likely many more as-yet-undiscovered functions of negative-arm proteins that lead to proper timing of circadian rhythms.
Macromolecular Complexes in the Plant Clock Repressive Arm Regulate Environmental Tuning and Feedback
The earliest recordings of a biological diurnal pattern come from Androsthenes, when he described the leaf movements of Tamarindus indicus during his time with Alexander the Great in the fourth century B.C. (McClung, 2006). Centuries later, the first demonstration of an anticipatory clock was completed in Mimosa pudica (de Mairan, 1729). While discoveries in these plants were the cornerstone for circadian research, it is Arabidopsis thaliana, the mouse-eared cress, that has become the key architype for molecular research into plant circadian rhythms as its molecular tools allowed for genetic manipulation (Meyerowitz, 2001; McClung, 2006; Leonelli, 2007). More complex than the TTFL of the fungal/animal lineage, the plant clock is timed using a series of inter-connected feedback loops, the morning, evening, and central loops. Morning-expressed proteins LATE ELONGATED HYPOCOTYL (LHY) and CIRCADIAN CLOCK ASSOCIATED1 (CCA1) are transcription factors that repress the expression of the evening-expressed pseudo response regulator TIMING OF CAB EXPRESSION1 (TOC1) gene. TOC1 represses most other genes in the clock, including CCA1, LHY, GIGANTEA (GI), and the evening complex genes EARLY FLOWERING 3 and 4 (ELF3 and 4), and LUX ARRHYTHMO/PHYTOCLOCK1 (LUX). CCA1 and LHY also regulate PRR gene expression (PRR9, PRR7, and PRR5), which in turn represses the expression of CCA1 and LHY. The evening complex relieves this repression by repressing the transcription of PRR9, PRR7, and PRR5 (Leonelli, 2007; Oakenfull and Davis, 2017). GI forms an additional negative-feedback loop with TOC1 (Figure 1). Due to the increased complexity of the plant clock, instead of “negative” and “positive” arm terminology, we will use “repressive” and “activating” arms, respectively, in our review of plant interactomes.
Studies of the macromolecular complexes centered on the repressive proteins in the central loop of the circadian clock in Arabidopsis established these complexes include proteins that regulate transcription, post-translational signaling, and inter-cellular communication. In a study that tracked ~8000 proteins (14 clock specific) in Arabidopsis for novel protein-protein interactions, central loop repressive arm proteins TOC1 and the TOC1 co-repressor of CCA1, CCA1 HIKING EXPEDITION (CHE), were found to interact with 10 and 26 proteins, respectively. These interactions included proteins involved in transcription (AP2-like ethylene-responsive transcription factor TOE2), RNA-regulation (RRM domain-containing protein), and protein modifications (E3 ubiquitin-protein ligase BOI; Chow and Kay, 2013) Beyond the central loop, the involvement of proteins from the repressive evening loop in extra-circadian complexes has been demonstrated. The interaction of the evening-loop protein ELF3 has been indicated in regulating plant growth, response to temperature changes, and flowering via ELF3s interaction with PHYTOCHROME INTERACTING FACTOR 4 (PIF4) and CONSTITUTIVE PHOTOMORPHOGENIC 1 (COP1; Figure 2; H. Huang and Nusinow, 2016). These core and evening-loop interactions reveal a pathway by which the clock can regulate output post-transcriptionally as well as provide a mechanism for environmental feedback into the clock.
Macromolecular Complexes that Coalesce Around the Positive Arm of the Circadian Clock
The positive arm of the circadian clock is a transcription factor complex that acts as the activator of the core clock mechanism by transcriptionally activating negative-arm gene promoters (Dunlap, 1999; Hardin, 2005; Kadener et al., 2008; Albrecht, 2012; Baker et al., 2012; Gustafson and Partch, 2015; Hurley et al., 2015). The positive arm is also thought to control the clock’s output through the transcriptional activation of genes beyond those directly involved in the core clock mechanism, dubbed clock-controlled genes (Yakir et al., 2007; Smith et al., 2010; Sancar et al., 2015). Inactivation of the positive arm occurs through an as yet undetermined mechanism, although it is known that repression of positive-arm activity is dependent upon the interaction with the negative arm of the clock (Yakir et al., 2007; Duong et al., 2011; Gustafson and Partch, 2015; Hurley et al., 2015; R. P. Aryal et al., 2017). However, recently published studies of positive-arm macromolecular complexes have identified interactors that may facilitate the transcriptional activation and repression of the positive-arm complex as well as allow the positive arm to control output on the post-transcriptional level. We review each of these findings and their potential impact on the model of circadian regulation below.
Circadian Interactomes in Fungi Facilitate Pioneer-like Transcription Factor Properties and Stability in the Positive Arm
In Neurospora, the positive arm is comprised of the transcriptional activators White Collar-1 (WC-1) and White Collar-2 (WC2), which interact to form the heterotetrametric WCC (Figure 1; Crosthwaite et al., 1997; Fuller et al., 2014). The WCC drives rhythmic transcription of approximately 40% of the Neurospora transcriptome, including the negative-arm protein FRQ, to initiate the circadian cycle and facilitate rhythmic output (Y. Liu and Bell-Pedersen, 2006; Baker et al., 2012; Beckwith and Yanovsky, 2014; Hurley et al., 2014; Sancar et al., 2015). The WCC is also the conduit through which environmental stimuli, for example, light and glucose levels, are integrated into the circadian circuit (Jankowski et al., 2020).
An interactomics approach in Neurospora revealed that to initiate transcription at the frq and ccg promoters, the WCC recruits chromatin remodelers to the positive-arm macromolecular complex. In a survey of WCC interactors, the WCC was found in complex with subunits of the SWItch/Sucrose NonFermentable (SWI/SNF) complex (B. Wang et al., 2014). SWI/SNF is an ATP-dependent chromatin-remodeling complex that alters chromatin structures for transcriptional activation. The SWI/SNF/WCC macromolecular complex is responsible for the transcriptional activation of frq, dictating proper circadian timing and allowing the WCC to impart its pioneer-like transcription factor function (Figure 3; Hurley et al., 2014; B. Wang et al., 2014). Another interactomics study established that the WCC is also regulated by the Not1-Ccr4 complex, a conserved multisubunit protein complex that regulates gene expression (Maillet et al., 2000; G. Huang et al., 2013). Not1-Ccr4 stabilizes the WCC, perhaps through phosphorylation events, to properly time WCC activity (G. Huang et al., 2013). These results are another example of how circadian macromolecular complexes are essential to proper circadian timing.
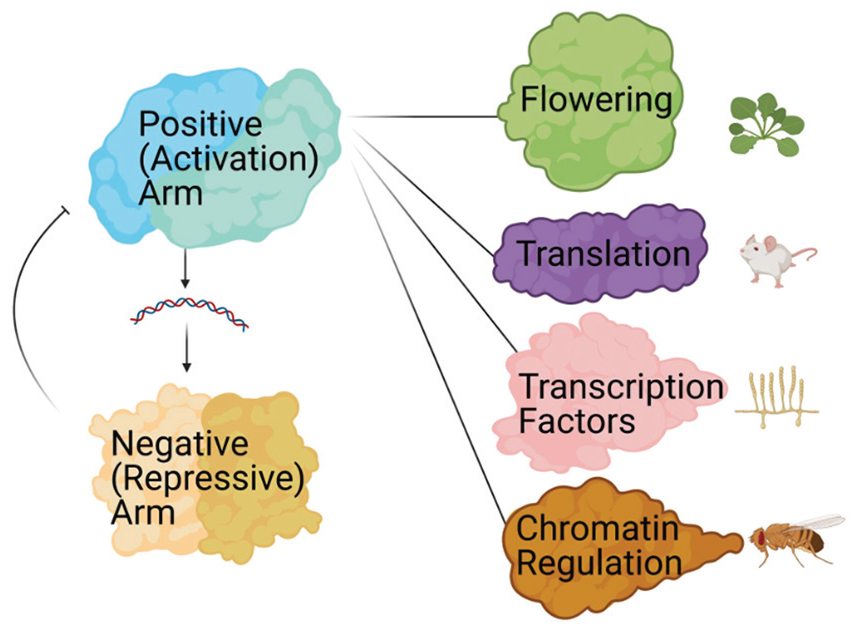
Proteins in the positive arm of the circadian clock form complexes that control circadian timing and output. A graphic depiction of the elements of cellular physiology that have been linked to the positive arm of the clock based on circadian protein interactomics studies. The organisms in which these discoveries were made are represented next to each cellular process.
Positive-Arm Macromolecular Complexes in the Drosophila Clock Impart Epigenetic Regulation
In Drosophila melanogaster, the positive arm of the TTFL comprises the transcriptional activating heterodimeric complex of CLOCK (CLK) and CYCLE (CYC; Blau, 2001). CLK/CYC activate the expression of period (per) and timeless (tim) which, when translated, heterodimerize and interact with CLK/CYC to repress per and tim transcription (Figure 1; Blau, 2001). Like positive-arm proteins in most other circadian model systems, while there is a clear understanding that Drosophila CLK and CYC play a key role in the initiation of the feedback loop, little is known about the mechanisms that allow for transcriptional activation or repression.
To elucidate the mechanics that time positive-arm activation and repression, and the protein factors that impart those mechanics, a recent interactomics study focused on identifying proteins in the macromolecular complexes centered on CLK. Over the circadian day, as many as 138 proteins were detected in complex with CLK, with functions ranging from kinases, to phosphatases, to protein degradation machinery components, to chromatin remodelers (Mahesah et al., 2020). Of particular interest, the CLK interactor chromatin remodeler NIPPED-A was found to regulate period length as well as rhythmicity (Murr et al., 2007; Helmlinger and Tora, 2017; Bu et al., 2020; Mahesah et al., 2020). The interaction and period determining phenotype of NIPPED-A, a scaffold protein in the transcriptional co-activator SAGA/Tip60 complex (Figure 2), suggests a mechanism to facilitate transcriptional activation, establishing an example of the potential importance of macromolecular complexes in the initiation of the circadian cycle.
In an independent analysis of Drosophila positive-arm macromolecular complexes, CLK was shown to interact with the chromatin-remodeling protein Brahma (Brm; Tamkun, 1992; Kwok et al., 2015). Similar to SWI/SNF in Neurospora, Brm was found to bind to the same promoting region of per as CLK, but unlike SWI/SNF, Brm acts to repress CLK transcriptional activation via the recruitment of repressive protein complexes such as Sin3-HDAC, NCOR1, and NuRD, and through the condensation of chromatin (Figure 3; Tamkun, 1992; B. Wang et al., 2014; Kwok et al., 2015). DOMINO (DOM), another chromatin remodeler, was also found to bind to CLK in this dataset (Z. Liu et al., 2019). DOM/CLK binding regulated per and tim levels in a splice-variant dependent manner. In total, these studies of circadian macromolecular complexes show the interconnection between circadian timing and chromatin states.
The Macromolecular Complexes of the Mammalian Positive Arm Imparts Circadian Regulation of Translation
The mammalian positive arm is centered on the bHLH-PAS proteins CLOCK and BMAL1, which form a complex to activate the transcription of the per and cry genes, triggering the initiation of the circadian cycle (Figure 1; Phifer-Rixey and Nachman, 2015; Takahashi, 2015). CLOCK/BMAL1 also initiates transcription of the nuclear receptors Rev-erba/b. Rev-erba/b act to repress Bmal1 expression, forming an ancillary feedback loop in the circadian circuit which increases both the robustness and environmental responsiveness of the clock, particularly in metabolic conditions (R. P. Aryal et al., 2017; Honma, 2018, 2020; Rosensweig and Green, 2020).
Like the PERs and CRYs, BMAL1 is found independently in macromolecular complexes in the cytoplasm (R. P. Aryal et al., 2017). In an unbiased investigation of the cytosolic macromolecular complexes centered on BMAL1, many pieces of translational machinery were identified. These interactors were dependent on the phosphorylation activity of the mTOR-effector kinase ribosomal S6 protein kinase 1 (S6K1; Lipton et al., 2015). In concordance with this, BMAL1 dependent circadian oscillations were detected in translation, demonstrating that the positive-arm macromolecular complexes directly affect protein translation (Figure 3; Lipton et al., 2015).
In addition to traditional mass-spec-based analyses of circadian macromolecular complexes, other approaches have been used to identify protein-protein interactions in the mammalian clock. Using a spectroscopic co-localization approach, BMAL1 was found to localize to the nucleolus where it interacts with NOP58 (Cervantes et al., 2020). NOP58 is a member of the nucleolar ribonucleoprotein complex, which aids in the processing of pre-rRNA (Boisvert et al., 2007). Through this interaction, BMAL1 regulates the levels of Snord118 in the nucleolus to facilitate the cleavage of specific pre-rRNAs (Cervantes et al., 2020). Using a yeast-2-hybrid approach to analyze direct interactions between murine clock components, over 150 direct protein-protein interactions were identified for core clock proteins. These interactions included proteins involved in chromatin modification, regulation of proliferation, DNA repair, and many other pathways (Wallach et al., 2013). One identified interaction, with the protein phosphatase 1 alpha catalytic subunit, reduced transactivation from the CLOCK/BMAL complex via the destabilization of BMAL1, adding a new layer to the model of how the core clock tunes circadian timing (Wallach et al., 2013). In total, non-mass spectrometric analyses of BMAL1 interactions demonstrated novel mechanisms of positive-arm control over the regulation of the clock.
Studies of the Macromolecular Complexes of the Plant Clock Activating Arm Suggest a Mechanism for Clock Regulation Over Flowering
The circadian timekeeping mechanism in plants has several points of transcriptional activation in its central, morning, and evening loops (H. Huang and Nusinow, 2016; Oakenfull and Davis, 2017). A key component of the central and evening loops in the plant clock is the chaperone protein GIGANTEA (GI), which affects the levels of TOC1 through the stabilization of its partner protein (W. Y. Kim et al., 2007; Singh and Mas, 2018). GI is known to be promiscuous in its binding and has been linked to flowering and carbon metabolism (Koornneef et al., 1991; Yu et al., 2008; Dalchau et al., 2011). However, a broad study of macromolecular complexes centered on GI over the circadian day found more interactors than predicted; 240 proteins were in complex with GI. These interactors were enriched for diverse ontologies (Krahmer et al., 2019). In particular, GI was shown to interact with CYCLING DOF FACTOR6 (CDF6), which regulates Arabidopsis flowering through transcriptional inhibition, demonstrating that clock proteins can act as regulators of cellular physiology beyond the canonical transcriptional activation of the circadian clock (Krahmer et al., 2019).
Conclusion
There is significant evidence to support the importance of the study of macromolecular protein complexes (via interactomics) in concordance with studying individual proteins. The circadian system is a quintessential example of this, as tracking the interactomes of the core clock proteins has demonstrated that these complexes involve many players, time the core clock, regulate output, and help the clock respond to environmental change. Proteins associated with the core clock macromolecular complexes have shown that the clock plays a role in regulating cellular processes such as transcription, translation, energy production, growth, and photoadaptation. Conversely, other interacting partners in core clock macromolecular complexes have been implicated in clock timing mechanisms, including initiation, subcellular localization, or the closing of the feedback loop. Major findings related to the functionality of circadian regulation based on these interactions include, but are not limited to, the regulation of output beyond the transcriptional activation from the positive arm and mechanisms by which the negative and positive arm both participate in the closing of the feedback loop. However, these findings highlight only a few of the hundreds of interactions that were identified in the above-described interactome analyses, which represent only a subset of the circadian interactomics studies that have been completed. The remaining unexplored components of the clock macromolecular complexes will surely provide even more biologically significant insights and identify novel questions.
Footnotes
Acknowledgements
The authors would like to thank Dr. Joanna Chiu for her advice on this manuscript. The figures were created with ![]() . This work was supported by an NIH-National Institute of Biomedical Imaging and Bioengineering Grant U01EB022546 (to J.M.H.), an NIH-National Institute of General Medical Sciences Grant R35GM128687 (to J.M.H.), an NSF CAREER Award 2045674 (to J.M.H.), Rensselaer Polytechnic Startup funds (to J.M.H.), and a gift from the Warren Alpert Foundation (to J.M.H.).
. This work was supported by an NIH-National Institute of Biomedical Imaging and Bioengineering Grant U01EB022546 (to J.M.H.), an NIH-National Institute of General Medical Sciences Grant R35GM128687 (to J.M.H.), an NSF CAREER Award 2045674 (to J.M.H.), Rensselaer Polytechnic Startup funds (to J.M.H.), and a gift from the Warren Alpert Foundation (to J.M.H.).
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
