Abstract
The fungal clock, especially that in Neurospora crassa, is composed of several proteins, notably FRQ, WC-1, and WC-2, which interact at the protein level and at the level of transcription. It is shown here that regions of the FRQ that are highly conserved in many fungal species show significant similarity to regions of proteins found in the amoebae Capsaspora and Acanthamoebae. These 2 amoebae were specifically explored because they have been suggested, based on extensive evidence, to be related to precursors of the modern fungi. Those proteins in Capsaspora/Acanthamoebae with some similarity to FRQ are LARP (an RNA-binding protein), ARNT (which has a PAS motif), and heat shock factor (HSF). These regions of LARP and HSF that show similarity to FRQ are highly conserved between plants, animals, and amoeba. This suggests that these regions were present at the time of the divergence of plants, fungi, insects, and animals, and therefore, they could be plausible precursors to regions of the fungal FRQ. These particular regions of FRQ that show similarity to LARP and HSF are also of functional significance since mutations in these regions of the Neurospora FRQ led to changes in the rhythm. The FRQ proteins from 13 different species of fungi were analyzed via motif analysis (MEME), and 11 different motifs were found. This provides some understanding as to the minimum requirements for an FRQ protein. Many of these FRQ motifs can be matched up with known domains in FRQ. In addition, these 13 different species of fungi were screened for the presence/absence of 7 additional genes/proteins that play some role in fungal clocks.
Keywords
Circadian rhythms have played a central and important role in the life of many of the known organisms. It is a widespread feature of almost all of the known eukaryotic organisms and endows them with a type of a clock mechanism that allows them to anticipate the earth’s daily light/dark cycle. An excellent textbook by Dunlap et al. (2004) relates these features. Even some bacteria, particularly the cyanobacteria, have a circadian mechanism (Mackey et al., 2011). All of the circadian rhythms have in common several elements: (1) a period of approximately a day (circadian); (2) a sensitivity to light and/or temperature changes that can shift the phase of the rhythm; (3) a mechanism for maintaining the period at widely different temperatures, termed temperature compensation; and (4) a mechanism that is inherited (i.e., gene based).
The purpose of this communication is 4-fold: (1) to serve as a partial update on certain aspects of this field; (2) to take a closer, detailed look at the structure/function relationship of some of the important clock proteins; (3) to reexamine the phylogeny of clocks focusing on known protein motifs and their structure; and (4) to introduce some new facts/ideas about the evolution of some of the important clock components found in fungi and their possible precursors.
Previous studies on fungal clock phylogeny have indicated that within the Sordariomycetes, the FRQ proteins were similar enough that the Sordaria frq gene was able to successfully restore the rhythm to a Neurospora FRQ-less mutant (Merrow and Dunlap, 1994). Subsequent work on clock phylogeny indicated that the presence of the FRQ protein was confined to 3 classes of fungi: the Sordariomycetes, the Leotiomycetes, and the Dothideomycetes (Salichos and Rokas, 2010). In that work, it was of interest that many species of fungi did not have the FRQ protein, despite some reports of a rhythm (Greene et al., 2003).
No previous studies were done to look for precursors to FRQ among single-celled organisms, such as Capsaspora or Acanthamoebae. These organisms, and others, were investigated because they have been postulated to be related to the ancestors of modern-day fungi (James et al., 2006). With these observations in mind, it was decided to take a more detailed look for potential precursors to just the important parts of FRQ (i.e., some of the motifs).
Materials And Methods
The standard BLASTP program was employed with the default parameters for the gap/extension penalties of 11,1, except where noted. The NCBI GenBank was the source for sequences, and Firefox was employed as the server. The Clustal Omega program supported by EMBL was employed. The website MEME-suite.org contained the motif discovery program (MEME) version 5.0.2.
Fungal FRQ(s) employed for comparative analysis (MEME): Neurospora crassa 595952; Sordaria fimicola 6016053; Fusarium oxysporum 342889466; Botrytis cinerea 347827659; Colletotrichum sublineola 640918779; Trichoderma virens 358386587; Acremonium chrysogenum 672797659; Sclerotonia sclerotiorum 156039415; Marssonina brunnea 597583253; Aureobasidium pullulans 662524541; Phaeosphaeria nodorum 169600449; Rhodotorula toruloides NP11 EMS19378; Taloromyces st. 242790999; Saitotella complicata 813215842; Magnaporthe oryzae 145610785; Zymosepteria tritica SMR54202; Pyronema tritica 189211261; Rhizophagus irregularis 595497380.
Table listing the comparison of 4 LARP(s): Acanthamoeba castellanii str. Neff 470512733; Capsaspora owczarzaki ATCC 30864 470292891; Arabidopsis thaliana 18420415; Danio rerio 326677994.
LARP(s) employed for the construction of LARP MEME(s): Branchiostoma floridae 260833404; Danio rerio 326677994; Drosophila pseudoobscura 198451090; Capsaspora owczarzaki 470292891; Amphimedon queenslandia 340381196; Ostreococcus tauri 308813524; Arabidopsis thaliana 18420415; Sordaria fimicola 336272473.
Others: Capsaspora owczarzaki ARNT 754351513; heat shock factor(s) (HSF[s]) for Acanthamoeba castellanii 470459745; for Danio rerio NP_001315137; for Arabidopsis thaliana 110738569. For the table listing the presence of clock components, the full species names and isolate numbers are: Sordaria macrospora K-hell; Leptospheria maculans JN3; Trichoderma virens GV29-8; Fusarium oxysporum f.sp.pisi HDV 247; Botrytis cinerea BcDW9?; Zymosepteria tritica IPO323; Magnaporthe oryzae Y34; Pyronema omph CBS100304; Saitoella complicata NRRL Y-17804; Talaromyces stipulatis ATCC 10500; Rhodotorula toruloides NP11; Rhizophagus irregularus DAOM 197198w; Aspergillus nidulans FGSC A4; Dimargaris RKP36089; Umbilicaria pustulata SLM37164; Drechselerella EWC45593.
The GI numbers that were employed for the queries were for Neurospora crassa: FRQ 595952; WC-1 5441498; WC-2 1835159; FRH 57019005; prd-1 91206541; prd-3 553139259; prd-4 107770479; prd-6 85101908.
Results
Database Screening
Three different set of analyses were performed: (1) screening of other fungi for FRQ homologues, (2) screening of certain single-celled organisms for possible precursors to clock proteins, and (3) screening for the presence of clock components in many different fungi.
Screening of Other Fungi
N. crassa FRQ was employed as the query, and the taxid FUNGI (4751) was the subject. Standard default conditions were employed for the BLAST analysis, except that the word size was 3 and the maximum target was 500. The similarities to the Neurospora FRQ ranged from 0.0 (Sordaria) along almost the entire length of the protein to just parts of the Neurospora FRQ. An example of the latter is the protein from Saitoella c., for which only 2 sections showed similarity, 2e−15(53) and 2e−6(76), where the numbers in parentheses indicate the lengths of the similarities.
Previous studies (Salichos and Rokas, 2010) indicated that FRQ was primarily found in the ascomycetes, and the authors postulated that the FRQ protein arose sometime after the divergence of the fungi into separate groups, the Sordariomycetes, Leotiomycetes, and Dothideomycetes. The results reported here are an update on those findings and indicate that FRQ can be found earlier in the phylogenetic diagram (i.e., in some Basidiomycota, Zoopagomycota, and Mucoromycotina; Fig. 1). A partial analysis of Basidiomycota is shown in Figure 2, and a partial analysis of the Ascomycetes is shown in Figure 3. Several FRQ-containing species have been added (i.e., Talaromyces, Saitoella, Rhizophagus, and Rhodotorula, as well as Dimargaris, Umbilicaria, and Drechselerella). The figures below incorporate the data from the excellent review of Montenegro-Montero et al. (2015) but in an expanded phylogenetic chart.
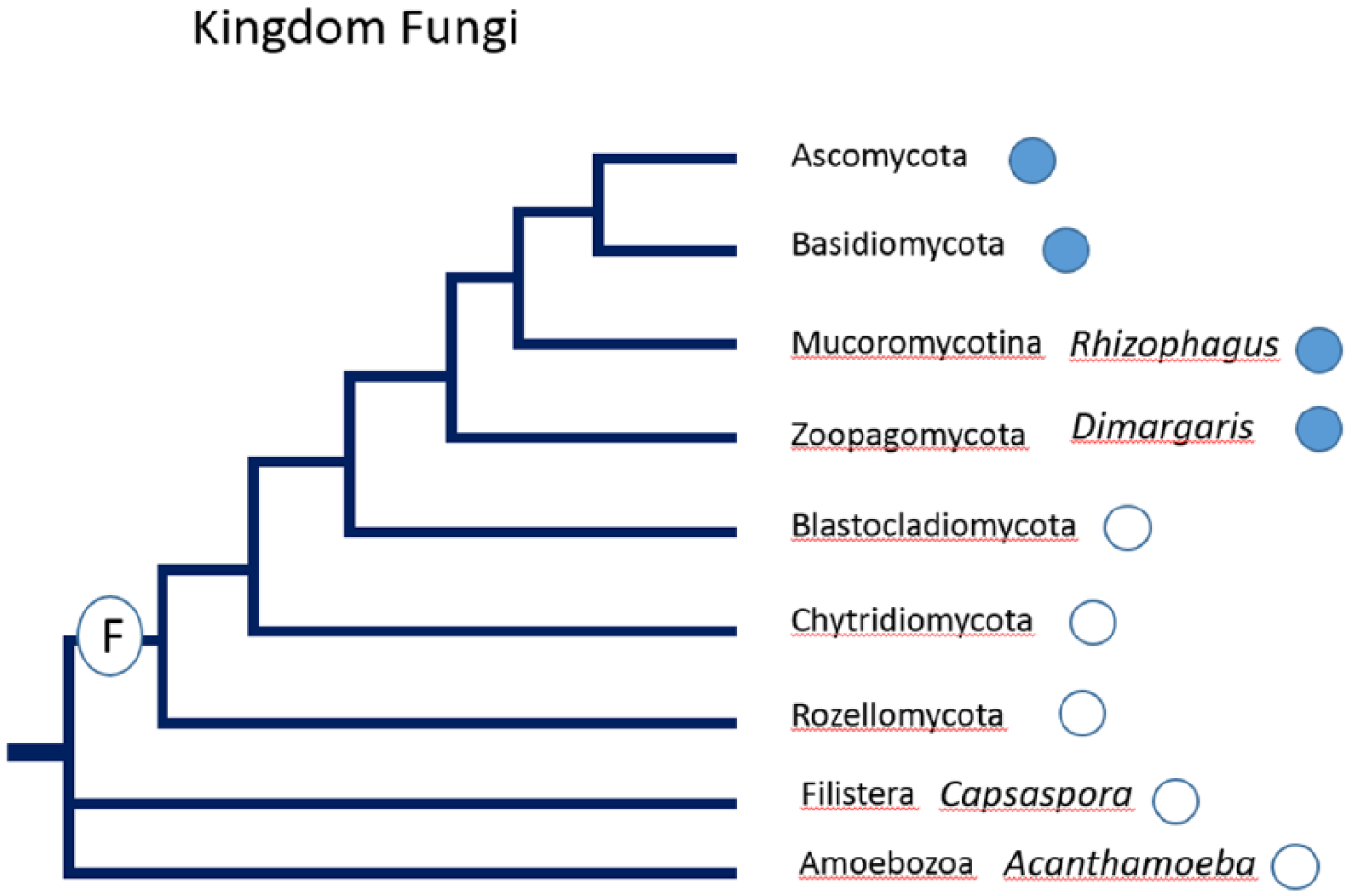

A phylogenetic diagram of the partial kingdom of fungi. Filled circles indicate the presence of FRQ; empty circles indicate the absence of FRQ. The circled “F” indicates the postulated time of the formation of fungi.

A schematic phylogenetic diagram of the Basidiomycetes. Filled circles indicate the presence of FRQ; empty circles indicate the absence of FRQ.
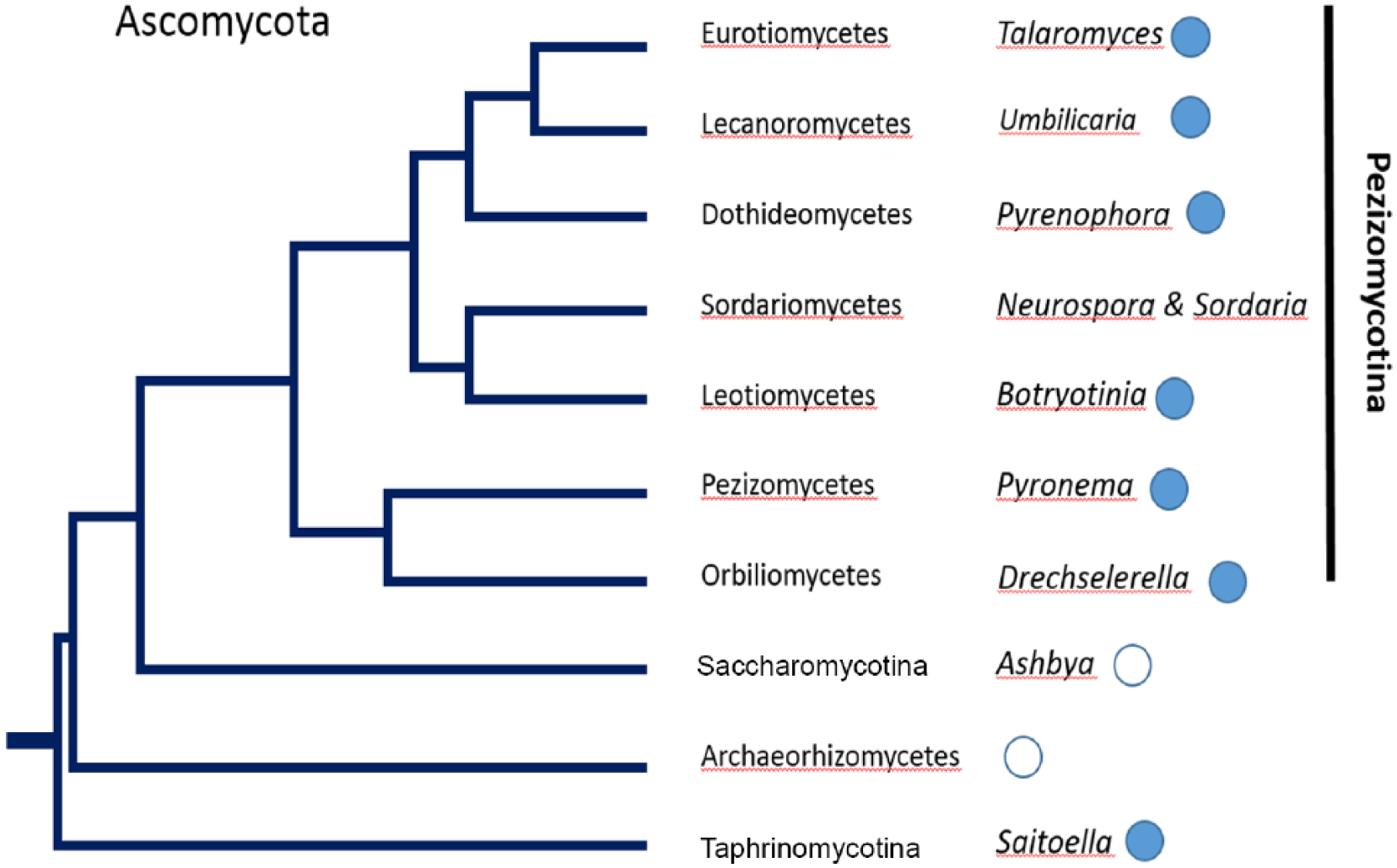
The finding of an FRQ or FRQ-like protein in Lecanoromycetes raises an interesting question, since this is the group of fungi that form lichens with other organisms, such as the cyanobacteria. Since certain cyanobacteria, such as Synechococcus, have been found to have a circadian clock mechanism, it raises the question of whether lichens have a clock and how and if the clock of the fungal partner interacts with the clock of the cyanobacterial partner.
Not all groups of fungi have FRQ. This is a tentative conclusion, since not all members of any given group have been tested. Even within a given FRQ-containing group, not all members of that group contain FRQ—only some species. The factors that determine the absence or presence of FRQ are not clear. It could be an ecological niche or a mode of propagation or some other factors. Further studies on the properties of these many organisms, such as done by Liu and Hall (2004), would be useful.
The FRQ protein is not found in all of the fungi (Salichos and Rokas, 2010; Montenegro-Montero, 2015; and the current article). It is not found even in species such as Aspergillus, which was shown to have some type of rhythm (Greene et al., 2003). The absence of FRQ in species such as Aspergillus could have several explanations: (1) there is no clock in this organism, which seems unlikely; (2) the clock employs a different type of oscillator; (3) the mechanism for responding to the environment is an “hourglass” mechanism, instead of an anticipatory mechanism; or (4) the FRQ is so altered that it cannot be easily found by the standard techniques employed. In that regard, this analysis was made a little more detailed by employing the 3 motifs (see Table 1) that are found in all FRQ(s) as queries versus the Aspergillus genome. However, no obvious FRQ-like protein was found. The general question of why some organisms in a given group have a certain protein while other closely related organisms in the same group do not have that protein is often debated. The general consensus is that there is selective loss of a gene in an organism in a given group rather than a gain of function for other organisms, which would require many events.
Distribution of motifs in different FRQ(s).
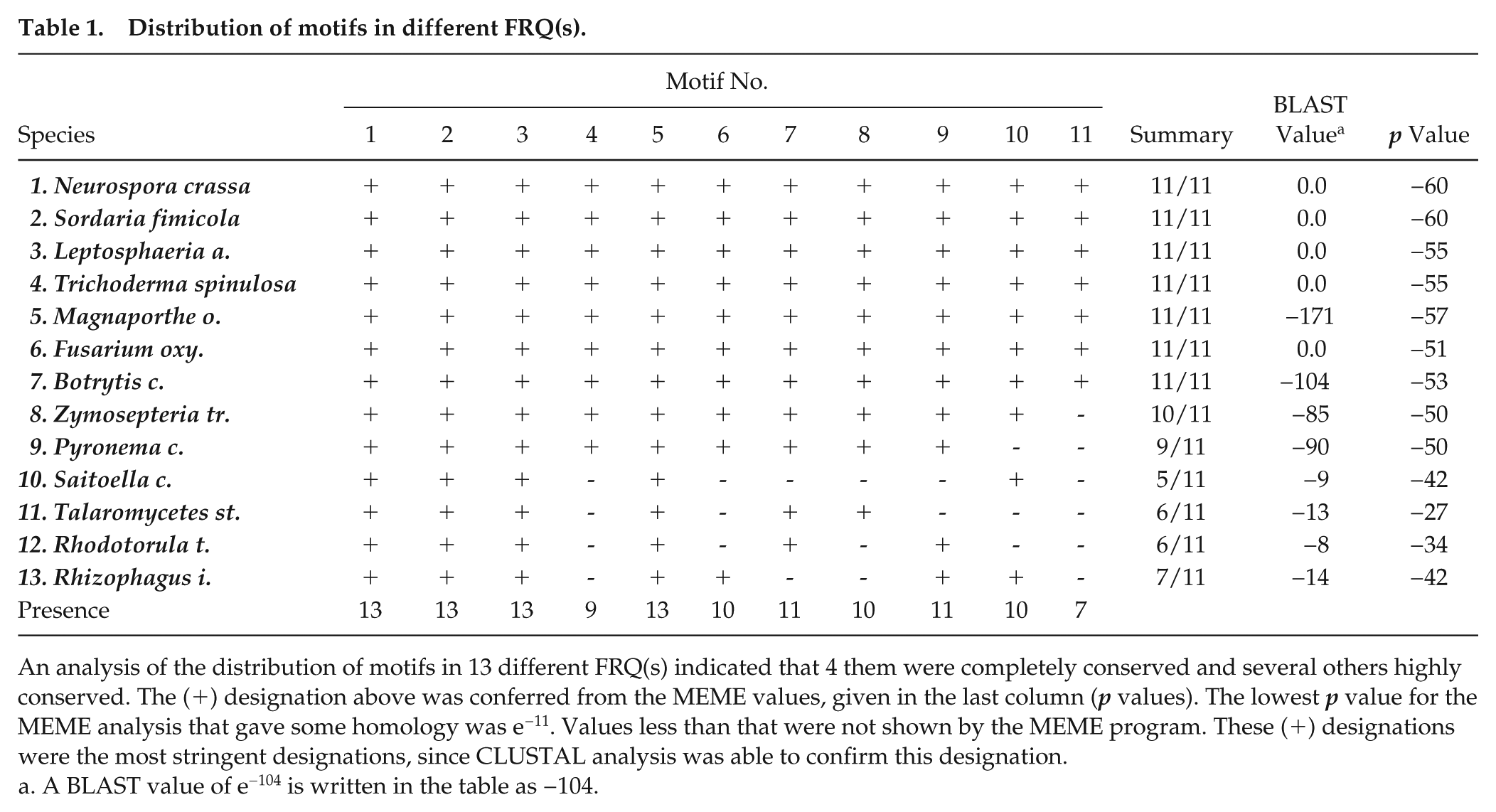
An analysis of the distribution of motifs in 13 different FRQ(s) indicated that 4 them were completely conserved and several others highly conserved. The (+) designation above was conferred from the MEME values, given in the last column (p values). The lowest p value for the MEME analysis that gave some homology was e−11. Values less than that were not shown by the MEME program. These (+) designations were the most stringent designations, since CLUSTAL analysis was able to confirm this designation.
A BLAST value of e−104 is written in the table as −104.
The photoreceptor proteins, WC-1 and WC-2, were more widely found than FRQ (Salichos and Rokas, 2010). Certain clock-controlled proteins, such as CCG-8, have also been found in almost all of the fungi analyzed so far (Lombardi and Brody, 2005).
Motif Analysis
MEME analysis was employed for FRQ(s) from different species of fungi since it has the potential to provide more information than just the BLAST analysis. It enables one to see which sections (motifs) of a sequence are important to the function of a protein and which sections may be less so in distantly related species. For instance, it might be able to link up an FRQ motif with a function such as an interaction with another protein. A similar line of analysis has been done for short linear motifs (Davey et al., 2015). Motif analysis could also provide information as to whether certain species acquired new functions in a protein during evolution (or lost functions). A study similar to this was done by Montenegro-Montero et al. (2015) by aligning the FRQ(s) from many ascomycetes by CLUSTAL. This study widens the analysis in 2 ways: by including more distantly related species, such as Rhodotorula and Rhizophagus, and by employing MEME.
The FRQ(s) from N. crassa and 12 others (listed in Table 1) were analyzed by this procedure, and 11 different motifs were found.
The motifs derived from these 13 FRQ(s) are listed in Supplementary Table S1 and are designated as FF-. Numbering is from the Sordaria FRQ sequence.
The results for just the FF-1 motif are diagrammed below.

The full height of a letter indicates that this amino acid, for example, W and N and others shown above, are the consensus amino acids at that position. The height of the other letters, for example, E, D, and H above, indicate the frequency of those amino acids at that one position.
These e values were obtained by a scan of “fungi” with N. crassa FRQ as the query. These e values are derived from a comparison of the entire Neurospora FRQ sequence versus the genome of another species. In some cases (i.e., the FRQ from Neurospora v. that of Rhizophagus), an individual comparison of just those 2 proteins gave greater similarity and indicated 2 areas of similarity e−23(213) and e−20(179).
Additional areas of similarity did show up when analyzed via CLUSTAL in that the motifs in the FRQ from one species aligned with the motifs from the FRQ from another species. The spacing between the motifs varied from one FRQ to another, but the order of the motifs was similar. The Saitoella FRQ is a good case in point, in which 3 additional motifs were found (numbers 3, 5, and 10) that were not found by MEME. In some cases (i.e., motif 9), the motif was so short that a (-) designation has less meaning and is not a definitive negative. In the cases in which there are (-) designations, these FRQ(s) were checked against many other FRQ(s) as well as the Neurospora and Sordaria FRQ(s), and no homology could be found. The FRQ(s) that were from species more distantly related to the Ascomycetes (Neurospora and Sordaria) are the last 4 in the list above. Not surprisingly, they showed the least number (5, 6, 6, 7) out of the 11 motifs found in the FRQ(s) of Neurospora/Sordaria. Certain motifs found in the FRQ(s) from the Ascomycetes have not been found in these 4 distantly related species. These are FF-4 and FF-11. It is not clearly known if the FRQ(s) without these motifs represent a primitive type of FRQ or a “streamlined” type that has lost some of these motifs. The FRQ from Rhizophagus is considerably shorter (572 amino acids) than most of the other FRQ(s), which are 900 to 1100 amino acids in length. This could account for the reason why this FRQ is missing 2 of the motifs located in the C-terminal area of FRQ. It is also possible that the genome scan did not pick up the complete sequence of this gene.
The FRQ(s) that appear to be missing some of the motifs found in the Neurospora FRQ bring up the question of what are the minimum protein structural requirements needed to operate effectively in the clock machinery. By analogy to the Neurospora FRQ, there would be (1) binding sites in the FRQ/WC protein complex for kinases; (2) binding sites for the WC proteins; (3) binding sites for the FRH protein; (4) binding sites for the protein degradation system (FWD), which would include PEST sequences; (5) a dimerization motif; (6) a nuclear localization sequence; and (7) a nuclear export signal (NES).
In addition, key serine residues and their surrounding sequences would probably be required. Thus, there would be a minimum of 7 or 8 structural areas of FRQ necessary to function in the clock circuit. Figure 4 shows many of these domains and motifs. It is not known how the circuits or the FRQ(s) found in distantly related fungal species, such as Rhodotorula, function. But it is interesting that some of the motifs found in Neurospora and Sordaria, such as the PEST motif or FRH binding, cannot be easily detected in them.
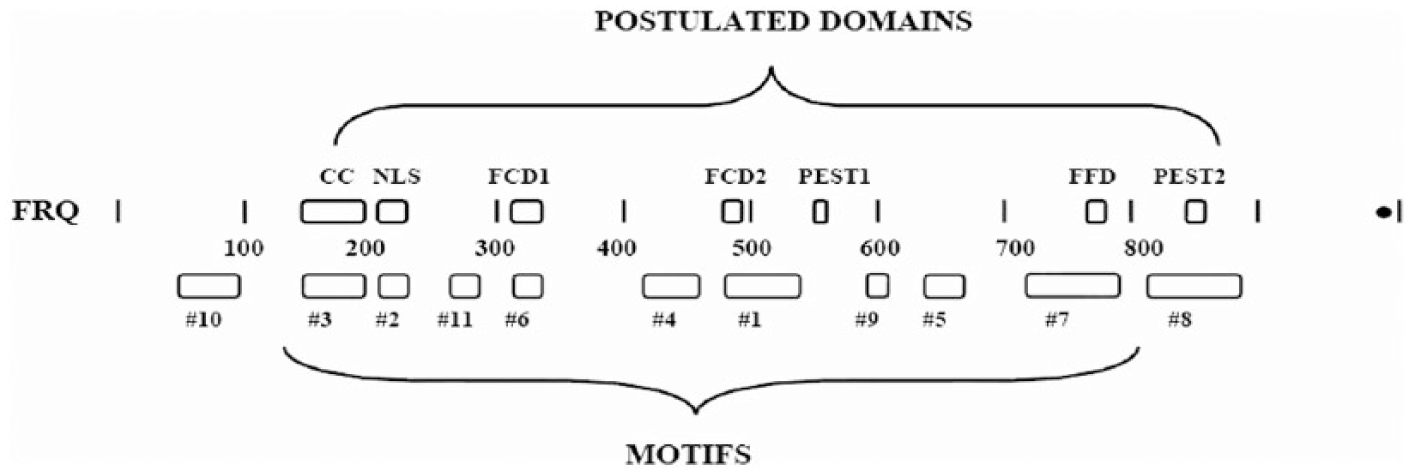
Schematic diagram of the 11 known motifs versus the postulated 7 domains in the Neurospora FRQ. Of the 7 domains listed, 6 match up to a motif. However, not all of the motifs have been matched up to a domain. Numbers 4, 5, 9, 10, and 11 are still to be explored. Domain CC = coiled/coil; NLS = nuclear localization signal; FCD1 = Frq casein kinase binding domain 1; FCD2 = domain 2; FFD = Frq Frh binding domain.
For the FRQ(s) more distantly related to the Neurospora FRQ (i.e., the last 4 in Table 1), the question becomes, are these FRQ(s) or FRQ-like?
Testable idea: If, in those fungi, it would be possible to set up a means of visualizing a putative clock, either by spore formation or a luciferase assay, this could be tested by mutation of these FRQ-like genes.
Another testable idea: Those organisms containing FRQ(s) that are missing in 1 or more motifs could be analyzed for many of the canonical properties of the Transcription Translational (TTL) oscillator (i.e., temperature compensation, light entrainment, etc.) to determine which, if any, properties are missing in these FRQ(s).
The Key Parts of the Neurospora FRQ Are Relatively Unique in the Neurospora Proteome
The N. crassa genome was screened employing these 11 motifs, and 3 items were of interest: (1) as expected, most of the motifs found FRQ with high similarity; (2) most of the motifs were not found in other proteins, suggesting that they were relatively unique to FRQ, with some exceptions as noted in the text; and (3) 2 motifs were of interest, that is, motif FF-1, which found the Neurospora LARP as expected, and motif FF-8, which found (with low similarity) an autophagy-related protein (EAA27491). This area of potential similarity is highly conserved in FRQ(s), and it is of interest that this autophagy-related protein has 8 of the 10 conserved FRQ residues. This area of FRQ was previously designated, based on its amino acid composition, as a PEST sequence (Gorl et al., 2001), so it is not surprising that it might also be found in other proteins involved in protein turnover.
Examination of the 13 FRQ homologues by MEME analysis showed that there were at least 4 distinct areas of FRQ that were highly conserved in all species (Table 1). In addition, there were several other areas that were conserved in 11 of 13, 2 of which were conserved in 10 of 13 of the species.
Testable idea: Since some of the conserved regions discovered by MEME analysis do not yet have any function assigned to them, it suggests that there is still some “biology” to find in the FRQ protein.
A schematic diagram indicating the postulate domains and the known motifs is shown as Figure 4. The FRQ domains are listed in figure 4 of Dunlap and Loros (2017).
Of the 7 domains listed above, 6 match up to a motif. However, not all of the motifs have been matched up to a domain. Numbers 4, 5, 9, 10, and 11 are still to be explored.
Some of the FRQ(s), such as those from Rhodotorula and Rhizophagus, do not show a clear indication of a second start site for the translation of the protein. In Neurospora FRQ, this area is (HDESHI
Testable idea: To determine if the species that contain those FRQ(s) that do not appear to have this second start site have temperature compensation or if they do not use this mechanism for temperature compensation.
Screening of Single-celled Organisms for Possible Precursors to Clock Proteins
The Neurospora or Sordaria FRQ was employed as a query versus the genomes of other organisms. Of particular interest are single-celled protists, such as Capsaspora, Dictyostelium, Acanthamoeba, and so on. Some of these have been postulated to be closely related to the precursors of the modern fungi (James et al., 2006) but probably have diverged after the animal/fungal split.
The Capsaspora genome was scanned with FRQ, employing BLAST default values, except the word size was 3. The result was that there was 1 protein of similar size that showed some similarity. It was a protein identified as LARP, an RNA-binding protein. When the Acanthamoeba genome was scanned, the LARP and a heat-shock protein were found to have significant enough similarity to show up on an unbiased genome scan (see the section on HSF/FRQ). The area of similarity for LARP is shown in Figure 5. This BLAST value was for only the comparison of these 2 areas, not a comparison of the 2 entire sequences.

A highly conserved area of the Neurospora FRQ shows similarity to a highly conserved area of the Capsaspora LARP.
It is interesting that just past the end of the FRQ in this match is a section of the Neurospora FRQ, that is, GTKFSSDSSEDKSQQS, * *
which shows some biological significance. Mutations of those serines* to alanine gave rise to cultures with longer periods of 4-5 h (Tang et al., 2009).
To confirm this area of similarity, a reciprocal analysis was performed, that is, screening of a fungal genome (Sordaria f.) using the Capsaspora LARP as the query. It was found that FRQ could be found in Sordaria f., but there was little similarity to any other protein, other than LARP. Several other fungal genomes besides Neurospora and Sordaria gave a “hit” of FRQ when screened with Capsaspora LARP. These initial results were followed up by broadening the search by employing the LARP(s) from 3 organisms (Neurospora, Acanthamoeba, Chlamydomonas) versus many different FRQ(s), some more distantly related to Neurospora and Sordaria. FRQ was also used to scan Acetabularia and Euglena, but no useful information was obtained.
The results can be summarized as follows: all genomes containing FRQ show some similarity between a part of FRQ to a part of one of the LARP(s), some slightly stronger than the original screen (Sordaria FRQ).
Comparison of the fungal FRQ with more distantly related LARP(s), such as the LA-related proteins from Drosophila, Amphimedon, Arabidopsis, or Aplysia, showed much less homology than expected. Screening of archaea genomes with FRQ did not show any obvious precursors to FRQ—only some regions found in HSF. Further studies on the LARP(s) from single-celled organisms involved yeasts, algae, and so forth. No useful information was obtained from the studies of yeasts, while the eukaryotic red algae Cyanidioschyzon had a LARP that gave a BLAST value of 1e−07 (51) versus an FRQ from Fusarium.
No known LARP proteins have been found to date in prokaryotes or archaea (Bousquet-Antonelli and Deragon, 2009) suggesting that this class of proteins is related to RNA/nuclear functions. The area of match (DAEGW….) was found only in LARP and FRQ. Its role is unknown, except that part of it in LARP is involved in RNA binding. In FRQ, part of it has been implicated in protein kinase binding (Querfurth et al., 2011). Other possibilities are a possible role in the NES. A testable idea would be to test this area in FRQ for RNA binding and/or nuclear export. This area of LARP contains part of the RNA-binding sequence of LARP (Teplova et al., 2006; reviewed in Bayfield et al., 2010), but the key residues of the RNA-binding sequence do not match the FRQ sequence. Thus, one cannot conclude that FRQ binds RNA directly from these data. FRQ may have been derived in part from an RNA-binding protein at some point. In the course of evolution, another RNA-binding protein, FRH (which binds to FRQ), may have taken over this function, so that an FRQ/RNA complex may have evolved into an FRQ/FRH/RNA complex.
An analysis of the LARP(s) from many widely different organisms (Table 2) showed that this area of the RNA-binding protein is very highly conserved in plants, animals, and single-celled organisms. Therefore, this section of the LARP could have been present at a very early time prior to the evolution of fungi. A highly schematic/speculative diagram illustrates this in Figure 6.
Comparison of 4 LARP(s) and Sordaria f. FRQ.

A comparison of LARP(s) from 4 different organisms showed the conservation of the active site amino acids in all 4 organisms and how the Sordaria f. FRQ matched up to those conserved residues. The symbol “#” signifies active site residues, derived from crystallography studies of RNA binding to LARP (Teplova et al., 2006). Identities and positives are shown in bold. LARP alignment via CLUSTAL Omega. The motif for LARP is divided into 2 parts, since those are the parts that match up to the parts of FRQ.
Presence of some clock components in different fungi.
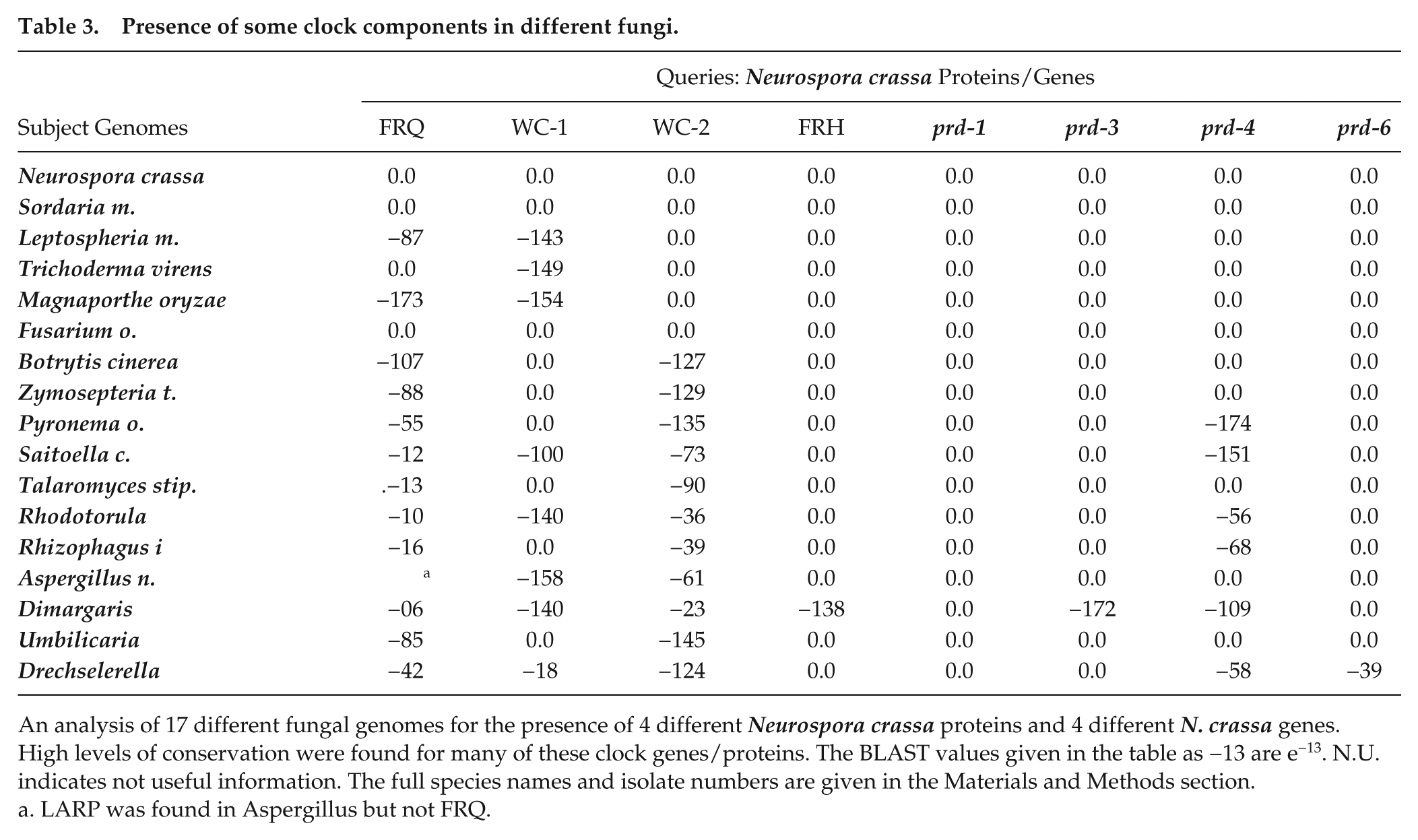
An analysis of 17 different fungal genomes for the presence of 4 different Neurospora crassa proteins and 4 different N. crassa genes. High levels of conservation were found for many of these clock genes/proteins. The BLAST values given in the table as −13 are e−13. N.U. indicates not useful information. The full species names and isolate numbers are given in the Materials and Methods section.
LARP was found in Aspergillus but not FRQ.
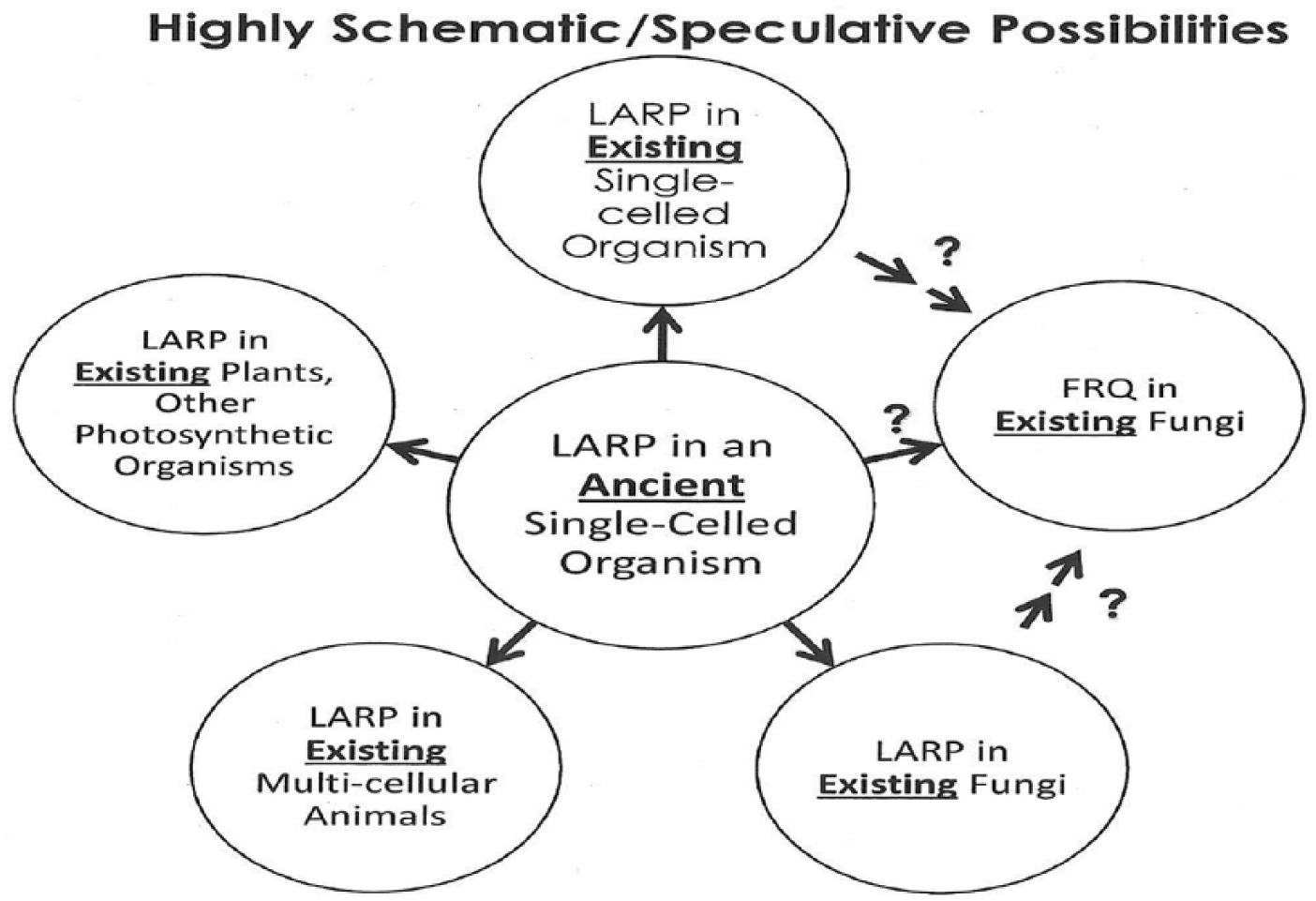
A highly schematic/speculative possible evolutionary relationship between LARP(s) and FRQ, indicating where parts of the existing areas of FRQ in fungi might have come from.
Alignment Analysis
CLUSTAL omega and BLAST were employed to determine if additional small area(s) of similarity exist between the Capsaspora LARP and the Sordaria or Neurospora FRQ. Analysis was done in both directions.
A few possible area(s) of low-level BLAST value were found, although these area(s) are not necessarily conserved in 1 or both of the sequences and are therefore not shown. One area was of interest and was highly conserved in both proteins. It is shown as section C in Figure 7. It was found by BLAST analysis and was a section of 72 amino acids, BLAST value of 1.2. Only part of that similarity is shown in Figure 7, and more discussion of this similarity is given under section C.

The Capsaspora LARP and Sordaria FRQ show 2 areas of similarity, both of which are highly conserved. The RNA-binding residues are shown as bolded amino acids. There are 10 known from the crystal structure (Teplova et al., 2006). Seven are in section C; 3 are in section A. The Sordaria FRQ protein shows some similarity only for 4 of the 10 amino acids needed for the RNA-binding found in LARP(s), while the Neurospora crassa FRQ shows similarity for 5 of the 10 amino acids. Neither protein is expected to show RNA binding.
A comparison of the LARP from Acanthamoeba to FRQ also showed several areas of low similarity. The FRQ from Sordaria fimicola or N. crassa was selected as the query for several reasons, including the BLAST value was 0.0 between these 2 proteins, and the Sordaria FRQ is similar enough to the Neurospora FRQ so that an FRQ deletion of Neurospora can be complemented by the FRQ gene from Sordaria (Merrow and Dunlap, 1994).
The parts of LARP and FRQ that show some homology to each other are special sections of each of these 2 proteins in several ways: (1) the 2 LARP sections are significant since they encompass the active sites of LARP; (2) these active site sections of LARP are highly conserved in plants, animals, amoebae, and so forth (see Table 2); (3) the corresponding FRQ sections are also highly conserved (see motifs 1 and 6); and (4) the 2 FRQ sections are of biological significance since mutations in either section affected the circadian rhythm of Neurospora.
Section A of FRQ
Section A is highly conserved, as seen in motif FF-1, shown in Supplementary Table S1. Section A is of biological significance for the following reasons: (1) It contains a key serine, designated with a
Section C of FRQ
Section C is also highly conserved, as seen in Figure 7, and it matches the motif FF-6, as listed in Supplementary Table S1 for FRQ(s). Section C is of functional significance in 2 ways: (1) part of the FRQ section C (i.e., VVRRLEQL) has been designated as FCD1 (FRQ-CK1 interaction domain) by Querfurth et al. (2011) and (2) a mutation of the L to N in the sequence RRLEQ led to an arrhythmic strain (Querfurth et al., 2011).
Only that part of the section C match that encompasses the RNA-binding residues is shown above. The actual complete match extends in both directions from that shown above. The complete section C sequences had a BLAST value of 1.2(72). Although there was a low BLAST value for section C between these 2 proteins, it is of some significance since these sections in both proteins are highly conserved and are of functional significance.
Sections A and C are contiguous and highly conserved in LARP(s) from Arabidopsis to humans, including amoeba such as Capsaspora (see Table 2). Sections A and C from FRQ(s) are also highly conserved in many fungal species, but they are separated by a stretch of 132 amino acids.
MEME analysis was also performed on many LARP proteins (see the Materials and Methods section), and the following was found, where the numbers refer to the Capsaspora LARP sequence (Suppl. Table S2). An analysis of just one of the common motifs found in the LARP(s) from widely different organisms when compared with the Sordaria FRQ is shown in Table 2.
For parts 1 and 2, Acanthamoeba LARP was compared with Sordaria FRQ via BLAST (FRQ was the subject). Part 2 had a BLAST value of e−5(59). The 4 LARP(s) had 32 identical or positive residues in common, only some of which are shown. The Sordaria f. FRQ sequence had 22 identities/positives to these 32 residues. Motif LA-3 is parts 1 and 2 combined.
Although the BLAST values are low for section C, both areas are highly conserved, as seen in Table 2. Both areas are of functional significance: section A in LARP encompasses the active site. Sections A and C in FRQ have been designated as kinase binding sites (Querfurth et al., 2011).
LARP/FRQ
The part (sections A and C ) of the Capsaspora LARP sequence (amino acids 403 to 480) shown in Table 2 that shows similarity to FRQ is interesting. That particular section of the LARP protein encompasses all of the currently known RNA-binding residues of LARP. Since FRQ is not known to bind RNA on its own, this similarity suggests that this part of the FRQ protein (amino acids 313 to 347 and 466 to 516) may have some different function other than RNA binding. It is proposed here that this part of FRQ may encompass a nuclear localization signal and possibly some other function, such as a site for protein kinase binding (Querfurth et al., 2011). A duplication of a LARP somewhere in the evolution of the fungi and a subsequent remodeling of this second copy to become FRQ could account for these data.
Nuclear Export Signal
A common NES in a protein is the sequence of LxxxLxxLxxL, where L can be replaced by other hydrophobic amino acids, and the spacing between the hydrophobic amino acids can also vary slightly. A good analysis of the PER NES was given by Vielhaber et al. (2001). A possible NES of FRQ is shown as a highly conserved section with asterisks designating the potential NES.
* * * * FF-1 487 GWVYLNLL[ci]N[ml]AQLH[im][il]NVTPDF[iv]RSA[vl]S[ed] [kl]STK[fl]Q[li]S[ps]DGRKIRWRGGT
The brackets, that is, [ci], indicate the 1 or more amino acids that can be found in this position.
Testable idea: To determine, via mutations, if this sequence was involved in nuclear export. If experimental evidence does not support the idea that this is the NES, then one would have to consider the possibility that FRQ leaves the nucleus via some other mechanism.
Localization
In the process of comparing LARP(s) to FRQ(s), it was seen that there is a human homologue to the LARP of lower organisms called La-antigen (GI#18089160). This protein is a shorter version of LARP but still contains the signature sequence “QIEYY . . . etc.,” which matches up to the active site of LARP. The comparison of the La-antigen to the Capsaspora LARP gave a BLAST value of 5e−10(75). It was noted that there is an area of the Homo sapiens La-antigen that gets phosphorylated and hence determines its localization. This area has the sequence

The LA-antigen, a homolog of LARP in humans, has a sequence area that shows similarity to a highly conserved area of the Sordaria FRQ.
The BLAST value of the LA-antigen versus the Sordaria f. FRQ is 3.9(20) for only the region listed in
Testable idea: Mutate this key serine or its flanking parts in the FRQ sequence to determine if it affects the localization of FRQ.
Evolutionary Considerations: Candidates for Partial FRQ Ancestry
Since the true ancestors to the fungi are either not known, not still alive, or not found yet, one can analyze genes only from existing organisms that might be closely related to these true ancestors. Capsaspora could be related to one of the candidates for this role, since it has been proposed by others in this regard (James et al., 2006) and since its genome is known at this time. Likewise, it is not known how closely related are the existing fungal genomes to the ancient fungal genomes. With these 2 important caveats in mind, the existing Capsaspora LARP protein shows similarities to the existing Sordaria FRQ in the following ways: the 2 proteins have approximately the same number of amino acids (1054 [LARP] vs. 989 [FRQ]), and the area of the strongest homology is approximately in the same position in the 4 proteins of both (starting at 431 LARP vs. 484 FRQ). The area of homology is almost a continuous one, with a gap of only 1 amino acid. This strong area of match is highly conserved in LARP(s) from Arabidopsis to Zebrafish and in fact is listed in the National Institutes of Health (NIH) GenBank as part of the LAM super family. Since this area of homology is part of a common motif for the LARP(s) from plants, animals, and protists, one can infer that much of this sequence area of LARP was present at the time of divergence of plants, fungi, and animals. It is plausible then that part of this protein could have been a precursor to part of the fungal FRQ protein. A highly schematic and speculative diagram of these possibilities is shown as Figure 6.
The area of FRQ that matches to LARP is also a highly conserved area (see FRQ motif in Suppl. Table S1).The BLAST value when just the 2 matched sections were compared with each other was 3e−9(50). There are other area(s) of low homology between the 2 proteins (not shown). These additional area(s) align well with the strongest match area but are not conserved in all LARP(s), nor do they have any known function, so they are not shown.
FRQ/HSF Comparison
As mentioned, a screen of the Acanthamoeba genome employing Sordaria FRQ found 2 proteins that showed some similarity. One was the LARP, illustrated above, and the other was an HSF, shown as section C in Figure 9. To look for more similarities between these 2 proteins, they were compared using CLUSTAL, starting with the N-terminal of each protein. In this way, sections A and B were found.

The Acanthamoeba heat shock factor (HSF) and the Sordaria FRQ show 3 sections of similarity. The query is Acanthamoeba heat shock factor (GI#470459745). The subject is Sordaria fimicola FRQ (GI#6016053). The numbers in parentheses (12) are the numbers of amino acids not shown in this figure. Section C was found by CLUSTAL in the same procedure that found sections A and B, as well as by BLAST. The BLAST result is shown in this figure. The amino acids in the 2 proteins that are highly conserved are shown in bold. For the HSF proteins, these are the amino acids conserved between the HSF from Arabidopsis, Danio, and Acanthamoeba (see Suppl. Fig. S3).
Sections A and B in the Acanthamoeba HSF are part of the DNA-binding motif for the HSF. A more detailed alignment of the HSF(s) from yeasts and Arabidopsis can be found in the NIH GenBank. Section C in the HSF is listed as part of an Nap-binding protein (NBP1) protein in the NIH GenBank, and an alignment of this area of HSF can be found both in the GenBank and in Supplementary Figure S3. The part of section C shown here is part of a coiled/coil domain found in many HSF(s). Section C is highly conserved in FRQ and is listed as FF-3 in Supplementary Figure S1 and is present in 11/13 FRQ(s) listed in Table 1. It should be noted that sections A, B, and C are in the N-terminal half of both proteins. There is no indication that these functions of an HSF, such as DNA binding, are still present in FRQ. It is possible that the original functions of the HSF(s) have been modified during evolution and now have different functions in FRQ.
An example of this would be the coiled/coil domain found in HSF that shows similarity to the coiled/coil domain found in FRQ (Chen et al., 2001), which is involved in binding of the white collar proteins.
For further analysis, the opposite was performed (i.e., the Acanthamoeba HSF was employed as a query vs. the Sordaria fimicola genome). Section C of FRQ was found. When compared with each other, the 2 proteins had a BLAST value of 4e−5(80). When just the matched sections C of each were compared with each other, the BLAST value was 1e−6(72).
Section C in the HSF was used as a query versus the genomes of Acanthamoeba, Capsaspora, and Neurospora. In Acanthamoeba and Capsaspora, only HSF(s) were found. But in Neurospora, HSF and FRQ were found. Thus, this part of the sequence is relatively rare among proteins in these genomes, making it potentially more interesting that FRQ shares this sequence with an HSF.
Since HSF(s) in fungi, Capsaspora, plants, and animals show high homology to each other, it is likely that an ancient form of an HSF was present at the time of the divergence into plants, animals, and fungi. To be thorough, these other HSF(s) were compared with the Sordaria FRQ, but they did not show as much similarity as did the Acanthamoeba HSF. The reciprocal was tested, that is, a comparison of the FRQ(s) from 30 different species of fungi (partial list in the Materials and Methods section) to the Acanthamoeba HSF. Many had BLAST values similar to that of the Sordaria FRQ; others had less similarity.
FRQ/ARNT Comparison
Since the ARNT protein has been shown to be similar to other clock proteins, such as PER and BMAL1, it was of interest to determine if there was any relationship to FRQ.
The following facts led to an investigation of possible similarities between FRQ and ARNT: it was already known that the Drosophila m. PER had a high similarity in its PAS sections to the Drosophila ARNT, with a BLAST value of e−27(289). It is reported here that the Drosophila m. PER also had a similarity in its PAS sections to the ARNT of Capsaspora, with a BLAST value of e−09(276). It was found by a BLAST comparison of FRQ to the Drosophila PER that there was a small section that showed some similarity between both. This section was the start of the PAS part of PER. Since these 3 proteins all had some similarity in some part of the PAS section, it seemed reasonable to investigate whether there were other sections of similarity between FRQ and ARNT.
To do this, a CLUSTAL analysis of all 3 proteins was performed, starting with the section that they all had in common. For the Neurospora FRQ, it was amino acid 646; for the Capsaspora ARNT, amino acid 450; and for Drosophila PER, amino acid 248. The results, shown just for FRQ and ARNT as Figure 10, indicated several additional sections of similarity. The CLUSTAL alignment found 4 sections of similarity.
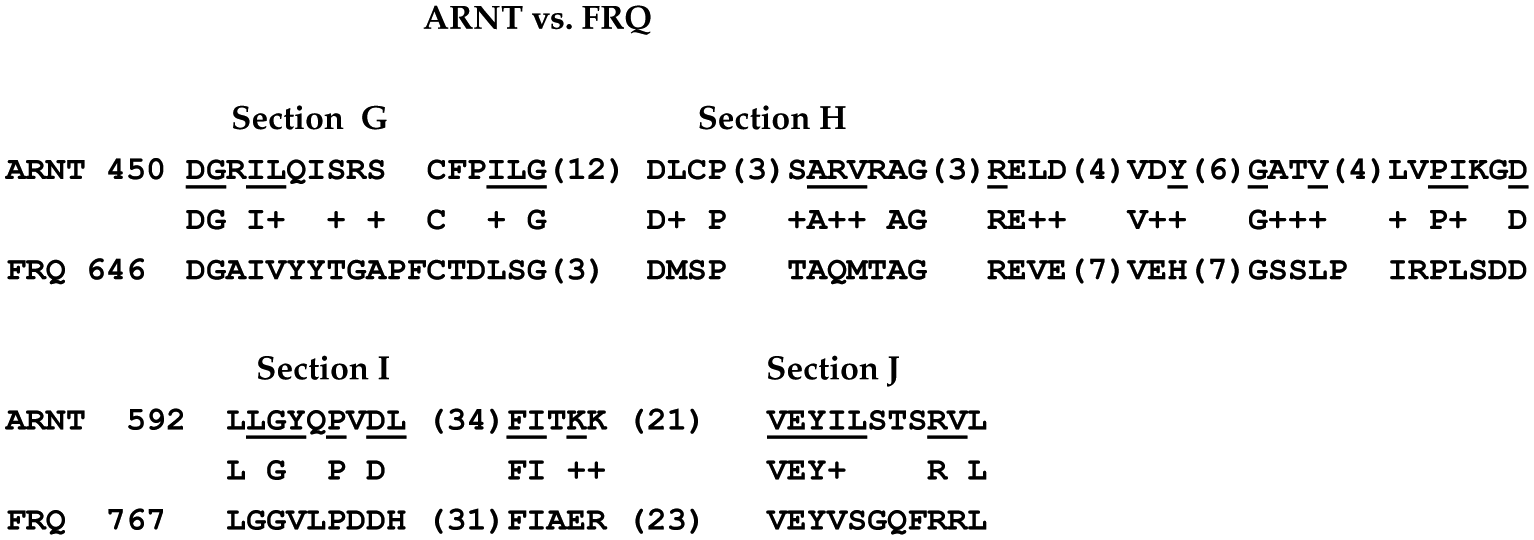

The Capsaspora ARNT and the Neurospora FRQ showed 4 sections of similarity. The underlined amino acids in ARNT (i.e.
This section of ARNT (and, by analogy, this section of PER) is known to bind other proteins. In Drosophila, this region is known to bind the clock protein, TIM, and to be involved with cytoplasmic localization. It is possible that these sections of FRQ shown in Figure 10 have evolved to take on some other function, such as providing a site for the binding of FRQ to the protein FRH.
FRQ Might Be a Composite
The fungal FRQ is suggested here to be a composite of sequences from ARNT, HSF, and LARP. Employing the sequences from Capsaspora, there are several sections from ARNT, listed in Figure 10 as sections G, H, I, and J. These are also diagrammed in Figure 11 (FRQ sections).
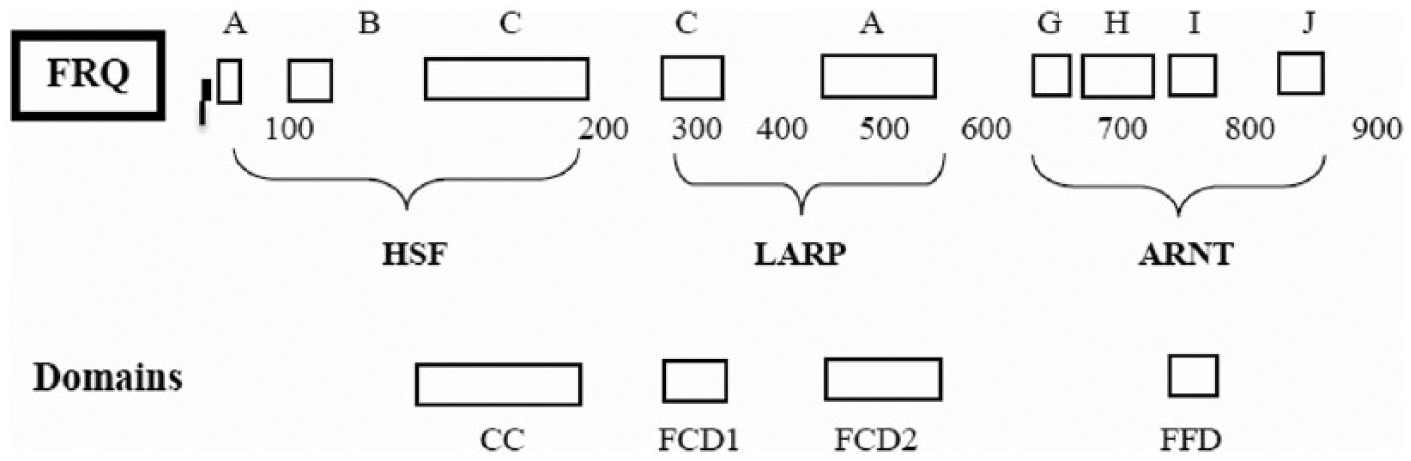
Diagrammatic representation of FRQ sections, indicating possible origin from Capsaspora ARNT, Capsaspora LARP, and Acanthamoeba HSF. Four of the known domains in FRQ are matched up to 4 of the known nine FRQ sections shown.
There are at least 2 sections of FRQ that could be derived from LARP. These are shown in Figure 11 as sections A and C. There are possibly others, which are not diagrammed here.
There are 3 sections of FRQ that show some homology to Acanthamoeba HSF as illustrated in Figure 11 as sections A, B, and C.
Clock Components
Salichos and Rokas (2010) have analyzed many different organisms for the presence or absence of FRQ, FRH, WC-1, and WC-2. Since many genomes have been sequenced since then, their analysis has been expanded and quantified as shown in Table 4.
Analysis of the genomes of 5 different organisms for the presence of 4 different Neurospora crassa proteins and 4 different genes.
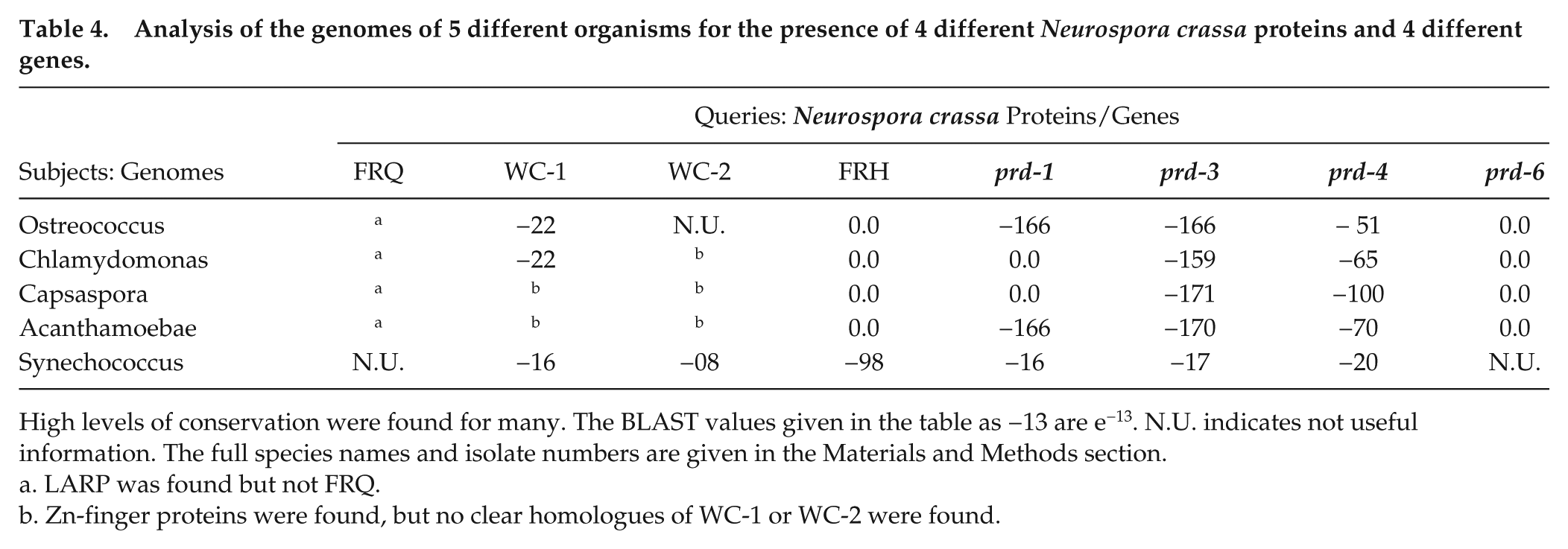
High levels of conservation were found for many. The BLAST values given in the table as −13 are e−13. N.U. indicates not useful information. The full species names and isolate numbers are given in the Materials and Methods section.
LARP was found but not FRQ.
Zn-finger proteins were found, but no clear homologues of WC-1 or WC-2 were found.
Compared with the previous work, this table has 3 improvements: (1) it provides actual BLAST values, instead of just + or −; (2) it contains many species of fungi not in the previous work; and (3) it contains 4 additional genes. Some proteins/genes, such as FRH, prd-1, prd-3, and prd-6, are highly conserved, even in distantly related species of fungi. Many of the other proteins from species furthest from Neurospora show the least homology to the Neurospora proteins, similar to what is noticed for the FRQ protein.
Clock Components in Nonfungal Species
More distantly related species, nonfungal species, such as Chlamydomonas or Ostreococcus, showed that homologues to WC-1, FRH, prd-1, prd-3, prd-4, and prd-6 could easily be found. For the cyanobacteria, Synechococcus elongatus, proteins for the 6 of the 8 were found, but no useful information was obtained for prd-6 or for FRQ. The finding of some similarity to these clock proteins in Synechococcus suggests that at least some parts of these clock components are quite ancient and that the emergence of the fungal clock then required some new genes/proteins, such as FRQ.
The same set of proteins/genes was scanned against the genomes of Acanthamoebae and Capsaspora, as shown in Table 4. Only proteins containing PAS motifs were found for the WC-1 or WC-2 in either genome. It is interesting to compare this list with the homologues found in the bacteria Synechococcus, as shown in Table 3. The presence of ARNT and LARP in the amoebae, but not in the bacteria, may represent an adaptation for cells containing a nucleus.
The Acanthamoeba genome only showed 4 proteins with enough similarity to WC-1 to give a significant BLAST score for the PAS motifs. This is consistent with the finding for the Neurospora genome; that is, highly homologous PAS motifs are not widespread in either organism.
Discussion
The finding that FRQ is a relatively unique protein brings up the general question of how genes/proteins with new functions, such as FRQ, are formed. The mechanisms for the formation of new genes are generally considered to be recombination of existing genes that have different functions, duplication/divergence of an “extra copy” of a gene to give a new function, and other mechanisms such unequal crossing over, translocations, crossing over within an inversion, and mutation.
To take just one of these examples, for instance, it can be proposed that the duplication of a LARP somewhere in the evolution of the fungi and a subsequent remodeling of this second copy to become FRQ could account for the data presented here. The duplicate LARP gene “copy” was then modified by recombination with the genes coding for HSF and ARNT to give a new gene coding for a new protein (FRQ). This new protein would have the motifs found in ARNT, LARP, and HSF.
Since this idea cannot be experimentally verified, one can determine only if the postulated ancient conversion of regions in LARP and HSF to regions in FRQ would be plausible employing the existing current sequences. In this regard, both the LARP and HSF have highly conserved regions that would make them plausible precursors to the regions currently found in FRQ. In addition, it is interesting that LARP, ARNT, and HSF can all be found in the nucleus as well as the cytoplasm, similar to what is found for FRQ.
Many of the gene/protein components found in the Neurospora clock system, such as kinases and so on, can be found in Capsaspora and other organisms (Table 4). In other words, the “context” for the appearance of a new protein, such as FRQ, would already be present. In addition, perhaps the gene components, such as LARP, ARNT, and HSF, were also present and were already rhythmic in nature? That is, they might have been oscillating under entraining (L/D) conditions. A selection process via gene recombination for a free-running, anticipatory oscillator could then have taken place to complete the clock system since many of the components of these were already present.
The motifs that are important for FRQ function can be found in the proteins listed here as follows (see details in text): (1) kinase binding, from LARP, sections A and C; (2) dimerization, from HSF, section C; (3) FRH binding, from ARNT, section 1; (4) NES, from LARP; and (5) nuclear import signal, from HSF.
The possible origins of other properties and motifs of FRQ, such as the PEST regions and key serines, are still to be determined.
The proposal that FRQ might have arisen from the existing proteins as listed here leads to some additional suggestions: (1) The early eukaryotic clock might have been based on 1 or more RNA-binding proteins that mutated over time to adapt to the needs/niches of different organisms or groups of organisms. (2) Those current organisms that have a circadian rhythm but do not have FRQ (fungi) or PER (insects, mammals) or TOC (plants) might have a clock based on RNA-binding proteins. This is a testable idea. (3) It might be worthwhile to investigate in more detail the role of the RNA-binding proteins in the existing clock machinery in those organisms that do have FRQ, TOC, and PER (i.e., fungi, plants, and animals). Some clues have already emerged, such as FRH in Neurospora, for example.
Supplemental Material
Supplementary_Figure_1_FRQ_motifs – Supplemental material for Circadian Rhythms in Fungi: Structure/Function/Evolution of Some Clock Components
Supplemental material, Supplementary_Figure_1_FRQ_motifs for Circadian Rhythms in Fungi: Structure/Function/Evolution of Some Clock Components by Stuart Brody in Journal of Biological Rhythms
Footnotes
Acknowledgements
The author acknowledges the assistance, encouragement, and helpful criticism from the following: Arnaud Tauton and Russell Doolittle of the University of California, San Diego; John Taylor of the University of California, Berkeley; and Carrie Partch of the University of California, Santa Cruz. The author is grateful to Mike Young for serving as a host for my many visits to his lab at Rockefeller University and for the interactions with him, Deniz Topf, and Leno Saez there over the past 7 years.
Conflict Of Interest Statement
The author has no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
