Abstract
Daily rhythms of physiology, metabolism, and behavior are orchestrated by a central circadian clock. In mice, this clock is coordinated by the suprachiasmatic nucleus, which consists of 20,000 neurons, making it challenging to characterize individual neurons. In Drosophila, the clock is controlled by only 150 clock neurons that distribute across the fly’s brain. Here, we describe a comprehensive set of genetic drivers to facilitate individual characterization of Drosophila clock neurons. We screened GAL4 lines that were obtained from Drosophila stock centers and identified 63 lines that exhibit expression in subsets of central clock neurons. Furthermore, we generated split-GAL4 lines that exhibit specific expression in subsets of clock neurons such as the 2 DN2 neurons and the 6 LPN neurons. Together with existing driver lines, these newly identified ones are versatile tools that will facilitate a better understanding of the Drosophila central circadian clock.
Keywords
Understanding circadian neural circuits has been a major challenge in the scientific field of circadian clock research. This is because animal circadian clocks are generally composed of a complex multicellular network that possess multiple input and output pathways. In the fruit fly Drosophila melanogaster, approximately 150 brain neurons act as the central pacemaker clock neurons that control behavioral rhythms (Top and Young, 2018; King and Sehgal, 2018). These 150 clock neurons are broadly subdivided into 9 groups: 4 dorsal neuron groups (anterior dorsal neuron 1 [DN1a], posterior dorsal neuron 1 [DN1p], dorsal neuron 2 [DN2], and dorsal neuron 3 [DN3]) and 5 lateral neuron groups (small ventral lateral neuron [s-LNv], large ventral lateral neuron [l-LNv], 5th small ventral lateral neuron [5th s-LNv], dorsal lateral neuron [LNd], and lateral posterior neuron [LPN]). These groups of clock neurons are distinct in their function, morphology, and neurotransmitter content. However, the functions of individual clock neurons and their network properties are still under investigation. The primary techniques for dissecting the function of clock neurons are the GAL4-upstream activating sequence (UAS) system and analogous gene expression systems such as the LexA and Q-systems, all of which enable expression of the gene of interest in a tissue-specific manner (Owald et al., 2015). GAL4 lines are engineered to contain the gal4 gene downstream of a desired gene promoter, enabling tissue-specific expression of the GAL4 protein. UAS lines are engineered to contain a gene of interest downstream to the UAS, a specific enhancer to which the GAL4 protein binds in order to activate transcription (Brand and Perrimon, 1993). Thus, the promoter upstream of the gal4 gene determines in which tissue the gene of interest (downstream of the UAS sequence) is expressed. For example, in circadian rhythm research, the Pigment-dispersing factor (Pdf)–GAL4 line has significantly contributed to the investigation of the ventral lateral clock neurons (s-LNv and l-LNv), because of the very specific expression pattern of Pdf in those neurons (Renn et al., 1999; Park et al., 2000). To date, several GAL4 lines that express in clock neurons have been generated or identified. However, these GAL4 lines do not cover all types of clock neurons, and there is a need for a greater variety of clock neuron GAL4 lines to facilitate further research.
In this study, we screened the databases of GAL4 lines that were established in the Janelia Research Campus and the Vienna Drosophila Resource Center, in which there are currently more than 10,000 GAL4 lines with different gene enhancers (Pfeiffer et al., 2008; Jenett et al., 2012; Tirian and Dickson, 2017). After screening the databases of Janelia-GAL4 and VT-GAL4 lines (http://flweb.janelia.org/cgi-bin/flew.cgi and https://stockcenter.vdrc.at/control/main, respectively), we identified approximately 300 lines as potential GAL4 lines that may exhibit expression in clock neurons. These GAL4 lines were crossed with a UAS-Green Fluorescent Protein (GFP) line. Their offspring were sampled at zeitgeber time 20 under a 12-h light:12-h dark cycle, and their brains were dissected and processed for triple immunostaining using antibodies to detect the expression of PAR domain protein 1 (PDP1), GFP, and PDF. As a result, we identified 63 GAL4 lines that exhibit expression in subsets of clock neurons. Some of these lines showed specific expression that was restricted to clock neurons, whereas many lines also exhibited expression in regions not specific to the neuronal clock network (Fig. 1). Table 1 lists the GAL4 lines that exhibited expression in clock neurons. We also show images of immunostained brain sections of all GAL4 lines in the supplemental data (Suppl. Fig. S1-S75), where the GAL4 expression patterns can be seen not only in clock neurons but also in nonclock neurons in the brain (for nonclock neurons, see also Janelia- and VT-GAL4 databases). While we were screening the GAL4 lines, some Janelia- and VT-GAL4 lines were already reported as clock neuron-specific lines (Table 2). Nevertheless, we included some of these in our list because GFP expression patterns can depend on which GFP reporter line is used. For example, the R18H11-GAL4 line has been previously identified as a GAL4 line specific for DN1 clock neurons (Kunst et al., 2014). However, we have additionally observed expression in a few LNd neurons, which may be due to the different GFP line or the sensitivity of the anti-GFP immunostaining protocol used in this study. In agreement with our result, at least 2 LNd neurons are clearly visible in the images of the R18H11-GAL4 expression pattern presented in the Janelia GAL4 Flylight database.
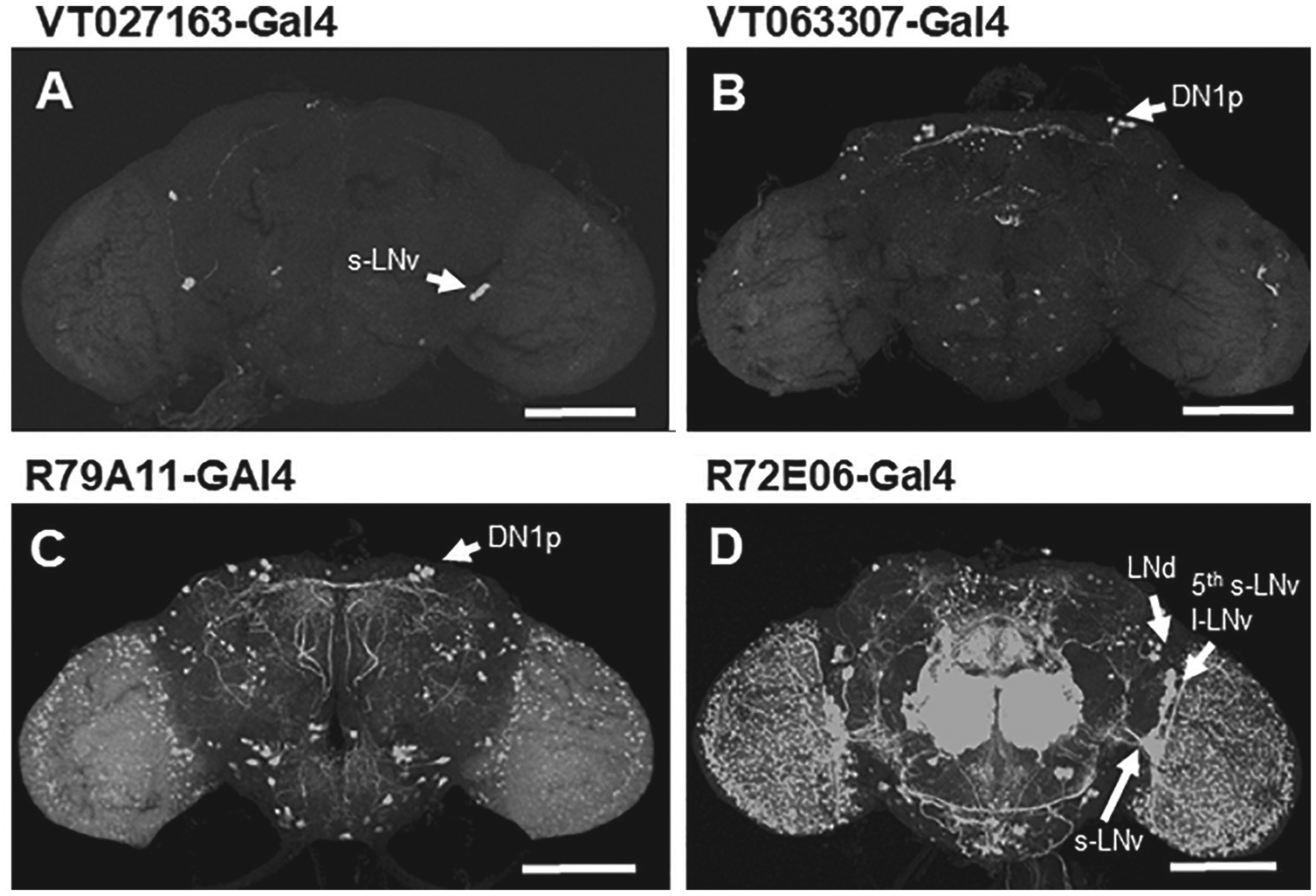
Representative GAL4 expression images in the Drosophila brain. Each GAL4 line was crossed with a UAS-GFP line, and their F1 offspring were processed for anti–green fluorescent protein GFP immunostaining. VT027163-GAL4 (A) and VT063307-GAL4 (B) lines exhibit rather specific expression in subsets of clock neurons, whereas the other GAL4 lines exhibit expression in many nonclock neurons (C, D). Arrows indicate clock neurons. Scale bar = 100 µm.
Clock neuron-specific Janelia-GAL4, VT-GAL4, and split-GAL4 lines. a
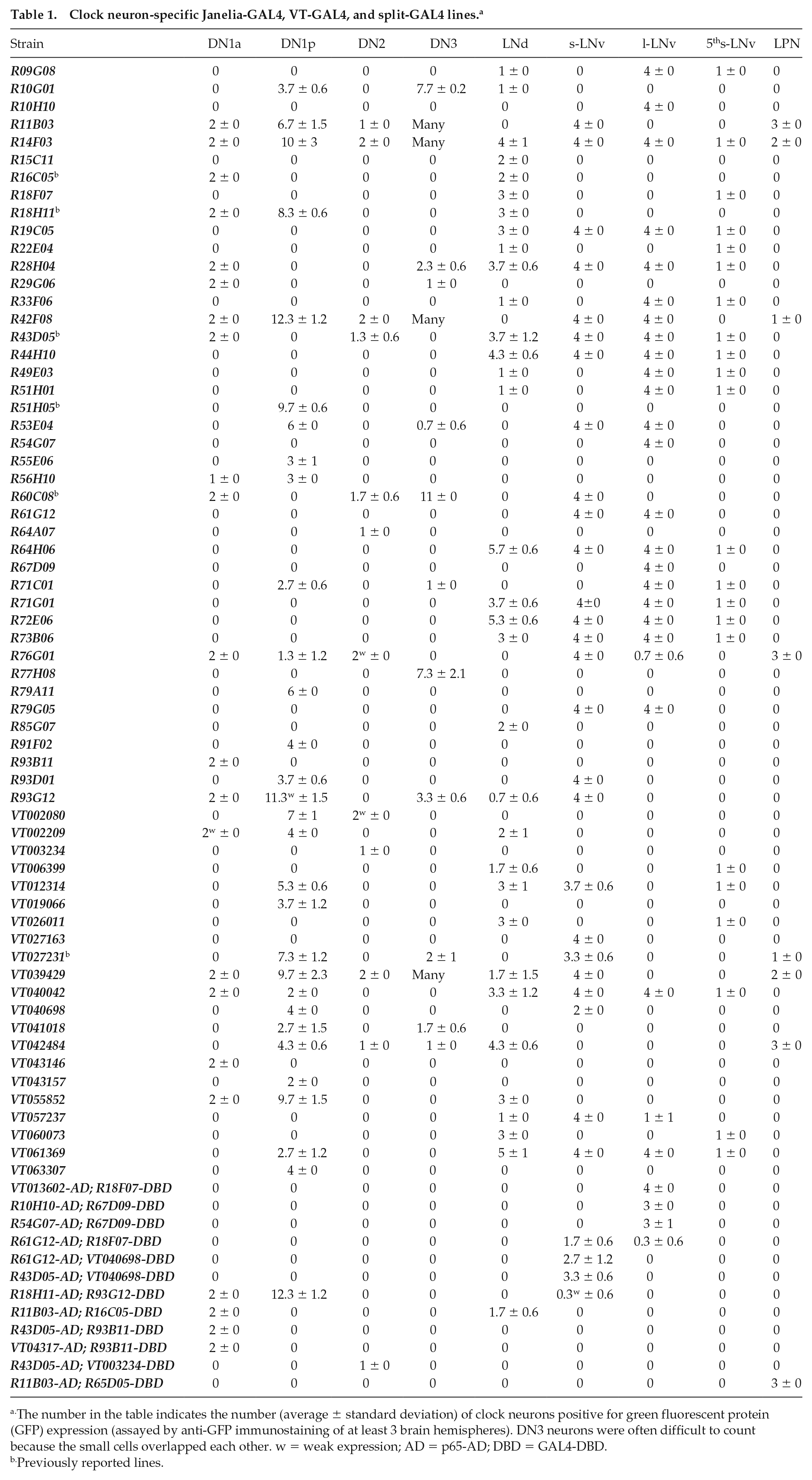
The number in the table indicates the number (average ± standard deviation) of clock neurons positive for green fluorescent protein (GFP) expression (assayed by anti-GFP immunostaining of at least 3 brain hemispheres). DN3 neurons were often difficult to count because the small cells overlapped each other. w = weak expression; AD = p65-AD; DBD = GAL4-DBD.
Previously reported lines.
Clock neuron-specific Gal4, LexA, or split-GAL4 lines that have been previously reported. a
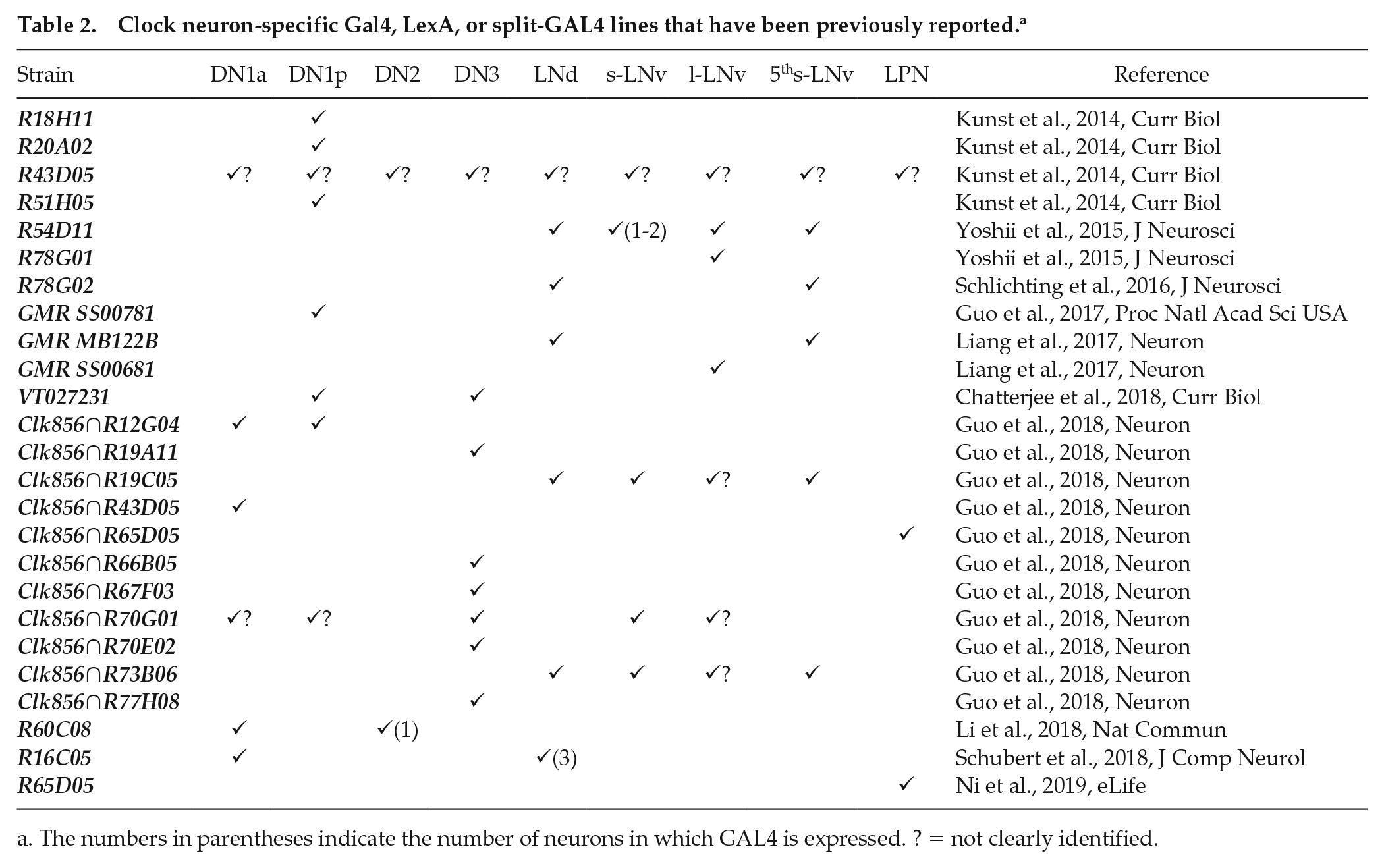
The numbers in parentheses indicate the number of neurons in which GAL4 is expressed. ? = not clearly identified.
Although the R18H11-GAL4 line exhibits very restricted expression in subsets of DN1 and LNd neurons, with only few nonclock cells in the brain, many cells in the ventral nerve cord express GAL4 (from the Flylight project database). Similarly, while R15C11 (LNd), R16C05 (DN1a and LNd), R51H05 (DN1p), R61G12 (s-LNv and l-LNv), R77H08 (DN3), R85G07 (LNd), VT027163 (s-LNv), VT055852 (DN1 and LNd), and VT063307-GAL4 (DN1p) lines exhibit rather restricted expression patterns in the brain, all express GAL4 in some cells of the ventral nerve cord. In addition, all GAL4 lines listed here may have expression outside of the nervous system, which was not examined in this study. Therefore, one should consider side effects when those lines are used to manipulate clock neurons. Several of the 63 GAL4 lines target the same group of clock neurons. Thus, it is better to use more than one GAL4 line to reduce the risk of side effects stemming from untargeted cells. Even for nonspecific GAL4 lines, clock neuron-specific effectors, such as the UAS-period line, can be used for a period rescue experiment (Blanchardon et al., 2001), UAS-dominant-negative clock and cycle lines for a clock disruption (Tanoue et al., 2004), or UAS-guide RNA line for cell-specific CRISPER-Cas9 knockouts (or UAS-RNAi lines) for genes specifically expressed in clock neurons (Martinek and Young, 2000; Delventhal et al., 2019; Schlichting et al., 2019).
LNd and DN1p neurons contain heterogeneous neuron types (Johard et al., 2009; Yoshii et al., 2008). We characterized several GAL4 lines that expressed in subsets of DN1p or LNd neurons, although not exclusively. Those lines could be further used to label and manipulate DN1p and LNd neurons. Moreover, the GAL4 lines identified here cover DN1a, DN2, DN3, and LPN neurons that are less studied and will be useful for functional analysis of those clock neurons.
The split-GAL4 system is a technique to refine GAL4 expression in cells by separating the DNA-binding domain (DBD) and the activation domain (AD) of GAL4 and inserting each domain downstream of 2 different gene promoters (Luan et al., 2006; Pfeiffer et al., 2010; Dionne et al., 2018). Previous studies have already reported split-GAL4 lines for s-LNv, 5th s-LNv, and LNd, and DN1p neurons (H. Dionne, G. Rubin, and A. Nern, unpublished data; Liang et al., 2017; Guo et al., 2017). A large collection of split-GAL4 hemi-drivers is now available in the Bloomington Drosophila Stock Center (donated by G. Rubin and B. Dickson at the Janelia Research Campus). We selected some hemi-drivers and crossed them to generate split-GAL4 lines. Eventually, we created 12 split-GAL4 lines that exhibit expression in DN1a, l-LNv, s-LNv, and LPN neurons, among others (please refer to Table 1 for the full list). V013602-AD; R18F07-DBD, R43D05-AD; R93B11-DBD and VT04317-AD; R93B11-DBD, R11B03-AD; and R65D05-DBD lines are specific for the l-LNv, DN1a, and LPN populations, respectively, and they fill in the gaps of already reported split-GAL4 lines. Interestingly, R10H10-AD; R67D09-DBD line labels 3 of the 4 l-LNv neurons, but the other l-LNv split-GAL4 line (R61G12-AD; R18F07-DBD) labels only 1 l-LNv neuron. A recent report demonstrated that one of the l-LNv neurons has a projection pattern that differs from those of the other 3 l-LNv neurons (Schubert et al., 2018), suggesting that the l-LNv neurons comprise 2 functionally distinct groups. Similarly, the R43D05-AD; VT003234-DBD split-GAL4 line labels only 1 of the 2 DN2 neurons, and it has been reported that one DN2 neuron exhibits a clock protein expression profile that is distinct from the other one (Yoshii et al., 2009). We observed nonspecific expression, even in the split-GAL4 lines; however, the level of expression was very low. This may be because GAL-DBD has weak transcriptional activity on its own (Luan et al., 2006). Thus, it is still worthwhile to create multiple split-GAL4 lines for a single clock population to minimize unwanted side effects. Our initial 63 GAL4 lines would provide many combinations of the 2 enhancers.
In the late 1990s, the discovery of PDF and PDF-positive LNv neurons caused a breakthrough in our understanding of the circadian clock at a neuronal level (Helfrich-Förster, 1995; Renn et al., 1999), which led to the functional analysis of the clock neuron network, such as its division into morning and evening oscillators (Grima et al., 2004; Stoleru et al., 2004). Recently, the role of DN1s neurons in sleep has received a lot of attention (Guo et al., 2016; Guo et al., 2018; Lamaze et al., 2018). In addition, several recent studies have performed functional analyses of relatively minor clock neuron groups (Fujiwara et al., 2018; Ni et al., 2019). An understanding of the entire clock circuit remains elusive, but the GAL4 lines presented in this study will provide a new genetic toolkit to facilitate further investigation.
Supplemental Material
Fig_Supplemental_v2 – Supplemental material for A Catalog of GAL4 Drivers for Labeling and Manipulating Circadian Clock Neurons in Drosophila melanogaster
Supplemental material, Fig_Supplemental_v2 for A Catalog of GAL4 Drivers for Labeling and Manipulating Circadian Clock Neurons in Drosophila melanogaster by Manabu Sekiguchi, Kotaro Inoue, Tian Yang, Dong-Gen Luo and Taishi Yoshii in Journal of Biological Rhythms
Footnotes
Appendix
The Janelia GAL4 and VT-GAL4 lines were obtained from the Bloomington Drosophila Stock Center (BDSC) and the Vienna Drosophila Resource Center, respectively. The y w; UAS-GFP S65T line (BDSC #1522) was used to cross with each GAL4 and split-GAL4 line.
Immunostainings were performed as described previously (Yoshii et al., 2015). The primary antibodies used were chicken anti-GFP (1:1000; Rockland, Limerick, PA), mouse anti-PDF (1:1000; Developmental Studies Hybridoma Bank; Cyran et al., 2005), and rabbit anti-PDP1 (1:9000; Cyran et al., 2003). We used the following fluorescent-dye conjugated secondary antibodies at a 1:500 dilution: Alexa Fluor 488 nm (goat anti-chicken), 647 nm (goat anti-mouse) antibodies (Life Technologies, Carlsbad, CA), and goat anti-rabbit Cy3 antibodies (Millipore, Billerica, MA). PDP1 immunostaining was used to visualize all clock neurons. The s-LNv and l-LNv neurons were characterized by PDF staining, which was also useful for distinguishing the 5th s-LNv and LNd groups that were also located within lateral brain regions. LPN neurons were located posteriorly; therefore, it was easy to distinguish them from other LN groups. DN1 neurons were located in the dorsal brain region. The DN1a group consisted of a pair of neurons that resided within the rim of the dorsal anterior side, compared with the DN1p group. The DN2 group was located deeper within the dorsal brain compared with the DN1p neurons and close to the s-LNv dorsal projections, as labeled by PDF staining. DN3 neurons were composed of approximately 30 small cells and were located within the rim of the dorsal brain but more lateral than the other DN1 group. At least 5 brains for each strain were used for immunostaining to characterize the number of clock cells stained by anti-GFP antibodies (GAL4 expression) in the first experiment, and the clock cells were briefly characterized. In the second experiment, we conducted the same immunostaining, but only for positive strains, to confirm the prior results. Images were taken from at least 3 different brains using laser scanning confocal microscopes (Olympus FV1200, Olympus, Tokyo, Japan). Finally, we counted the number of clock cells from the scanned brains. When the GFP immunostaining signals in clock cells were very weak and only slightly stronger than background signals, we added the annotation “weak expression,” as seen in Table 1.
Acknowledgements
This work was funded by the JSPS (KAKENHI 15H05600 and 19H03265) and National Natural Science Foundation of China (31671085, 31871058 and 31930043). We would like to thank C. Hermann-Luibl for preliminary work and comments on the article and the Janelia Flylight project team, the Bloomington Drosophila Stock Center, and the Vienna Drosophila Resource Center for providing fly lines. We are also grateful to J. Blau and the Developmental Studies Hybridoma Bank for providing antibodies.
Conflict Of Interest Statement
The authors have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
