Abstract
In Neurospora and other fungi, the protein frequency (FRQ) is an integral part and a negative element in the fungal circadian oscillator. In Drosophila and many other higher organisms, the protein period (PER) is an integral part and a negative element of their circadian oscillator. Employing bioinformatic techniques, such as BLAST, CLUSTAL, and MEME (Multiple Em for Motif Elicitation), 11 regions (sequences) of potential similarity were found between the fungal FRQ and the Drosophila PER. Many of these FRQ regions are conserved in many fungal FRQ(s). Many of these PER regions are conserved in many insects. In addition, these regions are also of biological significance since mutations in these regions lead to changes in the circadian clock of Neurospora and Drosophila. Many of these regions of similarity between FRQ and PER are also conserved between the Drosophila PER and the mouse PER (mPER2). This suggests conserved and important regions for all 3 proteins and a common ancestor, possibly in those amoeba, such as Capsaspora, that sits at the base of the phylogenetic tree where fungi and animals diverged. Two additional examples of a possible common ancestor between Neurospora and Drosophila were found. One, the white collar (WC-1) protein of Neurospora and the Drosophila PER, shows significant similarity in its Per/Arnt/Sim (PAS) motifs to the PAS motif of an ARNT-like protein found in the amoeba, Capsaspora. Two, both of the positive elements in each system (i.e., WC-1 in Neurospora and cycle [CYC] in Drosophila), show significant similarity to this Capsaspora ARNT protein. A discussion of these findings centers on the long-time debate about the origins of the many different clock systems (i.e., independent evolution or common ancestor as well as to the question of how new genes are formed).
Both the Neurospora and Drosophila clock systems have several basic properties in common. A simplified version of each indicates that they are composed of transcription translation feedback loops and have several other properties in common, as follows.
The Neurospora clock is composed of the positive elements, white collar 1 and 2 (WC-1 and WC-2), which dimerize and then translocate to the nucleus, where they activate the transcription of the frequency (frq) gene; the negative elements, such as FRQ, which interact with the positive elements to affect the levels of these positive elements; and the modulating elements, such as phosphorylation by 1 or more protein kinases, degradation by a ubiquitin-based process, and interaction with other proteins, which affect the level of the other elements as well as their location. A very detailed description of all of these processes as well as a useful diagram was published recently (Dunlap and Loros, 2017).
The Drosophila clock is composed of the positive elements clock (CLK) and cycle (CYC), which dimerize and then translocate to the nucleus, where they activate the transcription of the period (per) gene and the timeless (tim) gene; the negative elements, such as PER, which interact with the positive elements to affect the level of these positive elements; and the modulating elements, such as phosphorylation by 1 or more protein kinases, degradation by a ubiquitin-based process, and interaction with other proteins, which affect the level of the other elements as well as their location. A more detailed description of these processes was published recently (Lam and Chiu, 2017).
The discussion of the evolution of clocks has largely centered around 2 themes: What event or events led to the selection for a circadian oscillator? Did the eukaryotic circadian oscillator, found in plants, fungi, insects/animals, and so on, arise from a common ancestor, or were these independent events in different types of organisms?
The speculation on the first question largely consists of 3 parts: the circadian oscillator arose as a “flight from light,” the circadian oscillator arose to deal with a “great oxidation event,” and the circadian oscillator arose to synchronize the internal activities of a cell or group of cells (i.e., a temporal order to metabolism and synthesis).
Some of these questions have been the subject of speculation in Pittendrigh’s (1993) famous essay, “The Darwinian Clock-Watchers,” and more recently by Rosbash (2009) and in some detail by Lam and Chiu (2017).
The question of a common ancestor between the clock systems of Drosophila and mammals has been addressed by Rosbash (2009) and others. Some similarities are given in terms of the basic elements of the clock architecture. The similarities between the systems in terms of orthologs for the positive elements are striking. Additional evidence is found for the similarities for the kinases and other processes. But what about the similarities between the clock systems even further back in evolutionary time, that is, Neurospora versus Drosophila?
The clock systems found in Neurospora and Drosophila have many features in common, such as the formal architecture of their circuitry, as outlined in the positive and negative elements, the presence of temperature compensation, and so forth. Many of the similarities are found in the book The Genetics of Circadian Rhythms (Brody, 2011). Additional evidence for a common ancestor is the role of the effects on the clock due to protein kinases, such a CKI in Neurospora (He et al., 2006) and its homologue (DBT) in Drosophila (Price et al., 1998). In addition, the effects on the clock of Neurospora from ubiquitin-based proteolytic effects due to mutations in FWD-1 (He et al., 2003) and mutations in its homologue (SLIMB) has effects on the clock of Drosophila (Grima et al., 2002). However, these pieces of evidence are not compelling enough, as they can also be interpreted as evidence for independent evolution, that is, clocks just arose in the milieu of protein kinases, and so forth, and therefore, they share these components in common.
Many reviews of the Neurospora clock system have been published over the years. The most recent one by Dunlap and Loros (2017) is quite comprehensive and refers back to many of the earlier reviews. Reviews of the Drosophila clock system also have been plentiful and date back many years. Some recent reviews can be found published by King and Sehgal (2018), Young (2018), and Top and Young (2018).
Reviews comparing the different clock systems can be found by Rosbash (2009), Young and Kay (2001), and Dunlap and Loros (2017).
This communication differs from the others in that it deals with a comparison of the 2 clock systems at the level of the similarities of the protein sequences for the 2 key clock proteins in each system, FRQ in fungi and PER in Drosophila.
To better understand the structure of the key negative elements in each clock system (i.e., FRQ and PER), the domains in each, known so far, are shown in the 2 diagrams as Figure 1 and Figure 2. The postulated domains for the Neurospora crassa FRQ protein (Fig. 1) are adapted from a figure published previously (Dunlap and Loros, 2017).
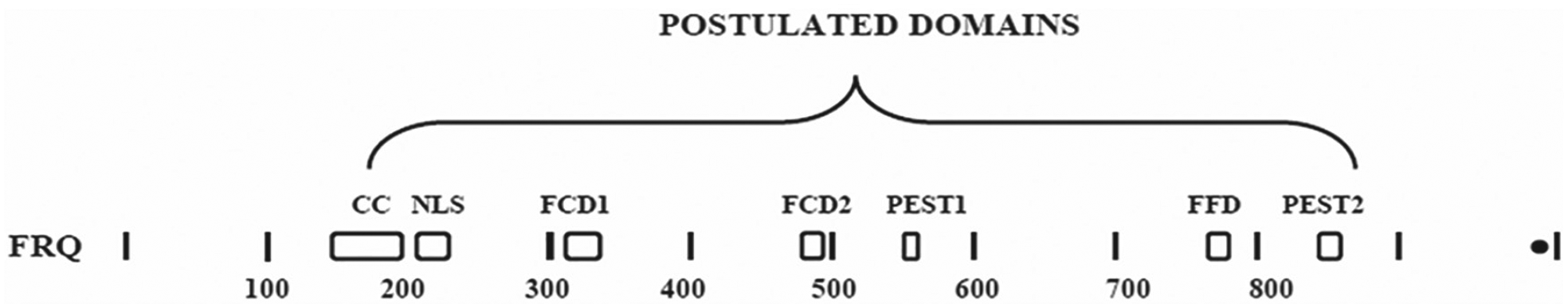
Diagrammatic illustration of the 7 FRQ domains postulated so far. Adapted from Dunlap and Loros (2017). Abbreviations: CC = coiled/coil; NLS = nuclear localization signal; FCD1 and FCD2 = FRQ/casein kinase binding sites 1 and 2; PEST 1 and 2 = postulated sequences containing the amino acids proline, glutamate(E), and serine/threonine; FFD = FRQ/FRH binding domain.

Diagrammatic illustration of the 8 PER domains postulated so far. Adapted from Li et al. (2019). Abbreviations: SLIMB-BINDING = site for the binding of a kinase; PAS-A and -B = Per/Arnt/Sim areas of similarity; CLD = cytoplasmic localization domain; PERs = area encompassing the site of the per-short mutation; DBT = site of the binding of Doubletime (protein kinase); NLS = nuclear localization signal; CLK = clock binding area.
The postulated domains for the Drosophila melanogaster PER (Fig. 2) are adapted from a study published previously (Li et al., 2019).
Methods
The standard BLASTP program was employed with the default parameters for the gap existence/extension penalties of 11,1, except where noted. The NCBI Genebank (blast.ncbi.nlm.nih.gov/Blast.cgi) was the source for sequences, and Firefox was employed as the server. The Clustal Omega program supported by EMBL (https://www.ebi.ac.uk/Tools/msa/clustalo) was employed. The website MEME-suite.org contained the motif discovery program (MEME) version 5.0.2. The numbers listed are the Genebank identity numbers (GI#..).
The fungal FRQ(s) employed for comparative analysis (MEME) were as follows: Neurospora crassa 95952, Sordaria fimicola 6016053, Fusarium oxysporum 342889466, Botrytis cinerea 347827659, Colletotrichum sublineola 640918779, Trichoderma virens 358386587, Acremonium chrysogenum 672797659, Sclerotonia sclerotiorum 156039415, Marssonina brunnea 597583253, Aureobasidium pullulans 662524541, Phaeosphaeria nodorum 169600449, Rhodotorula toruloides NP11 EMS19378, Taloromyces st. 242790999, Saitotella complicata 813215842, Magnaporthe oryzae 145610785, Zymosepteria tritica SMR54202, Pyronema tritica 189211261, and Rhizophagus irregularis 595497380.
Seven insect PER(s) were employed for the construction of PER MEME(s): Bombus 340723130, Apis 40645091, Tribolium 270008118, Drosophila m. 7290345, Nasonia XP_008209246, Anopheles KFB40661, and Acyrthosiphon 284005204.
Two other PER(s) employed for alignments of the PAS and cytoplasmic localization domain (CLD) sections were Danio 190337630 and Branchiostoma 163311886.
Other designations included mPER2 6174896, Capsaspora ARNT 754351513, Acanthamoeba HSF 470459745, Neurospora WC-1 5441498, and Ostreococcus LARP 308813524.
Results
FRQ versus PER
In the initial comparisons of FRQ versus Drosophila PER, it was noted that there were some regions of similarity, that is, tracts of threonine-glycine (T-G) repeats (McClung et al., 1989). These were of unknown significance at that time. Subsequently, it was found that many of the FRQ(s) from other species of fungi (Sordaria, Leptosphaeria, Magnaporthe, to name a few) have smaller or no T-G repeat regions, and some of the PER(s) from other insect species have variable T-G regions. For instance, the length of the T-G region in Drosophila virilis is ~20, in D. melanogaster is ~60, and in Drosophila pseudoobscura is ~200. In Drosophila, data have been presented suggesting that the T-G repeats form a type II or III beta-turn (Guantieri et al., 1999) and that these repeat regions may play a role in temperature compensation (Sawyer et al., 1997).
In this study, newer methods, such as CLUSTAL and MEME, were employed to reexamine the relationship of FRQ to PER. When the Sordaria or Neurospora FRQ were compared with the D. melanogaster PER via BLAST, only 2 sections showed even low levels of similarity. One in particular, shown in Figure 3 as section G, had a BLAST value of 0.16(24), where (24) is the length of the match. These regions of PER and FRQ were of interest since they have some biological significance. Section G of FRQ is conserved among the fungal FRQ proteins (listed in Supplementary Table S1 as motif FF-5).

Multiple FRQ/PER comparisons. CLUSTAL-derived comparisons of 11 sections of FRQ and PER. FRQ is that of Sordaria fimicola, PER is that of Drosophila melanogaster. Asterisks above the line of FRQ are some of the many serines or threonines that have been shown to be sites of phosphorylation in Neurospora (Tang et al., 2009), while the asterisks below the line of PER are some of the many serine or threonine residues that have been shown to be sites of in vivo phosphorylation in Drosophila (Li et al., 2019). The letters designating the amino acids that are in red (i.e., S) indicate the amino acids in Sordaria FRQ corresponding to where clock mutations have occurred in Neurospora. Likewise, the amino acids in red for Drosophila melanogaster PER are the sites of known clock mutations. The numbers in parentheses are gaps in the amino acid sequences.
This section of PER is 10 amino acids down from the beginning of a motif, designated some time ago (Nambu et al., 1991) as PAS-A (
To follow up on this small match of 2 conserved regions (section G), the Sordaria FRQ was compared with the Drosophila PER, employing CLUSTAL. The Sordaria FRQ has a BLAST value of 0.0 versus the N. crassa FRQ and can substitute for the Neurospora FRQ (Merrow and Dunlap, 1994). Instead of comparing the 2 proteins beginning at the N-terminus of each, the analysis began with the region of match shown as YDG.. for the FRQ protein and as HDG.. for the PER protein. This analysis showed several other regions of matching further down from the Y/HDG start site, and some of these regions are shown in Figure 3 as sections H, I, and J. Several of these regions are also of some biological significance. Section H is proposed to be part of the TIM-binding region of PER (Gekakis et al., 1995), section I is a conserved motif in the PAS-B domain (Suppl. Fig. S1), and section J of PER is part of the CLD as shown by Chang and Reppert (2003).
In addition, part of section G, most of section I, and most of section J are conserved in the PER(s) examined from 4 different species, as shown in Supplementary Figure S2.
For the FRQ(s), the equivalent sections (i.e., G, I, and J) are conserved, as exemplified as motifs FF-5, FF-7, and FF-8, respectively. To compare section G with motif FF-5, one needs to start with the amino acids DG in section G and DG in FF-5. The 3 areas of FRQ that are conserved match up to the 3 areas of PER that are conserved. In addition, the spacing between these three areas is roughly the same for both proteins.
Additional studies were performed employing the FRQ(s) from other fungal species versus the PER(s) from other Drosophila species. The same match shown in section G above was generally found. Section A was found when comparing N. crassa FRQ with D. pseudoobscura PER by BLAST. This section is shown in Figure 4. This region of FRQ is conserved (FF-10) among fungi. This region of Drosophila PER is also conserved in insects and is of biological significance; that is, mutations led to clock changes (Table 1). This region of PER in Drosophila is conserved in mouse PER2.
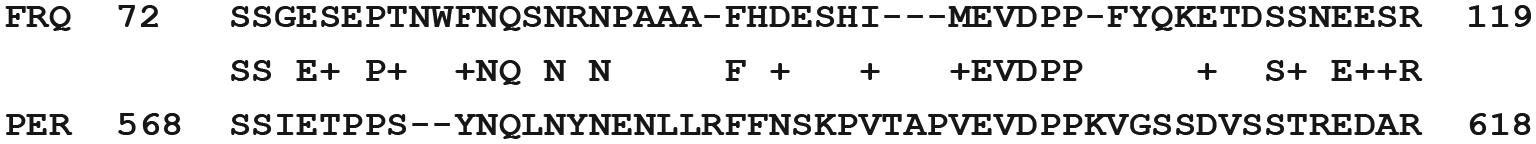
Similarity of Neurospora crassa FRQ and Drosophila pseudoobscura PER. When the gap/extension penalty was changed from 11,1 to 6,2, an additional area of low similarity, BLAST value 3.8(53), between the FRQ from N. crassa and the PER from D. pseudoobscura was found.
PER Sections a
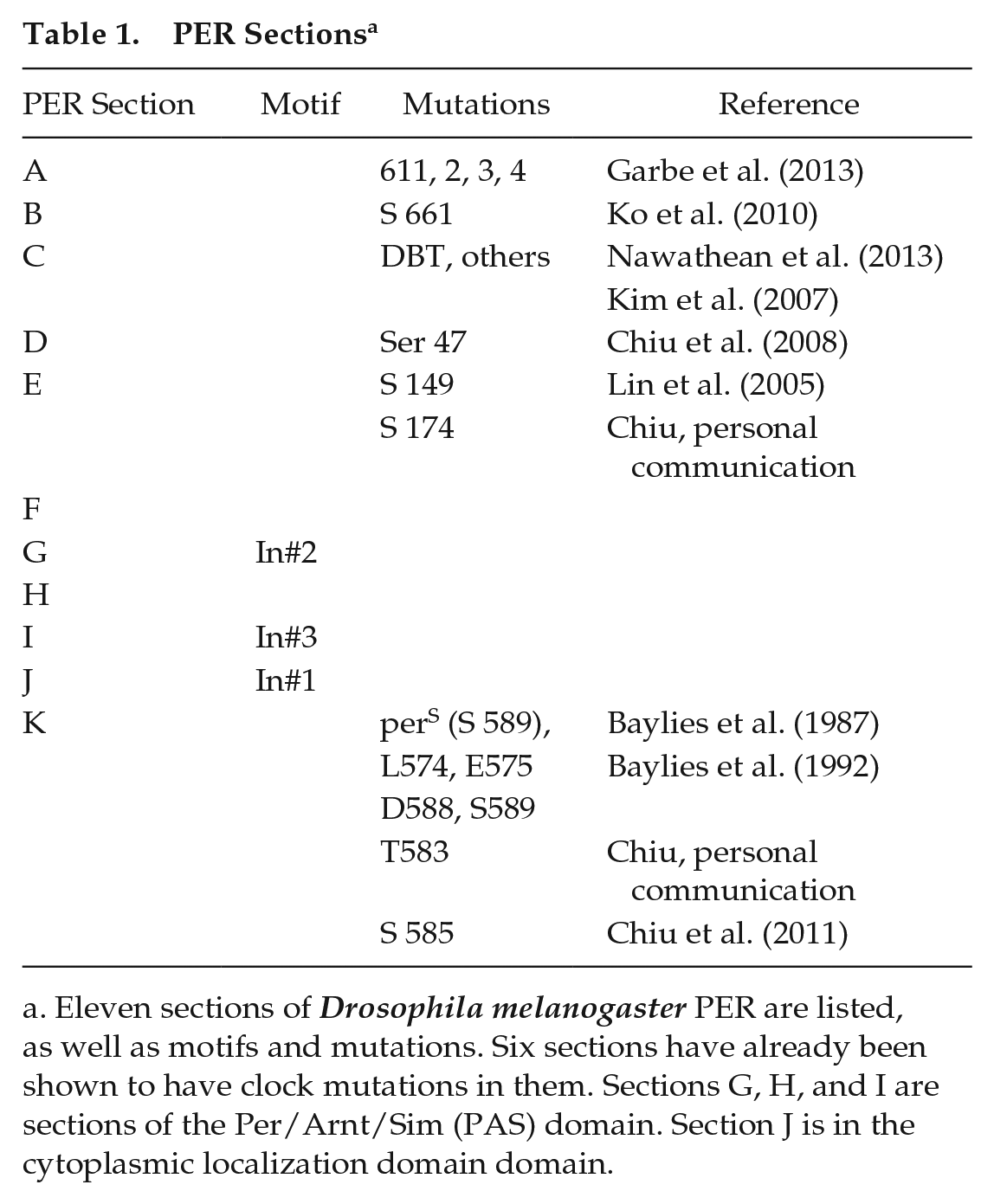
Eleven sections of Drosophila melanogaster PER are listed, as well as motifs and mutations. Six sections have already been shown to have clock mutations in them. Sections G, H, and I are sections of the Per/Arnt/Sim (PAS) domain. Section J is in the cytoplasmic localization domain domain.
Employing this sequence match, a very similar sequence comparison was then found for the FRQ from Sordaria f. and the PER from D. melanogaster, and this is illustrated as section A in Figure 3. The PER proteins from other Drosophila species also match up to this FRQ. The numbering for the amino acids in this section of Figure 3 is different than for the equivalent section shown in Figure 4 as section A, since Figure 4 was a comparison of D. pseudoobscura PER with Neurospora FRQ, while Figure 3 shows a comparison of D. melanogaster PER with Sordaria FRQ.
Another comparison of these 2 proteins was undertaken by BLAST analysis of Sordaria FRQ versus D. pseudoobscura PER, and another section of low similarity was found between the end of section H and the beginning of section I, when the gap existence/extension settings were changed from 11,1 to 6,2. This section is not shown, since it can easily be derived by a BLAST program and is part of the PAS-B motif/domain. Section I is also part of the PAS-B motif/domain. Another comparison was done by extending the search area just past section J employing CLUSTAL. The 2 proteins were aligned starting at Sordaria fimicola FRQ at 880 with the D. pseudoobscura PER at 486. The results were then formatted for Drosophila m. PER and then appear as section K in Figure 3.
A more detailed analysis of Drosophila PER versus Sordaria FRQ was undertaken employing CLUSTAL and is shown in Figure 3. A straightforward comparison of the 2 proteins, beginning with the N-terminal of each showed only a small section of potential match near the C-terminal area, listed as section K. There are several possible reasons for this. One, there is no similarity. Two, when one tries to align these 2 proteins beginning with the N-terminal of each, no useful information can be obtained. Three, the presence of large tracts of T-G repeats in both FRQ and PER affect the results so that those T-G tracts will preferentially match up. Thus, different alignments were employed. For instance, section D and parts of section E were found by beginning the CLUSTAL program with FRQ at 219 and PER at 4 and ending FRQ at 380 and PER at 149. Likewise, alignment of sections F, G, H, I, and J were found by beginning the CLUSTAL program with FRQ at 626 and the PER at 198. One can see that in addition to the possible match of PAS motifs, there are other sections, such as section D, in which more than half of the amino acids show identities or positives.
These individual sections were too small to give significant BLAST values, with the exception of section G, which gave a BLAST value of 0.56(24). Sections A, G, I, and J were conserved in many other fungal FRQ(s), as seen in motif(s) 10, 5, 7, and 8, respectively, as were much of sections D as motif 11. Section C contains motifs(s) 2 and 3. A summary of the FRQ sections, motifs, and mutations is given in Table 2.
FRQ Sections of Neurospora a
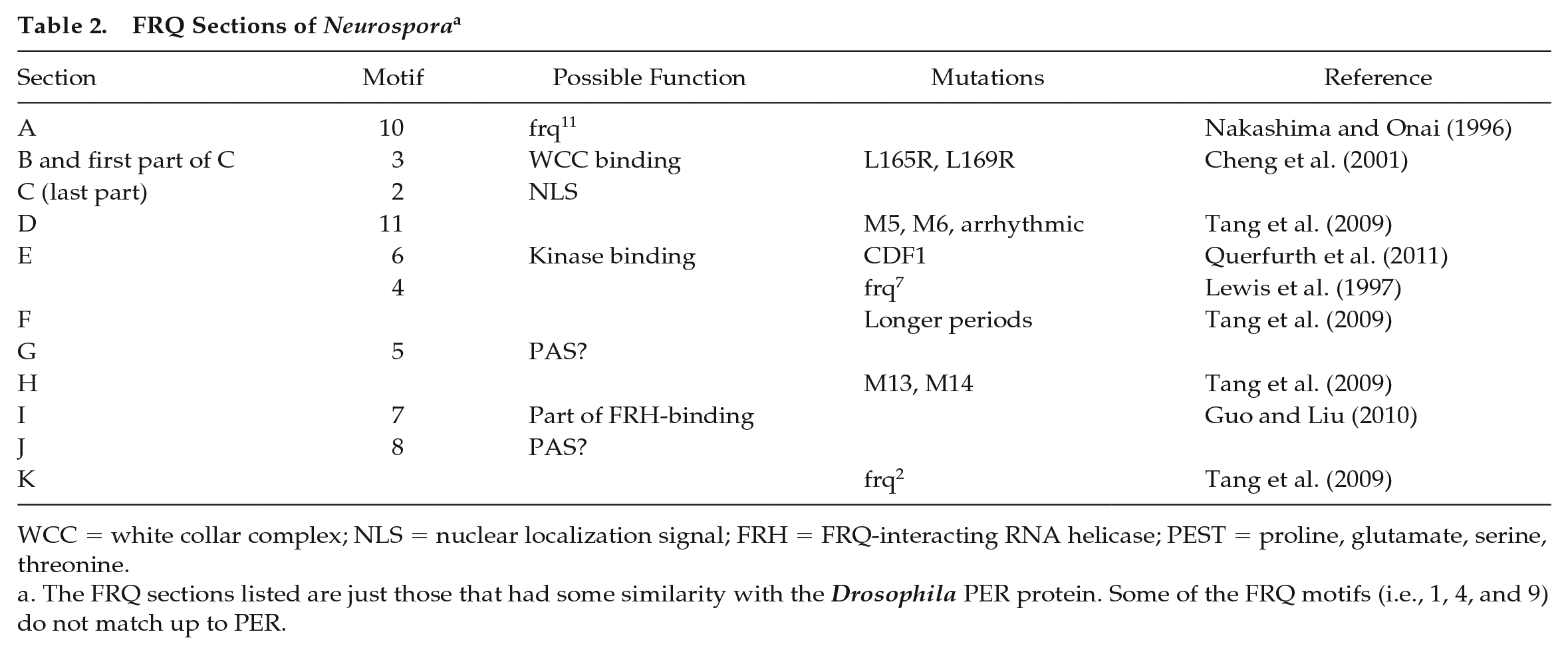
WCC = white collar complex; NLS = nuclear localization signal; FRH = FRQ-interacting RNA helicase; PEST = proline, glutamate, serine, threonine.
The FRQ sections listed are just those that had some similarity with the Drosophila PER protein. Some of the FRQ motifs (i.e., 1, 4, and 9) do not match up to PER.
Some of the FRQ motifs (i.e., 2, 3, and 5) are highly conserved in all of the 13 FRQ(s) listed in the Methods section.
In the Neurospora FRQ, there are 21 sites of amino acids changes that gave changes in the circadian rhythm. These are listed in Figure 3 in the Sordaria FRQ as the equivalents to the Neurospora FRQ. This is just a subset of the many that have been found. Likewise, there are 21 listed for the D. melanogaster PER, and this is just a sample of those known.
Eleven sections listed in Figure 3 were found to show some similarity between these 2 proteins. Approximately 240 identities/positives were found among the 11 sections. Since the total length of these proteins was 989 and 1218 amino acids, this is at least 20%. If one includes the matches of the T-G repeat regions (50-60 amino acids), it would be somewhat higher. There was no a priori reason to expect any similarities between FRQ and PER, considering that fungi and insects diverged hundreds of million years ago. Perhaps the finding of even low levels of similarity was surprising since the areas that do show some similarity in a protein from a given species show only partial conservation within that species. The nonconserved areas would be expected to show considerable divergence over this long time period.
If it is not just a coincidence that so many areas show even low levels of similarity, then it leads to several questions. One, did FRQ and PER have a common ancestor that was a part of a clock in a lower organism? Two, did FRQ and PER arise separately by different types of recombination of parts of a few genes that were not the “gears” of a clock mechanism? In this regard, FRQ and PER might have arisen from genes whose proteins were expressed with a rhythm due to entrainment and then there was a subsequent selection for new combinations of these genes/proteins that could have given rise to a free-standing oscillator mechanism.
It should be noted that the regions of FRQ that show some similarity to PER, as listed above, are the same regions of FRQ that are conserved in FRQ (Brody, 2019). In this regard, of the 11 regions of FRQ that were found by MEME analysis, listed in Supplementary Table S1, 8 were the regions that showed some similarity to PER. Of the 3 regions of PER found by MEME analysis (Suppl. Table S2), all were found to match up to regions of FRQ. Mutations in the amino acids P, P, S, Y (611, 612, 613, 614) all gave rise to longer periods (Garbe et al., 2013). When 27 amino acids of section C were deleted, strong effects on the rhythm were seen (Nawathean et al., 2013). Section D for the PER contains serine 47, which has been shown to be a site for phosphorylation (Chiu et al., 2008).
Some effort was put into matching the numerous other FRQ(s) versus the PER(s) from the numerous Drosophila species. There were a few small areas of additional areas of low homology between the Neurospora FRQ and the D. pseudoobscura PER, one of which is shown as Figure 4.
The alignment of the sections can be seen in Figure 5. The PAS motifs in PER are found in the N-terminal half, while the possible PAS motifs in FRQ are found in the C-terminal half. Vogt and Schippers (2015) have summarized the findings that show that PAS motifs are found in many different locations in proteins (i.e., N-terminal areas, mid-sections, C-terminal areas, etc.) depending on which particular protein contains them.
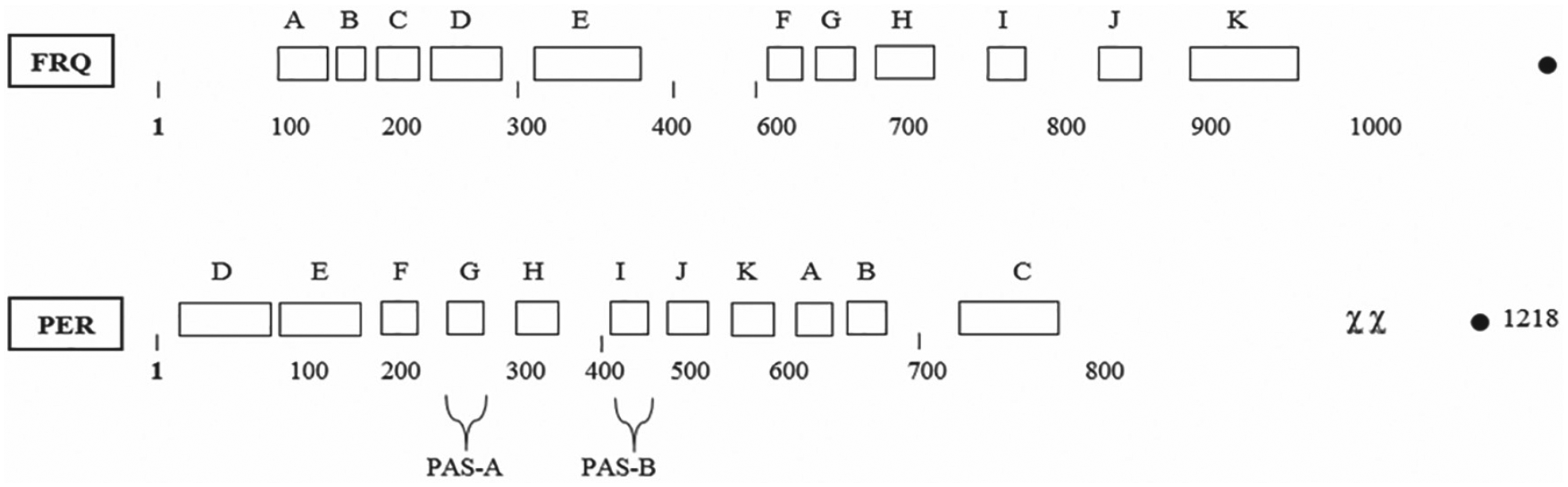
Diagrammatic representation of multiple potential similarities between Sordaria fimicola FRQ and Drosophila melanogaster PER. Note that the order of sections F through K is similar in both.
Does FRQ have PAS motifs found in Drosophila PER? The case for this idea is given above (i.e., several parts of the PAS conserved regions can be also found in FRQ). If this is not the case, then FRQ just has the remnants of all 3 of these PAS areas but with different functions. The possibility that FRQ(s) have PAS motifs is strengthened in several ways: (1) many of these regions of FRQ have not had any function assigned to them so far. Table 2 indicates the possible assigned functional areas of FRQ. (2) Section I in FRQ is part of the FRH-binding region. This function would be analogous with a protein (TIM) binding function for the equivalent PAS region of PER. (3) A scan of the Neurospora genome with PAS-containing proteins, such as ARNT or PER, found only “clock” proteins such as WC-1 and WC-2. Clearly, only experimental evidence will settle the question as to whether FRQ has some PAS-like functions.
In PER, section D includes serine 47, designated with an asterisk, which has been shown to be the site of a phosphorylation event necessary for the degradation of PER (Chiu et al., 2008). It would be interesting if the equivalent serine in an FRQ was also involved in this manner. This is a
The comparison of FRQ with Drosophila PER suggests the possibility, but clearly does not prove, that they had a common ancestor at the time of divergence of fungi from insects. One possibility would be a eukaryotic single-celled organism, such as the amoeba Capsaspora, Acanthamoebae, or others, which have been postulated, based on extensive evidence (James et al., 2006) to be related to the precursors of the modern day filamentous fungi. These organisms may have contained 1 or more proteins with PAS motifs that were then modified during evolution depending on the species. The ARNT protein from Capsaspora is shown in Supplementary Figure S2 to have PAS motifs with some similarity to the PER of Drosophila.
Figure 3 shows 11 regions of similarity between FRQ and PER. Of these 11 sections, 4 show similarity as follows to sections of a known protein. Sections G, H, I, and J of Drosophila PER show similarities to sections of Capsaspora ARNT. The BLAST value for Drosophila PER versus Capsaspora ARNT is e−09(276) and it is the PAS sections. A comparison of all 3 (i.e., FRQ, PER, and ARNT) is outlined in Supplementary Figure S2.
In addition, parts of FRQ and PER seen in section D of Figure 3 may also have a common origin in Capsaspora La-RNA domain-binding protein (LARP). CLUSTAL analysis of FRQ versus Capsaspora LARP and PER versus Capsaspora LARP gave rise to a CLUSTAL analysis of all 3 together, as shown in Supplementary Figure S3. This section of the existing Capsaspora LARP may not be the direct precursor to FRQ and PER, in that there could be possible other steps in between, other organisms, other LARP(s).
Many of these short-sequence similarities can be considered as short linear motifs (Davey et al., 2015). In addition, many of these regions in the Drosophila PER show similarity to regions found in the mouse homologue of the Drosophila PER (i.e., mPER2). The BLAST comparing these 2 PER proteins was e−33(276). Although all of these similarities were for the PAS regions, other regions of similarity were found by CLUSTAL as well. Since there were regions of FRQ that showed similarity to the PAS region of Drosophila PER, a comparison was made by CLUSTAL of FRQ/Drosophila PER/ARNT (Suppl. Fig. S2) as well as FRQ/Drosophila PER/mPER2 (Fig. 6).

Multiple alignments of FRQ, Drosophila PER, and mPER2. The FRQ listed is that of Sordaria fimicola FRQ, the PER is that of the Drosophila melanogaster, and mPER2 is the mouse homologue of the Drosophila PER. The hashtags (#) listed above each section designate those amino acids that are either identical or positive for each of the 3 proteins. At least for the sections shown above, the number of amino acids that are identical or positive between Drosophila PER and mPER2 is 109. The number of amino acids that are identical or positive for all 3 (i.e., FRQ, Drosophila PER, and mPER2) is 82.
Many of the FRQ/Drosophila PER similarities are also found further up the evolutionary scale (i.e., in the mouse PER; Fig. 6). This suggests that those FRQ/Drosophila PER similarities have important functions since they were conserved even further.
The mouse PER2 protein has a structure of 2 short regions that are approximately 130 amino acids apart. One region has the SYQQ sequence beginning with amino acid 583 S, and the second region has the sequence beginning with LTKE, amino acid 717 L. The Drosophila m. PER may have a similar structure, in which a sequence beginning with SYNQ (S613) is found about 150 amino acids before a sequence beginning with LTES (L764). It would appear from the data shown in Figure 3 that the Sordaria FRQ might have a similar structure, that is, a sequence that starts with FDQ (F88) and then has an LTVE (L164) sequence about 80 amino acids further down. Perhaps this structural motif was present in a common ancestor?
Both FRQ and PER Appear to Be Composites
The fungal FRQ was suggested (Brody, 2019) to be a composite of sequences from ARNT, heat shock factor (HSF), and LARP (an RNA-binding protein).
Drosophila PER is suggested here to also be a composite of sequences from these same 3 proteins as shown in Figure 7. Parts of the N-terminal half of PER could have been derived from an ARNT protein, such as that found in Capsaspora. The ARNT similarities are listed as the PAS sections shown in Supplementary Figure S2. Two other sections of PER show some similarity to ARNT but are not shown here since they were not highly conserved in other ARNT proteins.
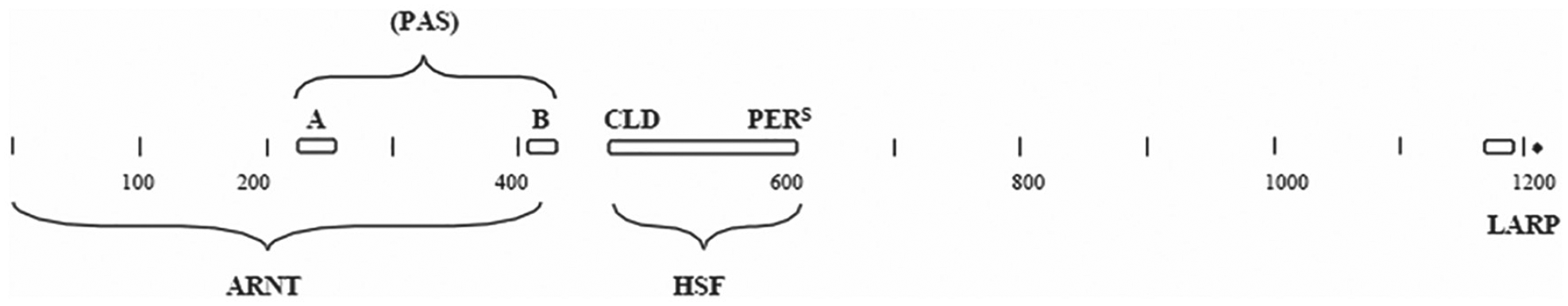
Diagrammatic representation of some of the sections of the Drosophila PER and the proposed similarities to other proteins. ARNT = aryl hydrocarbon receptor nuclear translocator; LARP = La-RNA domain-binding protein; HSF = heat shock factor. Other abbreviations are given in other legends.
The similarity to the HSF protein is also shown in Figure 7, in which a large region of PER shows similarity to HSF. These are the regions of PER given the label as CLD and pers. The complete match of these 2 PER regions with the Acanthamoeba HSF is shown in Supplementary Figure S4. Mutations in this pers region of PER showed changes in the circadian rhythm of Drosophila (Baylies et al., 1992). This region of PER and the area that it matched up to in HSF is shown as Supplementary Figure S4. Part 1 of this area of HSF has been shown to be involved in DNA-binding (NCBI Genebank superfamily cl12113). There is not enough similarity to propose that the corresponding area of PER has this same function, but it may be another example of a function in one protein being altered to take on another function in another protein.
Another section of PER, close to the C-terminal region, shows some similarity to the LARP protein C-terminal region (Suppl. Fig. S5). This similarity was found by CLUSTAL analysis of these 2 proteins. This region of LARP is highly conserved in LARP proteins and is close to the region of LARP(s) that has been shown to bind the 5′ end of certain RNA(s) (Bousquet-Antonelli and Deragon, 2009).
Analysis of WC-1
Although the original publication on the Neurospora WC-1 sequence (Ballario et al., 1996) did not indicate that there was some level of similarity to the Drosophila PER protein, subsequent studies (Linden and Macino, 1997) did. Examination of this similarity (BLAST value 0.001[303]) showed that the areas of PER that matched up to the WC-1 sequence were the PAS motifs. Additional studies of the WC-1 protein showed a similarity to the mouse BMAL1 protein (Lee et al., 2000) for these PAS motifs.
To follow up on this similarity, a search was made for a possible common ancestor to both WC-1 and PER. The search was focused on those amoeba-like species such as Capsaspora, Acanthamoeba, and so forth, which have been postulated to be at the base of the tree where fungi and animals diverged (James et al,. 2006). A scan of the Capsaspora genome employing the Neurospora WC-1 protein as the query and the word size of 3 indicated that there were several proteins with similarity. A protein of 848 amino acids in length had the third strongest “hit,” BLAST value e−13(199), GI# XP_004343694. This protein was the most interesting, since it also had the strongest “hit” in the Capsaspora genome, when the Capsaspora genome was scanned with the Drosophila PER, BLAST value e−9(276). This 848 protein showed high homology to the BMAL1b (
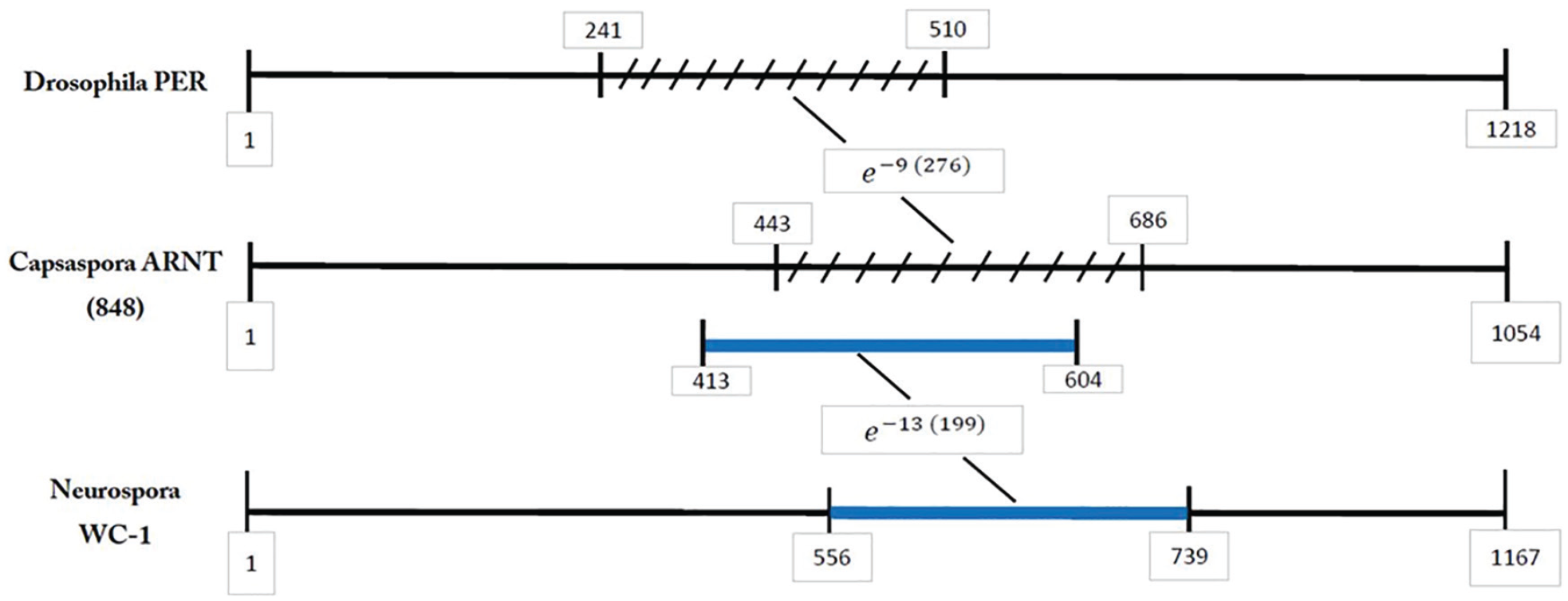
PER/ARNT/WC-1 similarities. A diagrammatic example showing that the same section of ARNT that matches up to the Drosophila PER also matches up to the Neurospora WC-1.
Although it cannot be known if the PAS domains of this Capsaspora 848 protein were the common ancestor for parts of both the Neurospora WC-1 and the Drosophila PER, it does raise this suggestion as a reasonable possibility. Clearly, an ancient species closely related to the modern Capsaspora should be considered as well.
Other genomes of existing micro-organisms were also scanned employing WC-1 and PER as queries. Of 6 possible fungal ancestors (James et al., 2006), only Capsaspora and Spizellomyces, but not Allomyces, Salpingoeca, Thecamonas, or Sphaeroforma, had proteins with significant homologous PAS regions to both WC-1 and PER. Proteins containing PAS domains were found in several other genomes (Chlamydomonas, Ostreococcus, Euglena, Acanthamobae) but not in Monosiga, Dictyostelium, or others. Many of these homologs appear to be blue-light receptors. The phylogeny of the WC-1 protein in fungi has been examined before (Salichos and Rokas, 2010), and this protein was found to occur in species belonging to many classes of fungi, as far back as the Zygomycetes. The analysis presented here suggests that the PAS motifs in the WC-1 protein can be found even further back in a phylogenetic tree (i.e., to the chytrids, such as Allomyces or Spizellomyces).
To be thorough, the WC-1 homologues from other fungal species were also tested versus the Capsaspora 848 protein, and all showed about the same amount of similarity to the 848 protein as did the N. crassa WC-1. Other insect PER(s) were tested versus the Neurospora WC-1 and/or the Capsaspora 848 protein. Most gave the same amount of similarity to WC-1 and/or the Capsaspora 848. However, some, such as the D. pseudoobscura PER versus N. crassa WC-1, gave slightly higher similarity e−5(308), and the Drosophila yakuba PER versus the Capsaspora 848 also had slightly higher similarity 3e−10(299).
The reciprocal analysis was performed, that is, a screen of the N. crassa genome employing the Capsaspora 848 (ARNT) protein as the query. Only 2 proteins were found in this Neurospora genome to have recognizable PAS domains by this analysis with high similarity. They were WC-1 and WC-2. The BLAST values were 2e−13(199) and 2e−07(57), respectively. The Neurospora genome was also screened by employing the N. crassa WC-1 as a query. A third PAS-containing protein was found, that is, the VVD protein, BLAST value e−42(147). However, the similarity to VVD was found to be only for the light, oxygen, voltage (LOV) domain not the PAS domains. A few other PAS-containing proteins were found with low levels of similarity, such as a protein kinase or a histidine kinase, but a protein with high similarity to ARNT was not found. An additional screen was performed, employing the Drosophila PER. No additional proteins containing the PAS domains were found beside those mentioned above. Several other PAS-containing proteins have been found in the Neurospora genome and are detailed in a review (Borkovich et al., 2004). It is interesting that of the few PAS-containing proteins in Neurospora, 3 of them play a role in the conidiation rhythm (Lakin-Thomas et al., 2011).
Additional studies were performed to follow up on the PAS matchup between the Neurospora WC-1 and the Capsaspora 848 (ARNT). The presence of other PAS-containing proteins in organisms such as Chlamydomonas and Ostreococcus was examined further. The Chlamydomonas genome contains a light receptor–type protein that has a high similarity to the Neurospora WC-1, BLAST value e−26(113). However, this protein showed low similarity to the Drosophila PER. Likewise, a protein found in Ostreococcus, designated as a histidine kinase, showed some similarity to WC-1, BLAST value e−18(120), but not to the Drosophila PER. These findings do not rule out these proteins as possible precursors to Drosophila PER, since there could have been many intervening steps between them. But it does enhance the possibility that the ARNT from Capsaspora, or a close relative, could be a common ancestor. It should be kept in mind that the PAS motifs that match between WC-1 and ARNT are complex; that is, there are 3 separate regions, originally designated in the Drosophila PER as PAS-A, PAS-B, and CLD. It is not clear if all 3 of these motifs have a common ancestor or if each one comes from a different eukaryotic gene or even a bacterial gene. In addition, there are many species that contain proteins with PAS domains. Capsaspora is of particular interest since it has been proposed to be one of the organisms that might be a common ancestor to fungi and animals. A good example of this (Pare et al., 2012) is a transcription factor found in Capsaspora (XP_004347648) that shows a BLAST value of e−23(232) versus a Neurospora transcription factor (CSP-2) and a BLAST value versus a Drosophila transcription factor (grainyhead) of e−42(194).
A screen of higher organisms, such as mice or Arabidopsis, indicated that the PAS domains are easily found in mouse proteins but only in a few plant proteins. A mouse ARNT-like protein, BMAL1, shows strong similarity to the Drosophila PER, e−27(312) but less so to the Neurospora WC-1, BLAST value e−8(59), and another section, e−4(215).
The LOV Motif/WC-1
The LOV domain can be widely found (i.e., in bacteria, plants, microbes, animals, etc.; Vogt and Schippers, 2015). The Neurospora WC-1 shows similarity to many of these LOV-containing proteins. Several examples are (1) the similarity to a Synechococcus response regulator, WP Synechococcus _011242514, BLAST e−17(145); (2) the Halobacteria bat protein, (167727267), BLAST value e−18(128); (3) the ZTL protein of Arabidopsis (81170304), BLAST value of e−23(113); and (4) a photoreceptor found in Chlamydomonas (XP_001693387), BLAST value of e−26(113). These examples appear to be proteins involved in light responses. Since there are so many LOV-containing proteins, it is difficult to cite a common ancestor for the LOV domain found in WC-1 and ZTL. But it is still interesting that these proteins share a common domain, as pointed out previously by others (Cheng et al., 2003). This would suggest that at least for the LOV domain, and for light sensing, that Neurospora and Arabidopsis share a common ancestor. This is of some interest in determining whether clocks as we know them now evolved separately or from a common origin. Obviously, more examples will be required on this point.
In N. crassa, a scan of the genome with ZTL showed that the 3 proteins with the highest similarity were WC-1, VVD, and a kelch domain–containing protein (GI# XP_965023). This third protein showed similarity in a domain different from the LOV domain. Thus, the LOV domain appears to be mostly confined in Neurospora to proteins involved with light responses. Both the LOV domain found in Neurospora WC-1 and the LOV domain found in Arabidopsis ZTL are central to their respective clock machinery. This is evidenced by the fact that mutations in or near the LOV domain as well as other mutations of Arabidopsis ZTL lead to altered periods (Somers et al., 2000) while deletions of WC-1 in Neurospora lead to arrhythmic cultures (Crosthwaite et al., 1997).
Comparison of Positive Elements
The Drosophila CYC protein is a positive element in its clock system. Its sequence has been known for quite some time and consists of a helix-loop-helix (HLH) section and PAS sections (Genebank GI# 24667005). The Drosophila CYC was then compared with a protein that is a positive element in the Neurospora clock system (i.e., the WC-1 protein). A low level of similarity was found, BLAST value of 8e−06(96), but only in 1 of the PAS sections and not in the HLH section. In Figure 8, the Neurospora WC-1 protein showed some similarity to the Capsaspora ARNT protein. The BLAST value was e−13(199), but again, the similarity was only in the PAS sections. Therefore, it was of interest to determine if the Drosophila CYC protein also showed some similarity to the Capsaspora ARNT protein. The results are that these 2 proteins showed similarity in 2 regions: the HLH region, BLAST value of 3e−11(70), and the PAS sections, BLAST value of 1e−16(310). These 2 sections comprise about 90% of the CYC protein. Comparison of these 2 proteins via CLUSTAL confirmed these findings.
In the mouse clock system, one of the positive elements is the BMAL1 protein. A comparison of that protein with the Capsaspora ARNT showed a BLAST value of 7e−23(312) for the PAS sections and a value of 5e−12(61) for the HLH section. From these results, it can be suggested that the Capsaspora ARNT would be a plausible precursor to all three (i.e., the CYC protein of Drosophila, the WC-1 protein of Neurospora, and the BMAL1 protein in the mouse). Obviously, other proteins and other organisms could be considered for this role. However, a search of the genomes of the 6 other possible fungal ancestors (James et al., 2006) employing CYC as a query did not show any proteins that had significant similarities to CYC. Therefore, it would appear that amongst the existing genomes known so far, the Capsaspora ARNT protein should be considered as the best candidate for a common ancestor to the positive elements in the Neurospora, Drosophila, and mouse clock systems.
Discussion
The FRQ/PER matched sections shown in Figure 3 can be classified in several ways. One, the sections G, H, I, and J of both FRQ and PER appear to have the Capsaspora ARNT in common as seen in Supplementary Figure S2. These 4 sections of FRQ were previously shown to have similarities to ARNT (figure 10 in Brody, 2019). Two, the FRQ sections A and C in Figure 3 show similarities to the Acanthamoeba HSF (figure 9 in Brody 2019), but no clear similarities could be found to PER. Three, some of the sections (B, D, and F) in Figure 3 have no obvious precursor found so far for either protein. Four, in Figure 3, the initial 10 amino acids of section E for FRQ shows similarities to the Capsaspora LARP as shown (figure 7 in Brody, 2019) but shows limited similarities to PER. Five, for section K in Figure 3, the last 22 amino acids in PER as listed in Figure 3, in addition to showing some similarity to FRQ, also show some similarity to the HSF as seen in Supplementary Figure S4.
In the Results section above (a comparison of positive elements), a potential common ancestor to the Neurospora WC-1, the Drosophila CYC, and the mouse BMAL1 proteins is suggested to be the Capsaspora ARNT protein. This ARNT protein also shows similarities to the Drosophila PER, as shown in Supplementary Figure S2. This is another indication of the significance of this amoeba as a branch point in the evolution of fungi from insects/animals and has not been pointed out so far in the literature on the evolution of circadian clocks.
It is interesting that some similarities can be found between the fungal clock system and the Drosophila clock system in terms of the positive elements (WC-1 and CYC) and for the negative elements (FRQ and PER). Similarities between the modulating elements (protein kinases, etc.) have been pointed out previously by many, and some of these are listed in the introduction section to this article.
Where do the new “functions” of a circadian system that are not found in precursor organisms come from? On a mechanistic level, this means how do new genes/proteins arise? The mechanisms for the appearance of a new gene/protein, which gives rise to a new function, have been cited many times and appear in genetics textbooks. Intron conversion and horizontal gene transfer are two of the mechanisms often cited. Perhaps the most interesting is the idea of duplication/divergence? If a second “copy” of a gene mutated, a new function could then arise while preserving the function of the first copy. A more detailed discussion can be found in Snustad and Simmons (2009).
Gene duplication can occur by many mechanisms and can be found in many genetic textbooks (see Snustad and Simmons, 2009). A common mechanism is via a crossover event within a pericentric (including the centromere) inversion. This allows each chromatid to have a centromere, one chromatid to have some duplicated material, and one chromatid that is deficient in some genetic material. Subsequent “ rounds” of mutation in the second copy of a gene can then produce a new function. This altered new copy will usually still retain significant sequence similarity to the original copy.
How might a new gene be formed from pieces of several genes? Again, there are several possible mechanisms. A widely discussed one is the idea of some type of transposon element. Another mechanism, the more classical one, is the idea that chromosomal translocations are the sources of new genes. There are many types of chromosomal translocations. A common type is the insertional type (i.e., in which a piece of one chromosome is inserted into a different chromosome). A second type is the reciprocal type, in which chromosomal arms are swapped and new linkage groups are formed. A third type is intrachromosomal, in which one part of a chromosome becomes inserted elsewhere in the same chromosome. A more detailed discussion can be found in Snustad and Simmons (2009). The ability to form new genes depends on where the breakpoints are in the chromosome. Breakpoints within genes, followed by recombination of the parts, can lead to some new types of hybrid genes. It is likely that a random process such as this would not lead to genes with useful new functions. However, a small subset of these hybrid genes could be new genes with a useful function. It is not likely to be known what the probability of each one of these events (translocations, recombination, formation of a productive recombinant) was. But it is likely to be low, and the overall probability of all of these events occurring one after the other is likely to be very low. In any event, a new hybrid gene would then have only partial sequence similarity to any one of its precursor genes.
As pointed out in the Results section, FRQ and PER appear to be a composite of elements found in other proteins. One could account for these data by proposing that the ARNT gene recombined with the genes for LARP and HSF to give FRQ. In the case of PER, perhaps the ARNT gene recombined with the gene for HSF to give most of PER. Clearly the possible derivation of some of the other regions of FRQ and PER is still unaccounted for. It should also be pointed out that the recombination events do not have to have the same topography in every organism (i.e., the N-terminal region of one protein could end up being the C-terminal region of another protein).
It is interesting that the regions of FRQ that show some similarity to other proteins are all regions of these other proteins that are highly conserved in these other proteins and have functional significance. For example, the pertinent region of HSF that shows some similarity to FRQ is part of the DNA-binding region of HSF (NCBI Genebank pfam Cl12113). The pertinent region of LARP is part of the active site of the RNA-binding region of LARP (Bousquet-Antonelli and Deragon, 2009). The pertinent region of ARNT is part of a PAS domain, known to bind proteins or other molecules. Likewise, several of the regions of PER that show similarity to other proteins are regions that are highly conserved and have well-defined functions in these other proteins, such as ARNT and HSF. There is a precedent for clock proteins to have been derived from proteins with another function. This is the case of a key clock protein in plants, TOC-1, for which the evidence strongly suggests a derivation from response-regulator proteins (Strayer et al., 2000).
Independent Evolution versus Common Ancestor?
There are several possibilities relevant to this question. One, a prototype clock protein was formed in a single-celled organism (amoeba), and after much shuffling of motifs and so forth, it subsequently evolved into FRQ. This original prototype protein was similar enough to PER that it required only some minor subsequent modifications to form the current Drosophila PER. This scenario would require only one major further type of event (i.e., the formation of FRQ). But it would fit with the common ancestor hypothesis.
A second possibility would entail an early clock protein that required 2 independent shuffling of motifs: one to form FRQ and one to form PER. This would be a modified common ancestor idea.
A third possibility would be some type of convergent evolution, in which key clock genes were independently formed from pieces of other genes but without a common ancestor. It is not clear what the mechanism for this would be, and there is no evidence in these data for such a possibility. Many independent events would have had to occur.
The data presented here cannot clearly distinguish these 3 possibilities. However, it can be stated that the evidence favoring the common ancestor idea has these 2 parts: (1) that these 2 clock genes in fungi and Drosophila may have both originated from genes in those single-celled amoebae that are precursors to both fungi and insects (i.e., the base of phylogenetic tree), and (2) it is also proposed here that both PER and FRQ arose from the same parts of other genes, such as the ARNT and HSF genes.
Among the evidence favoring the independent evolution side is the observation that the component pieces of the PER gene are not in the same order as the pieces of the FRQ gene. This would suggest independent events for the formation of each.
It is possible that some might find the data presented here less than compelling for the common ancestor idea, or for the hybrid theory, and stick with the independent evolution idea or be agnostic for the entire discussion. However, perhaps these data tip the scales somewhat in favor of a common ancestor between fungi and insects? Clearly, additional evidence is required. Even then, there is still an open question about the relationship between the fungal/insect clock systems with that of other eukaryotes, such as those found in higher plants (Arabidopsis) and lower photosynthetic organisms (Ostreococcus, Chlamydomonas). An even bigger question exists as to possible connections to prokaryotic clocks (Synechococcus).
Supplemental Material
A_comparison_of_Neurospora_and_Drosophila_Supplementary_material – Supplemental material for A Comparison of the Neurospora and Drosophila Clocks
Supplemental material, A_comparison_of_Neurospora_and_Drosophila_Supplementary_material for A Comparison of the Neurospora and Drosophila Clocks by Stuart Brody in Journal of Biological Rhythms
Footnotes
Acknowledgements
The author thanks the following individuals for assistance, encouragement, and helpful criticism: Barry Grant, W. McGinnis, and Chris Wills of the University of California, San Diego; Carrie Partch of the University of California, Santa Cruz; Jason Stajich of the University of California, Riverside; Joanna Chiu of the University of California, Davis; and Mike Young of Rockefeller University.
Conflict Of Interest Statement
The author has no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
