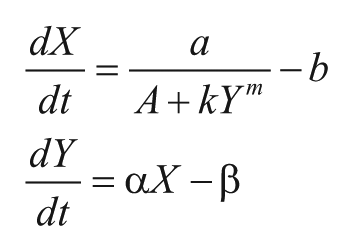Abstract

Since Bünning’s seminal work on the physiological clock (Bünning, 1967), the study of circadian and biological rhythms has developed into a well-established interdisciplinary research field, where the use of genetic and molecular biology methods has given us a wealth of new information about the biological clock’s “ticking” components; the 2017 Nobel Prize in Physiology or Medicine was awarded to Jeffrey C. Hall, Michael Rosbash, and Michael W. Young for their discoveries of molecular mechanisms controlling the circadian rhythm.
Despite these advancements, we still know surprisingly little about how organisms can keep time—how, for example, plants measure day-night lengths or keep their circadian period compensated against environmental influences. To understand, for example, what makes a set of chemical reactions oscillate, we need kinetic/mathematical techniques that allow us to describe and analyze them in more quantitative (kinetic/mathematical) terms. Here Forger’s book comes in.
Forger’s book is aimed at giving readers a practical guide, a mathematical toolbox, so to speak, on how to set up and deal with the equations describing oscillatory biochemical systems. The book is divided into three parts with a total of 10 chapters. A nice aspect of the book is that each chapter contains a Frontiers section, which covers timely topics by discussing or commenting on techniques but also deals with unresolved issues. Another nice feature is that each chapter contains Matlab code to illustrate a certain aspect covered in the chapter. Each chapter ends with exercises.
Part I (Models, chapters 2-4) covers basic modeling techniques, how to set up rate equations, model examples, Michaelis-Menten dynamics, the Goodwin oscillator, the Hodgkin-Huxley equations, and models for neuronal dynamics. Part I also covers stochasticity by using the Gillespie method and takes up the conditions when feedback loops start to oscillate. There is a slight confusion in terminology when treating Michaelis-Menten kinetics. The way the Michaelis-Menten equation is derived on page 51 is based on a steady-state approximation of the enzyme-substrate complex and not by a rapid equilibrium between substrate and the enzyme-substrate complex. There is no need to assume a rapid binding between the enzyme and the substrate.
In Part II (Behaviors, chapters 5-7), Forger gives an outline of biochemical oscillators and classifies them as type 1 and type 2 dependent based on the type of bifurcation (SNIC or Hopf, respectively). This part also covers phase response curves and coupled oscillators. There is a rich literature on the kinetics and “taxonomy” of chemical oscillators, unfortunately not mentioned by Forger, which provides alternative classifications of chemical/biochemical oscillators. The chemical oscillator literature (e.g., Eiswirth et al., 2007; Epstein and Showalter, 1996; to start with) provides a nice supplement to Forger’s book.
The final part of the book (Analysis and Computation, chapters 8-10) deals with problems of model parametrization and optimization and multiscale modeling. Statistical and computational tools for model building are reviewed and applied, such as maximum likelihood, least squares, and Kalman filtering.
I very much like the use of the Goodwin oscillator to illustrate different topics throughout the book, even that chaos can be generated! Forger provides a historical note about the Goodwin oscillator on page 108, about which I would like to comment. While Goodwin assumed in his original equations an inhibition cooperativity of one, later analysis showed that there is a problem with such an assumption. While it is correct, as Forger (and several others) have pointed out, that the cooperativity m needs to be larger than 8 in order to obtain oscillations, the condition m > 8 applies to models only where degradation terms are described by first-order reactions with respect to the degrading species. However, in Goodwin’s initial formulation (1963), the Goodwin equations were described with zero-order degradation terms, that is, of the form
with m = 1. The two variables X and Y show conservative oscillations. 1 The equations can be extended to show limit-cycle oscillations, even for m = 1, when three (or more) dynamical variables are included, in addition to the use of Michaelis-Menten degradation terms (Kurosawa and Iwasa, 2002; see also exercise 7 on page 129 of Forger’s book). The change from zero-order to first-order degradation terms (while maintaining the m = 1 condition) was clearly a mistake made in the later papers by Goodwin, but it illustrates the influence that degradation kinetics have on the system’s oscillatory behavior.
Forger’s book is an excellent choice for students/researchers who are interested to model biological rhythms and already have a working knowledge of differential equations/Matlab, chemical kinetics, and/or nonlinear dynamics. Readers with a more (molecular) biology background may find the book more difficult at certain stages but may find it useful to first cover some of the mathematical basics from other texts as those recommended by the author.