Abstract
The circadian clock is an evolutionarily conserved mechanism that generates the rhythmic expression of downstream genes. The core circadian clock drives the expression of clock-controlled genes, which in turn play critical roles in carrying out many rhythmic physiological processes. Nevertheless, the molecular mechanisms by which clock output genes orchestrate rhythmic signals from the brain to peripheral tissues are largely unknown. Here we explored the role of one rhythmic gene, Achilles, in regulating the rhythmic transcriptome in the fly head. Achilles is a clock-controlled gene in Drosophila that encodes a putative RNA-binding protein. Achilles expression is found in neurons throughout the fly brain using fluorescence in situ hybridization (FISH), and legacy data suggest it is not expressed in core clock neurons. Together, these observations argue against a role for Achilles in regulating the core clock. To assess its impact on circadian mRNA rhythms, we performed RNA sequencing (RNAseq) to compare the rhythmic transcriptomes of control flies and those with diminished Achilles expression in all neurons. Consistent with previous studies, we observe dramatic upregulation of immune response genes upon knock-down of Achilles. Furthermore, many circadian mRNAs lose their rhythmicity in Achilles knock-down flies, suggesting that a subset of the rhythmic transcriptome is regulated either directly or indirectly by Achilles. These Achilles-mediated rhythms are observed in genes involved in immune function and in neuronal signaling, including Prosap, Nemy and Jhl-21. A comparison of RNAseq data from control flies reveals that only 42.7% of clock-controlled genes in the fly brain are rhythmic in both males and females. As mRNA rhythms of core clock genes are largely invariant between the sexes, this observation suggests that sex-specific mechanisms are an important, and heretofore under-appreciated, regulator of the rhythmic transcriptome.
Our planet completes one rotation every 24 h, giving daily rhythms of temperature and light. Consequently, organisms have evolved endogenous biological timing systems to adapt to these predictable environmental changes. Termed circadian rhythms, every kingdom of life on Earth demonstrates 24-h rhythmic processes in behavior, metabolism, and physiology (Hastings et al., 2003). For example, the efficacy of immune function rhythmically oscillates, as both mice and flies show time-dependent survival after infection in a circadian manner (Labrecque and Cermakian, 2015; Lee and Edery, 2008; Scheiermann et al., 2018). These rhythmic physiological processes are delicately regulated to maintain their homeostasis.
At the molecular level, there is a self-sustaining core clock system that cycles every 24 h and forms the basis of circadian clocks (Allada and Chung, 2010; Hardin, 2005; Ko and Takahashi, 2006). It also drives the expression of hundreds of other genes. Some of these genes encode regulatory proteins that drive the rhythmic expression of further downstream genes. Named clock-controlled genes (CCGs), these circadian genes regulate many aspects of physiology and maintain organismal homeostasis in a tissue-specific manner (Ceriani et al., 2002; Hastings et al., 2003; Panda et al., 2002; Storch et al., 2002; Xu et al., 2011).
Extensive work has been done to identify rhythmic mRNAs in animals as well as cultured cell lines (Ceriani et al., 2002; Du et al., 2014; Duffield et al., 2002; Filichkin and Mockler, 2012; Hughes et al., 2009; Hughes et al., 2012; Menet et al., 2012; Menger et al., 2007; Ruben et al., 2018; Rund et al., 2011; Zhang et al., 2014). Further studies also focus on identifying post-transcriptional rhythms, like rhythmic proteins, post-translational modifications, and metabolites (Baggs and Green, 2003; Masri et al., 2016; Mauvoisin et al., 2014; Panda, 2016; Reddy et al., 2006; Robles et al., 2017; Wang et al., 2018). Nevertheless, little is known about the function of most rhythmic genes in an organismal setting.
In Drosophila, the principal oscillator resides in a cluster of about 150 dispersed yet interconnected neurons (Allada and Chung, 2010; Nitabach and Taghert, 2008). These neurons integrate environmental signals and transmit the synchronized timing cues to other brain regions as well as peripheral tissues (Albrecht, 2012; Guo et al., 2017; Liang et al., 2016; Yoshii et al., 2016).
Achilles (Achl) is a high-confidence CCG with an amplitude that rivals or exceeds that of most core CCGs in the brain (Claridge-Chang et al., 2001; Ceriani et al., 2002; Hughes et al., 2012; Keegan et al., 2007; Li et al., 2017; Lin et al., 2002; McDonald and Rosbash, 2001; Wijnen et al., 2006). It is predicted to encode an RNA-binding protein, making it a strong candidate for regulating downstream mRNA rhythms. For these reasons, we sought to determine its role in sculpting the rhythmic transcriptome. Previous analyses demonstrate that it plays critical roles in regulating the expression of immune response genes as well as in resistance towards pathogenic infection (Li et al., 2017). Moreover, Achl participates in signaling from neurons to immunological tissues, thus establishing a compelling model system for studying how neural circuits regulate rhythmic signals to the peripheral systems (Li et al., 2017).
To further explore how Achl regulates circadian output rhythms, we collected mRNA samples from fly heads every 2 h over 2 consecutive days and performed high-throughput RNAseq. We examined overall transcriptome rhythmicity in control and Achl RNAi flies. We focused on CCGs that lose their rhythmicity in Achl RNAi flies using 3 different analytical methods. As Achl regulates the immune system in both male and female flies, we further examined such candidate CCGs that lose rhythmicity in Achl RNAi in both males and females. The inclusion of samples from both males and females afforded us the opportunity to examine sex specificity in overall mRNA rhythmicity by comparing rhythmic genes identified in our control male and female flies. To our surprise, there is considerable sex-specificity in circadian mRNA rhythms, suggesting the importance of a heretofore under-studied biological variable in studies of circadian clock output.
Materials and Methods
Fly Stocks and Maintenance
Flies were maintained on Nutri-Fly BF food (Cat #66-113, Genesee Scientific; San Diego, CA) at 25°C in 12-12 h LD conditions. Humidity was maintained at roughly 50%. Groups of flies from which females were collected for RNAseq were maintained at 29°C for maximal Gal4 activity. Flies subject to RNAseq were entrained to LD conditions until they were 1-2 days old; they were released into free-running conditions of constant darkness (DD) 24 h before sample collection. All fly stocks used were acquired from the Bloomington stock center: Elav-Gal4 strain: P{w[+mW.hs]=GawB}elav[C155] w[1118]; P{w[+mC]=UAS-Dcr-2.D}2. CG17386 (Achl) RNAi strain: y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF01976}attP2. Control strain: y[1] v[1]; P{y[+t7.7] v[+t1.8]=UAS-GFP.VALIUM10}attP2. Elav-Gal4; UAS-Dcr2 virgins were crossed with UAS-Achl RNAi/control males to get pan-neuronal knock-down of Achl in their F1 offspring.
Fluorescence In Situ Hybridization and Immunostaining
Adult male fly heads collected at ZT8 or ZT20 were fixed in 4% formaldehyde in PBS for 30 min on a shaker at room temperature, then transferred to 4°C Dent’s fixative (80% methanol, 20% DMSO) and stored at −20°C overnight. The samples were washed in 100 mM Tris/HCl (pH 7.4) containing100 mM NaCl on a shaker for 1 h at room temperature, then transferred to 15% fish gelatin in 15% sucrose for overnight incubation on a shaker at room temperature. This step was repeated the following day with 25% fish gelatin in 15% sucrose. Samples were positioned dorsal side up in plastic molds containing 20% fish gelatin in 15% sucrose, flash-frozen with dry ice and stored at −80°C. The blocks were sectioned at 10-μm-thick along the dorsal-ventral axis, mounted on Superfrost Plus slides (Fisher Scientific, 12-550-15) and dried. The sections were re-permeabilized with 70% ethanol for 10 min, and then incubated with 300 μL rat monoclonal anti-ELAV primary antibody (1:1000) (DSHB, 7E8A10) at 4°C overnight. After washing, Alexa488-conjugated anti-rat secondary antibody (1:500) (A-11006; Thermo Fisher Scientific, Waltham, MA) was added to the slide and incubated for 2 h at 4°C. Fluorescence in situ hybridization (FISH) probes for Achl mRNA were used as previously described (Cvetkovska et al., 2013). Forty-eight custom Stellaris FISH Probes conjugated to Quasar670 (SMF-1065-5) were designed against the Achilles mRNA sequence by utilizing the Stellaris RNA FISH Probe Designer (Biosearch Technologies, Inc.; Petaluma, CA) available online at www.biosearchtech.com/stellarisdesigner (Version 4.2). Probe Sequences are as follows: CACGGGACTCACAAGGAACG CTTGGTCTTGTCCATTGAAG ATCGATTCCATTTGGTGGAG GGAGCAGAGGGTGAAGTATC TCGGCTATGATGTGATTGTC CAAATTCGCCTCACTCAAGA ATCTGGTGCTTACGGAACTT GGAGAAGCATCCAGAGTGAA GGTGAGGAACGAACGATGGG TGTGTCTATCTCATTGGGTC TGGGAATGAGTGAGGGATCT GGAGAAGTAGAACTCCACGG TGTTTCAGCAGGAATGCATC TTTAAGGCTGACGAAACCCT ACATCATTCAGCTCGATCTT AGGGCATGGTTTCATCGAAA CAGATCACAGGCAACGATCG CTTTTCGATGGTTAGCTTGT CATGGCGAGAATAGATCCGA AATGCGTATGAGGGCAATCT ATGAACTGACGCACATCCAC ACTAGAGCACACTCCTTTTG GGCGGAAGAGGATTCTAAGT CCATCTCGTAGACCTGAAAG TAGTTGGCATCGTTGTAGTA TTGTTGCGCTTGAGTTTGAA CCGTTTGTTGGTAGTAACTG TGATACGGATTGGCATGGTA CTGGGACCATAGCCGAAAAA TTGTCGATGTGTTGCTGTAT GATATAGGCGAAACATCGCC ATTACCGCCTGAATTGACAG CAGACTGCTTATACCAGACA ACTTCTGGAGTGTAGTTCTG CCTTCGTGACATCGAAGGAT CTCTCCGAACAGTTCGAATC TCTGTCGTCTGAGCAAAGTC CCGAGGTCTTTCTGTAACTA GAGAAGGTTCGCTCGAACTG TCGACGGGCGACGATAGAAG TCCCGCTCGATATTGAAATC TGCTGGAAGTTGTTGGAGTA CTGAGCATTATCATATGCCG ATATACGTCTTCCTCCGATA CAAACGAGCAGCGACGATCG TGTAGTTGCTCAGGAAGCTA CAAACGAACCGGAACGCGAC CGATATGGTCTGTCCGAATT.
The probes were diluted to 125 nM with hybridization buffer and 300 μL was added to each slide, which was placed in a humidified chamber and incubated at 37°C in the dark for 12 h. The slide was washed with wash buffer provided by the Stellaris kit (Biosearch Technologies, SMF-WA1-60) and incubated at 37°C in the dark for 30 min. Two more washes were performed, and the slide was mounted with SlowFade Diamond Antifade Mountant with DAPI (Thermo Fisher Scientific, S36973). Fluorescence microscopy was performed using an Olympus laser-scanning confocal microscope FV1000. Images were acquired using 20× (N.A. 0.85) or 60× oil objective (N.A. 1.4).
RNA Preparation
Flies between 3 and 5 days old were used for RNAseq. Seven fly heads per sample were manually dissected in PBS, transferred into 100 μL of Trizol (Life Technologies; Carlsbad, CA) and homogenized using RNase-free pestles (Thermo Fisher Scientific). After 5 min of incubation at room temperature with an additional 400 μL of Trizol, total RNA was prepared with phase lock gels (5Prime; Gaithersburg, Maryland) and an RNeasy mini kit (Qiagen; Hilden, Germany), according to the manufacturer’s protocol. RNA quantity and quality were assessed using a Qubit 2.0 Fluorometer (Life Technologies) and a 2100 Bioanalyzer (Agilent Technologies; Santa Clara, CA).
Library Preparation and RNA Sequencing
RNAseq libraries were prepared from 200 ng total RNA per sample using TruSeq Stranded mRNA LT Sample Prep Kit (Illumina; San Diego, CA) following the manufacturer’s protocol, with 13 rounds of PCR amplification. Libraries were quantified and qualified using a Qubit 2.0 Fluorometer (Life Technologies) and a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA). Prepared libraries with unique indexes were pooled and sequenced on an Illumina HiSeq-4000 platform with H4K Single End 50 Cycle run type at the University of Michigan DNA Sequencing Core.
RNAseq Alignment and Quantification
Spliced Transcripts Alignment to a Reference (STAR) algorithm v2.5.3a (Dobin et al., 2013) was used to align sequenced reads to the genome and transcriptome of Drosophila melanogaster (build dm6) with a 2-pass mapping strategy and following the standard protocol. Option “–quantMode TranscriptomeSAM” was used for the second 2-pass run.
RNAseq by Expectation Maximization (RSEM) package v1.3.0 was used to quantify transcript-level and gene-level expression after 2-pass STAR alignments (Li and Dewey, 2011).
Statistical Analyses
Analysis of rhythmicity
JTK_CYCLE is an algorithm implemented in R that analyzes the rhythmicity of time-series data (Hughes et al., 2010). JTK_Cycle was performed using a period length window precisely equal to 24 h. TPM values were used for the analysis. Genes with median TPM < 1 were excluded from the JTK_CYCLE analysis. LAG indicates the phase of expression, and amplitude was calculated as previously described (Miyazaki et al., 2011).
DODR analysis
DODR is an algorithm implemented in R that analyzes the differential rhythmicity between groups of time-series data (Thaben and Westermark, 2016). All methods were used in DODR. TPM values were used for analysis. P values obtained from the robustDODR method, and the lowest p values obtained after multiple testing correction were chosen for further analysis.
LimoRhyde analysis
LimoRhyde is an algorithm implemented in R that analyzes the differential rhythmicity between groups of time-series data (Singer and Hughey, 2018). Log transformed TPM + 1 values were used for analysis. When performing LimoRhyde analysis, RAIN was used to identify rhythmicity. Differential expression and differential rhythmicity were identified using Limma.
GO term analysis
GO enrichment analysis was performed using DAVID software available online (Eden et al., 2007; Eden et al., 2009). We show results named “simplified GO enrichment analysis,” for which we manually curated GO terms when several enriched terms were very similar.
Data expression
Heatmaps were generated using Matlab. Expression levels for each sample within the group were median-normalized before generating heatmaps. Volcano plots were generated during LimoRhyde analysis.
Differential rhythmicity analysis
DODR and LimoRhyde analyses were performed as previously described (Thaben and Westermark, 2016; Singer and Hughey, 2018), resulting in p- and q-values to determine differential rhythmicity. In JTK_CYCLE, we compared p values from each condition and considered p < 0.001 in one condition and p > 0.01 in the other condition as differential rhythmicity.
Analysis of Legacy Datasets
Data pertaining to Achl expression in the brain were downloaded from NCBI Gene Expression Omnibus (GEO) under accession number GSE36108 (Hughes et al., 2012). Achl expression data in the fatbody were downloaded from NCBI GEO under accession number GSE24503 (Xu et al., 2011). Achl expression data from single-cell RNAseq in the brain were visualized online at http://scope.aertslab.org (Davie et al., 2018). Achl expression in the core clock neurons was visualized as shown previously (Kula-Eversole et al., 2010) and downloaded from NCBI Gene Expression Omnibus (GEO) under accession number GSE77451 (Abruzzi et al., 2017).
Data and Resource Availability
All custom software is available on demand. As recommended for circadian profiling experiments (Hughes et al., 2017), all raw sequencing data as well as qualified expression data have been submitted to the NCBI Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo/) under accession number: GSE120100.
Results
To knock-down Achl specifically in neurons, we used the UAS-GAL4 system in flies (Brand and Perrimon, 1993) and crossed Elav-Gal4; UAS-Dcr2 flies with TRiP UAS-Achl RNAi flies to drive pan-neuronal knock-down of Achl in their offspring (Perkins et al., 2009). Previously, we found that this manipulation dramatically upregulates various genes involved in immune defense pathways (Li et al., 2017). Elav is transiently expressed in glial cells during early development, raising the possibility that glia may be involved in the effects of Achl on the immune system (Berger et al., 2007). Therefore, we performed FISH to examine the expression pattern of Achl. As shown in Figure 1, Achl was expressed in a diffuse, neuron-specific manner, overlapping with ELAV-positive cells in the adult brain. There was no obvious enrichment of Achl expression in core clock neurons (Liang et al., 2018; Schlichting et al., 2016). These results were consistent in brains collected at both ZT8 and ZT20, and the FISH negative control confirmed the specificity of the neuron-specific expression pattern (Supplemental Figure S1). These observations are in agreement with recent single-cell RNAseq data from the brain that suggest that Achl is expressed in neurons but not glial cells (Davie et al., 2018). Moreover, our results showing that Achl is not expressed in all neurons in the brain (Figure 1) are also in agreement with previously published microarray/RNAseq data from sorted neuron populations (Abruzzi et al., 2017; Kula-Eversole et al., 2010). We further note that Achl mRNA cycles robustly in both fly heads and manually dissected brains (Hughes et al. 2012), decreasing the likelihood that its circadian function is exclusive to expression in the compound eye, a major contributor to RNA samples collected from fly heads. Overall, these data support the conclusion that Achl is expressed in most neurons; although, it is most likely not expressed in core clock neurons.
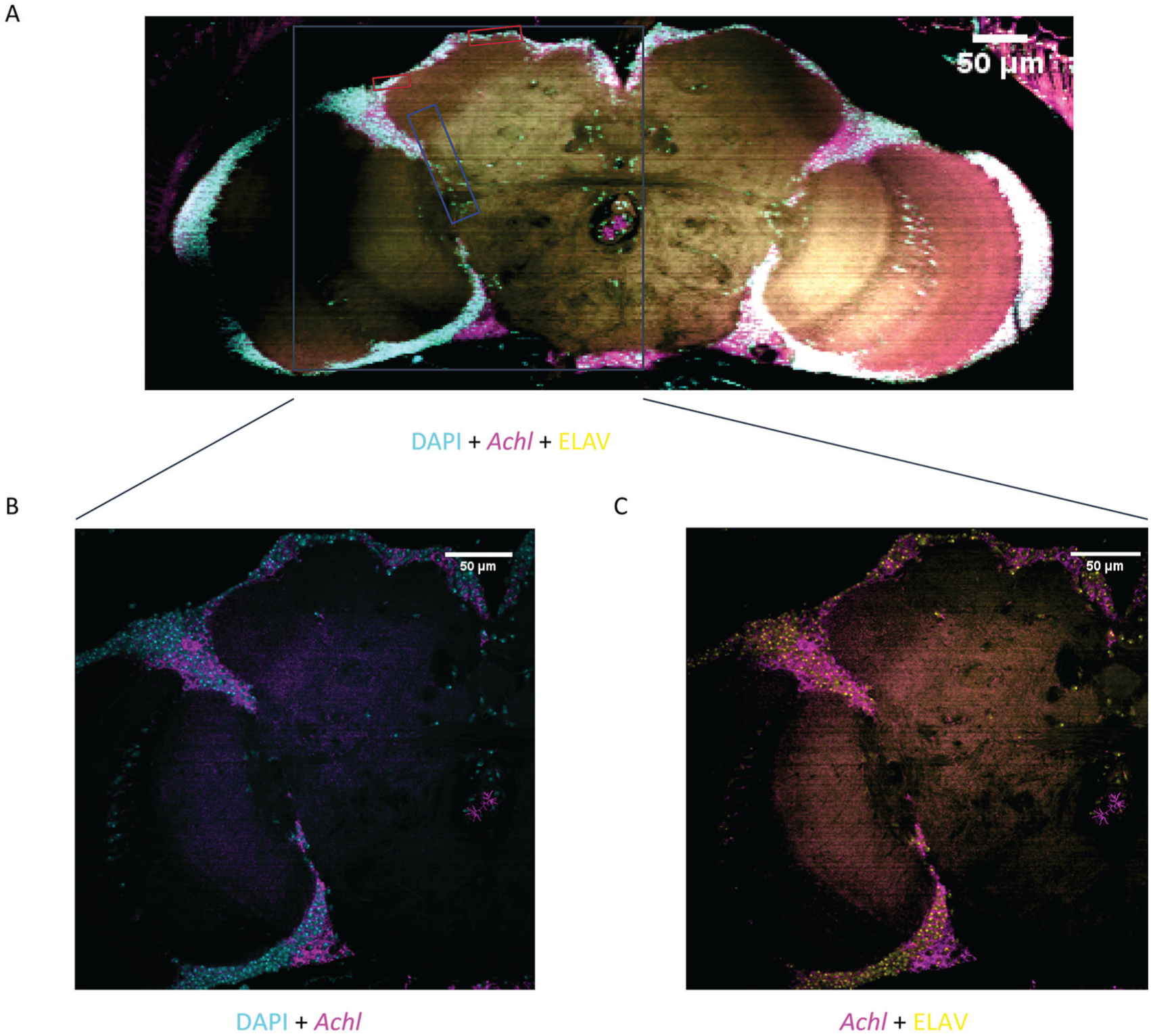
Achl FISH and ELAV immunostaining in Drosophila brain suggests that Achl is expressed broadly in most neurons. (A) Whole male fly brains were collected at ZT20 for staining. Squares indicate regions where clock neurons reside: red squares indicate the region of dorsal neurons; the blue square indicates the region of lateral neurons. The gray squared region was zoomed in with a 60× objective. (B) Visualization of DAPI and Achl co-staining under 60× objective. (C) Visualization of Achl and ELAV co-staining under 60× objective. DAPI staining is in cyan, Achl staining is in magenta, and ELAV staining is in yellow. Note that whereas most ELAV-positive neurons express Achl, some ELAV-positive neurons show no detectable Achl expression.
To identify downstream CCGs regulated by Achl, we collected both control and Achl RNAi fly heads in constant conditions every 2 h for 2 consecutive days from both sexes and performed RNAseq. On average, 16.4 million reads per sample were obtained from RNAseq (Supplemental Table S1). This sampling resolution and RNAseq depth is adequate to detect CCGs (Hughes et al., 2017; Li et al., 2015). Among these reads, about 98% were uniquely mapped to the Drosophila genome/transcriptome.
We confirmed that our RNAi effectively reduced Achl expression at all time points (Figure 2A). Although low amplitude 24-h rhythmicity persists, Achl expression is lower at every time point in RNAi flies than the trough level in the corresponding negative controls. We also looked at the expression pattern of the CCGs. In agreement with previous observations, CCGs maintain their rhythms (Figure 2B and Supplemental Table S2) (Li et al., 2017). To identify genes differentially expressed in Achl knock-down flies, we performed LimoRhyde analysis followed by Limma (Singer and Hughey, 2018). In agreement with the transcriptional profiling at ZT2 (Li et al., 2017), Achl RNAi dramatically upregulated immune response genes (Figure 2C, Figure 2D, Supplemental Table S3 and Supplemental Table S4).
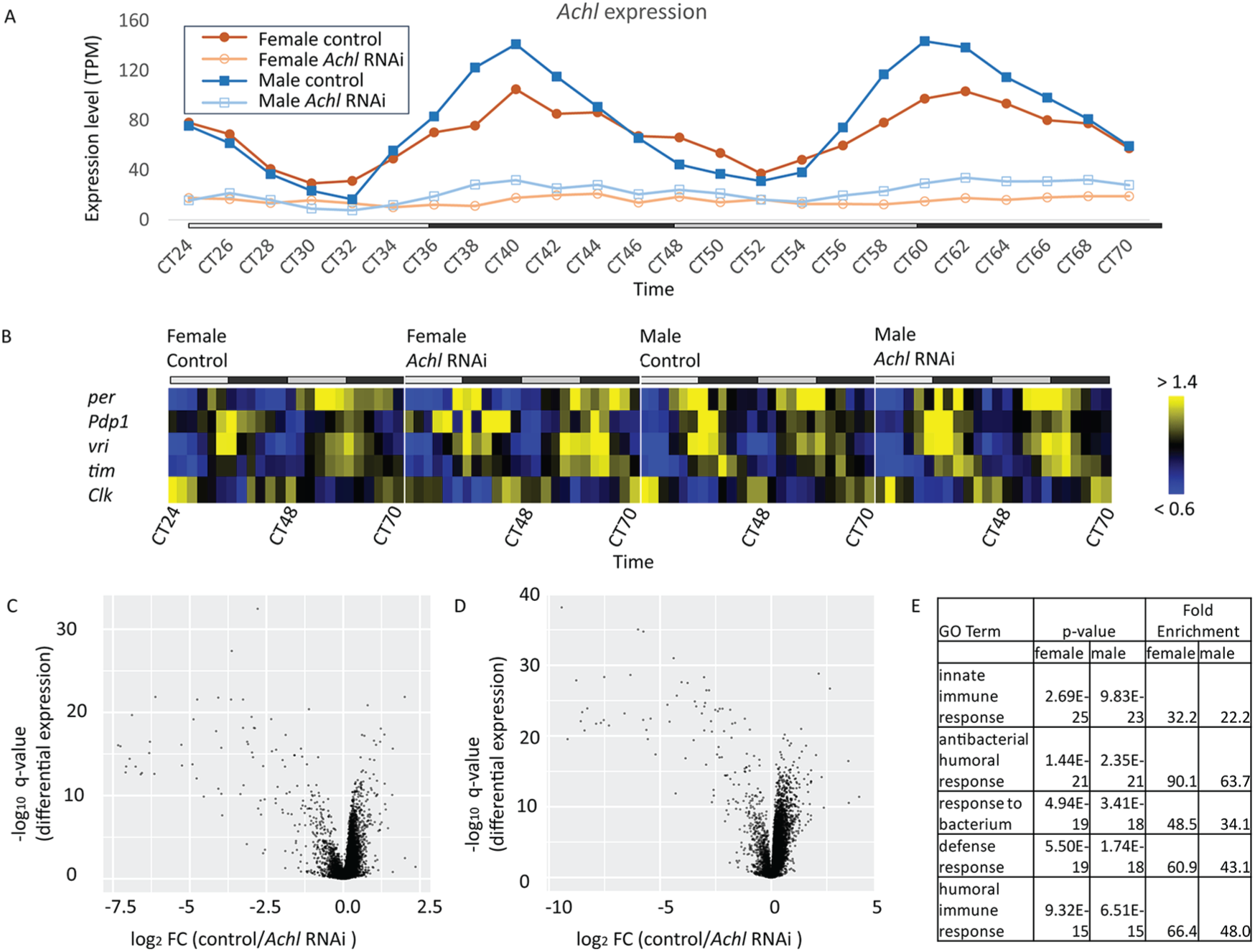
Achl RNAi upregulates immune response genes with no effect on core clock genes (CCGs). (A) Achl expression is downregulated at all time points in Achl RNAi flies (i.e., Elav-Gal4/+; UAS-Dcr2/TRiP UAS-Achl RNAi line). (B) Heatmap of the median-normalized expression of key CCGs suggests that the core clock is not affected in Achl RNAi flies. Yellow indicates higher expression, and blue indicates lower expression (intensity scale on the right). The top gray and black bars represent subjective day and night, respectively. Detailed JTK_CYCLE analysis results can be found in Supplemental Table S2. (C and D) Volcano plot of RNAseq expression data in female (C) and male (D) flies showing that Achl specifically regulates the immune system in the fly head. Each dot represents a single gene. (E) Manually curated GO enrichment of genes differentially expressed in control and Achl RNAi flies. A complete GO table is available as Supplemental Table S4.
We used JTK_CYCLE (Hughes et al., 2010) to identify circadian genes in all 4 RNAseq datasets: 1) female control, 2) female Achl RNAi, 3) male control, and 4) male Achl RNAi flies. Since q-values are calculated based on the distribution of p values within the dataset and depend on the total number of cycling genes within the dataset, we used p value thresholds for our inter-group analysis, with q-values as a supplemental statistic to estimate the total number of cycling genes in any given genotype. Using a p-value threshold of less than 0.001 (corresponding q-value of 0.06 in male control dataset), we found 94 rhythmic genes in female control flies, 41 rhythmic genes in female Achl RNAi flies, 163 rhythmic genes in male control flies, and 107 rhythmic genes in male Achl RNAi flies (Supplemental Figure S2). Overall, fewer genes are rhythmic in flies of both sexes with knocked-down Achl expression.
We used 3 different methods to compare the rhythmicity pattern of all transcripts expressed in control and Achl RNAi flies. First, we compared cycling parameters estimated by JTK_CYCLE. We picked a p value of <0.001 (corresponding to a q-value of <0.06 in male, control flies) as the rhythmic threshold, and a more relaxed p value of >0.01 as the non-rhythmic threshold. Different thresholds here were chosen to avoid false positives that are likely rhythmic in Achl RNAi flies but fall just outside the statistical threshold for being declared “rhythmic.” Second, we ran the DODR algorithm to detect differential rhythmicity (Supplemental Table S5) (Thaben and Westermark, 2016). Third, we ran the LimoRhyde algorithm to detect differential rhythmicity (Table S6) (Singer and Hughey, 2018). We compared differentially rhythmic genes under the 3 conditions for each sex. All 3 analytical methods agree that a substantial number of genes have altered rhythmicity in Achl knock-down flies (Supplemental Figure S3); although, we note that there was weak overlap between the 3 methods. The amplitude of Achl diminished in Achl knock-down flies in both males and females (Figure 2A), whereas Jhl-21 lost rhythmicity in both sexes (Supplemental Figure S4). Nevertheless, both DODR and LimoRhyde detected Achl and Jhl-21 as differentially rhythmic in females but not in males. We provide summary statistics for DODR and LimoRhyde in Supplemental Tables S5 and S6.
With the listed cutoff thresholds, 62 genes lost rhythmicity in female flies in Achl RNAi (Figure 3A) and 81 genes in males (Figure 3B). We examined the phase distribution of these genes and found different phase enrichment in male and female flies. The enriched phase is early morning in males but roughly 8 h later in females (Supplemental Figure S5A). This observation suggests sex-specific differences in the generation of circadian transcriptional outputs, motivating our subsequent analysis of sex-specific RNA rhythmicity. Furthermore, because Achl regulates the immune system in both sexes (Li et al., 2017), we speculate that there are both sex-invariant and sex-dependent functions performed by Achl.
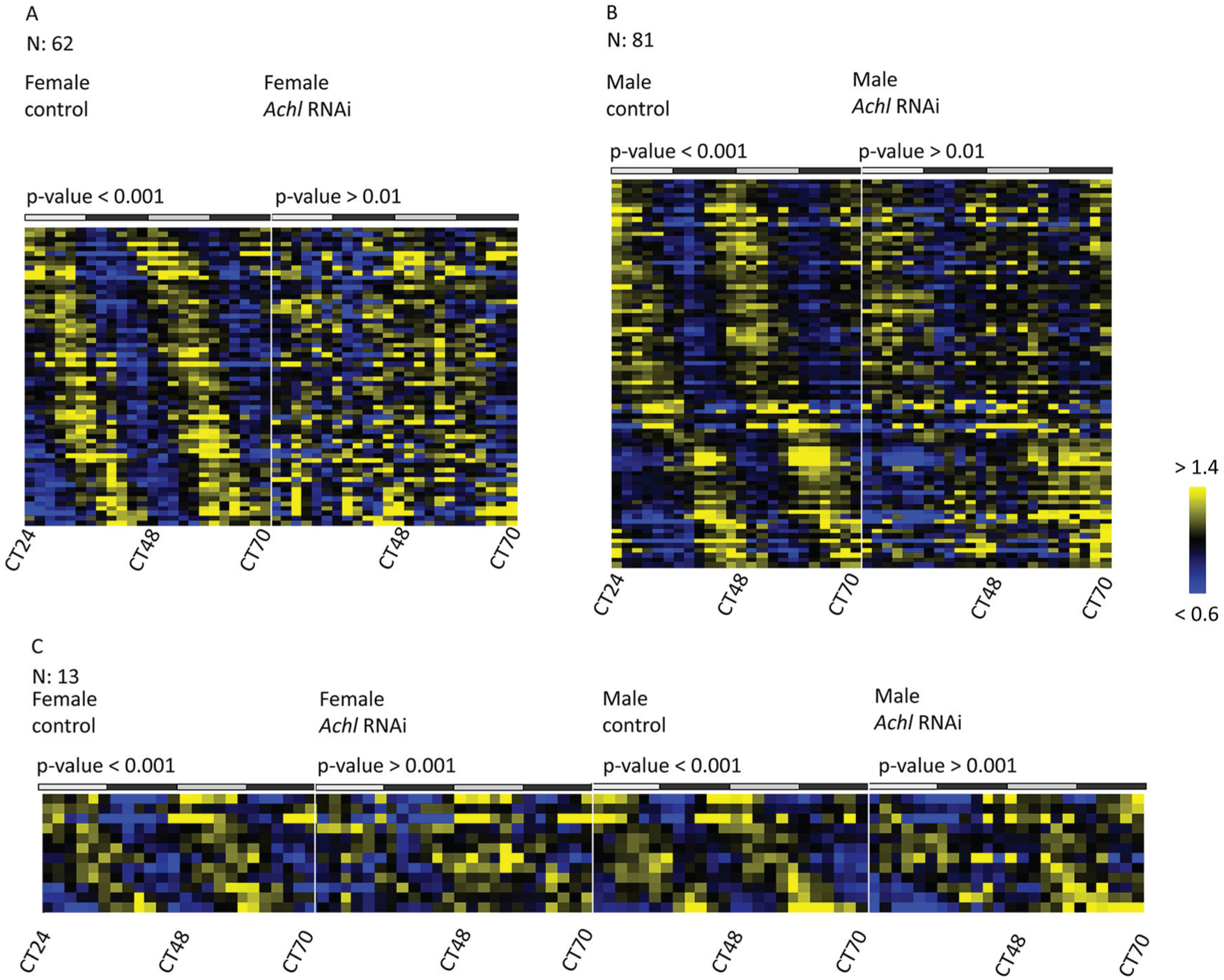
Dozens of clock-controlled genes (CCGs) become arrhythmic in Achl RNAi flies. (A) Genes losing rhythmicity in female flies upon Achl RNAi. (B) Genes losing rhythmicity in male flies upon Achl RNAi. (C) Genes losing rhythmicity in both sexes upon Achl RNAi. The top gray and black bars represent subjective day and night, respectively. Yellow indicates higher expression, and blue indicates lower expression (intensity scale on the right).
To identify CCGs under the control of Achl regardless of sex, we picked a p-value threshold of 0.001 and a –log10 p-value foldchange threshold of 1.2. We found 12 genes that lost rhythmicity in both male and female flies: Prosap, CG16978, CG14329, GstE2, CG5326, JhI-21, nemy, CG5789, inaC, eys, CG2121 and Cpr72Ec (Figure 3C, Supplemental Table S7 and a representative example in Supplemental Figure S4). We found no obvious bias in the phase distribution for these genes between control male and female flies; although, the peak expression of these genes was restricted to a range between CT0 and CT16 (Supplemental Figure S5B).
In addition to genes that lost rhythmicity in Achl RNAi, we found a small group of genes that gained rhythmicity in Achl RNAi: 14 genes in female flies (lowest JTK q-value: 0.004) and 31 genes in male flies (lowest JTK q-value: 0.0005). We did not find any gene that gained rhythmicity in Achl RNAi in both sexes. As there was no consistency between the sexes in terms of gain, we speculate that these results are either false-positives or the result of an unknown compensatory mechanism downstream of Achl signaling.
As mentioned above, we observed sex-specific differences in the response of the rhythmic transcriptome to Achl knock-down. Most circadian transcriptome studies have been performed exclusively in male animals (Hughes et al., 2007; Hughes et al., 2009; Keegan et al., 2007; Zhang et al., 2014). To our knowledge, there are no direct comparisons of rhythmic transcriptomes between males and females. Therefore, we compared the overall mRNA rhythmicity pattern in male and female control flies. CCGs were rhythmic in both sexes with essentially identical phases; though, with greater statistical significance in male than female flies (Figure 2B). Consistent with this, there were fewer total rhythmic genes identified by JTK_CYCLE in females than in males (Supplemental Figures S2A and S2C). To compare these identified rhythmic genes in male and female flies, we again set a p-value threshold of less than 0.001 (q < 0.06 in males) and a more relaxed p value of < 0.01 in the other sex as being rhythmic. We found 90 (43%) rhythmic genes in both male and female flies (Figure 4A and Supplemental Table S8), which were enriched in genes involved in circadian rhythms, metabolism, signaling transmission, and light response (Supplemental Figure S6A and Supplemental Table S9). There was no obvious absolute difference in the phase distribution for rhythmic genes common to male and female flies (Supplemental Figure S6B). We also used DODR and LimoRhyde algorithms to detect differential rhythmicity in male and female flies (Supplemental Table S10). As previously, there was weak overlap between the 3 methods; however, all 3 analytical methods agreed that a substantial number of genes have altered rhythmicity in male and female flies (Supplemental Figure S7).
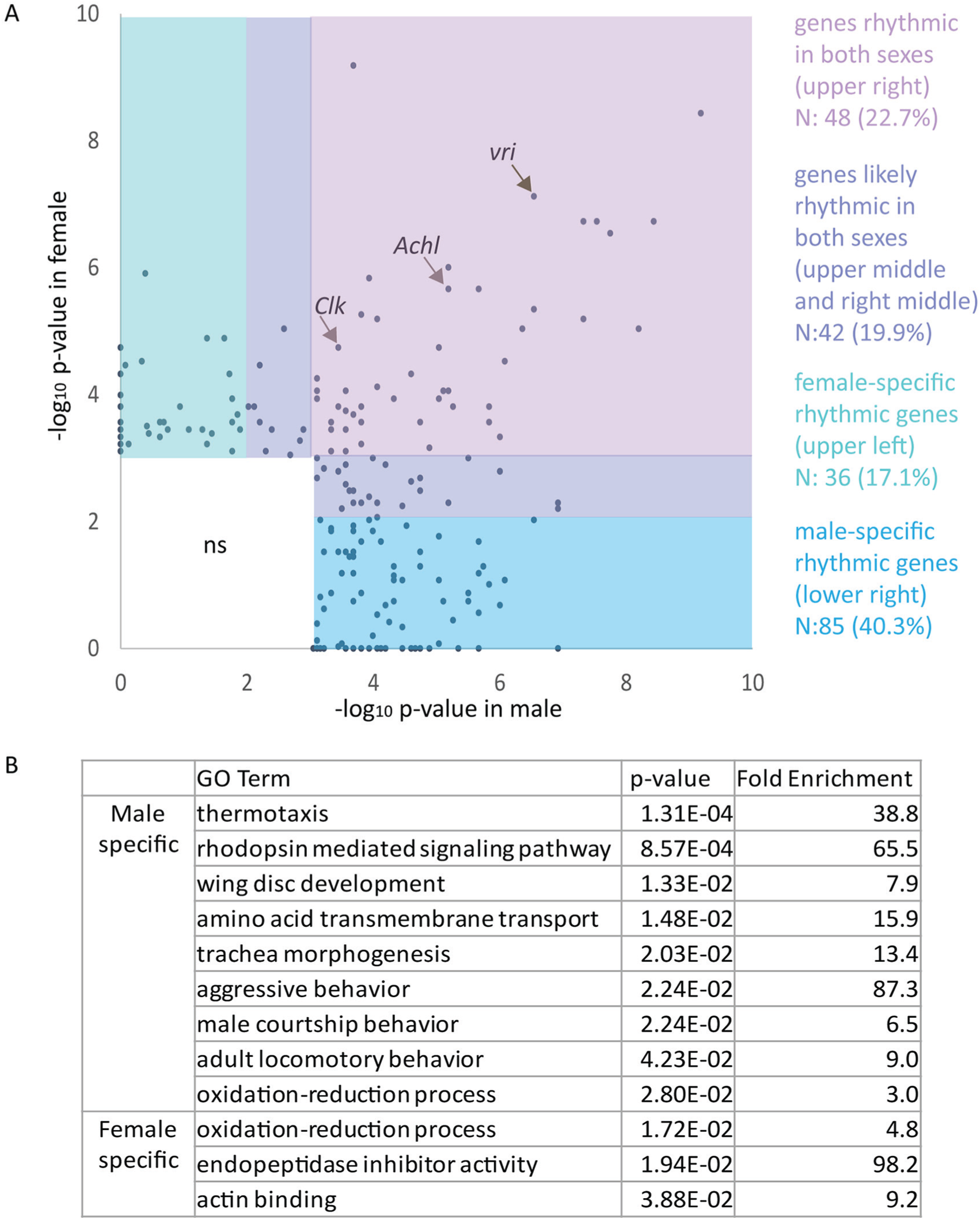
Over half of all rhythmic genes in the fly head are sex-specific. (A) A scatter plot of JTK_CYCLE p values showing different groups of genes identified as rhythmic in males and females. Pink: genes rhythmic in both sexes. Purple: genes likely rhythmic in both sexes. Green: female-specific rhythmic genes. Blue: male-specific rhythmic genes. Detailed gene names and JTK_CYCLE p values are available in Supplemental Table S8. (B) Manually curated GO enrichment analysis of genes rhythmic specifically in either male or female flies. A complete GO table is available as Supplemental Table S11.
We then examined genes that were rhythmic in only one sex. In all, 121 (57%) genes were rhythmic in only one sex (85 genes are rhythmic specifically in males and 36 in females). Most of these were expressed at detectable levels in both sexes (N = 118; 97.5%), suggesting that sex-specific rhythmicity is independent of overall transcription level. We performed GO analysis for genes that were rhythmic specifically in one sex (Figure 4B and Supplemental Table S11). We found several categories of rhythmic genes enriched in males, such as courtship behavior and aggressive behavior, trachea and wing disk development, and light/environmental signaling processing processes. Courtship and aggression are notable sexually dimorphic behaviors, suggesting a mechanism by which the circadian clock may regulate these behaviors. However, all other enriched terms that are more rhythmic in males are involved in both males and females. We note that, although the GO term “oxidation-reduction process” is enriched in both male and female flies, there are different rhythmic genes involved in this process for both sexes. Overall, these results indicate considerable variation between the CCGs found in males and females.
Discussion
Although the core time-keeping mechanism is understood in great biochemical and genetic detail (Allada and Chung, 2010; Hastings et al., 2003; Ko and Takahashi, 2006), little is known about the mechanism by which individual rhythmic genes drive downstream circadian output mRNAs. Achl is a robust CCG with a large amplitude. It is expressed broadly and exclusively in neurons, and we believe that it acts indirectly to alter the sensitivity of anti-microbial peptide-secreting cells in the fatbody (Li et al., 2017). The mechanisms by which neurons regulate gene expression in immunological tissues such as the fatbody are presently unknown. Furthermore, we speculate that circadian rhythmicity of Achl or its downstream targets ultimately influences daily rhythms in the sensitivity of organismal immune responses (Lee and Edery, 2008). We note that more sophisticated manipulations of Achl expression or its target genes may resolve these questions.
RNA-binding and -processing proteins, such as LARK, Nocturnin and CIRBP, likely orchestrate the rhythmicity and function of downstream targets (Gotic et al., 2016; Huang et al., 2007; Stubblefield et al., 2012). Achl knockdown is therefore an excellent model for studying how RNA-binding proteins influence the rhythmic transcriptome. Using RNAseq, we identified dozens of CCGs whose mRNA rhythmicity depended on Achl expression. Consistent with this observation, we find that neuron-specific knock-down of Achl is sufficient to reduce the total number of rhythmic genes in the fly head. These data strongly suggest that Achl is a key upstream CCG and is partially responsible for establishing the overall pattern of normal rhythmic gene expression.
Rhythmicity was lost in over a dozen genes in Achl RNAi flies in both males and females (Figure 3). Because Achl loss-of-function has immunological consequences in both sexes (Li et al., 2017), we speculate that these genes are the likely mediators of Achl’s effects on the immune system. Some of these genes are involved in the immune response, albeit with presently undefined functions, such as CG16978, CG14329, and CG2121 (De Gregorio et al., 2001; De Gregorio et al., 2002; Pal et al., 2008; Paparazzo et al., 2015). We also noticed an enrichment of synapse- and neurotransmitter-related genes. Prosap encodes a major scaffolding protein of the neuronal postsynaptic density; it plays roles in synapse functions in a dose-dependent manner (Harris et al., 2016). Nemy plays a role in the regulation of neurotransmitter secretion as well as synaptic plasticity (Knight et al., 2015). Jhl-21 (Supplemental Figure S4) is an amino acid transporter that modulates the secretion of neurotransmitters including serotonin and glutamate (Ro et al., 2016; Ziegler et al., 2016). Moreover, JhI-21 regulates the immune system, as knock-down of JhI-21 diminishes the induction of antimicrobial peptides like Dpt. It is also involved in resistance towards both fungi and bacteria (Jin et al., 2008; Paik et al., 2017). All these data suggest that Achl-mediated regulation of the immune system relies largely on neuronal signaling, most likely through multiple independent circuits. These data suggest the necessity of mapping the output pathways using neurobiological tools, which will help define the detailed signaling process from Achl-expressing neurons to immune tissues.
We note that 2 genes, inaC and eys, are both highly expressed in the compound eye and were differentially rhythmic in Achl knock-down flies. As Achl is known to be rhythmic in photoreceptor cells (Wijnen et al., 2006), this result suggests that Achl may play a role in visual perception. Since Achl is rhythmically expressed in brains isolated from the compound eye (Hughes et al. 2012), we favor the interpretation that Achl’s regulation of immune pathways is independent of its role in the eye; although, we do not exclude the possibility that a more complex mechanism is at work.
It is unknown whether genes losing rhythmicity after Achl knock-down are directly or indirectly regulated by Achl. Recent technical advances, like RIP-seq and hyperTRIBE, have made it easier to identify targets of RNA-binding protein (Rahman et al., 2018; Zambelli and Pavesi, 2015). Combining our RNAseq data with the identification of Achl target mRNAs will improve our understanding of the entire regulatory process.
Our data show weak overlap between CCGs losing rhythmicity in male and female Achl RNAi flies (Figure 3). Moreover, most CCGs regulated by Achl in both sexes show different phase distributions (Supplemental Figure S5A), suggesting that Achl may regulate its targets in a sex-specific manner. As the expression level Achl and its rhythmic profile are largely invariant between the sexes (Figure 2A), it is of great interest to determine the mechanism by which Achl influences the sex-specific rhythmic transcriptome.
Females and males have different circadian behavioral outputs (Astafev et al., 2017; Krizo and Mintz, 2014; Mong et al., 2011); although, the underlying molecular mechanisms are not well understood. Most published studies regarding rhythmic genes are biased towards males, creating an unfortunate blind spot in the circadian literature. To address this gap, we provide the first time-dependent transcriptome analysis of both male and female flies. Male flies have more rhythmic genes than female flies, and over half (57.3%) of all rhythmic genes are specific to one sex. Moreover, male-specific rhythmic genes are enriched in ontology terms related to courtship and behavior (Figure 4). These observations suggest the compelling possibility that circadian control of sexually dimorphic behavior may be established at the level of rhythmic mRNA accumulation (Menet et al., 2012; Rodriguez et al., 2013). Since the expression of CCGs is largely similar between males and females (Figure 2B), these results indicate that sexual differentiation exerts enormous control over circadian clock output pathways through mechanisms currently unknown. Moreover, these results provide strong motivation for testing whether sexual dimorphism in clock mRNA output is conserved in higher animals, particularly humans.
Supplemental Material
Figure_S1._Negative_control_for_in-situ_hybridization – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Figure_S1._Negative_control_for_in-situ_hybridization for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Figure_S2._Achl_loss-of-function_decreases_the_number_of_rhythmic_genes_in_male_and_female_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Figure_S2._Achl_loss-of-function_decreases_the_number_of_rhythmic_genes_in_male_and_female_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Figure_S3._overlap_between_three_methods_used_to_determine_differential_rhythmicity_in_control_and_Achl_RNAi_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Figure_S3._overlap_between_three_methods_used_to_determine_differential_rhythmicity_in_control_and_Achl_RNAi_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Figure_S4.__Jhl-21_is_a_CCG_regulated_by_Achl – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Figure_S4.__Jhl-21_is_a_CCG_regulated_by_Achl for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Figure_S5._Phase_distribution_of_genes_lose_rhythmicity_in_Achl_RNAi – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Figure_S5._Phase_distribution_of_genes_lose_rhythmicity_in_Achl_RNAi for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Figure_S6._Comparison_of_genes_rhythmic_in_both_male_and_female_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Figure_S6._Comparison_of_genes_rhythmic_in_both_male_and_female_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Figure_S7._overlap_between_three_methods_used_to_determine_differential_rhythmicity_in_male_and_female_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Figure_S7._overlap_between_three_methods_used_to_determine_differential_rhythmicity_in_male_and_female_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
JBR830845_Supplemental_Material_CLN – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, JBR830845_Supplemental_Material_CLN for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S1._STAR_statistics – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S1._STAR_statistics for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S10._Differential_rhythmicity_in_male_and_female_flies_detected_by_DODR_and_LimoRhyde – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S10._Differential_rhythmicity_in_male_and_female_flies_detected_by_DODR_and_LimoRhyde for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S11._Full_Gene_Ontology_enrichment_table_for_simplified_table_shown_in_Figure_4B – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S11._Full_Gene_Ontology_enrichment_table_for_simplified_table_shown_in_Figure_4B for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S2._Detailed_JTK_results_for_core_clock_genes_displayed_in_Figure_2B – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S2._Detailed_JTK_results_for_core_clock_genes_displayed_in_Figure_2B for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S3._Genes_differentially_expressed_in_control_and_RNAi_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S3._Genes_differentially_expressed_in_control_and_RNAi_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S4._Full_Gene_Ontology_enrichment_table_for_genes_differentially_expressed_in_control_and_Achl_RNAi_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S4._Full_Gene_Ontology_enrichment_table_for_genes_differentially_expressed_in_control_and_Achl_RNAi_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S5._DODR_statistics_of_differential_rhythmicity_in_control_and_Achl_RNAi_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S5._DODR_statistics_of_differential_rhythmicity_in_control_and_Achl_RNAi_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S6._Limorhyde_statistics_of_differential_rhythmicity_in_control_and_Achl_RNAi_flies – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S6._Limorhyde_statistics_of_differential_rhythmicity_in_control_and_Achl_RNAi_flies for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S7._JTK_CYCLE_statistics_of_genes_lose_rhythmicity_in_Achl_RNAi_flies_in_both_sexes – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S7._JTK_CYCLE_statistics_of_genes_lose_rhythmicity_in_Achl_RNAi_flies_in_both_sexes for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S8._Detailed_gene_names_and_JTK_CYCLE_p-value_for_Figure_4A – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S8._Detailed_gene_names_and_JTK_CYCLE_p-value_for_Figure_4A for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Supplemental Material
Table_S9._Full_Gene_Ontology_enrichment_table_for_simplified_table_shown_in_Figure_S6 – Supplemental material for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila
Supplemental material, Table_S9._Full_Gene_Ontology_enrichment_table_for_simplified_table_shown_in_Figure_S6 for Achilles-Mediated and Sex-Specific Regulation of Circadian mRNA Rhythms in Drosophila by Jiajia Li, Renee Yin Yu, Farida Emran, Brian E. Chen, and Michael E. Hughes in Journal of Biological Rhythms
Footnotes
Acknowledgements
We thank Jeanne Geskes, Robert Lyons, and the University of Michigan DNA sequencing core facility for assistance with next-generation sequencing. We thank members of the Hughes and Chen laboratories for helpful discussion and technical support throughout this project as well as revisions on the manuscript. We thank especially Erin Terry, Ayesha Baig, Jason Bedwinek, Christy Hoffman and Kylie Mink for technical support during the course of this work. We thank Dr. Nicholas Lahens (University of Pennsylvania) and Dr. Malcolm Tobias (Washington University in St. Louis) for bioinformatics technical support. We thank the Holtzman, Nerbonne, Skeath and Taghert labs (Washington University in St. Louis) for help with equipment and reagents. Jiajia Li was supported by dissertation fellowship (University of Missouri-St. Louis). Work in the Hughes Lab is supported by an award from NIAMS (1R21AR069266).
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
