Abstract
The principal circadian pacemaker in mammals, the suprachiasmatic nucleus (SCN), expresses a number of neuropeptides that facilitate intercellular synchrony, helping to generate coherent outputs to peripheral clocks throughout the body. In particular, arginine vasopressin (AVP)– and vasoactive intestinal peptide (VIP)–expressing neurons have been recognized as crucial subpopulations within the SCN and have thus been the focus of many chronobiological studies. Here, we analyze the neuropeptide expression of 2 popular transgenic mouse strains commonly used to direct or restrict Cre-mediated recombination to AVP- and VIP-ergic neurons. The Avp-IRES2-Cre (JAX #023530) and Vip-IRES-Cre (JAX #010908) “driver” mouse strains express the Cre recombinase under the control of the endogenous Avp or Vip gene, respectively, allowing scientists either to ablate their gene of interest or to overexpress a transgene in a cell type–specific manner. Although these are potentially very powerful tools for chronobiologists and other scientists studying AVP- and VIP-ergic neurons, we found that neuropeptide expression in these mice is significantly decreased when an IRES(2)-Cre cassette is inserted downstream of the neuropeptide-encoding gene locus. The impact of IRES(2)-Cre cassette insertion on neuropeptide expression may be a confounding factor in many experimental designs. Our findings suggest that extreme caution must be exercised when using these mouse models to avoid misinterpretation of empirical results.
Mouse genetic models have been indispensable for improving our understanding of circadian timekeeping mechanisms in mammals. In particular, the Cre-LoxP system is a powerful approach for determining the contributions of a gene of interest (GOI) to circadian timekeeping by ablating its expression in a cell- or tissue type–specific manner (Utomo et al., 1999). Despite the availability of numerous mouse strains that carry loxP-flanked alleles of clock-relevant genes (Billon et al., 2017; Debruyne et al., 2006; Delezie et al., 2016; Fu et al., 2005; Johnson et al., 2014; Meng et al., 2008; Storch et al., 2007), only a handful of tissue-specific Cre driver mouse strains that have been used in chronobiological studies provide Cre-mediated recombination in the suprachiasmatic nucleus (SCN; Brancaccio et al., 2017; Brancaccio et al., 2019; Cheng et al., 2019; Husse et al., 2011; Izumo et al., 2014; Lananna et al., 2018; Lee et al., 2015; Mehta et al., 2015; Smyllie et al., 2015; Tien et al., 2012; Tso et al., 2017; van der Vinne et al., 2018; Weaver et al., 2018), distinguishing between cell types (neurons versus astrocytes) but not SCN subregions. The SCN is anatomically and functionally divided into the shell (dorsomedial [dm]) and core (ventrolateral [vl]) compartments, the former being the highly rhythmic region and the latter serving as the retinorecipient, or light-responsive, area of the SCN (Abrahamson and Moore, 2001; Moga and Moore, 1997). The dmSCN and vlSCN compartments are defined by their expression of the neuropeptides arginine vasopressin (AVP) and vasoactive intestinal peptide (VIP), respectively, both of which mediate interneuronal coupling (Aton et al., 2005; Edwards et al., 2016; Maywood et al., 2011). Recently, 2 Cre strains were generated that expressed Cre under the control of the endogenous Avp or Vip gene (Taniguchi et al., 2011; Harris et al., 2014) and that were made commercially available through The Jackson Laboratory (Bar Harbor, ME; stock #023530 and #010908). These strains are potentially valuable tools for chronobiologists who are interested in teasing apart the functions of a GOI in different SCN compartments or to direct the expression of a transgene to a particular compartment. To evaluate the utility of these 2 Cre driver strains in chronobiological studies, we assessed circadian locomotor rhythms and expression of VIP and AVP in Cre-expressing mice.
Methods
Animals
Heterozygous Avp-IRES2-Cre-D (stock #023530; Avpcre/+), homozygous Vip-IRES-Cre (stock #010908; Vipcre/cre), and C57BL/6J (stock #000664) mice were purchased from The Jackson Laboratory. Avpcre/+ and Vipcre/cre mice were bred with C57BL/6J mice to establish an ultimate breeding scheme of Avpcre/+ or Vipcre/+ mice crossed with C57BL/6J animals. Heterozygous mice used for experiments were at least 8 generations (for Avpcre/+) or 3 generations (for Vipcre/+) backcrossed to C57BL/6. Only wild-type littermates (Avp+/+ or Vip+/+) were employed as controls, with the exception of the LD→DD behavioral paradigm, in which C57BL/6J mice were used as controls for Vipcre/+ mice. The C57BL/6J mice that were used for the LD→DD behavioral paradigm were generated by in-house breeding. All procedures were performed at the University of Toronto Mississauga (UTM) Animal Facility and were approved by the UTM Animal Care Committee in accordance with guidelines established by the University of Toronto and the Canadian Council on Animal Care.
Circadian Behavioral Paradigms and Analyses
Starting at 5 to 8 weeks of age, male mice were singly housed in running-wheel cages within ventilated, light-tight cabinets under computer-controlled lighting schedules (Phenome Technologies, Skokie, IL). Illumination within the cabinets was provided by white light-emitting diodes, with intensity set at 40 Lux (0.50 µE) at cage level unless otherwise indicated. Wheel revolutions were recorded and analyzed using ClockLab software (Actimetrics, Wilmette, IL). In all experiments, mice were initially entrained to a 12:12 LD schedule for at least 2 weeks prior to indicated changes in the light schedules. Free-running rhythms were examined by releasing mice from 12:12 LD into constant darkness (DD) for 2 weeks. To determine the effects of constant light (LL), mice were exposed to LL in which the light intensity increased stepwise (10 Lux [0.13 µE], 40 Lux [0.5 µE], 80 Lux [1 µE], and 160 Lux [2 µE]) every 2 weeks. Times of activity onset were determined by ClockLab and corrected when necessary following visual inspection of the actograms. Onset error was defined as the standard deviation of daily onsets over 7 consecutive days. Period length was measured by fitting a regression line through daily activity onsets or by using the χ2 periodogram. Amplitude was measured by the χ2 periodogram and fast Fourier transformation. Phase angles of entrainment were measured by calculating the difference between onset of activity and lights-off.
Tissue Harvest
Mice (both sexes, 5 to 8 weeks of age) were maintained on a fixed 12:12 LD schedule and released in DD for 2 consecutive cycles prior to tissue harvest at prescribed CTs on the third day of DD. Mice were killed by cervical dislocation, and their brains were rapidly dissected under dim red light. Brains were sectioned in ice-cold oxygenated media or diethyl pyrocarbonate–treated phosphate-buffered saline (PBS), pH 7.4 (for RNAscope in situ hybridization experiments) with an oscillating tissue slicer (Electron Microscopy Sciences, Hatfield, PA) to obtain an 800-µm-thick coronal slice containing the SCN or other brain regions. For immunohistochemistry, tissue slices were fixed in 4% paraformaldehyde (PFA) in PBS (pH 7.4) for 6 h at room temperature (RT), cryoprotected in 30% sucrose in PBS at 4 °C overnight, cut into 30-µm-thin sections using a freezing microtome (Leica Microsystems, Buffalo Grove, IL), and stored in 30% sucrose at 4 °C until further use. For RNAscope in situ hybridization experiments, tissue slices were fixed in 4% PFA in RNase-free PBS (pH 7.4) for 24 h at 4 °C, cryoprotected in 30% sucrose in PBS at 4 °C overnight, and then frozen in Tissue Tek Optimal Cutting Temperature (OCT) embedding media (Sakura Finetek, Torrance, CA) with dry ice. Tissue blocks were cut into 14-µm-thin sections using CryoStar NX50 Cryostat (Thermo Scientific, Waltham, MA) and thaw mounted on SuperFrost Plus slides (Fisher Scientific, Waltham, MA). Slides were air dried for 2 h at −20 °C and then stored at −80 °C until further use.
Immunohistochemistry
Coronal brain sections were washed 5 × 5 min in PBST (PBS with 0.1% Triton X-100, pH 7.4), treated with 0.3% H2O2 in PBS for 20 min at RT, washed 5 × 5 min in PBST, and incubated for 1 h at RT in blocking solution (10% horse serum in PBST). Tissues were incubated overnight at 4 °C in rabbit anti-AVP (1:30,000; AB1565, EMD Millipore, Burlington, MA), rabbit anti-VIP (1:10,000; 20077, ImmunoStar, Hudson, WI), or rabbit anti–peptide histidine isoleucine (PHI; 1:4000; H-064-07, Phoenix Pharmaceuticals, Burlingame, CA) primary antibodies diluted in fresh blocking solution. The next day, sections were washed 5 × 5 min in PBST, incubated for 2 h at RT with the appropriate biotinylated secondary antibody diluted in blocking solution, washed 5 × 5 min in PBST, and incubated for 45 min with avidin-biotinylated horseradish peroxidase (HRP) complexes (ABC) from the VECTASTAIN ABC-HRP kit (PK-4000, Vector Laboratories, Burlingame, CA). After washing 5 × 5 min in PBST, sections were developed with the 3,3′-diaminobenzidine (DAB) HRP substrate (SK-4100, Vector Laboratories) according to the manufacturer’s instructions. For each experiment, all sections were processed at the same time and with the same DAB development time. Sections were mounted onto gelatin-coated glass microscope slides, dehydrated, and coverslipped with Permount Mounting Media (SP15-500, Thermo Fisher Scientific).
RNAscope In Situ Hybridization
The in situ hybridization (ISH) probes that were used to visualize Avp and Vip mRNA expression were purchased from Advanced Cell Diagnostics (Newark, CA; Mm-Avp-C2, 401391-C2; Mm-Vip-C3, 415961-C3). The RNAscope Multiplex Fluorescent Reagent Kit v2 (Cat #323100, Advanced Cell Diagnostics) was used according to the manufacturer’s instructions for fixed frozen sections. Briefly, slides were washed in PBS for 5 min, baked at 60 °C for 30 min, fixed with 4% PFA in PBS for 90 min at RT, dehydrated in increasing concentrations of ethanol (50%, 70%, 100%, and 100%), and baked again at 60 °C for 10 min. Slides were then treated with RNAscope hydrogen peroxide for 10 min at RT. Antigen retrieval was performed by placing slides into mildly boiling RNAscope target retrieval reagent for 5 min and then washed in distilled water for 15 sec and 100% ethanol for 3 min. Slides were dried and permeabilized with RNAscope Protease III for 30 min at 40 °C. Next, slides were rinsed with distilled water followed by probe incubation for 2 h at 40°C. Afterward, slides were incubated with amplifier probes and the appropriate HRP solution at 40 °C (AMP1, 30 min; AMP2, 30 min; AMP3, 15 min; HRP-C2 or HRP-C3, 15 min) with 2 × 2 min of washes with RNAscope wash buffer in between each step. Slides were incubated with Opal 570 dye (Akoya Biosciences, Menlo Park, CA) diluted in TSA buffer for 30 min at 40 °C. HRP activity was then blocked, and sections were counterstained with DAPI and coverslipped with VECTASHIELD Mounting Medium (Vector Laboratories).
Image Acquisition and Quantification
Images were acquired using a Zeiss Axio Observer Z1 inverted microscope equipped with a laser scanning microscope 700 module for confocal images and an AxioCam MRm Rev.3 monochromatic digital camera (Zeiss, Oberkochen, Germany) for bright-field pictures, with the Zen 2010 software (Zeiss). Identical settings were used for imaging samples within each experiment (gain, pinhole, filter sets for confocal and brightness for light microscopy). Confocal images were acquired in separate channels for each fluorophore. All image analyses were performed on ImageJ.
Intensity
For quantification of signal intensity, 10× bright-field images (for immunohistochemistry) or 20× confocal images (for ISH) of the region of interest (ROI) were acquired. The area of each ROI was delineated using the polygon selection tool, and the average optical density was obtained using the “measure” function. The background signal was measured in an unlabeled region adjacent to each ROI and subtracted from the average intensity within the ROI.
Cell Counts
For Avp+ and Vip+ cell counts and density measurements, 20× confocal images were acquired that covered the entire surface of the bilateral SCN from two to four 14-µm-thick coronal sections. Avp+ or Vip+ cells at each threshold were counted manually through all z-planes of these confocal images. For Avp, high and medium thresholds were set to 210 and 180, respectively; for Vip, high and medium thresholds were set to 180 and 150, respectively. Cells that exceeded the medium threshold were denoted as Avpmed and Vipmed cells, which included cells with medium-to-high expression, whereas Avphigh and Viphigh cells denoted cells that surpassed the high threshold and were of high expression only.
Statistical Analysis
Data were analyzed using linear mixed effects modeling fit, 2-way analysis of variance, and Student’s t test with R (version 3.5.0). Post hoc significance of pairwise comparisons was assessed using Tukey’s honest significant difference test with α set at 0.05.
Results
The design of both Cre driver strains, Avp-IRES2-Cre-D and Vip-IRES-Cre, is similar. Each carries a knock-in allele in which an internal ribosomal entry site (IRES) sequence and the Cre recombinase transgene were inserted downstream of the translational STOP codon of the Avp or Vip gene locus (Figs. 1A and 2A). For the Avp- or Vip-specific Cre strains, gene targeting was performed in (129S6/SvEvTac x C57BL/6)F1–derived G4 embryonic stem (ES) cells or (C57BL/6 x 129S4Sv/Jae)–derived V6.5 ES cells, respectively. The Cre activity of both driver strains was previously characterized by crossing these mice to the Ai14 reporter strain in which expression of tdTomato is induced by the excision of an upstream loxP-flanked (fl) STOP cassette (Madisen et al., 2010). The transgenic characterization experiment conducted by the Allen Institute for Brain Science (e.g., experiments 293795159 and 183132842) revealed that each driver strain induces Cre-mediated excision in the expected AVP- and VIP-positive cell populations in the brain.
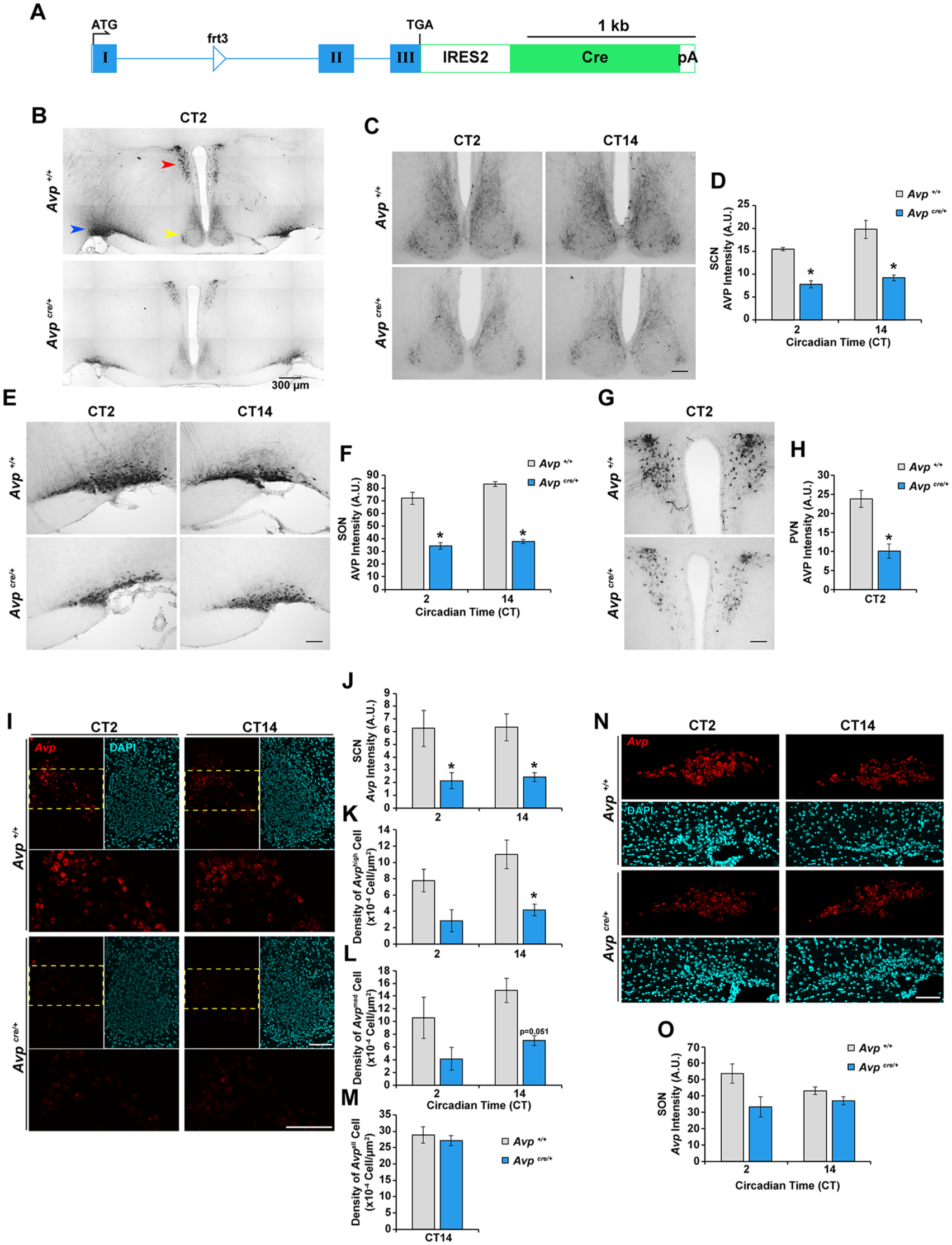
Avpcre/+ mice exhibit reduced expression of arginine vasopressin (AVP) at the transcript and protein levels. (A) Schematic representation of the Avp-IRES2-Cre-D knock-in allele. Roman numerals denote exons, blue boxes represent the exonic regions encoding the prepro-AVP-neurophysin 2-copeptin precursor, and the green box represents the region encoding Cre recombinase. ATG, start codon; TGA, stop codon; Cre, Cre recombinase gene; pA, polyA sequence; IRES2, internal ribosome entry site 2; frt3, FLP recombinase target site. (B, C, E, G) Representative photomicrographs of AVP immunoreactivity in the (B) hypothalamic region, (C) SCN, (E) supraoptic nucleus (SON), and (G) paraventricular nucleus (PVN) of Avpcre/+ and Avp+/+ mice at the indicated circadian time. The SCN, PVN, and SON are indicated by the yellow, red, and blue arrowheads, respectively, in (B). (D, F, H) Quantification of AVP immunoreactivity in the (D) SCN, (F), SON, and (H) PVN. Values are given as arbitrary units of background-subtracted mean gray-scale intensity. (I, N) Representative photomicrographs of Avp mRNA expression (red) in the (I) SCN and (N) SON of Avpcre/+ and Avp+/+ mice at the indicated circadian time. DAPI (cyan) is used as a nuclear stain. In (I), regions outlined by the yellow dashed lines are shown in higher magnification in the lower panels. (J, O) Quantification of Avp mRNA expression intensity in the (J) SCN and (O) SON. Values are given as arbitrary units of background-subtracted mean gray-scale intensity. (K-M) Quantification of the density of (K) Avphigh, (L) Avpmed, and (M) Avpall cells in the SCN of Avpcre/+ and Avp+/+ mice. All values are represented as mean ± SEM. n = 3 to 5 per group. *p < 0.05 versus Avp+/+ littermate control mice. Scale bars are 100 µm unless otherwise specified.
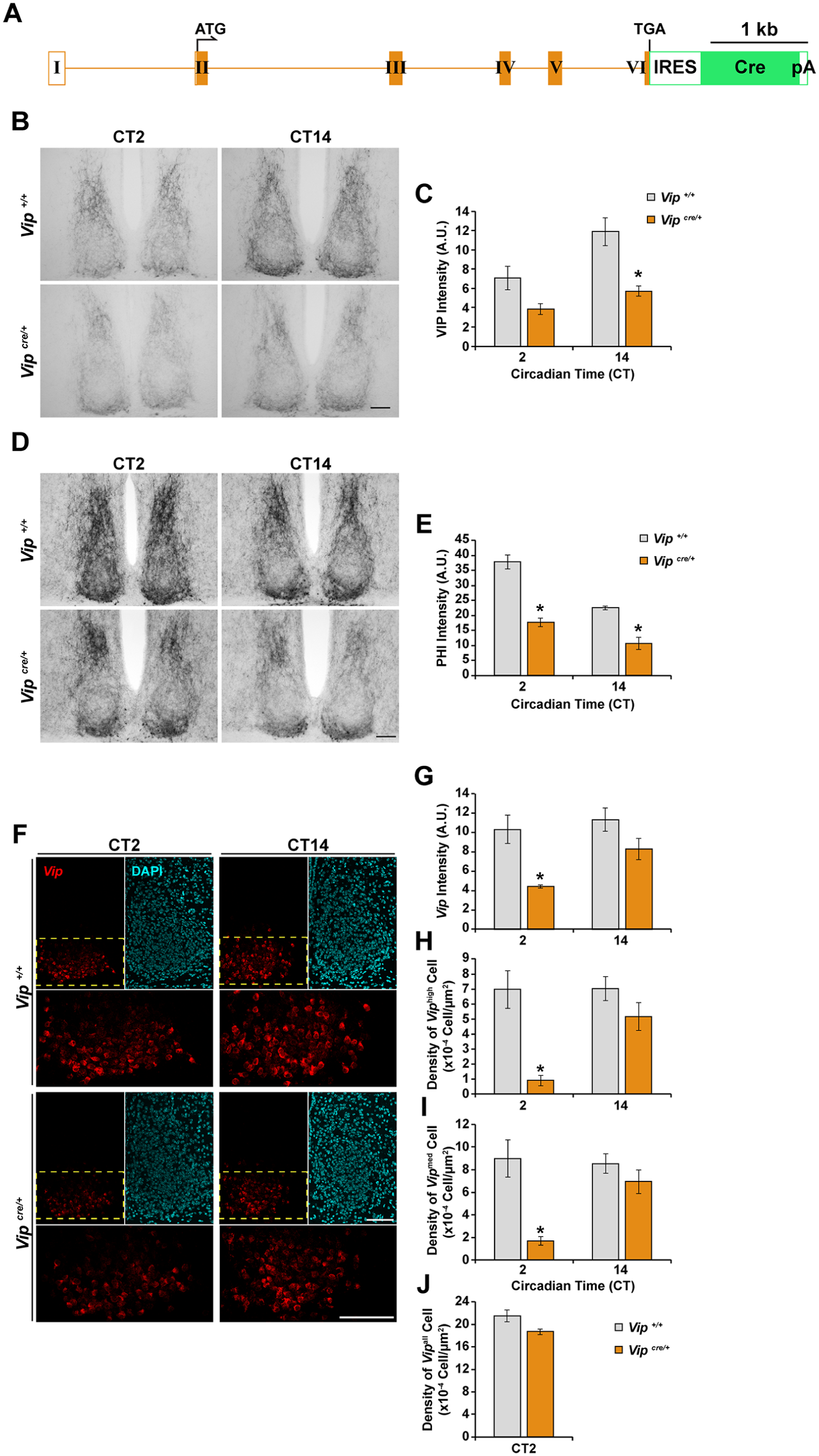
Vipcre/+ mice exhibit reduced expression of vasoactive intestinal peptide (VIP) and peptide histidine isoleucine (PHI) at the transcript and/or protein levels. (A) Schematic representation of the Vip-IRES-Cre knock-in allele. Roman numerals denote exons, orange boxes represent the exonic regions encoding the prepro-VIP precursor, and the green box represents the region encoding Cre recombinase. ATG, start codon; TGA, stop codon; Cre, Cre recombinase gene; pA, polyA sequence; IRES, internal ribosome entry site. (B, D) Representative photomicrographs of (B) VIP and (D) PHI immunoreactivity in the SCN of Vipcre/+ and Vip+/+ mice at the indicated circadian time. (C, E) Quantification of (C) VIP and (E) PHI immunoreactivity in the SCN. Values are given as arbitrary units of background-subtracted mean gray-scale intensity. (F) Representative photomicrographs of Vip mRNA expression (red) in the SCN of Vipcre/+ and Vip+/+ mice at the indicated circadian time. DAPI (cyan) is used as a nuclear stain. Regions outlined by the yellow dashed lines are shown in higher magnification in the lower panels. (G) Quantification of Vip mRNA expression intensity in the SCN. Values are given as arbitrary units of background-subtracted mean gray-scale intensity. (H-J) Quantification of the density of (H) Viphigh, (I) Vipmed, and (J) Vipall cells in the SCN of Vipcre/+ and Vip+/+ mice. All values are represented as mean ± SEM. n = 3 to 5 per group. *p < 0.05 versus Vip+/+ littermate control mice. Scale bars are 100 µm.
The insertion of the IRES2-Cre cassette into the Avp gene locus markedly attenuated AVP immunoreactivity (IR) in hypothalamic nuclei (Fig. 1B), including the SCN (Fig. 1C), supraoptic nucleus (SON; Fig. 1E), and paraventricular nucleus (PVN; Fig. 1G). Compared with wild-type (Avp+/+) littermates, Avpcre/+ mice exhibited significantly lower AVP-IR intensity in all 3 nuclei (Fig. 1D, F, H). The reduction in AVP protein abundance in the SCN of Avpcre/+ mice was mirrored in a similar decrease at the transcript level, as determined by ISH using the RNAscope® technology (Fig. 1I; Wang et al., 2012). Attenuated levels of Avp mRNA were reflected in the lower intensity of the fluorescent label within the SCN (Fig. 1J) as well as fewer SCN neurons expressing high (Fig. 1K) or intermediate (Fig. 1L) levels of Avp. The total number of Avp-positive neurons within the SCN was not different between Avpcre/+ and Avp+/+ mice (Fig. 1M), ruling out a potential loss of AVPergic neurons as a result of the IRES2-Cre insertion. We did not detect a statistically significant difference in fluorescence label intensity in the SON of Avpcre/+ mice compared with Avp+/+ controls (Fig. 1N and O).
Similarly, we observed that the neuropeptide products encoded by the Vip gene, VIP (Fig. 2B and C) and PHI (Fig. 2D and E), were reduced in abundance in the SCN of Vipcre/+ mice compared with Vip+/+ controls. Vip/Phi transcript levels were also attenuated in the SCN of Vipcre/+ mice, reaching statistical significance at CT2 (Fig. 2F and G). In these animals, there were fewer SCN neurons that expressed high (Fig. 2H) or intermediate (Fig. 2I) levels of Vip/Phi at CT2, although the total number of Vip/Phi-positive neurons remained unchanged compared with Vip+/+ controls (Fig. 2J).
Importantly, the insertion of the IRES(2)-Cre cassette perturbed only the expression of the targeted gene locus and did not affect the expression of neuropeptides that are not encoded by that locus. Specifically, VIP-IR was unaltered in the SCN of Avpcre/+ mice (Fig. 3A and B), as was AVP-IR in the SCN (Fig. 3C and D) and SON (Fig. 3E and F) of Vipcre/+ mice.
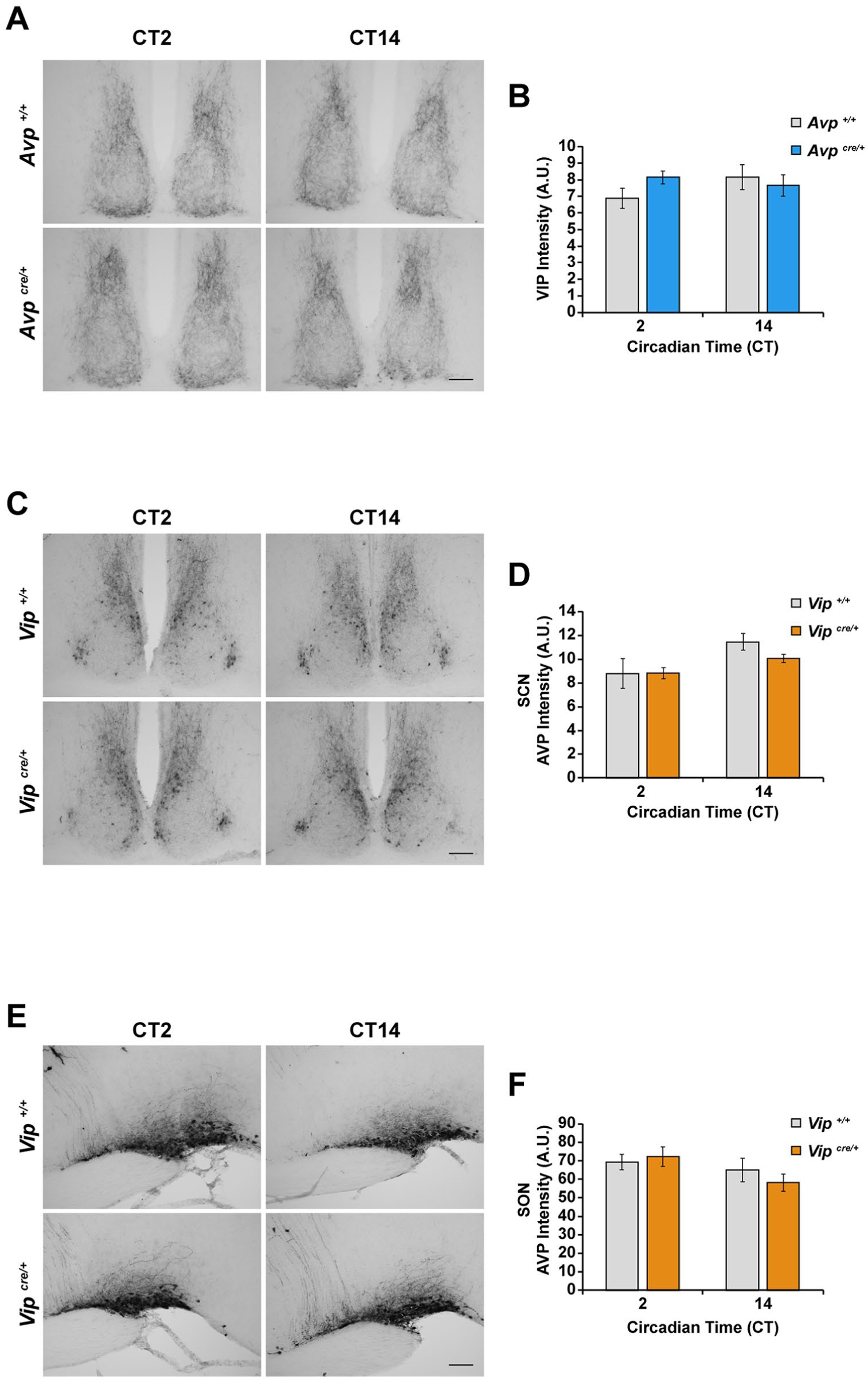
Insertion of the IRES(2)-Cre cassette into the Avp or Vip gene locus does not perturb the expression of vasoactive intestinal peptide (VIP) or arginine vasopressin (AVP), respectively. (A) Representative photomicrographs of VIP immunoreactivity in the SCN of Avpcre/+ and Avp+/+ mice at CT2 and CT14. (C, E) Representative photomicrographs of AVP immunoreactivity in the (C) SCN and (E) supraoptic nucleus (SON) of Vipcre/+ and Vip+/+ mice at CT2 and CT14. (B, D, F) Quantification of (B) VIP and (D, F) AVP immunoreactivity in the SCN or SON of (B) Avpcre/+ and (D, F) Vipcre/+ mice and their respective wild-type littermate controls. Values are given in arbitrary units of background-subtracted mean gray-scale intensity. All values are represented as mean ± SEM. n = 3 to 5 per group. Scale bars are 100 µm.
Finally, we examined the impact of the IRES(2)-Cre insertion into the Avp and Vip gene loci on wheel-running activity rhythms under 12:12 LD-entrained, constant dark, and constant light conditions (Fig. 4; Suppl. Figs. S1 and S2). Avpcre/+ (Fig. 4A and G) and Vipcre/+ (Fig. 4B and 4H) mice did not differ significantly from respective controls in their phase angle of LD entrainment (Fig. 4K and O), error of activity onset under DD (Fig. 4C and E), or period under DD (Fig. 4D and F). When exposed to LL conditions of escalating light intensities, Avpcre/+ (Fig. 4G) and Vipcre/+ (Fig. 4H) mice were indistinguishable from control animals in terms of mean amplitude (Fig. 4I, J, M, and N; Supplementary Tables S2, S3, S5, and S6) and period (Fig. 4L and P; Supplementary Tables S1 and S4) of behavioral rhythms at every intensity examined. Despite the lack of a statistical difference between Vipcre/+ and control mice on the abovementioned parameters, the LL behavior of Vipcre/+ mice appeared to be somewhat more variable than Vip+/+ animals, particularly at the intermediate light intensities (Suppl. Fig. S2), although a larger cohort of animals will be required to confirm this tentative observation.
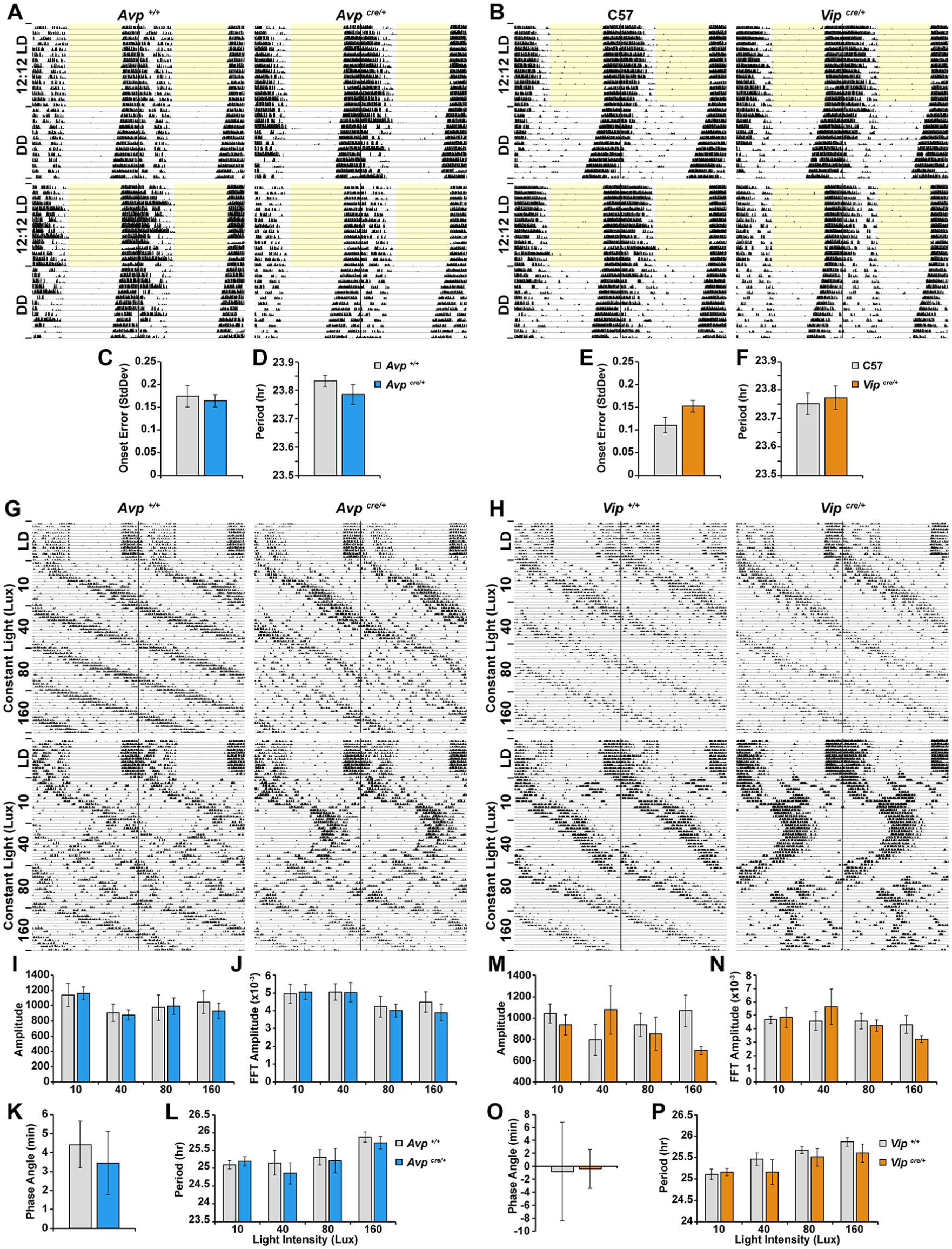
Insertion of the IRES(2)-Cre cassette into the Avp or Vip gene locus does not grossly perturb wheel-running activity rhythms under constant conditions. (A, B) Representative actograms of wheel-running activity of (A) Avpcre/+ and Avp+/+ and (B) Vipcre/+ and C57 mice. Periods of light are shaded in yellow. (C, E) Error of activity onsets under DD. (D, F) Period length under DD. (G, H) Representative actograms of wheel-running activity of (G) Avpcre/+ and Avp+/+ and (H) Vipcre/+ and Vip+/+ mice under LL conditions of increasing light intensities. Quantification of the (I, M) amplitude determined from the χ2 periodogram, (J, N) Fast Fourier transformation amplitude. (K, O) Phase angle of entrainment under LD. (L, P) Period length under LL conditions. Values represent mean ± SEM; n = 6 to 10 per group.
Discussion
Our results reveal that the IRES(2)-Cre insertion into the Avp and Vip gene loci interferes with the expression of the encoded neuropeptide(s) in the SCN and other hypothalamic nuclei, raising concerns about the utility of these Avp-IRES2-Cre-D and Vip-IRES-Cre mouse models for the study of circadian rhythms. Even with a single copy of the wild-type Avp or Vip allele, heterozygous Cre knock-in mice exhibited reduced expression of the targeted locus at both the transcript and protein levels. Immunohistochemical analyses suggest that the reduction in neuropeptide abundance may be as much as 50% in Avpcre/+ and Vipcre/+ mice compared with wild-type controls. Although we did not evaluate neuropeptide expression in homozygous Avpcre/cre and Vipcre/cre mice, we presume that there will be an even greater reduction in the absence of a wild-type allele. The IRES(2)-Cre insertion might interfere with gene transcription, transcript stability, or mRNA translation, all of which would contribute to reduced neuropeptide abundance. Changes in transcription and mRNA stability are consistent with the observed decrease in Avp and Vip/Phi transcript levels. Given that independent targeting of the IRES(2)-Cre cassette into the Avp and Vip gene loci had similar effects at the level of neuropeptide expression, it seems unlikely that the effects are the consequence of a random mutation introduced during construct generation or ES cell targeting.
The insertion of the IRES(2)-Cre cassette into a single allele of the Avp or Vip gene did not markedly perturb wheel-running activity rhythms, although we evaluated circadian behavior under only a limited set of environmental light conditions. Other paradigms such as reentrainment to a shifted LD schedule and acute phase shifts to nocturnal light pulses were not examined. We did notice that Vipcre/+ mice demonstrated slightly more variable behavior compared with controls under LL conditions, despite the lack of a group difference between Vipcre/+ and Vip+/+ mice. Although, on its own, the IRES(2)-Cre insertion appears to have minimal impact on circadian activity rhythms, it may render mice more susceptible to circadian perturbations when a second genetic manipulation is introduced. The most obvious scenario would be the use of the Cre driver strain to conditionally ablate a GOI in a specific cell population. A behavioral phenotype may arise because of the combined effects of the IRES(2)-Cre insertion and the loss of function of the GOI, rather than solely the latter. Hence, use of these Cre driver strains may confound the interpretation of behavioral phenotypes, even when both control mice—Avpcre/+ (or Vipcre/+) and GOIfl/fl—are employed.
In addition to the potential confound of the IRES(2)-Cre insertion on behavior, its demonstrated effects on expression of the neuropeptide(s) encoded by the targeted locus are problematic from a mechanistic perspective, particularly for conditional knockout studies. One may wrongly interpret the change in neuropeptide expression, or other phenotypes, to be the consequence of ablating a GOI, when in fact it may be the IRES(2)-Cre insertion that is producing such an effect. Given that most conditional knockout studies use GOIfl/fl mice, and not Cre-bearing GOI+/+ mice, as controls, they may be vulnerable to this erroneous interpretation.
Our examination of the Avp-IRES-Cre-D (JAX #023530) and Vip-IRES-Cre (JAX #010908) mouse models raises concerns about their use in conditional knockout studies of the circadian timing system as well as the interpretation of experimental findings. However, there are specific scenarios in which these models may be useful. For instance, one might ask how altering AVP or VIP expression (using either Cre/+ or Cre/Cre mice) affects certain organismal behaviors or physiological processes. These models may also be employed to target the expression of a transgene (e.g., fluorescent reporter gene, opto- or chemogenetic gene) to AVP- or VIP-ergic cells, using Cre/+ animals as controls where relevant to account for the change in neuropeptide expression caused by the IRES(2)-Cre insertion. Finally, it is important to note that other models that target the AVP or VIP cell populations may not suffer from the same drawbacks as the ones examined in this study (Mieda et al., 2015; Pei et al., 2014). For example, Mieda et al. (2015) used bacterial artificial chromosome transgenesis to drive Cre expression in Avp neurons, a method that does not require manipulation of the endogenous Avp locus.
Supplemental Material
Cheng_et_al_supplemental – Supplemental material for Limitations of the Avp-IRES2-Cre (JAX #023530) and Vip-IRES-Cre (JAX #010908) Models for Chronobiological Investigations
Supplemental material, Cheng_et_al_supplemental for Limitations of the Avp-IRES2-Cre (JAX #023530) and Vip-IRES-Cre (JAX #010908) Models for Chronobiological Investigations by Arthur H. Cheng, Samuel W. Fung and Hai-Ying Mary Cheng in Journal of Biological Rhythms
Footnotes
Acknowledgements
The authors thank L. Martin for sharing equipment. Many thanks to S. Hegazi, C. Lowden, S. Poulson, and J. Rios Garcia for advice and feedback. This work was supported by operating grants to H.-Y.M.C. from the Canadian Institutes of Health Research and the Natural Sciences and Engineering Research Council (NSERC) of Canada. H.-Y.M.C. was supported by an NSERC Tier II Canada Research Chair in Molecular Genetics of Biological Clocks. A.H.C. was supported by NSERC postgraduate scholarships.
Conflict of Interest Statement
The authors have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
