Abstract
The mammalian circadian clock system, which includes the master clock and peripheral clocks, times different biological processes in reaction to external cues, such as the light-dark cycle. However, the chronobiology of prokaryotic cells is less well understood, except for in cyanobacteria. The recent blooming of gut microbiome research has revealed a critical role for the trillions of microbes residing in the vertebrate gut in determining both health and disease in the host. The question of whether the gut microbiome exhibits circadian oscillation and how it synchronizes with the host circadian clock has attracted considerable interest. In this review, we discuss the time-of-day–dependent compositional and functional structure within the gut microbial community, how it is regulated by the host, and how it reciprocally influences the host circadian clock.
The Human Gut Microbiota
The human body is home to a large collection of microbes, including bacteria, archaea, viruses, and unicellular eukaryotes, collectively referred to as the microbiota. Among them, bacteria are more intensely studied. It has been estimated that the human intestinal tract harbors over 1014 bacteria grouped into about 1000 species (Bengmark, 1998; Ley et al., 2006; Zhu et al., 2011), accounting for up to 98% of the intestinal microbiota (Manichanh et al., 2012). This collection of microbes has a combined genome size exceeding that of humans by at least 2 orders of magnitude (Ley et al., 2006).
Although virtually all surfaces of the human body are colonized by bacteria, the most heavily colonized organ is the gastrointestinal tract, and bacterial density in the colon reaches 1011 to 1012 per mL (Whitman et al., 1998). Intestinal bacteria contribute greatly to the host physiology: in the maintenance of the mucosal structure (Stappenbeck et al., 2002); host defense against pathogens (Bengmark, 1998); the activation of host immune responses (Littman and Pamer, 2011); the fermentation of dietary fiber (Sekirov et al., 2010), vitamin production, including biotin and vitamin K (Stevens and Hume, 1998); the metabolism of peptides and proteins (Farthing, 2004); and the metabolism of xenobiotics, including drugs (Illing, 1981; Bakke and Gustafsson, 1986; Rowland et al., 1986).
Despite controversy over the presence of an in utero microbiome (Aagaard et al., 2014; Lauder et al., 2016), colonization of the gut begins at birth. The early colonizers in infant guts are the microbes they first encounter, either from their mothers’ vaginal or skin microbiome, depending on the mode of their delivery (Adlerberth and Wold, 2009; Dominguez-Bello et al., 2010). Later, the gut microbiota goes through temporal compositional and functional changes, and fully resembles the adult microbiota by 2.5 years of age (Koenig et al., 2011). Thereafter, the gut microbiota remains stable throughout adulthood. Nevertheless, the composition is subject to changes induced by the use of antibiotics (Eckburg et al., 2005; Jernberg et al., 2010), dramatic dietary changes (David et al., 2014), and concomitant disease (Wu et al., 2013; Zhao, 2013). Eventually, as we reach old age, the microbial structure in the gut shifts, leading to an altered abundance of some taxa (Claesson et al., 2011; Claesson et al., 2012).
The gut microbiota is dominated by Bacteroidetes and Firmicutes (Eckburg et al., 2005; Turnbaugh et al., 2007; Arumugam et al., 2011). However, significant inter-individual variability in microbial composition exists. While common species have been found across individuals by both 16S and shotgun sequencing (Tap et al., 2009; Qin et al., 2010), variability in abundance of common species ranges from 12- to 2,200-fold (Qin et al., 2010). This variability stems from the combined influence of host genetics (Benson et al., 2010), ageing (Biagi et al., 2010; Agans et al., 2011), geographic locations (Yatsunenko et al., 2012), lifestyle (Annalisa et al., 2014), diet (Wu et al., 2011; Zoetendal and de Vos, 2014), and pet ownership (Song et al., 2013); many of these factors are confounded.
A balanced gut microbial composition is critical to host physiology, and compositional disruption has been associated with many diseases. For instance, obesity is associated with a significant reduction in Bacteroidetes and a corresponding increase of Firmicutes (Ley et al., 2005; Turnbaugh et al., 2008; Zhao, 2013). Partially explained by host genetics, the etiology of Crohn’s disease also has an environmental component: patients have altered microbial structure compared with their healthy identical twin (Dicksved et al., 2008). Likewise, adherent-invasive E. coli derived from spondyloarthritis patients confer aspects of the human inflammatory bowel disease phenotype in mouse models (Viladomiu et al., 2017). In addition, dysbiosis has been linked to malnutrition (Million et al., 2016), ulcerative colitis (Frank et al., 2007; Garrett et al., 2010), neurological disorders (Gonzalez et al., 2011; Sampson et al., 2016), autoimmune diseases (Wen et al., 2008; Giongo et al., 2011), hypertension (Li et al., 2017; Marques et al., 2017), thrombosis and atherosclerosis (Wang et al., 2011; Koeth et al., 2013; Tang et al., 2013; Zhu et al., 2016), and cancer (Lampe, 2008; Zhu et al., 2011; Chen et al., 2012).
Circadian Rhythms
The rotation of the earth results in the oscillation of light during the 24-h cycle. Organisms adapted to this cycle by developing circadian rhythms, an endogenous and entrainable mechanism that times daily events such as feeding, temperature, sleep-wakefulness, hormone secretion, and metabolic homeostasis (Green et al., 2008; Saini et al., 2011).
In mammals, the rhythm is controlled by a master clock that resides in the suprachiasmatic nucleus (SCN) of the hypothalamus. It is regulated tightly by interlocking feedback and feed-forward loops at transcriptional, translational, and post-translational levels (Bass and Takahashi, 2010; Xu et al., 2013) (Figure 1). Classically, the BMAL-1/CLOCK heterodimer binds to E-box motifs in the promoter regions of Per and Cry genes, forming the positive limb of the clock. PER and CRY dimerize and relocate to the nucleus, where they repress the activity of BMAL-1/CLOCK complex, and inhibit the transcription of downstream genes, including their own. This forms the negative limb of the clock. The retinoic acid receptor (RAR)-related orphan receptor gene (Ror) and the nuclear receptor-encoding Rev-erbα gene form additional positive and negative loops.
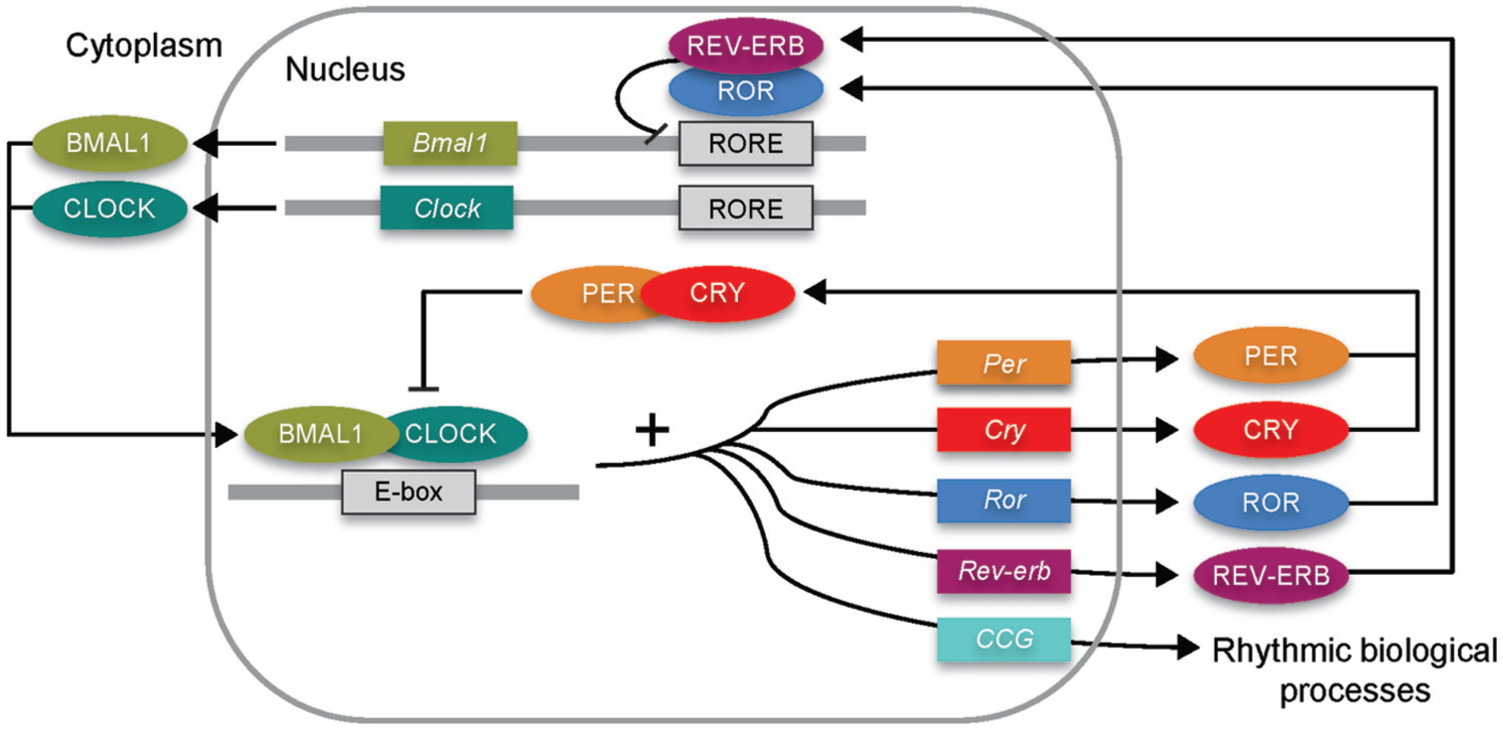
The mammalian molecular clock.
Besides the master clock, peripheral clocks exist in most, if not all tissues (Mohawk et al., 2012), such as the liver, gut, heart, and retina (Lee et al., 2001; Reppert and Weaver, 2002; Young, 2006; Froy and Chapnik, 2007). Cultured cells and explants from these tissues exhibit robust circadian oscillations in gene expression (Panda et al., 2002; Storch et al., 2002; Yamamoto et al., 2004; Yoo et al., 2004). For example, intestinal cells not only transcribe central clock genes but also translate central clock proteins with circadian periodicity that are out of phase with the SCN (Pardini et al., 2005; Hoogerwerf et al., 2007; ladek et al., 2007). The gut clock, dependently or independently of the SCN, regulates a number of gut functions, such as nutrient absorption, xenobiotic and endobiotic detoxification, and colonic motility (Gachon et al., 2006; Pan and Hussain, 2009; Hoogerwerf et al., 2010). This peripheral circadian clock also feeds back to the SCN via humoral or metabolic cues. For instance, the peripheral clock in adipocytes influences central feeding behavior by modulating lipid signaling to the feeding centers of the brain (Paschos et al., 2012).
The circadian clock network is fundamental to maintaining many physiological processes, especially metabolism and immune function. Disruption of this system is associated with various disease phenotypes in patients and in mice, such as obesity (Froy, 2010), diabetes (Turek et al., 2005; Marcheva et al., 2010), atherosclerosis (McAlpine and Swirski, 2016), cancer (Masri et al., 2015), neurodegeneration (Videnovic et al., 2014), and ageing (Froy, 2011). Many human diseases vary in their expression or severity with clock time, including asthma, myocardial infarction and stroke, suicide and sudden cardiac death. However, the role of the molecular clock in these phenomena remains to be elucidated (Yang et al., 2013).
Circadian rhythms also exist in prokaryotes, and are well-characterized in Cyanobacteria (Grobbelaar et al., 1986; Hut and Beersma, 2011). Since free-living bacteria are exposed to daily cycles of light and temperature, their circadian clock allows them to anticipate and prepare for changes in environmental conditions. In Cyanobacteria, a cluster of 3 genes – kaiA, kaiB, and kaiC – encodes central oscillators with a periodicity of ~24 h, and this is essential for the regulation of nitrogen fixation, photosynthesis, and cell division (Ishiura et al., 1998). Incubating KaiC with KaiA, KaiB, and adenosine triphosphate results in self-sustainable oscillation of KaiC phosphorylation in vitro (Nakajima et al., 2005; Tomita et al., 2005). This non-transcriptional rhythm is sufficient to generate oscillations in vivo, yet the presence of transcriptional-translational feedback regulation stabilizes the clock (Teng et al., 2013; Johnson et al., 2017). Recently, the KaiA, KaiB, and KaiC oscillator has been successfully transplanted to the noncircadian bacterium Escherichia coli and can drive transcriptional output for 3 days (Chen et al., 2015).
The Circadian Rhythmicity of Gut Microbiome
Although not exposed to the light-dark cycle, the gut bacteria are still subject to environmental changes, such as nutrient availability, and the level of host-derived, anti-microbial peptides and autoantibodies. Indeed, over the past 3 years, evidence has emerged revealing compositional oscillation of the fecal microbiota during the 24-h light-dark cycle – the gut microbiome exhibits compositional and functional structures at different times of day (Thaiss et al., 2014; Zarrinpar et al., 2014; Liang et al., 2015).
Thaiss et al. (2014) reported the time-of-day–dependent composition of the murine fecal microbiota (Thaiss et al., 2014) (Figure 2a). More than 15% of detected OTUs (Operational Taxonomical Units) undergo daily oscillation in their relative abundance, including Lactobacillus reuteri, Dehalobacterium spp., and other species belonging to Clostridiales, Lactobacillales, and Bacteroidales. Additional support was added later by another 2 groups: Zarrinpar et al. (2014) reported that 17% of OTUs are cyclical (Zarrinpar et al., 2014), and we reported oscillation of the relative abundance of genera belonging to Bacteroidetes, Firmicutes, and Proteobacteria (Liang et al., 2015). In addition to the fluctuation in relative taxonomic abundance, the total biomass of the gut bacterial community in mice is higher during the dark phase when the animals are active (Liang et al., 2015). The discovery of microbial oscillation in the gut was detectable despite the dramatic variability of microbiota composition from different cages and animal facilities (Deloris Alexander et al., 2006; Rausch et al., 2016).
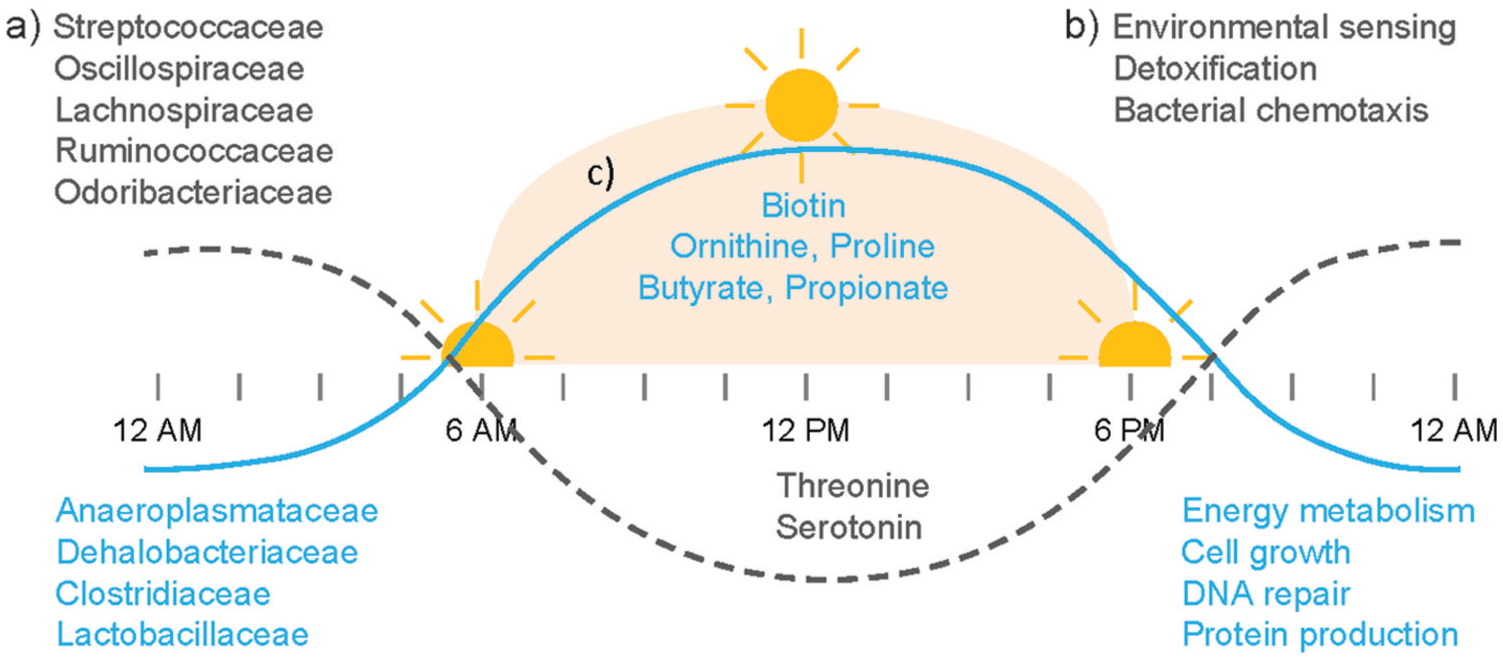
Circadian oscillation of the gut microbiome, such as a) bacteria, b) functional pathways, and c) bacterial metabolites. Dotted line, peaks at resting/dark phase. Solid line, peaks at active/light phase.
These compositional oscillations imply translation into oscillations of functional consequence (Thaiss et al., 2014) (Figure 2b). Thus, about 23% of identified KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways encoded in bacterial genomes fluctuate with a 24-h rhythm. For example, genes for pathways involved in energy metabolism, DNA repair, and cell growth peak in abundance during the dark (active) phase, whereas those involved in detoxification, motility, and environmental sensing peak during the light (resting) phase (Figure 2b). In humans, ~10% of OTUs and ~20% of KEGG pathways oscillate (Thaiss et al., 2014). Temporal changes in such pathways may assist the bacteria to anticipate changes in gastrointestinal function. For example, pathways involved in energy metabolism, such as nucleotide, carbohydrate, and amino acid metabolism are enriched in the active phase (Thaiss et al., 2014). It is possible that some bacteria can anticipate and prepare for the increased availability of nutrients, such that they are ready to utilize them for growth. On the other hand, the abundance of genes involved in chemotaxis and flagellar assembly peaks in the resting phase (Thaiss et al., 2016) (Figure 2b). This drives increased bacterial penetration into the mucus layer, facilitating the use of mucin as a nutrient.
Furthermore, such daily oscillation of pathways leads to temporal changes in the level of microbiota-generated metabolites (Figure 2c). For example, the concentrations of butyrate and propionate are elevated in the cecum and feces in mice at the beginning of the light phase, and remain lower throughout the rest of the day (Leone et al., 2015). Metabolomics analyses have revealed significant oscillations across diverse chemical groups, such as serotonin, ergothioneine, lysine, xylose, glucose, isovalerate, and others (Thaiss et al., 2016). The biosynthesis of biotin, a bacteria-derived vitamin, shows rhythmicity in the conversion of dethiobiotin and the subsequent production of biotin.
Daily Oscillation in the Gut Microbiome is Regulated by Host
While the gut microbiota composition is highly influenced by the host, this relationship is bi-directional.
First, the daily oscillation of microbial structure requires a functional circadian clock in the host (Figure 3). For example, disruption of the backward limb of the clock in Per1/2−/− mice abolishes daily oscillations in both bacterial abundance and metagenomics gene contents (Thaiss et al., 2014), and disruption of the forward limb of the clock by Bmal-1 deletion, which leads to arrhythmicity without pleiotropic effects, also abolishes the circadian rhythmicity of gut microbiota composition in mice (Liang et al., 2015). Mice subjected to simulated jet-lag shift the composition of their gut microbiome, enhancing dysbiosis (Thaiss et al., 2014). Collectively, the host circadian clock is fundamental to communicating cues to the gut microbiota, leading to synchronization, co-ordination, and adaptation between the host and the bacteria. What remains to be elucidated is whether the effects of Per1/2 and Bmal-1 deletions, and jet-lag are mediated through the SCN and/or the intestinal clock system.
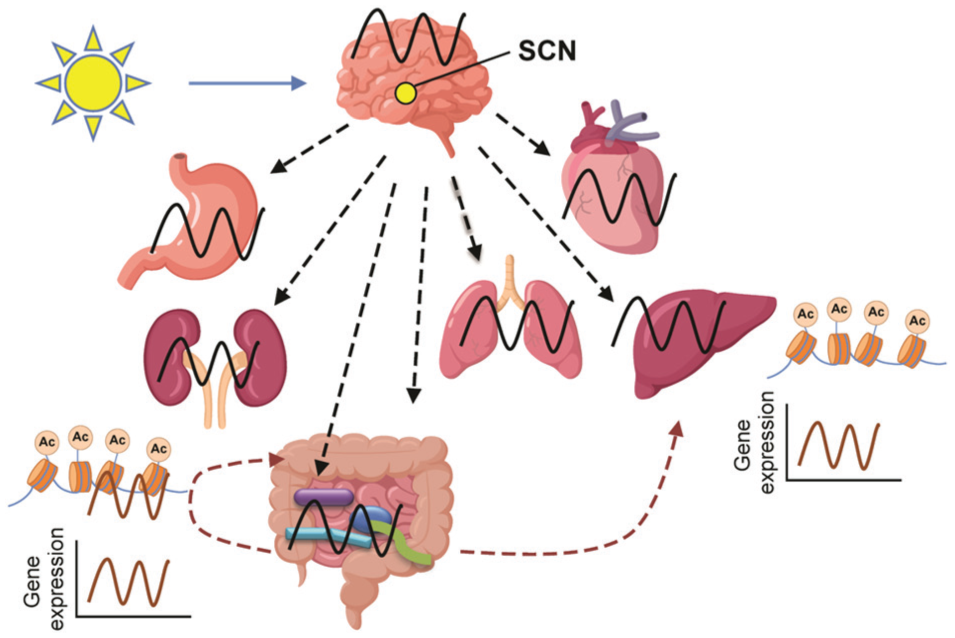
The mutual influence between the host clock network and the gut microbiome. The host central clock integrates light signals and regulates the circadian clock in the stomach, kidney, intestine, lung, liver, and heart, as well as in the gut microbial community (black oscillation). The gut microbes reciprocally influence circadian clock gene expression and post-translational regulation (acetylation) in the intestine and liver. Ac, acetylation.
A recent study has revealed host-derived melatonin and body temperature as possible direct regulators of gut microbiome oscillation (Paulose et al., 2016). Here, a clinical isolate of the Enterobacter aerogenes strain expressed circadian behavior in swarming and motility. This was enhanced by physiological levels of melatonin, a rhythmically controlled circulating hormone, in a temperature-compensated fashion. It is likely that bacteria within the gut microbiome have endogenous circadian clocks that interact with the host circadian clock via signaling molecules, such as hormones.
Second, diet and feeding influences the rhythmicity of the gut microbial composition. For instance, mice that have ad libitum access to a high-fat diet lack daily fluctuation in the relative abundances of Bacteroidetes and Firmicutes. However, consolidation of access to a high-fat diet to the dark phase only partially restores microbial oscillation (Zarrinpar et al., 2014). Moreover, restricting feeding to either the light phase or the dark phase rescues the loss of fluctuations in the gut microbial composition of Per1/2−/− mice (Thaiss et al., 2014). Therefore, not only the dietary components but also the timing of meals play an important role in the circadian oscillation of the gut microbiome. This may reflect the time-dependent metabolic profile and timed availability of nutrients to the gut bacteria. Nonetheless, since the circadian rhythms of host behavior and physiology are entrainable by daily cycles of restricted feeding (Stephan, 1984), it remains to be determined whether the restoration of microbial oscillation is due to a direct influence of feeding, restoration of the host circadian clock, or a combination of the two.
Third, the host gender interacts with the host clock to regulate gut microbial oscillation (Liang et al., 2015). Although both male and female mice show circadian rhythmicity, oscillation in females is more pronounced than in males. However, Bmal-1 deficiency abolishes the rhythmicity irrespective of sex, indicating the dominant regulatory role of a functional host circadian clock.
How these integrated signals communicate with the gut microbiome largely remains to be elucidated. Additionally, given the control over the gut microbial community by mucosal immunity—as exemplified by circadian oscillation of defensins, such as Cryptdin (Froy et al., 2005)—it is intuitive that the clocks of the host immune cells, such as macrophages, lymphocytes and dendritic cells, also contribute to regulating oscillation of the gut microbiome (Santisteban et al., 2017).
The Gut Microbiome Influences the Peripheral Host Circadian Clock
Metabolism, now recognized as influenced both by factors in the host and in the microbiome, is intertwined with the circadian clock. For example, Toll-like receptors (TLRs) on gut epithelial cells can recognize bacterial metabolites and integrate these signals into the rhythmic activities of c-Jun N-terminal kinase (JNK) and inhibitor of nuclear factor kappa-B kinase subunit beta (IKKβ), preventing peroxisome proliferator-activated receptor alpha (PPARα)-mediated activation of the nuclear receptor RevErbα and subsequent feedback suppression of the circadian clock (Mukherji et al., 2013). Additionally, ablation of the gut microbiome induces alterations in the expression of circadian genes, such as RORα, RevErbα, Bmal1, Per1, and Per2, resulting in ileal corticosterone overproduction and a prediabetic phenotype. A similar disrupted gene expression profile has been observed in hepatocytes from mice lacking a gut microbiome, leading to shifts in downstream clock-controlled gene expression, especially nuclear receptors involved in metabolic regulation (Leone et al., 2015; Montagner et al., 2016). Changes in cholesterol, free fatty acids, bilirubin, and lactate reflect a disordered hepatic metabolome in the absence of gut microbiota (Montagner et al., 2016). Dietary fiber intake and microbiome-derived acetate also regulate circadian gene expression in heart and kidney, which might play a role in the prevention of hypertension (Marques et al., 2017).
Rhythmic bacterial adherence or proximity to the mucus layer serves as an environmental factor to regulate the oscillating epigenome and transcriptome in colonocytes (Thaiss et al., 2016). Additionally, the gut microbiome regulates the circadian transcriptome in the liver, likely through the daily oscillation of microbiota-derived metabolites, including lipids, amino acids, carbohydrates, vitamins, nucleotides, and xenobiotics (Thaiss et al., 2016). Leone et al. suggested that butyrate, a short-chain fatty acid produced by bacterial fermentation of fiber in the colon, functions as a histone deacetylase inhibitor in liver to regulate oscillating chromatin modifications in hepatocytes (Leone et al., 2015). Furthermore, deoxycholic acid (DCA) and chenodeoxycholic acid (CDCA), both of which are unconjugated bile acids derived from the gut microbiome, altered the expression level of clock and clock-regulated genes in ileum, colon, and liver of mice when administered orally (Govindarajan et al., 2016). Thus, DCA and CDCA significantly decreased the expression of Bmal1 and increased that of Per2 and Per3 in the ileum and the liver.
The host immune system exhibits circadian fluctuations (Curtis et al., 2014) that coincide with time-dependent susceptibility to infections. Mice challenged with pneumococci or Coxsackie B3 virus in the early morning survive longer than those challenged later in the day (Feigin et al., 1969; Feigin et al., 1972). A recent study reported better colonization by the oral pathogen, Salmonella enterica, during the resting phase than the active phase and a stronger pro-inflammatory response of the mice (Bellet et al., 2013). Viral titers of herpes and influenza A were higher when mice were inoculated at the beginning of resting phase, and both replication and infection were enhanced by Bmal1 depletion (Edgar et al., 2016). Given that gut microbiota maintain mucosal barrier function and enhance innate and adaptive immunity (Kamada et al., 2013), it is important to learn more about whether and how the rhythmicity of the gut microbial community would influence the rhythmicity of the host immune homeostasis and defense. A recently described mouse intestine culture system (Yissachar et al., 2017) may facilitate investigations of the microbial influence on intestinal circadian rhythms.
Perspective
Evidence for oscillation of the gut microbiome and for a bidirectional interaction between the gut microbiome rhythm and the host circadian clock has now been well-established (Mukherji et al., 2013; Thaiss et al., 2014; Leone et al., 2015; Liang et al., 2015; Montagner et al., 2016; Thaiss et al., 2016). Therefore, it is very important to consider the timing of sample collection in all studies of the gut microbiome.
The gut microbiome has emerged as a regulator of brain development, function, and behavior (Collins et al., 2012; Cryan and Dinan, 2012; Sampson and Mazmanian, 2015), including microglial maturation and function (Erny et al., 2015) and astrocyte activity (Rothhammer et al., 2016). Additionally, alterations to the gut microbiome have been associated with disorders in the central nervous system (CNS), such as inflammation, autism spectrum disorder, stroke, spinal cord injury, depression, pediatric autoimmune neuropsychiatric disorders associated with streptococcal infection, Alzheimer’s disease, and Parkinson’s disease (Gonzalez et al., 2011; Sampson et al., 2016; Fung et al., 2017). However, the functional consequences of such associations have yet to be elucidated.
Many intriguing questions about the mechanisms regulating interactions between the host and gut microbiome clocks remain to be answered. For example, what are the molecular mechanisms driving circadian oscillations in bacteria and other organisms in the gut? Are there immunological, neural, and/or hormonal regulators? Does the gut microbiome influence the host circadian clock in the SCN? What are the signaling molecules emanating from these interactions that are communicating with the host circadian clocks, both in the SCN and in peripheral tissues? What are the host signals downstream of the SCN or the peripheral clocks that talk to organisms in the gut? What are the influences of the presence and/or rhythmicity of the gut microbiome on the SCN clock? How does the host integrate signals from the gut microbial metabolome and from its own metabolome to modulate various biological processes, including metabolic homeostasis, immunity, and metabolism of xenobiotics? How do microbial rhythmicity, host peripheral clocks, and the host SCN clock communicate and synchronize with each other?
It is also worth noting that most of these studies are conducted in coprophagic mice with an inverted day-night cycle of activity: our understanding of the oscillations and connections in humans is still modest. The inter-species difference can be significant, as a recent study reported that sleep restriction alters one single bacterial species in rat gut microbiota but has no influence on the diversity or abundance of fecal microbiota in healthy human subjects (Zhang et al., 2017). In mice, disruption of circadian homeostasis and a western high fat diet can both shape the gut microbiome and its function (Voigt et al., 2014). It remains to be established whether these features of modern life contribute to our susceptibility to disease.
Currently, up to 30% of the working population are employed in shift work and about one-third of adults have less than 6 h of sleep every night. Both shift work and chronic sleep deficiency are associated with an increased incidence of cancer, cardiovascular disease, and metabolic disorders, including obesity and diabetes (Wang et al., 2013; Wang et al., 2014; Zhang et al., 2017). Elucidating the role of the gut microbiome is now fundamental to understanding the pathogenesis of morbidities due to circadian misalignment.
Footnotes
Acknowledgements
This work was supported by the Clinical and Translational Science Award 1U54TR001623. G.A.F. is the McNeil Professor in Translational Medicine and Therapeutics.
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
