Abstract
In mammals, the molecular circadian clockwork is comprised of interlocked transcriptional-translational feedback loops (TTLs). Three Period (Per1-3) and 2 Dec (Dec1/2) genes interact in regulating the activity of the transcriptional activators CLOCK/NPAS2 and BMAL1. While deletion of Per1 and Per2 in mice results in behavioral arrhythmicity, Dec deletion has less dramatic effects on activity rhythms, affecting primarily phase of entrainment and free-running period. In intact animals, clock gene mutant phenotypes are often masked due to intercellular coupling mechanisms that stabilize cellular rhythms. Therefore, to study Per/Dec genetic interaction at the cellular level, we isolated fibroblasts from different tissues of Per1, Per2, and Dec2 single and double mutant mice. We show that in the cellular TTL, Pers and Dec2 act in a principally synergistic way, but tissue-specific differences in this interaction are seen. A rescue of rhythmicity in Per2 mutant cells after additional deletion of Dec2 was observed, indicating that in the absence of Per2, DEC2 destabilizes TTL function. Rhythm power in Per1/Dec2 and Per2/Dec2 double mutants was strongly reduced, suggesting that interaction of Dec2 with both Per genes is important for stabilizing clock period. Contrary to what was observed for behavior, nonsynergistic effects of Dec2 and Per1/2 mutations were observed on cellular clock phase regulation that do not correlate with period effects. Our data reveal cell type-specific interactions of Per1/2 and Dec2 in the regulation of period, phase, and rhythm sustainment, emphasizing the differential organization of the mammalian clock machinery in different tissues.
In most organisms, endogenous timekeeping systems have evolved that organize physiological and behavioral adaptation to the 24-h day (Brown et al., 2012). In mammals, these so-called circadian clocks are based on cellular oscillators driven by a set of clock genes organized in a system of interlocked transcriptional-translational feedback loops (TTLs) (Oster, 2006; Takahashi et al., 2008). Positive TTL elements include the transcription factors CLOCK, NPAS2, and BMAL1 (ARNTL) that drive expression of negative elements such as Period (Per1-3) and Cryptochrome (Cry1/2) genes via E-box regulatory promoter elements during the day (Bunger et al., 2000; Gekakis et al., 1998). In the night, PER/CRY protein complexes translocate into the nucleus where they interfere with CLOCK/NPAS2/BMAL1, thereby inhibiting their own transcription (Griffin et al., 1999; Kume et al., 1999; Sun et al., 1997). Toward morning, successive degradation of PER/CRY proteins releases the inhibitory pressure on CLOCK/NPAS2/BMAL1, and a new round of E-box-mediated transcription resumes (Lee et al., 2001; Yoshitane et al., 2009). The periodicity of this feedback is believed to be regulated by posttranslational mechanisms and accessory loops that stabilize the necessary delay between production and inhibitory action of negative TTL elements. One such loop involves the transcriptional modulator DEC2 (SHARP-1, BHLH-42) (Honma et al., 2002). Dec2 transcription is controlled by CLOCK/NPAS2/BMAL1 (Hamaguchi et al., 2004; Noshiro et al., 2005), and DEC2 proteins can—depending on time and tissue—act as either activators or inhibitors of E-box transcription (Bode et al., 2011b; Rossner et al., 2008). Very similar characteristics have been observed for the PER proteins (Preitner et al., 2002; Zheng et al., 2001). In previous experiments we have shown that Per and Dec genes interact closely and in a time-of-day dependent way in regulating activity rhythms and clock gene expression in the mammalian circadian pacemaker of the suprachiasmatic nucleus (SCN) (Bode et al., 2011a, 2011b).
Unlike other tissues, the SCN is characterized by tightly coupled neurons generating a highly stabilized rhythm at the tissue level (Aton et al., 2005; Liu et al., 2007; Liu and Reppert, 2000; Maywood et al., 2006; Yamaguchi et al., 2003). Previous studies have shown that by this coupling, SCN cells become highly resistant to the destabilizing effects of clock gene mutations (Jakubcakova et al., 2007; Liu et al., 2007; Pando et al., 2002; Robles et al., 2010). Moreover, the organization of the clock machinery and, thus, the phenotype of specific clock gene mutations can differ between different tissues (Cermakian et al., 2001; DeBruyne et al., 2007b; Jakubcakova et al., 2007; Nakashima et al., 2008; Reick et al., 2001; Robles et al., 2010).
To analyze the interactivity of PER and DEC feedback in molecular clock regulation at the cellular level and in different tissues, we here compared molecular rhythms of primary fibroblasts from ear and lung tissue preparations of Per1, Per2, and Dec2 single and double mutant mice. Our data suggest a close, cell type–specific interaction of Per1/2 and Dec2 in the regulation of period, phasing, and sustainment of molecular circadian rhythms, emphasizing the multiform organization of the mammalian clock machinery in different tissues.
Materials and Methods
Primary Cell Cultures
Mouse ear and lung fibroblasts were obtained from adult male and female Per1 mice (Per1tm1Brd) (Zheng et al., 2001), Per2 mice (Per2tm1Brd) (Zheng et al., 1999), Dec2 mutant mice (Bhlhe41tm1Mrjo) (Rossner et al., 2008), and PER2::LUC mice (Per2tm1Jt) (Yoo et al., 2004) on a C57B/6J background. Six or 7 independent preparations were done per genotype spread over several days, and always 2 or more genotypes were processed in parallel to minimize unspecific effects due to experimental variations. Tissues were extracted, rinsed with ice-cold HBSS, and subjected to mincing and subsequent enzymatic digestion. Ear tissues (2 ears per preparation) were digested with 100 µg/mL Liberase TM (Roche Applied Science, Mannheim, Germany) in the presence of 20% FBS at 37 °C and 5% CO2 overnight. Lung tissues (2 lung wings per preparation) were digested with 100 µg/mL Liberase TM in the presence of 1% BSA at 37 °C with constant agitation for 1 h. Dissociated cells were rinsed with PBS and plated onto 60-mm cell culture dishes containing culture medium (DMEM with 20% FBS, 100 U/mL penicillin, 0.1 mg/mL streptomycin, 2 mM stable glutamine, 2.5 µg/mL amphotericine B) with medium change every second day. Upon confluence, fibroblast cells were trypsinized, and 5 × 105 cells were seeded to a 35-mm cell culture dish containing culture medium as described above, but without amphotericine B. Viral transduction was conducted on the next day with ~1.68 × 108 infection units (IFUs) per dish in the presence of 8 µg/mL polybrene. One day after transduction, cells were rinsed with PBS and restored with fresh culture medium.
Bioluminescence Recording
Cells were synchronized by 2 h of incubation with 100 nM dexamethasone. After that, cells were changed to recording medium (same as above, but without phenol-red, plus 20 mM HEPES, 1 mM pyruvate, 100 µM D-luciferin; Biosynth AG, Staad, Switzerland). Dishes were sealed with silicon grease and cover glasses. Luminescence was measured using a LumiCycle luminometer (Actimetrics, Evanston, IL) at 32.5 °C. To assess dexamethasone sensitivity, ear- and lung-derived fibroblasts of PER2::LUC mice were isolated and plated into a 96-well plates with recording medium. Forty-eight hours after plating, plates were sealed with transparent film, and luminescence was recorded at 34 °C using a Berthold TriStar LB 941 (Berthold Technologies, Wildbach, Germany). Luminescence was measured over 1 h to determine reference baselines. After that, cells were treated with various concentrations of dexamethasone, and recording was resumed for additional 24 hours. Sensitivity was determined by dividing the 24-h peak value of each trace by the respective pretreatment baseline.
Lentivirus Production
Bmal1:luc encoding pBluF-puro plasmid was a kind gift from Prof. Steven Brown, University of Zurich, Switzerland. To produce Bmal1:luc lentiviral particles, each 10-cm dish of HEK293T cells was cotransfected with 10 µg psPAX2 (Addgene plasmid #12260, Prof. Didier Trono, EPFL, Switzerland), 5 µg pMD2.G (Addgene plasmid #12259, Prof. Didier Trono, EPFL, Switzerland), and 15 µg pBluF-puro using Xfect transfection reagent (Clontech, Saint-Germain-en-Laye, France). Cells were rinsed with PBS on the second day and restored with fresh culture medium. Viral containing medium was harvested at 36 h after transfection and stored at 4 °C. Cells were restored with fresh medium, and further medium collections were performed on the next day. Two collections were pooled and concentrated using LentiX concentrator reagent (Clontech) according to the manufacturer’s protocol. Virus titers were determined by transducing HEK 293T cells with a serial dilution of GFP-encoding lentiviral particles (produced as described above but with pWPI (Addgene plasmid #12254, Prof. Didier Trono, EPFL, Switzerland) instead of pBluF-puro). At 72 h after transduction, GFP-positive cells were counted under a fluorescence microscope. The same serial dilution was also subjected to real-time quantitative PCR (qPCR) analysis using a primer pair amplifying the viral WPRE element (forward: 5′-GGCACTGACAATTCCGTGGT-3′; reverse: 5′-AGGGACGTAGCAGAAGGACG-3′) to determine viral genome abundance. qPCR was also done for Bmal1-luc virus-containing medium, and the titer was estimated by comparison with the pWPI abundance versus GFP calibration curve.
Gene Reporter Assays
Dbp:luc plasmid was a kind gift from Prof. Ueli Schibler, University of Geneva, Switzerland. HEK 293T cells were plated into 96-well plates coated with poly-D-lysine and transfected with the following expression plasmids: 10 ng Dbp-luc and 2 ng CMV-Renilla luciferase with various combinations (20 ng each) of the following clock gene constructs: HA-Clock, HA-Bmal1, HA-Dec1, FLAG-Dec2, Per1-GFP, Per2-V5, and pcDNA3.1 (mock transfection). We used Lipofectamine LTX transfection reagent (Life Technologies, Darmstadt, Germany). Forty-eight hours after transfection, luciferase activity was measured using Dual-Glo Luciferase Assay System (Promega, Mannheim, Germany) using the Berthold TriStar.
Statistics
For statistical comparisons, 6 or 7 independent fibroblast preparations for each tissue and genotype were used. Rhythmicity and period length were assessed by χ2 periodogram analysis over the first 3 days on 24-h moving average baseline-subtracted luminescence recordings. Dampening times (D) were determined by fitting a dampened sine wave to baseline-subtracted luminescence recordings over 5 days using a Levenberg-Marquardt algorithm on the following equation: Y(t) = Amplitude * sin (2 * π * Frequency * t + Phase) * e–(t/D) + Offset. Dampening times are expressed relative to rhythm period length (1 CT = (2 * π * Frequency)/24 h). To determine phase, the first peak of the baseline-subtracted luminescence curve was measured relative to the time of synchronization. All measurements were done using the Lumicycle analysis software (Actimetrics, Evanston, IL). Group statistics were done with GraphPad Prism 5 (GraphPad, La Jolla, CA).
Results
Rhythmic Bmal1:luc Expression in Per/Dec-Deficient Fibroblasts
We isolated and cultured primary fibroblasts of ear and lung tissues from adult wild-type (WT) and homozygous Per1 (P1), Per2 (P2), Dec2 (D2), Per1/Dec2 (P1D2), and Per2/Dec2 (P2D2) single and double mutant mice. We chose 2 similar cell types from 2 different organs as this would allow us to compare tissue-specific effects under comparable culture conditions. At 60% confluence, cells were infected with Bmal1:luc lentivirus. Transduction efficiency was determined on parallel infections using a GFP-expressing lentivirus. More than 95% of cells were expressing the transgene in all tested preparations (Suppl. Fig. S1). Upon confluency, cells were synchronized by dexamethasone treatment and transferred into a luminometer for luciferase activity measurements. Cells of all genotypes and tissues showed at least 2 full cycles of circadian rhythmicity, with the exception of P2 (both tissues) and P1D2 (lung). Representative luminescence recordings and corresponding normalizations are shown in Figure 1A for ear and Figure 1B for lung preparations. To exclude that a difference in sensitivity to dexamethasone treatment might underlie the observed differences in rhythmicity in the 2 cell types, we stimulated PER2::LUC fibroblasts from both tissues with increasing amounts of dexamethasone and monitored the direct luciferase response over the next 24 hours. Luciferase activity was dose-dependently and similarly induced by dexamethasone in both tissues (Suppl. Fig. S2).
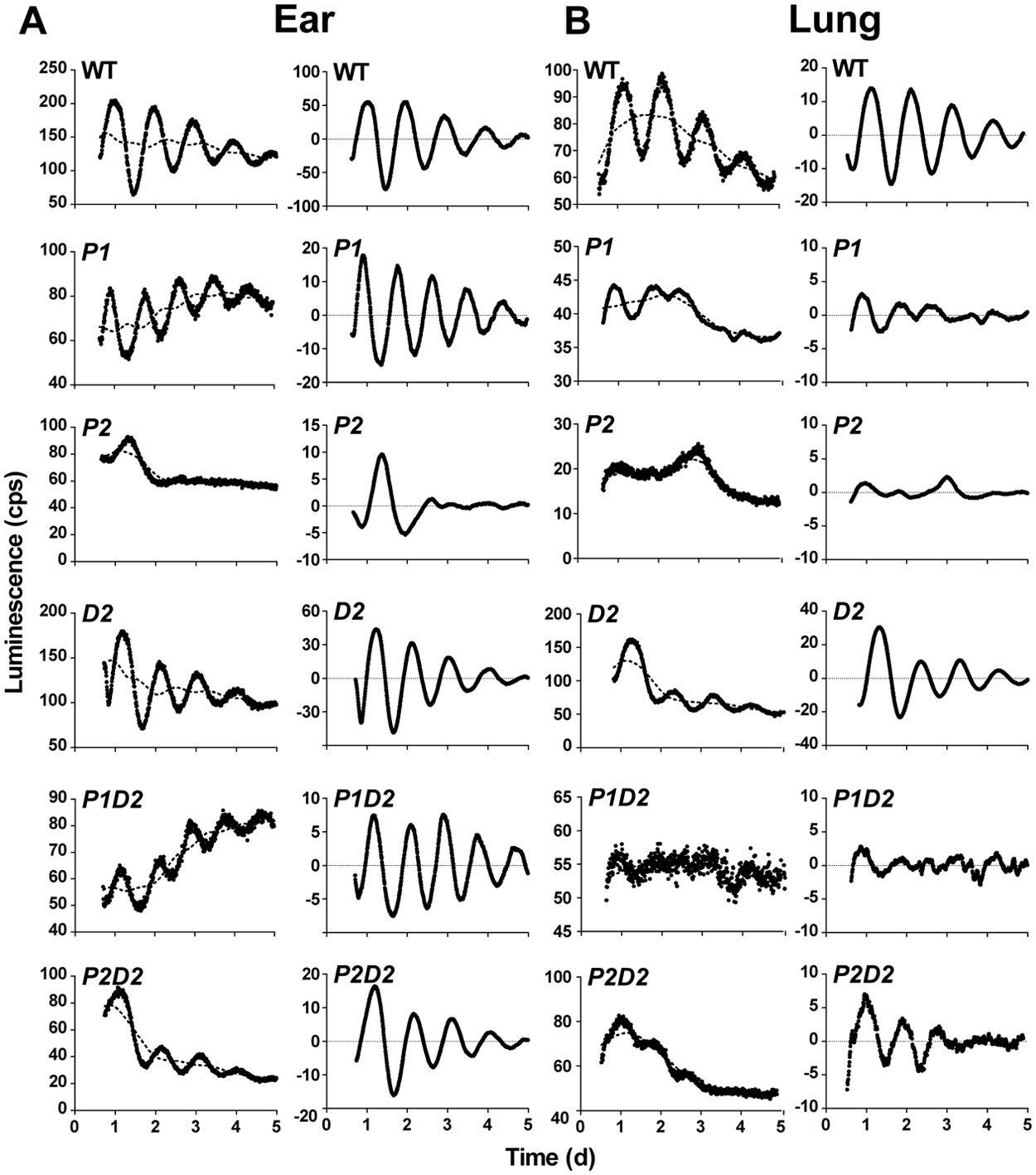
Cellular circadian rhythms in ear- and lung-derived Per/Dec2 single and double mutant fibroblasts. (A, B) Representative luminescence recordings from wild-type (WT), Per1 (P1), Per2 (P2), Dec2 (D2) single, and Per1/Dec2 (P1D2) and Per2/Dec2 (P2D2) double mutant fibroblasts derived from adult mouse ear (A) and lung tissue (B). Left panels show raw data (black circles) and 24-h moving average baselines (dotted lines). Right panels show the same curves after baseline subtraction.
Tissue-Specific Period Regulation in Per/Dec-Deficient Fibroblasts
Period length was shortened in P1 relative to WT cells from both tissues (Fig. 2A, B). D2 cellular rhythms were significantly shortened in ear-derived (Fig. 2A) but not in lung-derived fibroblasts (Fig. 2B). P1D2 ear fibroblasts showed short periods comparable to those seen in P1 single mutant cells (Fig. 2A), indicating that period length is affected less by Dec2 than by Per1 in this tissue. In contrast in lung cells, arrhythmicity was observed after simultaneous mutation of Per1 and Dec2 (Fig. 2B). Surprisingly, circadian rhythmicity was rescued in Per2 mutant fibroblast preparations after additional deletion of Dec2 (P2D2). While P2D2 ear fibroblasts showed short periods comparable to those of D2 cells (Fig. 2A), in lung-derived P2D2 cells periods were shorter than those of either WT or D2 single mutant fibroblasts (Fig. 2B). Comparable results were obtained when sine wave fits were used instead of periodogram analyses (Suppl. Fig. S3).
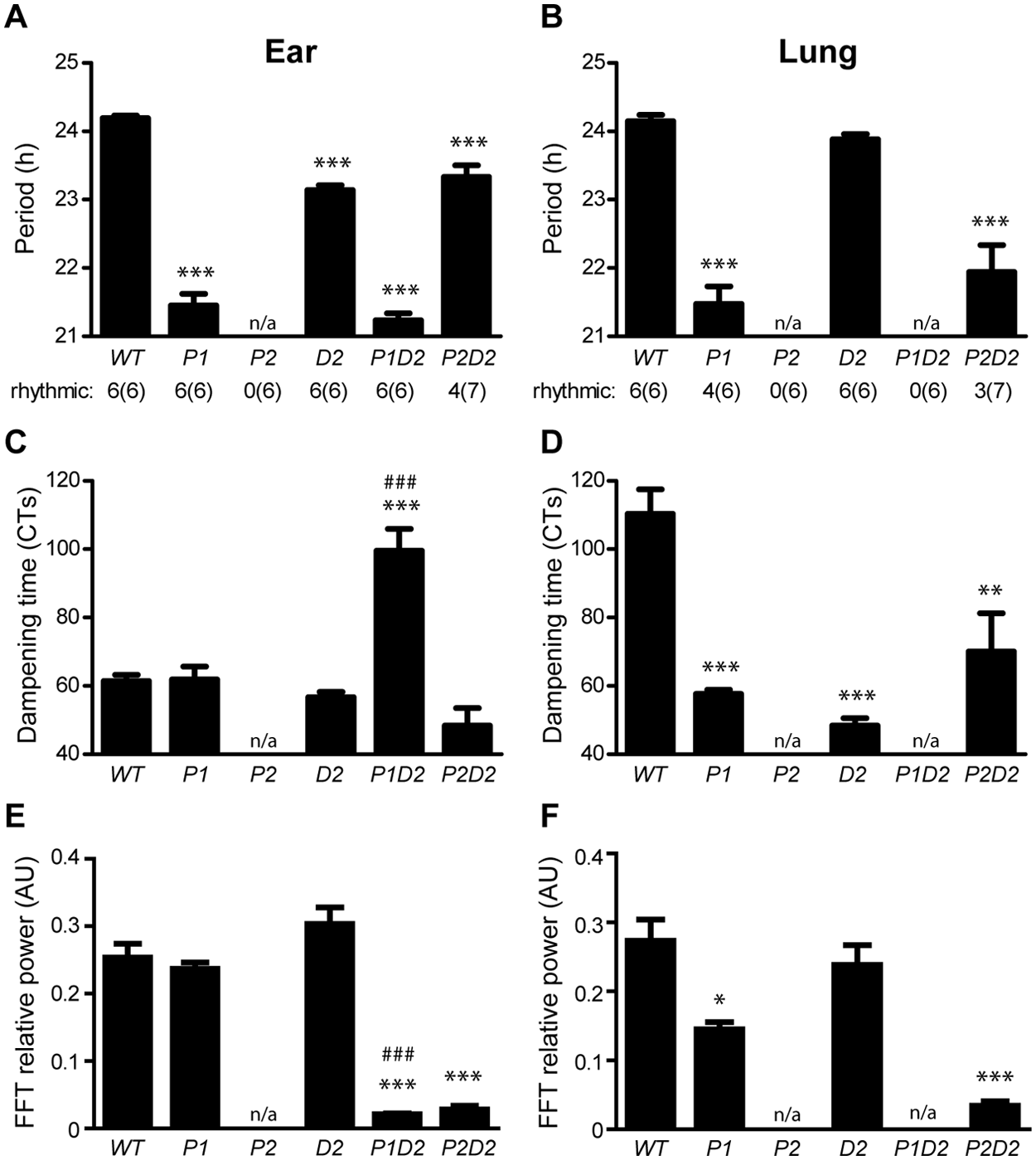
Period length and rhythm dampening in ear- and lung-derived Per/Dec2 single and double mutant fibroblasts. (A, B) Period length as determined by χ2 periodogram analysis on baseline-subtracted recordings from ear-derived (A) and lung-derived cells (B). Values below depict numbers of rhythmic and total (in parentheses) preparations. (C, D) Dampening times as determined by dampened sine wave fitting on baseline-subtracted recordings from ear-derived (C) and lung-derived cells (D). Dampening times were normalized to internal period to allow a better comparison of rhythm stability between cells with different internal periods. (E, F) Fast Fourier transformation (FFT) power analyses of luciferase activity during the first 3 days of the recordings from ear-derived (E) and lung-derived cells (F). All data are presented as means
Tissue-Specific Regulation of Dampening Time and Rhythm Strength in Per/Dec-Deficient Fibroblasts
The restoration of circadian molecular rhythmicity in P2D2 double mutant cells was very reminiscent of the rescue in behavioral locomotor rhythmicity observed in P2D2 double mutant animals (Bode et al., 2011a). To further assess interactive effects of Per1, Per2, and Dec2 on cellular rhythmicity, we quantified Bmal1:luc activity rhythm sustainment by determining dampening times and rhythm stability after fitting a dampened sine wave to baseline-subtracted luminescence recordings. For ear tissue-derived fibroblasts, dampening times were unaltered in P1, D2, and P2D2 while, surprisingly, an increase in amplitude stability was observed in P1D2 cells (Fig. 2C). In contrast, in lung-derived cells, rhythm sustainment was strongly and consistently decreased in P1, D2, and P2D2 compared with WT (Fig. 2D). Rhythm stability values correlated well with dampening times (Fig. 2E, F), with faster dampening correlating with reduced rhythm power. Notable exceptions were seen in ear-derived P1D2 and lung-derived D2 cells. In ear P1D2 cells, reduced rhythm power rates suggested that the extended amplitude sustainment was compromised by low period stability, making the oscillation rather unreliable. At the same time, in D2 lung fibroblasts, short dampening times were combined with relatively high rhythm power, indicating that early rhythm cessation may not derive from unstable period regulation per se.
Tissue-Specific Regulation of Circadian Phase in Per/Dec-Deficient Fibroblasts
One of the most prominent interactive phenotypes observed in Per/Dec double mutant mice was seen in the regulation of locomotor activity onset (i.e., phase angle) under entrained conditions and after release into free-run (Bode et al., 2011a, 2011b). We did not measure rhythm entrainment in our cells (e.g., by ambient temperature cycles). However, because dexamethasone synchronization represents a type 0 resetting paradigm, release into free-run can be simulated—and thus Bmal1:luc phase differences between genotypes estimated—in vitro by comparing the time of the first peak of luminescence in rhythmic fibroblasts after synchronization, and it has been shown in cultured fibroblasts that after synchronization, the intrinsic phase relationship among the expressions of different clock genes are well preserved and comparable to the in vivo situation (Saini et al., 2012). WT ear fibroblasts peaked at 13.4 ± 0.4 h after synchronization (Fig. 3A). In P1 cells—and similar to what was seen for period length (Fig. 2A, B)—phasing of the first peak was advanced (11.0 ± 0.6 h), while in D2 (17.4 ± 0.1 h), P1D2 (17.3 ± 0.6 h), and P2D2 (16.3 ± 0.7 h), cells peak phase was delayed relative to WT fibroblasts (Fig. 3A). In lung-derived cells, the situation was similar with WT cells peaking at 11.2 ± 1.2 h after synchronization, P1 cells showing highest luciferase activity significantly earlier at 2.9 ± 0.1 h, and D2 cells showing a delayed peak (17.6 ± 0.2 h) compared with WT cells. Unlike what was seen in ear-derived cells, phasing in P2D2 lung cells (9.5 ± 1.0 h) was not delayed relative to WT fibroblasts (Fig. 3B).
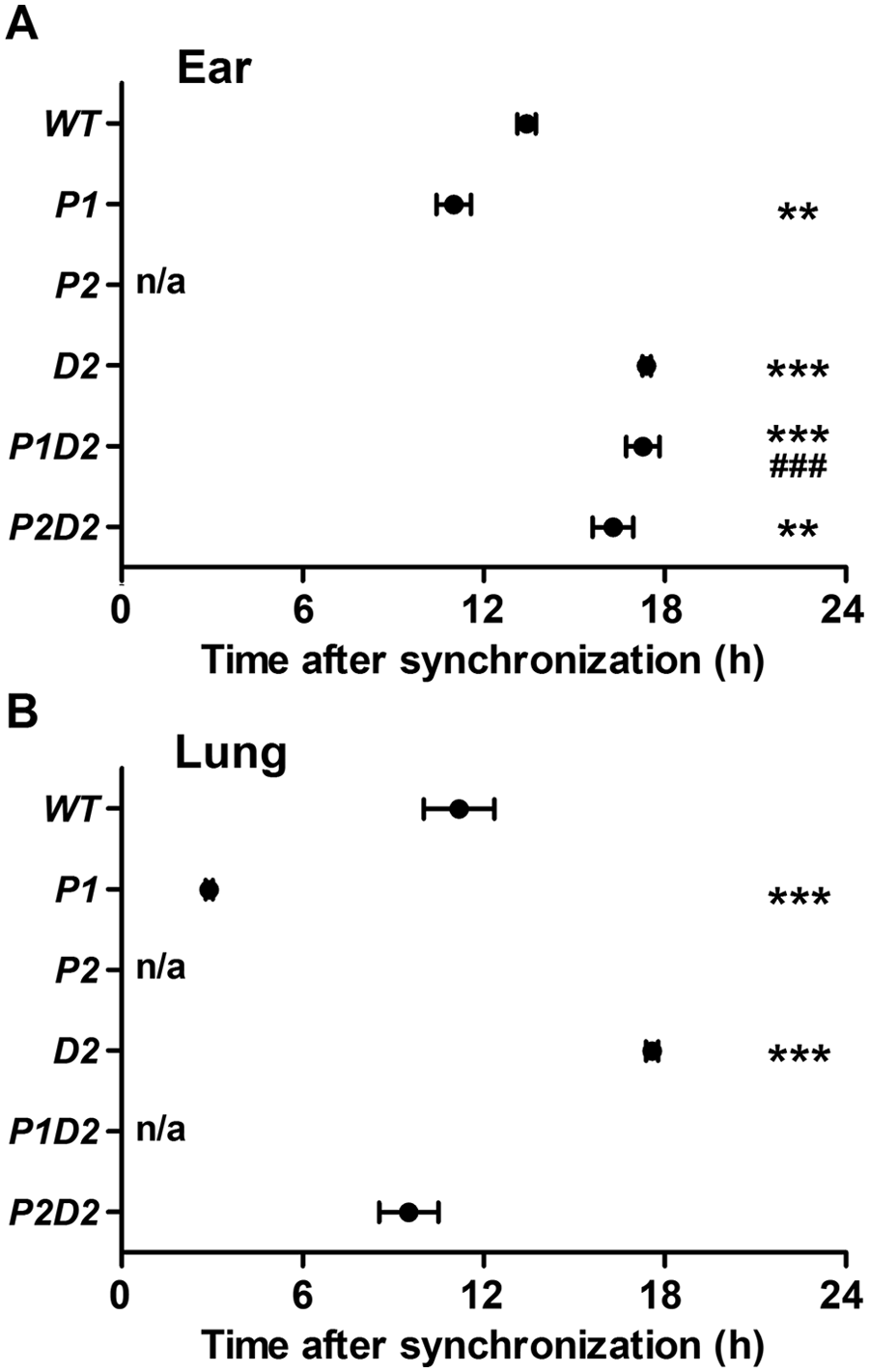
Bmal1:luc peak phase in ear- and lung-derived Per/Dec2 single and double mutant fibroblasts. Timing of first peaks of baseline-subtracted luminescence recordings after dexamethasone synchronization in ear-derived (A) and lung-derived Bmal1:luc-transduced fibroblasts (B). All data are presented as means
Correlation between Period Length and Circadian Phase
Previous data from animals and humans suggest a strong correlation between period length and phasing of behavioral and molecular rhythms (Brown et al., 2008; Liu et al., 2007; Meng et al., 2008; Oster et al., 2003; Ralph and Menaker, 1988; Zheng et al., 1999). In our study, we also observed some genotypes in which shortened period coincided with advanced phasing (e.g., P1 for both tissues; compare Figs. 2 and 3). To further analyze this relationship, we plotted peak time against period length for all preparations. When both tissues were pooled, no significant correlation between the 2 measures was found (data not shown). However, when we compared genotype effects separated for tissues, phasing did not correlate with period in ear-derived fibroblasts (Fig. 4A), whereas a strong correlation (p < 0.001) between the 2 parameters was seen for lung (Fig. 4B). In all cases, the extent of advance in phasing was much higher than that of period shortening, suggesting that period variations may contribute but are not sufficient to explain the observed phase advances.
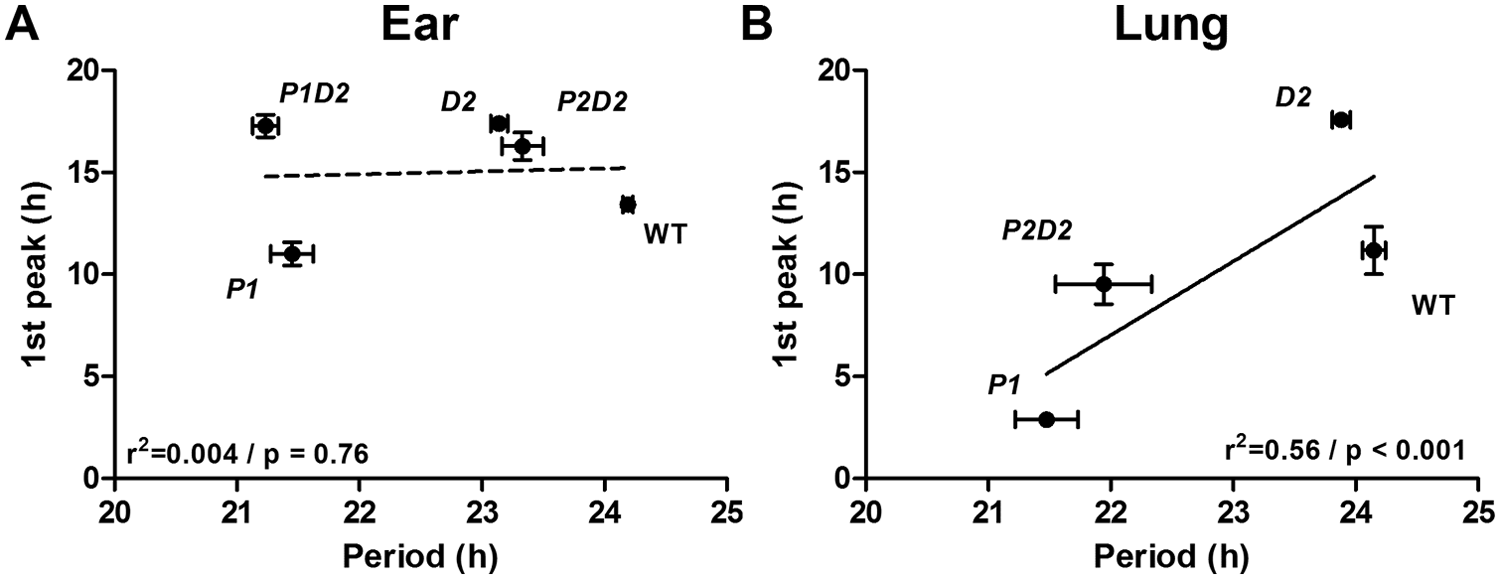
Correlation of phase vs. period length in ear- and lung-derived Per/Dec2 single and double mutant fibroblasts. (A, B) Correlation of phase and period in ear-derived (A) and lung-derived synchronized fibroblasts (B) of wild-type (WT), Per1 (P1), Per2 (P2), Per1/Dec2 (P1D2), and Per2/Dec2 (P2D2) mutant mice. All data are presented as means
Interaction of PER and DEC Proteins in the Regulation of Dbp Transcription
Our cell culture data suggested that Per1/2 and Dec2 have tissue type-independent synergistic roles in period regulation. So far, physical interaction of PER and DEC proteins has not been demonstrated, but it is known that they can both act on CLOCK/BMAL1 activity at E-box promoters (Bode et al., 2011b; Honma et al., 2002; Jin et al., 1999; Rossner et al., 2008). Genetic and modeling experiments suggest that relieving E-box repression, for example by enhanced destabilization of PER proteins, shortens free-running period (Gallego et al., 2006; Maywood et al., 2011). We therefore hypothesized that synergistic de-repression of CLOCK/BMAL1 activity may underlie the interactive effect of Per1/2 and Dec2 deletions on period regulation. The opposite effect would be expected upon overexpression of PER and DEC proteins. To test this we measured luciferase activity in HEK 293T cells transfected with a Dbp-luc reporter, Clock, and Bmal1 and various combinations of Per1, Per2, and Dec2 expression plasmids. As expected, expression of Clock and Bmal1 resulted in a prominent increase in luciferase activity (Fig. 5A). Coexpression of Per1, Per2, or Dec2 alone led to a moderate inhibition of CLOCK/BMAL1 (although significance was not reached in a 1-way ANOVA). In contrast, cotransfection of Per1 or Per2 with Dec2 produced a significant repression of CLOCK/BMAL1 activity with luciferase activity levels comparable to control conditions. A second tissue type–independent finding from our fibroblast experiments was that P2D2 mutant cells had consistently longer periods than P1D2 mutants. Assuming that this effect was also mediated by differential regulation of CLOCK/BMAL1 transactivation and following the same argument as above, this would mean that PER1 and DEC1 together (i.e., resembling the P2D2 mutant situation) should have a stronger inhibitory effect on CLOCK/BMAL1 than PER2/DEC1 (resembling P1/D2). We tested this by cotransfecting Dec1 with Per1 or Per2 in HEK 293T cells as described above. While Dec1 alone had a moderate effect on CLOCK/BMAL1 activity, cotransfection with Per1, but not with Per2, caused further repression of Dbp-luc activity (Fig. 5B). Together, these data suggest that the interaction of Per1/2 and Dec2 in the regulation of the circadian TTL may—at least with respect to period—be mediated via E-box regulation.
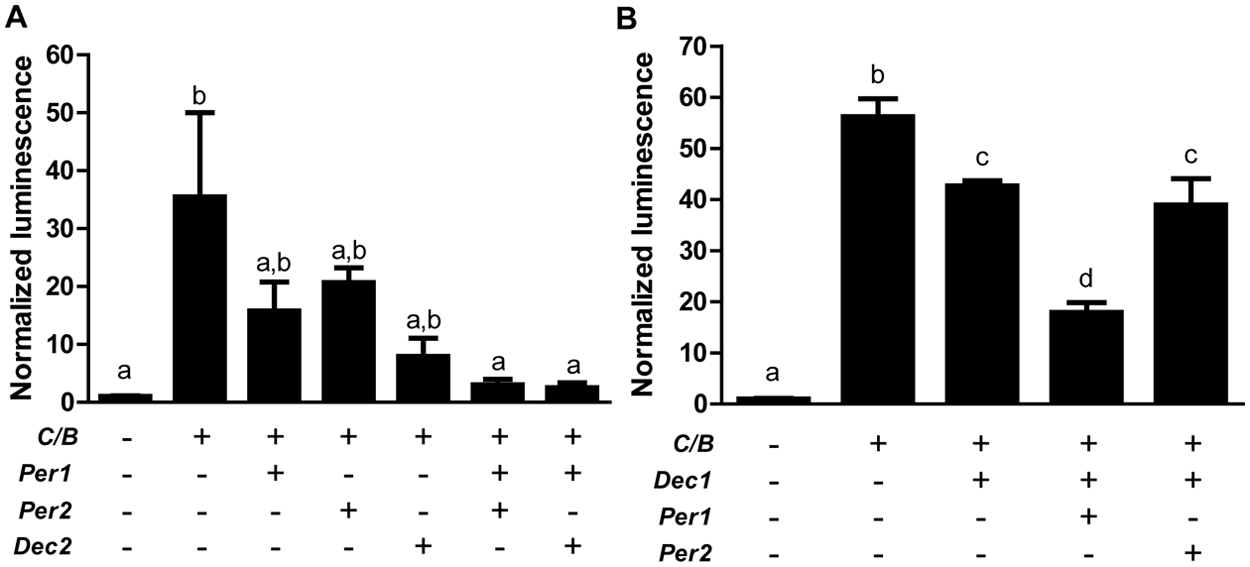
Interaction of Pers and Decs in the regulation of CLOCK/BMAL1-induced Dbp-luc activity. (A, B) Luciferase activity in HEK 293T cells after transfection with different combinations of expression plasmids depicted below. Data are presented as means
Discussion
Our data suggest a predominantly synergistic interaction between Per1, Per2, and Dec2 in cellular clock regulation. This interaction is tissue-specific, affecting rhythmicity itself as well as rhythm sustainment, period length, and stability. In contrast, nonsynergistic interactions were seen in the regulation of cellular clock phase. Luciferase promoter assays suggest that Per/Dec interaction is mediated by interaction at the level of E-box regulation.
Synergistic but Tissue-Specific Interaction of Dec2 and Per1/2
It has previously been shown that the effects of clock gene mutations on circadian rhythmicity may differ between tissues (Cermakian et al., 2001; DeBruyne et al., 2007b; Jakubcakova et al., 2007; Nakashima et al., 2008; Reick et al., 2001; Robles et al., 2010). Due to its unique intercellular coupling, the SCN pacemaker is particularly robust against genetic perturbation (Hogenesch and Herzog, 2011; Liu et al., 2007). This seems also true for Per/Dec mutations. When one compares the cellular data obtained from this study with previous reports on Per/Dec mutant behavior and SCN clock gene expression (Bode et al., 2011a, 2011b; Rossner et al., 2008; Zheng et al., 2001; Zheng et al., 1999), overall larger effects on cellular rhythmicity than on SCN-controlled locomotor behavior are observed (Table 1). For example, deletion of Dec2 had little effect on running-wheel activity rhythm period, phase, and stability, whereas period (ear), phase (ear and lung), and sustainment (lung) were affected in D2 fibroblasts. The absence of communication between single oscillators in cell culture, however, may not be the only reason for the observed differences. Tissue-specific alterations in the clock gene machinery itself seem to also play a role as cells from ear or lung preparations react differently to genetic perturbation. For example, Bmal1:luc peak phase was delayed in P2D2 ear fibroblasts, whereas no change relative to WT was seen in lung-derived cells. In contrast, rhythm sustainment was unaltered—although rhythm power was strongly reduced—relative to WT cells in P2D2 ear fibroblasts whereas lung-derived cells showed accelerated dampening in their luminescence rhythm. Along this line, a mutation of Per1 has previously been shown to result in circadian arrhythmicity in dispersed SCN neurons and in tail tissue-derived fibroblasts (Liu et al., 2007), whereas rhythmicity was preserved in our setup (though with accelerated dampening and reduced rhythm power in lung). Together these data indicate a certain degree of variability in the composition of the clock gene machinery between tissues or, alternatively, the existence of tissue-specific modulators of clock function. A tissue-specific nature has previously been postulated for the role of Clock in circadian regulation (Kennaway et al., 2006). Interestingly, a mutation of Clock also differently affects Dec2 expression in different tissues (Noshiro et al., 2005).
Comparison of behavioral and cellular phenotypes.
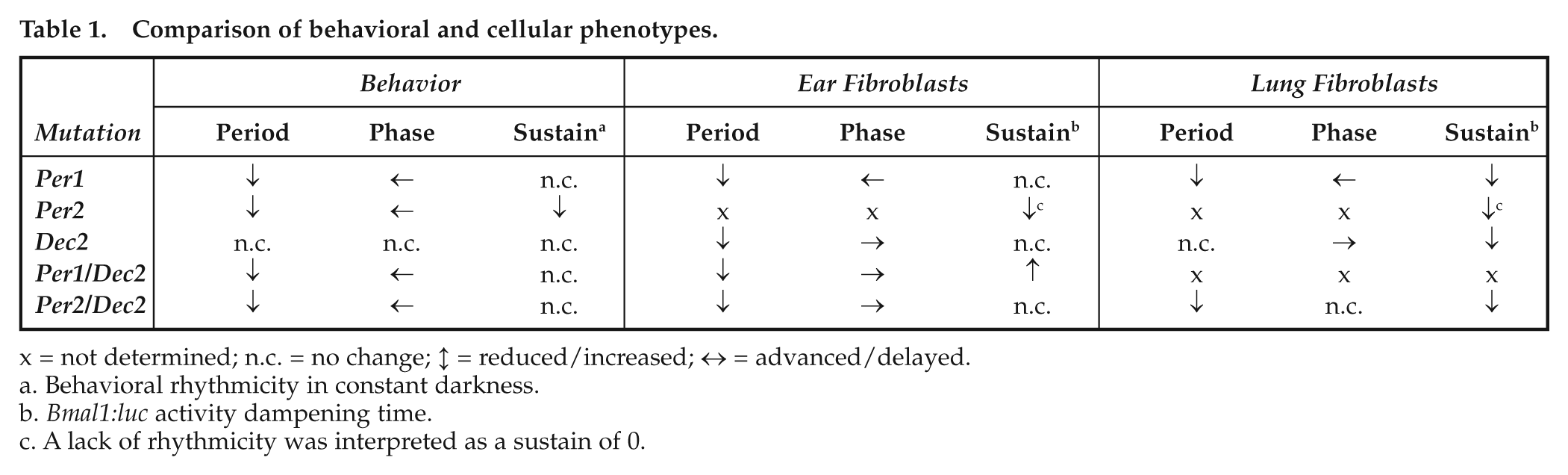
x = not determined; n.c. = no change; ↕ = reduced/increased; ↔ = advanced/delayed.
Behavioral rhythmicity in constant darkness.
Bmal1:luc activity dampening time.
A lack of rhythmicity was interpreted as a sustain of 0.
Dec2 Deletion Rescues Rhythmicity in Per2 but Not in Per1 Mutant Cells
A phenotypic rescue of clock rhythmicity after additional mutation of Dec2 in a Per2 mutant background was seen both at the behavioral level (Bode et al., 2011a) and in fibroblasts (Table 1), similar to what had previously been observed for circadian behavior and in vivo clock gene rhythms after deletion of Cry2 and Per2 (Oster et al., 2002). Together, these data suggest that Per2 function in the circadian TTL is strongly modulated by other clock regulators and that under certain conditions, Per2 may even be dispensable for circadian rhythm generation. This interpretation is further supported by recent studies that report normal rhythmicity and period regulation in Per2ldc mutant mice (Pendergast et al., 2010; Xu et al., 2007). Surprisingly, our data suggest that an additional mutation of Dec2 in a Per1 mutant background can stabilize cellular rhythm sustainment, while at the same time rhythm power is strongly reduced. It is generally believed that cellular oscillators in culture do not communicate with each other (Liu et al., 2007). Therefore, rhythm dampening in cell culture is mainly based on 2 factors: individual differences in endogenous cellular period lengths and rhythm destabilization at the molecular level. Imaging studies in WT fibroblast cultures show that cellular clocks are self-sustained and single cells robustly cycle over extended periods of time while overall phase coherence is rapidly lost (Leise et al., 2012; Nagoshi et al., 2004). Our data support the hypothesis that individual period variation may contribute to rapid dampening in Per1 mutant cells and that—specifically in ear-derived fibroblasts—this effect is compensated after additional mutation of Dec2. The finding that in both P1D2 and P2D2 double mutant cells rhythm sustainment does not correlate with increased rhythm power indicates a rather high variability in day-to-day period regulation, rendering the remaining rhythm highly unreliable. Per1 mutant mice show a high variability in locomotor activity, rhythm period, and length regulation under free-running conditions (Zheng et al., 2001), and this effect is preserved in P1D2 double mutant animals (Bode et al., 2011b) and cells (this article), suggesting that the absence of Per genes may confer intrinsic instability to the clock without principally affecting rhythm sustainment per se. However, lacking single-cell recordings, we cannot distinguish between inter- and intracellular effects in our setup.
The observation that rhythm sustainment in ear-derived P1D2 mutant cells seems improved—although with reduced power—when compared with WT cells appears surprising but may emphasize 2 important aspects. First, circadian clocks have not evolved under free-running conditions. Although for physiological entrainment it is important that clocks are capable of predicting time under zeitgeber free conditions and as a counterbalance against external noise, for most species optimization of entrainment under natural zeitgeber cycles might have been evolutionarily much more relevant (Hut and Beersma, 2011). Second, cell culture conditions only poorly recapitulate the native environment to which a cell is exposed inside the body. Thus, interactions between the altered genetic machinery of a mutant cell and components of the culture medium may critically interfere with the regulation and stability of cellular circadian rhythms. Some components of the clock gene machinery have been shown to interact with external signaling pathways. For example, CRY proteins regulate glucocorticoid receptor signaling and cellular energy metabolism in murine fibroblasts (Lamia et al., 2011). Such implications should be taken into consideration when interpreting circadian phenotypes under in vitro conditions.
Nonsynergistic Interaction of Per1 and Dec2 in the Regulation of Cellular Clock Phase
Our analyses of clock phase reveal 2 major findings. First, loss of Per1 has an opposite effect on Bmal1-luc phase regulation than loss of Dec2. In ear-derived P1D2 double mutant cells, phase regulation resembles that of D2 mutants, whereas in lung-derived cells, loss of both genes results in arrhythmicity. Second, peak phase relations between different genotypes are not always correlated to period length. Theoretical considerations suggest that entrainment phase of an oscillatory system to an external zeitgeber is influenced by internal period (Roenneberg et al., 2008). Experimental data from human and animal studies support these postulations (Brown et al., 2008; Ralph and Menaker, 1988; Toh et al., 2001; Xu et al., 2005; Zheng et al., 1999). The extent of this effect, however, is highly dependent on zeitgeber strength. Activity rhythms in rodents, for example, can be stably entrained by extremely low light intensities (Foster et al., 2007), and standard laboratory light regimens are usually far beyond this entrainment threshold, thus minimizing the observed effects of internal period on phase adjustment. In cell culture conditions, the determination of phase will further depend on the selected readout. In this project we used luciferase activity as a real-time reporter of Bmal1 promoter activity. However, with deletion of TTL components in clock gene mutant cells, the normal phase relationship between different TTL transcripts may be lost. Thus, phase relationships between genotypes may be different if another readout (e.g., another clock gene) is chosen. Of note, such shifts in clock gene mRNA profiles were also observed in Per/Dec mutant SCN (Bode et al., 2011a, 2011b). Third, genetic mutations may differentially affect resetting pathways and period regulation (Jakubcakova et al., 2007). The fact that in Per/Dec mutant fibroblasts phase differences are much more pronounced than those observed in free-running period lengths suggests that this may represent a major factor in this context. This assumption is supported by the fact that both Per genes have been implicated in resetting of the TTL in both the SCN and the periphery (Dibner et al., 2010).
Per1/2 and Dec2 Interact at the Level of E-Box Regulation
Our reporter gene assays show that Per1/2 and Dec2 interact in regulating CLOCK/BMAL1 activity on the Dbp promoter. Such mode of interaction has been suggested before for mammals (Rossner et al., 2008) and for the Per/Dec orthologs period and clockwork orange of the Drosophila clock (Kadener et al., 2007). In vitro experiments on nonsynchronized cells, however, only poorly reflect the dynamics of an oscillatory system, and it has indeed been suggested that Dec function in E-box regulation may depend on clock phase (Rossner et al., 2008). A direct physical interaction of PER and DEC proteins has so far not been shown, but even though our transcription assay data indicate that E-box effects are sufficient to explain the period effects observed in Per1/2-Dec2 mutant cells, other mechanisms may play a role or become even more important for the regulation of clock phase and stability or for explaining tissue-specific effects of Per/Dec mutations.
Conclusion
The analysis of oscillatory behavior after combinatorial deletion of different genetic components of the biological clock, in particular in cells, may help us to better understand the complex interactions of the numerous components of the circadian clockwork (for a successful example in plants, see Locke et al., 2005). In mammals this approach is hampered by the fact that multiple mutations frequently result in arrhythmicity, which then prevents further measurements (Cho et al., 2012; DeBruyne et al., 2007a; Oster et al., 2002). We show that in cells, Per/Dec2 interaction is either synergistic (period and rhythm power) or nonsynergistic (phase) and highly tissue-specific. Given that in the SCN, Dec1 and Dec2 have partly redundant function (Honma et al., 2002; Rossner et al., 2008), it would be interesting to extend these studies to Dec1 mutant cells. Many clock components specifically regulate biological processes outside circadian timekeeping (Bradshaw and Holzapfel, 2010; Franken et al., 2007; Rosenwasser, 2010; Sidman et al., 1962). Therefore, describing the modes of clock gene interactions in different tissues may be particularly useful to identify pharmacological targets for the tissue-specific intervention into clock-controlled as well as noncircadian physiological systems.
Footnotes
Acknowledgements
We would like to thank Jana Husse for critical comments on the manuscript. This project was funded by the German Research Foundation (DFG; OS 353/2-1 & KFO 241). H.O. is an Emmy Noether fellow of the DFG and a Lichtenberg fellow of the Volkswagen Foundation. A.H.T. is supported by a GGNB Excellence Fellowship from the University of Göttingen. M.J.R. is supported by the DFG Research Center Molecular Physiology of the Brain (CMPB).
Conflict of Interest Statement
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
