Abstract
We used real-time imaging to detect cAMP levels in neurons of intact fly brains to study the mechanisms of circadian pacemaker synchronization by the neuropeptide pigment dispersing factor (PDF) in Drosophila. PDF receptor (PDF-R) is expressed by both M (sLNv) and E (LNd) pacemaker subclasses and is coupled to Gsα in both cases. We previously reported that PDF-R in M pacemakers elevates cAMP levels by activating the ortholog of mammalian adenylate cyclase 3 (AC3) but that AC3 disruptions had no effect on E pacemaker sensitivity to PDF. Here, we show that PDF-R in E pacemakers activates a different AC isoform, AC78C, an ortholog of mammalian AC8. Knockdown of AC78C by transgenic RNAi substantially reduces, but does not completely abrogate, PDF responses in these E pacemakers. The knockdown effect is intact when restricted to mature stages, suggesting a physiological and not a development role for AC78C in E pacemakers. The AC78C phenotype is rescued by the overexpression of AC78C but not by overexpression of the rutabaga AC. AC78C overexpression does not disrupt PDF responses in these E pacemakers, and neither AC78C knockdown nor its overexpression disrupted locomotor rhythms. Finally, knockdown of 2 AKAPs, nervy and AKAP200, partially reduces LNd PDF responses. These findings begin to identify the components of E pacemaker PDF-R signalosomes and indicate that they are distinct from PDF-R signalosomes in M pacemakers: we propose they contain AC78C and at least 1 other AC.
Neural circuits that coordinate daily locomotor rhythms utilize cell-intrinsic molecular oscillators that produce robust 24-hour rhythms. Studies of such circadian neural circuits in both mice and in Drosophila suggest that they are composed of multiple oscillators that are interconnected by synaptic and modulatory interactions that promote their sensitivity to environmental conditions and their synchronization (Welsh et al., 2010; Allada and Chung, 2010). Specific neuropeptides are critical synchronizing agents: in the mouse, about 10% of the approximately 20,000 neurons in the suprachiasmatic nucleus express vasoactive intestinal polypeptide (VIP); in the fly brain, a similar percentage of the approximately 150 circadian pacemaker neurons express its functional ortholog, pigment dispersing factor (PDF). Genetic analyses suggest that both VIP and PDF are instrumental for the proper display of cellular and behavioral 24-hour rhythms. Thus, definition of the neuropeptide signaling pathways that mediate pacemaker synchronization represents an important experimental goal for the field.
In Drosophila, pacemakers are categorized into M or E subgroups based on the preponderant actions they show with respect to the 2 prominent peaks of daily locomotor activity (Grima et al., 2004; Murad et al., 2007; Stoleru et al., 2005; Yoshii et al., 2009). Small LNvs principally are associated with morning activity, while LNds (along with several other subgroups) are associated with evening activity. These behavioral assignments are not rigid and can change under different environmental conditions (Rieger et al., 2009; Zhang et al., 2010; Peschel and Helfrich-Förster, 2011). Understanding how M and E cells interact and how their hierarchical relationships can change due to network properties are key issues to help understand how a complex neural circuit controls locomotor behaviors. M cells release the 18–amino acid peptide PDF, which targets its receptor, PDF-R, in subsets of both M and E pacemaker subgroups (Shafer et al., 2008; Im and Taghert, 2010), but between M and E pacemakers, PDF:PDF-R signaling has different consequences.
For example, loss of the PDF peptide alters molecular oscillations in multiple clock cells, but M and E cell subgroups respond differently. Loss of PDF signaling desynchronizes the daily rhythms of intensity and subcellular localization of PER staining in M cells but produces an advanced phase and reduced amplitude of PER staining in E cells (Lin et al., 2004, Lear et al., 2005). In addition, mutations that increase overall PDF levels in the brain and expression of a membrane-anchored PDF (to effect constitutive autoactivation) both result in complex (i.e., multiple) behavioral rhythms (Choi et al., 2009; Wulbeck et al., 2008). This suggests that PDF can accelerate some clock cells and slow others down (Wu et al., 2008; Yoshii et al., 2009). The interaction between PDF signaling and cryptochrome signaling pathways differs between M and E cells: double mutants of pdf and cry show normal molecular oscillations in M cells, while E cells show severely disrupted clock oscillations (Cusumano et al., 2009; Zhang et al., 2009; Im et al., 2011). Finally, we recently reported that PDF-R in M cells is coupled to the adenylate cyclase 3 (AC3) isoform but that disruptions of AC3 have no such effect on PDF responsiveness in E cells (Duvall and Taghert, 2012). This result suggested that PDF-R signalosomes in E cells must employ a different AC isoform(s); here, we explore the identification of the E cell AC isoform(s).
Previous work suggests that the E cell subgroup is the primary driver of circadian locomotor behavior. Broad expression of tethered PDF in the clock network produces complex rhythms in pdf01, even when it is not expressed in M cells (Choi et al., 2009). Lear and colleagues (2009) reported that the expression of PDF-R exclusively in E cells provides a rescue of locomotor behavior in pdf-r mutants, although it is incomplete (Im et al., 2011). These studies suggest that the disruption of E cell PDF signaling results in predictable disruptions of circadian behavior. Based on the importance of PDF-R activation in E cells, the nature of PDF-R signalosomes is therefore a significant issue to understand the mechanisms of circadian synchronization in the Drosophila brain.
Materials and Methods
Fly Rearing and Stocks
Drosophila were reared on cornmeal/agar supplemented with yeast at 25 °C, unless otherwise indicated by experimental design. Male flies (age, 2-5 days) were moved to 29 °C for 24 to 48 hours before imaging to increase UAS transgene expression. For temperature shift (tubulin gal80ts) experiments, crosses were maintained at 18 °C to maintain gal80ts suppression of gal4, and males were collected and moved to 29 °C for 24 to 48 hours before imaging to allow UAS transgene expression. All gal4 and UAS lines used in this study have been described previously: UAS-Epac1camps50A (Shafer et al., 2008), Mai179gal4 (Cusumano et al., 2009), tim(UAS)gal4 (Blau and Young, 1999), and UAS-ACXD, -AC3, -AC78C, and -AC76E (Duvall and Taghert, 2012). The TRiP RNAi (UAS-TRiPAC3RNAi, UAS-TRiP-nervyRNAi, UAS-TRiPAKAP200RNAi, UAS-TRiP:CG33958, UAS-TRiP:CG31183), UAS-rutabaga, UAS-dicer2, tubulin gal80ts, and Df(3)LBSC553 lines were obtained through the Bloomington Drosophila Stock Center (thanks to the Harvard TRiP RNAi project) (Bloomington, IN), and the UAS-GDAC3RNAi, UAS-AC13ERNAi, UAS-AC78C, UAS-rutRNAi, UAS-ACXARNAi, UASACXBRNAi, UAS-ACXCRNAi, UASACXDRNAi, UAS-yuRNAi, UAS-rugoseRNAi, UAS-KK:CG32305, and UAS-GD:CG32301 lines were obtained through the Vienna Drosophila RNAi Center (Vienna, Austria).
Live Imaging
For epifluorescent FRET imaging, living brains expressing gal4-driven uas-Epac1-camps were dissected under ice-cold calcium-free fly saline (46 mM NaCl, 5 mM KCl, and 10 mM Tris [pH 7.2]). All genotypes include 1 copy of each transgene unless otherwise indicated. Dissected brains were placed on poly-L-lysine–coated coverslips in an imaging chamber (Warner Instruments, Hamden, CT), and HL3 saline was perfused over the preparation (0.5 mL/min) as described in Duvall and Taghert (2012). Exposure times were 20 milliseconds for YFP-FRET and 500 milliseconds for CFP donor emission. Live FRET imaging was performed on individual cell bodies; YFP-FRET and CFP donor images were captured every 5 seconds, with YFP and CFP images captured sequentially at each time point. Following 45 seconds of baseline YFP/CFP measurements, the PDF peptide was injected into the perfusion line to result in a final concentration of 10–6 M. FRET readings were then continued to result in a total imaging time course of 10 minutes. PDF was synthesized by Neo-MPS (San Diego, CA).
FRET Imaging Data Analysis
For all experiments reported, we collected responses from at least 10 cells that were found in at least 5 brains for all genotypes. A region of interest defined each individual neuron, and for each, we recorded background-subtracted CFP and YFP intensities. The ratio of YFP:CFP emission was determined after subtracting CFP spillover (SO) into the YFP channel from the YFP intensity, as in Shafer and colleagues (2008). The CFP SO into the YFP channel was measured as 0.397. For each time point, FRET was calculated as (YFP – [CFP*SO CFP])/CFP. To compare FRET time courses across different experiments, FRET levels were normalized to initial baseline levels and smoothed using a 7-point boxcar moving average over the 10-minute imaging time course, as in Duvall and Taghert (2012). Statistical analysis was performed at maximal deflection from the initial time point by performing ANOVA analysis, followed by post hoc Tukey tests using Prism 5.0 (GraphPad Software, La Jolla, CA).
Results
AC78C Scores Positive in an In Vivo RNAi Screen that Targets Responses to PDF
We previously reported that PDF responses in LNd are completely abrogated by the mutation of PDF-R and that PDF-Rs are coupled to the Gsα subunit (Duvall and Taghert, 2012). However, while manipulations of AC3 alter PDF responsiveness in M pacemakers, they have no such effect on E cell PDF responses. That suggests that the 2 pacemaker subgroups use different signaling components (Duvall and Taghert, 2012). To pursue E cell PDF-R signaling components, we therefore performed an RNAi screen directed against 13 of the 14 predicted ACs in the Drosophila genome. At the time of our experiments, no available RNAi line targeted ACXE, a cyclase that is predicted to be expressed in the male germline exclusively (Cann et al., 2000). We used the Mai179gal4 driver to express UAS-RNAi transgenes in the 3 (of 6) LNds that are known to express PDF-R and cryptochrome (Im and Taghert, 2010; Yoshii et al., 2009). By focusing on neurons that express PDF-R, we reasoned that a negative result would be more likely explained by an effect on PDF signaling. When individual AC isoforms were knocked down in LNds, AC78C RNAi significantly reduced PDF responses, although not completely (Fig. 1A). Five ACs are broadly expressed in Drosophila tissues (Flybase, 2013; DroID.org, 2010)—AC78C, AC76E, AC3, rutabaga, and AC13E—and they arguably represent the strongest initial candidates for an LNd AC coupled to PDF-R. We therefore screened RNAi constructs for these 5 ACs (Fig. 1B) with the addition of a UAS-dicer2 transgene to increase the efficiency of RNAi knockdown (Dietzl et al., 2007). However, this screen did not implicate any additional candidate ACs (Fig. 1B): Expression of the UAS-dicer2 transgene itself partially reduced PDF responses, and only AC78C RNAi produced a further significant reduction in PDF response. However, that reduction was once again not complete, suggesting the involvement of an additional and, as yet, unidentified AC.
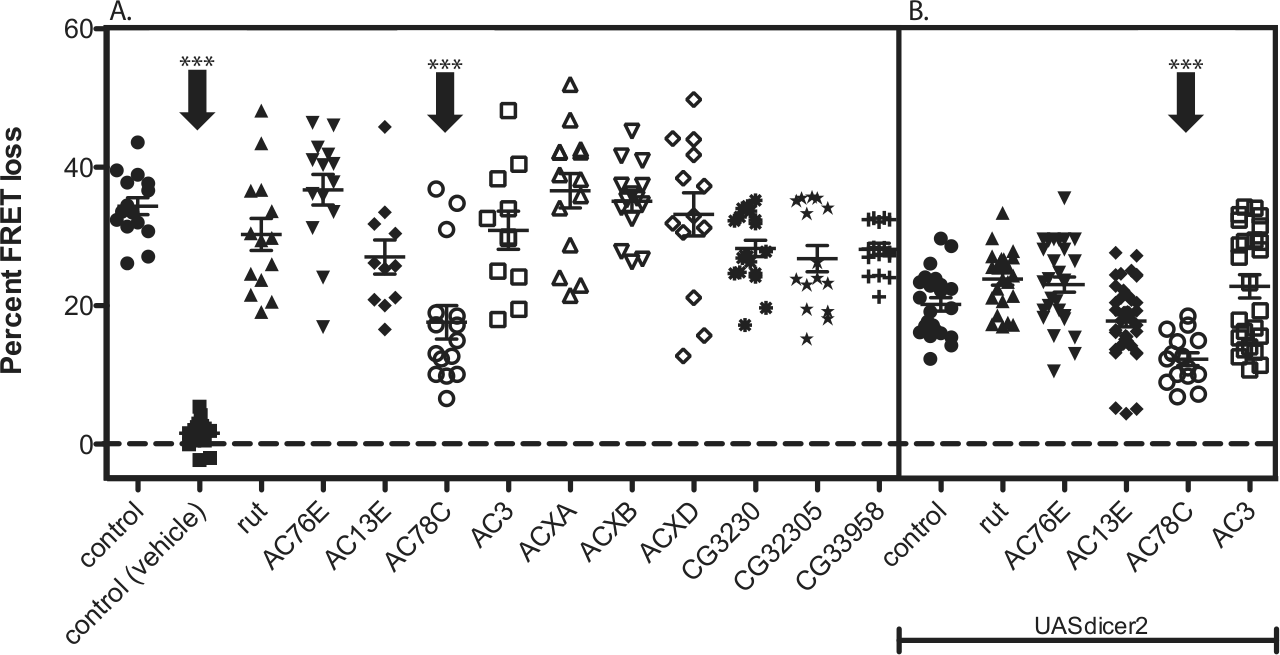
Knockdown of AC78C reduced PDF cAMP responses in LNds. (A) RNAi screen directed against most of the predicted ACs in Drosophila. Only the AC78C RNAi significantly reduces PDF responses in LNd pacemakers. (B) Addition of UAS-dicer2 to increase RNAi knockdown does not reveal additional candidates for ACs mediating the PDF response and does not enhance the AC78C RNAi phenotype. All genotypes include Mai179-gal4;Epac1camps and 1 copy of UASRNAi (except for control). Error bars denote SEM. ***p < 0.001 (compared with control). Statistical analysis was performed at maximal deflection from the initial time point by performing ANOVA analysis, followed by post hoc Tukey tests. Detailed information about RNAi lines is included in the Materials and Methods section.
Genetic Confirmation of a Role for AC78C in E Cell PDF Responses
We evaluated the AC78C RNAi phenotype in 2 ways. First, we asked whether the loss-of-function phenotype could be explained by developmental, as opposed to physiological, effects of AC78C in E cells. Thus, we employed a conditional genetic design (via tub-gal80ts), which permits normal development, followed by the induction of RNAi only in the adult fly (McGuire et al., 2003). When induction of AC78C RNAi was restricted to adult stages, it nevertheless produced a reduction in PDF responses that was quantitatively similar to results after continuous knockdown (Fig. 2A). Expression of the tub-gal80ts transgene alone had no effect on PDF responses (Fig. 2A). This suggests that the reduction of E cell PDF responsiveness is not due to developmental defects and that, in mature LNds, AC78C plays a substantial role in mediating PDF responses in these pacemakers.
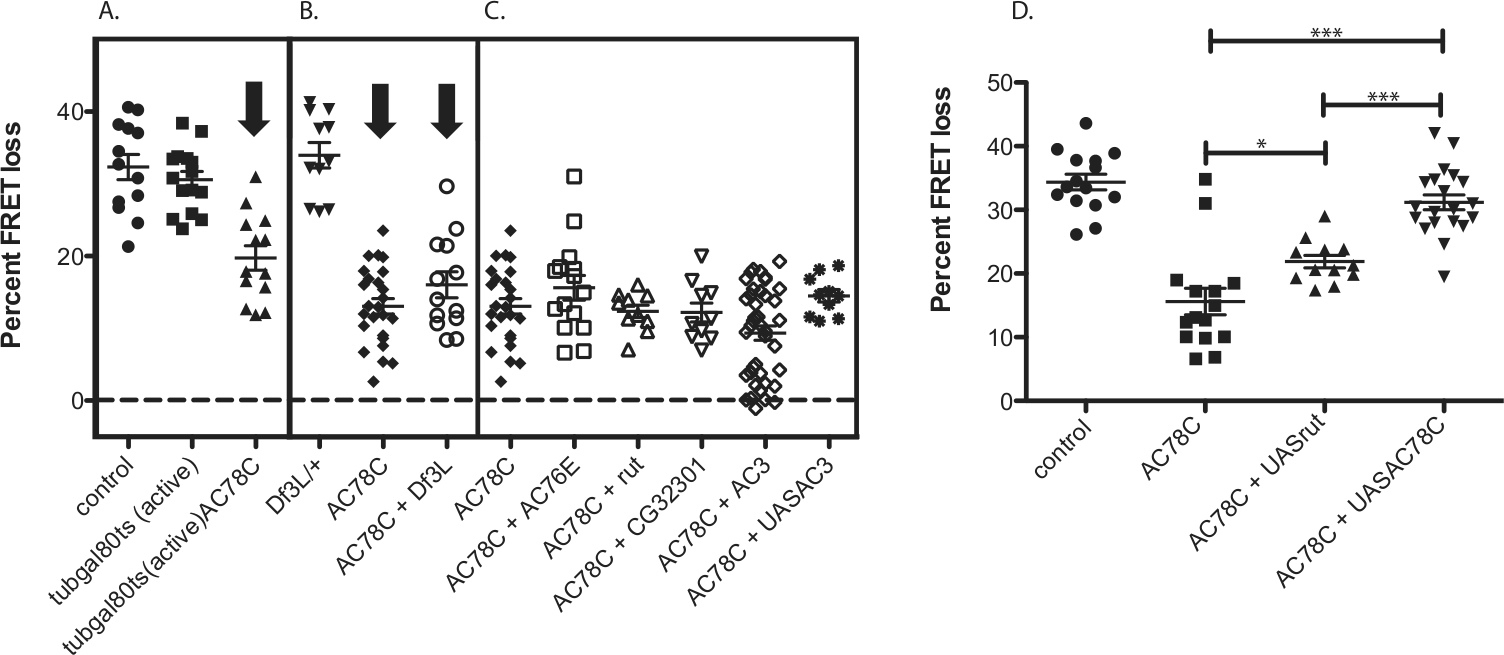
AC78C RNAi partially reduces PDF responses in adult LNds. (A) AC78C RNAi expressed only in adult cells reduced the PDF response in LNds (expression of tubgal80ts alone has no effect). Animals with “active” GAL4 were transferred to 29 °C as described in Materials and Methods to inactivate inhibition by the tub-GAL80 transgene. (B) The AC78C RNAi phenotype is not enhanced by a combination with a cognate deficiency. (C) Pairwise combinations of RNAi constructs for AC78C and other ACs do not enhance the AC78C RNAi phenotype (none is significantly different from AC78C RNAi alone by the Tukey multiple comparison test). (D) The AC78C RNAi-mediated reduction in PDF cAMP responses in LNd is rescued by the overexpression of AC78C but not of rutabaga. All genotypes include Mai179-gal4;Epac1camps and 1 copy of UASRNAi (except for control). Error bars denote SEM. ***p < 0.001 and *p < 0.05 (compared with control, internal comparisons are shown and brackets between genotypes compared). Statistical analysis was performed at maximal deflection from the initial time point by performing ANOVA analysis, followed by post hoc Tukey tests.
As a second test, we combined AC78C RNAi with different UAS-AC overexpression constructs to determine if the loss-of-function phenotype could be reversed. Because the VDRC AC78C RNAi transgene targets a portion of the AC coding region (Vienna Drosophila RNAi Center), both the endogenous AC78C as well as the overexpression of RNA are targeted in this design. Nevertheless, overexpression of AC78C cDNA fully restored PDF responsiveness in E cells expressing AC78C RNAi. In contrast, overexpression of the rut AC did not fully rescue PDF responses in such E cells (Fig. 2D). The latter observation provides a measure of rescue specificity and also rules out the possibility that the reversal was due to a simple dilution of GAL4’s efficacy because of the presence of an additional UAS transgene. Overexpression of rutabaga provides a small increase in PDF responsiveness in the AC78C RNAi background, suggesting that rutabaga partially substitutes for AC78C. This is in contrast to the situation that we found earlier in small LNvs in which rutabaga showed no level of substitution for the relevant cyclase (AC3). This observation highlights another difference between M and E cell signaling.
Compound AC RNAi Knockdowns Do Not Further Reduce E Cell PDF Responses
To further evaluate the candidacy of AC78C in E cell PDF responses, we combined AC78C RNAi with a deficiency for the AC78C region. We did not observe any further decrement in the E cell PDF response, suggesting the AC78C RNAi knockdown is effectively complete (Fig. 2B). That finding again raises the possibility that 1 or more additional ACs normally contributes to E cell PDF responses. Using this logic, we therefore tested several pairwise combinations of AC RNAi lines. However, no combination of RNAi lines significantly reduced PDF responses beyond the level found with AC78C knockdown alone, although the combination of AC78C and AC3 knockdown produces a subpopulation of cells that fails to respond to PDF (Fig. 2C). This suggests that there may be a subset of LNds that depend specifically on AC78C/AC3. We also combined UAS-AC3 with AC78C RNAi; AC3 overexpression totally and specifically disrupts PDF responses in M cells (Duvall and Taghert, 2012). However, this combination did not further reduce the E cell PDF-generated cAMP responses either (Fig. 2C).
Overexpression of AC Isoforms Had No Effect on LNd PDF Responses
We previously reported that in small LNvs, PDF-R is associated with AC3 and that AC3 overexpression in that M pacemaker type specifically disrupted PDF signaling (Duvall and Taghert, 2012). On that basis, we anticipated a finding wherein overexpression of the relevant AC in LNds (possibly AC78C) would abrogate PDF responses in that E pacemaker cell type. We therefore tested the effects of UAS-mediated overexpression of UAS-ACXD, -AC78C, -AC76E, -rut, and -AC3 on PDF responses in E pacemakers, but we saw no effects (data not shown). This suggests that LNd signaling pathways are less sensitive to the overexpression of cognate ACs and may use some compensatory mechanism(s) to maintain PDF responsiveness. Regardless of the exact mechanism, this observation highlights another difference between PDF responses in small LNvs versus LNds.
Knockdown of AKAPs Nervy and AKAP200 Reduced LNd PDF Responses
Scaffolding proteins such as AKAPs can bind to signaling components to organize efficient signaling (Dessauer, 2009). Knockdown of the AKAP nervy reduced PDF responses in small LNvs (Duvall and Taghert, 2012). In a similar fashion, we tested the possible involvement of 4 candidate AKAPs as scaffolding proteins for PDF-R in LNd cells using gene-specific RNAi constructs. Knockdown of either AKAP200 or nervy reduced LNd PDF responses (Fig. 3). Knockdown of AKAP200 reduces PDF responses to about 40% of their original levels, whereas knockdown of nervy only reduced PDF responses to about 65% of their original levels. These findings are consistent with the possibility that multiple signalosomes normally mediate PDF responses in LNds.
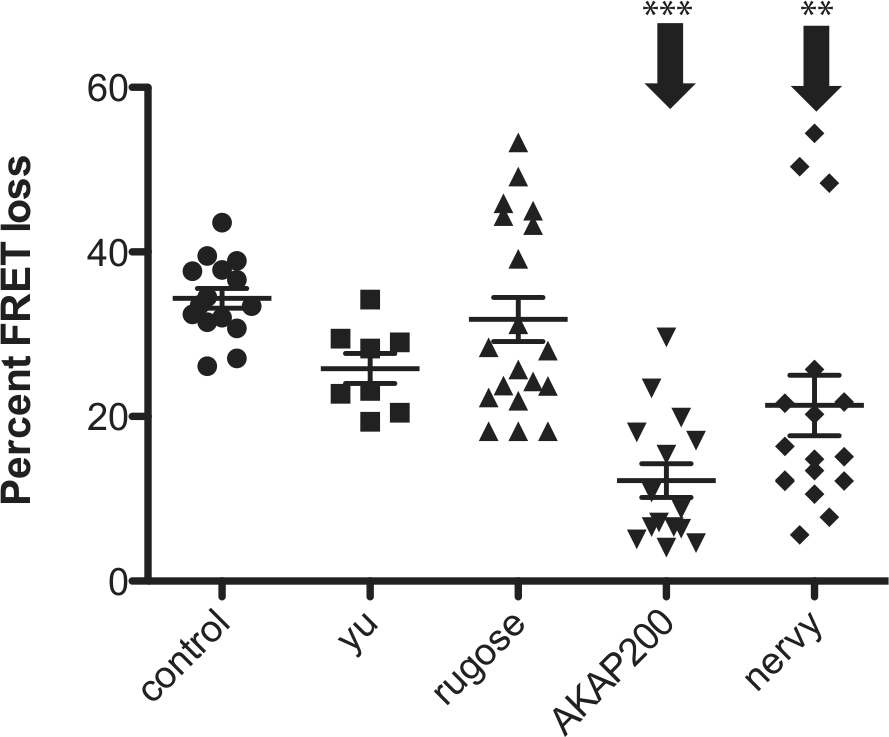
RNAi-mediated knockdown of 2 potential AKAPs reduces cAMP elevation by PDF in LNd pacemakers. Knockdown of AKAP200 and nervy significantly but partially reduced LNd PDF responses. All genotypes include tim(UAS)-gal4;Epac1camps and 1 copy of UASRNAi (except for control). Error bars denote SEM. ***p < 0.001 and **p < 0.01 (compared with control). Statistical analysis was performed at maximal deflection from the initial time point by performing ANOVA analysis, followed by post hoc Tukey tests.
AC78C Manipulations Did Not Result in a Circadian Locomotor Phenotype
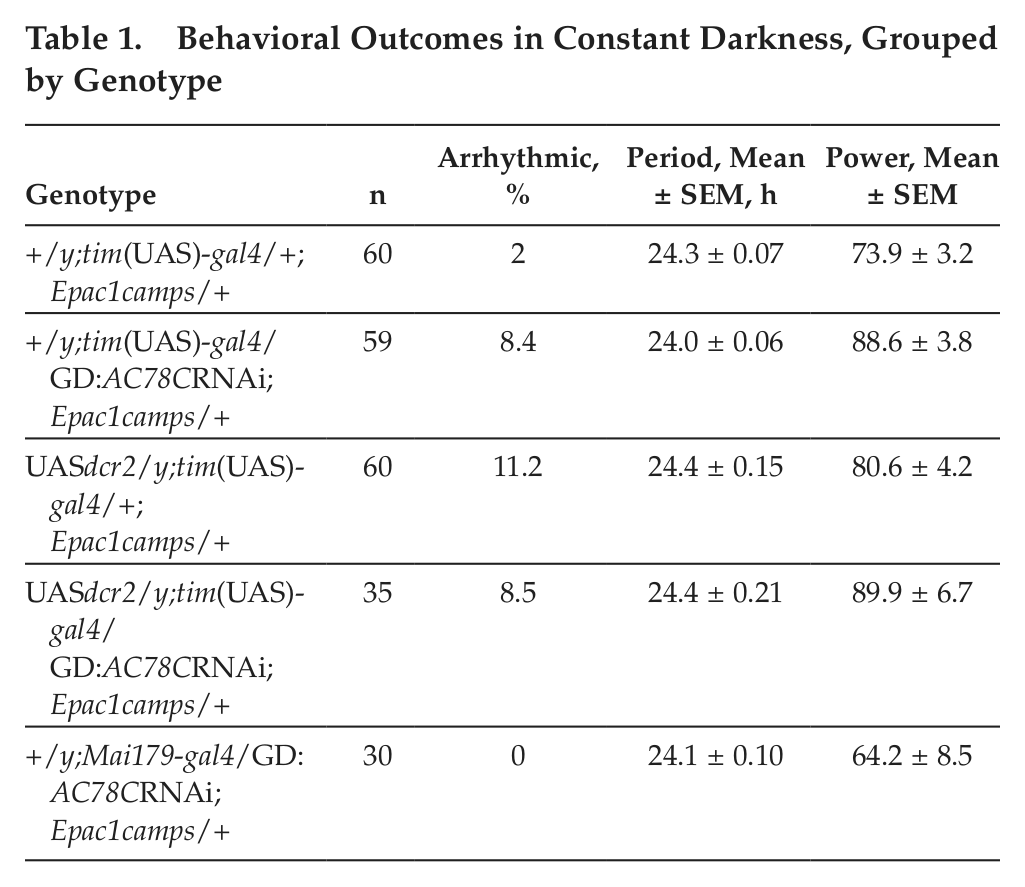
Previous work suggests that PDF signaling within the E cell subgroup is largely responsible for normal circadian locomotor behavior (Choi et al., 2009; Lear et al., 2009; Im et al., 2011). Because LNds make up only a subset of E cells in the Drosophila brain, and because the Mai179gal4 driver is relatively weak and restricted to a small subset of cells, we primarily used a stronger driver, expressed broadly throughout the clock network (tim(UAS)gal4), to drive UAS transgenes in locomotor assays. We expected that disruptions to PDF responses by AC knockdown in these cells might phenocopy animals with PDF-R expressed only in the M cell subgroup (small LNvs). Specifically, we looked for an advance in the evening peak in LD conditions and weak short rhythms in DD (cf. Hyun et al., 2005). However, AC78C manipulations that partially reduced PDF responses in E pacemakers did not result in behavioral deficits, even with the addition of a UAS-dicer2 transgene to increase RNAi knockdown efficiency (Fig. 4). A single experiment using the Mai179gal4 driver to drive AC78C RNAi gave similar behavioral results (Table 1 and Suppl. Fig. S1). This result is consistent with the view that partial reductions in PDF responsiveness are not sufficient to produce behavioral changes. Indeed, our previous results suggested that only AC knockdowns that produce severe reductions in PDF responsiveness also affect locomotor behavior (Duvall and Taghert, 2012). Additionally, to date, no combination of genetic elements that partially reduce LNd PDF responses (AC RNAi/AKAP RNAi/AC78C Df, etc.) shows circadian disruptions consistent with a total loss of PDF signaling (data not shown).
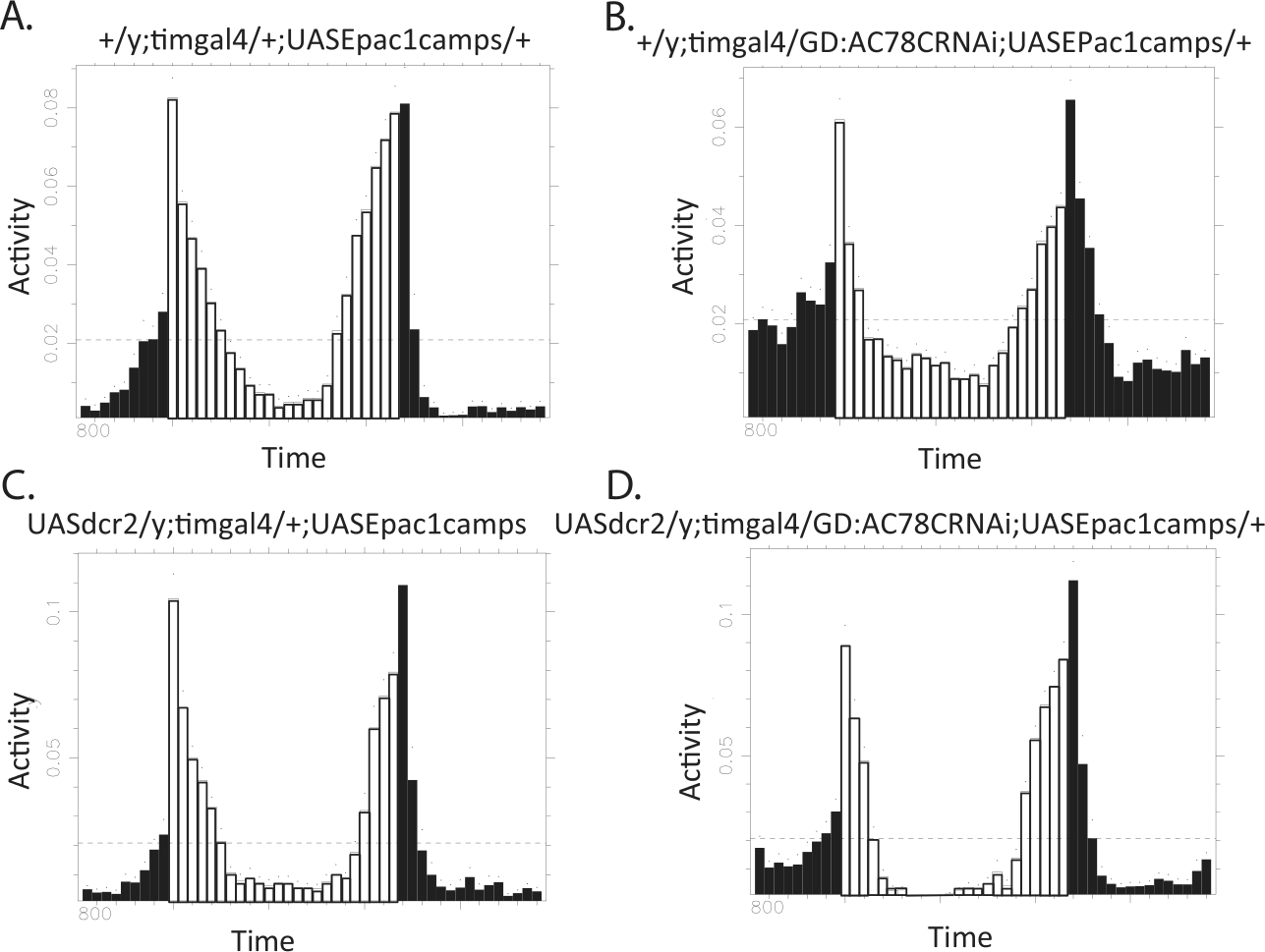
Knockdown of AC78C does not result in locomotor defects in circadian behavior. (A) Representative locomotor behavior of control flies expressing the Epac1camps sensor under control of the tim(UAS)-gal4 driver. (B) Representative locomotor behavior of flies with the knockdown of AC78C in pacemaker cells. (C) Representative locomotor behavior of control flies expressing the UAS-dicer2 transgene in addition to Epac1camps under tim(UAS)-gal4 driver. (D) Representative locomotor behavior of flies expressing RNAi directed against AC78C in addition to the UAS-dicer2 transgene to increase RNAi efficiency.
Behavioral Outcomes in Constant Darkness, Grouped by Genotype
Discussion
Multiple lines of evidence suggest that PDF signaling differs between clock cell subgroups. 1) Loss of PDF has distinct effects on PERIOD protein cycling in LNv (M cells) versus non-LNv cells (E cells) (Lin et al., 2004). Both cell groups continued to show cycling in PER immunostaining levels and localization, but while M cells become phase-dispersed in PER cycles, E cells remain synchronized with the altered phase and amplitude of PER accumulation. 2) In pdf/cry and pdfr/cry double mutants, a subset of E cells shows a phase advance and/or severe attenuation (Im et al., 2011) of the PER protein molecular rhythm, while M cells continue to cycle normally (Zhang et al., 2009; Park et al., 2000; Im et al., 2011). 3) Knockdown of AC3 disrupts small LNv PDF responses but has no such effect in LNds. These earlier findings are consistent with the hypothesis that there are at least 2 functionally different PDF signaling pathways that normally operate in different pacemaker cell types.
The present results provide strong evidence that AC78C, an AC previously associated with sugar perception in Drosophila (Ueno and Kidikoro, 2008), is a candidate AC for mediating a portion of PDF responses in LNds. Importantly, the partial reduction in PDF responsiveness in LNds is fully rescued by the overexpression of AC78C but only partially rescued by the overexpression of rutabaga. However, we emphasize that there likely remains 1 or more additional ACs that contribute to PDF responses in these pacemakers and that they remain as yet uncharacterized. The partial reduction in LNd PDF responses is still observed when AC78C is knocked down only in adult stages, and this reduction remains incomplete when AC78C RNAi is combined with a number of other UAS-AC RNAi lines. Hence, we have been unable to completely abrogate the PDF responses in E pacemakers. The addition of the deficiency for the AC78C genetic region does not further reduce the PDF response, which is yet more evidence that suggests the contribution of another AC(s). The additional LNd PDF-R–associated AC(s) may have been missed among the isoforms that we tested because it produced a false-negative result(s): it is technically possible that 1 or more AC RNAi lines does not appropriately target its cognate cyclase for degradation.
A number of studies suggest that the basic unit of AC function is a dimer (Rodbell, 1980) and that AC homodimerization may be required for normal trafficking to the plasma membrane (Gu et al., 2001; Seebacher et al., 2001; Gu et al., 2002). Previous work indicated that ACs may also form heterodimers: these may exhibit characteristics different from either of the single isoforms or of homodimers. For example, AC2 and AC5 form heterodimers that show higher sensitivity to Gsα signaling than either of the components individually (Baragli et al., 2007). Although ACs can form specific and preferred interactions with other signaling molecules, it is likely that more passive mechanisms of coupling are also used and that signaling components are not directly interacting but localized nearby each other in the plasma membrane (Dessauer, 2009). It is possible that LNds use less stringent coupling of PDF-R to specific ACs than do small LNvs. Thus, although there is a preference for AC78C coupling in LNds, it is possible that PDF signaling can in fact access 2 or more AC isoforms to generate some sufficient level of cAMP. Alternatively, the neurons may assemble and segregate signalosomes bearing different ACs, whose activation instigates different functional outcomes. In sum, we favor the hypothesis that AC78C and at least 1 additional AC together mediate PDF responses in E pacemakers like the LNds (Fig. 5). Whether they do so jointly, for instance, as heterodimers or in a distinct fashion, remains to be determined.
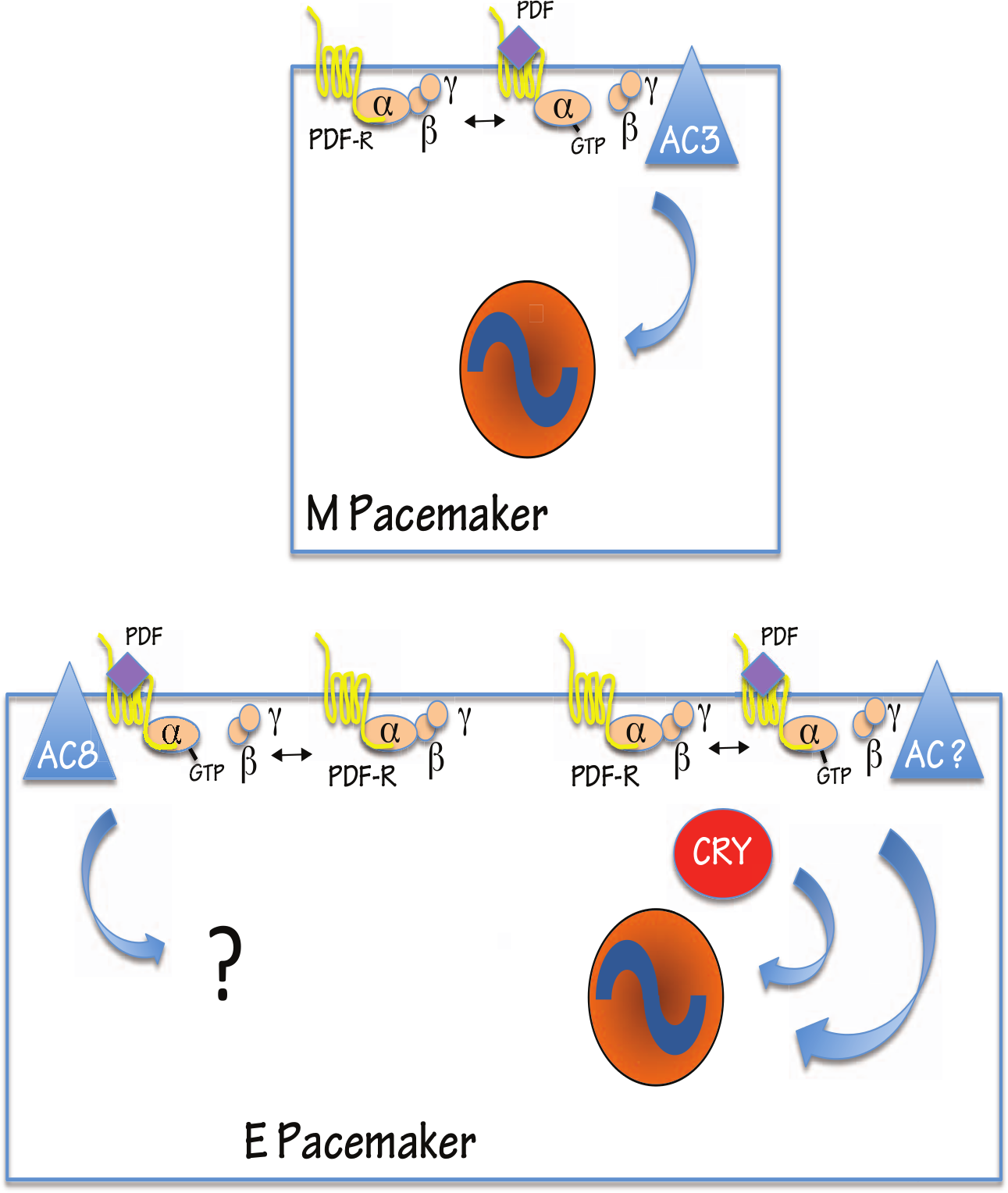
Model for PDF-R signaling in M versus E cells in Drosophila. PDF-R associates with AC3 in small LNv cells, but signalosome components differ in LNds. In this E cell subgroup, PDF signaling relies on AC78C (AC8) and at least 1 other AC.
To what extent do different signaling components actually result in differences in downstream PDF signaling between clock cell subgroups? If the considered end point is simply the production of cAMP, then it is possible that signalosomes with different components may nevertheless perform the same function. We observe that both M and E cell subgroups respond to PDF with an increase in cAMP and that LNds respond with slightly higher amplitudes (Duvall and Taghert, 2012). Our analysis has not revealed any temporal differences in PDF responses or recovery between these 2 subgroups, although more subtle differences, subsecond differences in kinetics or microdomain differences, are not easily detected by the imaging techniques that we have used (Nikolaev et al., 2003). Alternatively, PDF-R signalosome differences between pacemaker cell types, as highlighted by AC isoform differences, may indicate fundamental differences in signaling networks, within which cAMP elevations may play important local roles. The extent to which differences in the networks that generate cAMP account for the differences in downstream effects of PDF in different clock cell subgroups helps define a testable hypothesis.
PKA is important for maintaining normal locomotor rhythms, although PKA alterations do not affect core clock proteins themselves (Majercak et al., 1997; Park et al., 2000). This has been taken to mean that a likely role for cAMP and PKA in the Drosophila circadian timing system might be in the flow of information between pacemaker cells and output pathways. However, these previous studies were performed with mutant flies that make it difficult to disambiguate the roles of individual pacemaker subgroups.
Why would a single neuropeptide receptor utilize multiple signaling pathways in different target cells? Recent studies have implicated feed-forward mechanisms in neuropeptide circuit modulations, often as a method of regulating competing circuit elements (Jing et al., 2010; Wu et al., 2010; reviewed by Taghert and Nitabach, 2012). Differential signalosome compositions may provide a mechanism for a single neuropeptide like PDF to differentially modulate components of the circadian circuit. In fact, the connections between Drosophila clock cells suggest that PDF may be working through such a mechanism. There are 2 sources of PDF in the Drosophila brain: the large and small LNvs (Helfrich-Förster, 1998). Previous work suggests that there are direct anatomic connections between some nodes of the circadian circuits; small LNvs and large LNvs are directly connected, and the small LNvs send projections to the dorsal brain, where the E cells, including LNds, are located (Helfrich-Förster et al., 2007; reviewed in Sheeba, 2008). Additionally, previous studies have also suggested that PDF released from the large LNvs can act on dorsal cells over a longer distance (Cusumano et al., 2009; Shafer and Taghert, 2009). This suggests that PDF from large LNvs can act directly on dorsal (E) clock cells and indirectly through the small LNv (M) cells. A feed-forward model provides a mechanism for PDF to act differentially in its actions to promote synchrony of the circadian neural circuit in Drosophila and to allow flexibility in the regulation of multiple competing oscillators.
Footnotes
Acknowledgements
The authors thank members of their laboratory and Aaron DiAntonio and Erik Herzog for helpful comments and advice. They also thank Weihua Li for technical assistance. They received fly stocks from the Bloomington Drosophila Stock Center, from the Vienna Drosophila RNAi Center, and from the Harvard TRiP collection. L.B.D. was supported by National Institutes of Health Training Grant 5-T32-GM08151-27 and by Institutional National Research Service Award 5-T32-EY013360-10 from the National Eye Institute. This work was supported by National Institutes of Health (NIMH) Grant R01MH067122 to P.H.T.
Conflict of Interest Statement
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
