Abstract
Surgical lesion of the suprachiasmatic nuclei (SCN) profoundly affects the circadian timing system. A complication of SCN ablations is the concomitant scission of SCN afferents and efferents. Genetic disruption of the molecular clockwork in the SCN provides a complementary, less invasive experimental approach. The authors report the generation and functional analysis of a new Cre recombinase driver mouse that evokes homologous recombination with high efficiency in the SCN. They inserted the Cre recombinase cDNA into the Synaptotagmin10 (Syt10) locus, a gene strongly expressed in the SCN. Heterozygous Synaptotagmin10-Cre (Syt10Cre) mice have no obvious circadian locomotor phenotype, and homozygous animals show slightly reduced light-induced phase delays. Crosses of Syt10Cre mice with β-galactosidase reporter animals revealed strong Cre activity in the vast majority of SCN cells. Cre activity is not detected in nonneuronal tissues with the exception of the testis. The authors demonstrate that conditionally deleting the clock gene Bmal1 using the Syt10Cre driver renders animals arrhythmic.
In the mouse, gene targeting in embryonic stem (ES) cells as well as N-ethyl-N-nitrosourea (ENU)–mediated mutagenesis plays a key role in elucidating the molecular basis of the mammalian circadian clock (Bunger et al., 2000; van der Horst et al., 1999; Vitaterna et al., 1994; Zheng et al., 2001; Zheng et al., 1999). The mammalian circadian clockwork rests on a limited number of core clock proteins that form interlocked transcriptional-translational feedback loops creating a 24-h rhythm. This machinery drives rhythmic expression of hundreds of clock-controlled genes that regulate a wide range of rhythmic physiological functions (Ko and Takahashi, 2006). Clocks have been found in almost all cells and tissues of the body. They are equipped with the complete molecular clockwork and exhibit a self-sustained circadian rhythm even when kept in explant culture (Balsalobre et al., 1998; Tosini and Menaker, 1996; Yamazaki et al., 2000; Yoo et al., 2004). The mammalian circadian system is organized in a hierarchical manner with the central pacemaker in the suprachiasmatic nucleus (SCN) controlling numerous peripheral clocks (Dibner et al., 2010). Pioneering lesion studies carried out in the early 1970s established the SCN as the central pacemaker (Moore and Eichler 1972; Stephan and Zucker 1972).
Over the past decade, mice deficient for each of the core clock genes have been generated. Typically, such animals are characterized by smaller or greater defects in locomotor circadian rhythmicity and, additionally, may also show a number of phenotypes, including metabolic defects, changes in the reward system, or memory impairments (Abarca et al., 2002; Garcia et al., 2000; Turek et al., 2005). It is as yet unclear whether these defects are clock mediated and, if they are, which tissue clocks exert control. In the adrenal, for example, transplantation experiments have been used to address the function of a tissue clock (Kiessling et al., 2010; Oster et al., 2006). An alternative to transplantation and surgical ablation is the tissue-specific deletion of core clock genes using the Cre-loxP system (Storch et al., 2007; Lamia et al., 2008; Marcheva et al., 2010).
In view of the overarching role of the SCN in circadian timekeeping, we attempted to knock-out clock genes such as Bmal1 specifically in this nucleus. To achieve this goal, we generated a SCN Cre driver that can delete conditional alleles of genes of interest in this nucleus. The Allen Brain Atlas (Lein et al., 2007) has uncovered numerous genes that are expressed in the SCN. However, genes solely expressed therein were not found. In situ hybridization on brain sections identified Synaptotagmin10 (Syt10) as a gene that is strongly expressed in the SCN with relatively few other expression sites in the CNS, even during development (www.genepaint.org, Genepaint ID MH808). Synaptotagmins are involved in regulated exocytosis of synaptic vesicles and are thought to function as calcium sensors (Gustavsson and Han, 2009). Most Synaptotagmins are widely expressed throughout the nervous system, and Syt1, 2, 4, 5, 9, 11, 13, 14, and 16 are also found in the SCN, albeit much less enriched therein than Syt10. The presence of multiple Synaptotagmins in the SCN is beneficial for the design of a Cre driver mouse, as it makes it less likely that inserting the Cre recombinase gene into the Syt10 locus would result in a major impairment of neuronal function as paralogs of Syt10 should compensate for a loss of this particular Synaptotagmin.
Here we report the generation and characterization of a Syt10Cre driver line that enables SCN targeting without targeting of peripheral, nonneuronal clocks. We verified the usefulness of the Syt10Cre driver line by knocking out a conditional Bmal1 allele. Depending on the dosage of Cre recombinase, we obtained mice with phenotypes ranging from minimal circadian perturbation to complete arrhythmicity.
Materials and Methods
Cloning of Syt10Cre Targeting Vector
We replaced the ATG in exon 1 of the Syt10 gene by a Cre cassette. Nts 92-229 (NM_018803.2) were replaced with the Cre cassette composed of the iCre (Shimshek et al., 2002), an internal ribosomal entry site (IRES), followed by an enhanced green fluorescent protein (EGFP) reporter cDNA and a Flippase recognition target (FRT)–flanked PGK-neomycin selection marker. An 8.7-kb genomic region (5.3 kb upstream of exon 1 and 3.4 kb down-stream of exon 1) was cloned by recombineering (Liu et al., 2003) from a BAC clone (BMQ295d20). Exon 1 was then replaced by the Cre knock-in cassette.
Generation and Genotyping of Syt10Cre Knock-in Mice
Gene targeting of Syt10Cre knock-in mice was performed by GenOway using 129Sv/Pas embryonic stem cells. DNA from G418-resistant clones was digested with SphI and analyzed by Southern blotting. Targeted cells were identified using a 474-bp probe generated by PCR (P1; forward primer: 5′-CAAGATGGCTTCTTTAATGACCCCAG-3′, reverse primer: 5′-AGAGTTGCACACACTTCGGTGCAC-3′) that hybridizes with the 3′ homology arm. SphI digestion of the targeted allele resulted in an 8.9-kb band in addition to the 14.7-kb wild-type band (Fig. 1A). For positive clones, the 5′ homology arm was independently tested using a suitable PCR probe. Targeted clones were injected into blastocysts. Chimeric offspring were backcrossed to C57BL/6. The resulting F1 generation was screened for germline transmission by Southern blotting as described for ES cells and PCR genotyping using the following primers: Syt10 F: 5′-AGACCTGGCAGCAGCGTCCGTTGG-3′, Syt10 R: 5′-AAGATAAGCTCCAGCCAGGAAGTC-3′, and Syt10 KI R: 5′-GGCGAGGCAGG CCAGATCTCCTGTG-3′. PCR was performed for 38 cycles with an annealing temperature of 65 °C. A wild-type band of 426 bp and a mutant band of 538 bp were separated on a 1.5% agarose gel (Fig. 1C). In order to delete the FRT-flanked neomycin cassette, mutant mice were crossed to an ubiquitously expressing Flippase line on C57BL/6 background (Farley et al., 2000). Offspring were tested for neomycin deletion by PCR. The Flippase allele was out-crossed in the next generation by back-crossing to C57BL/6.
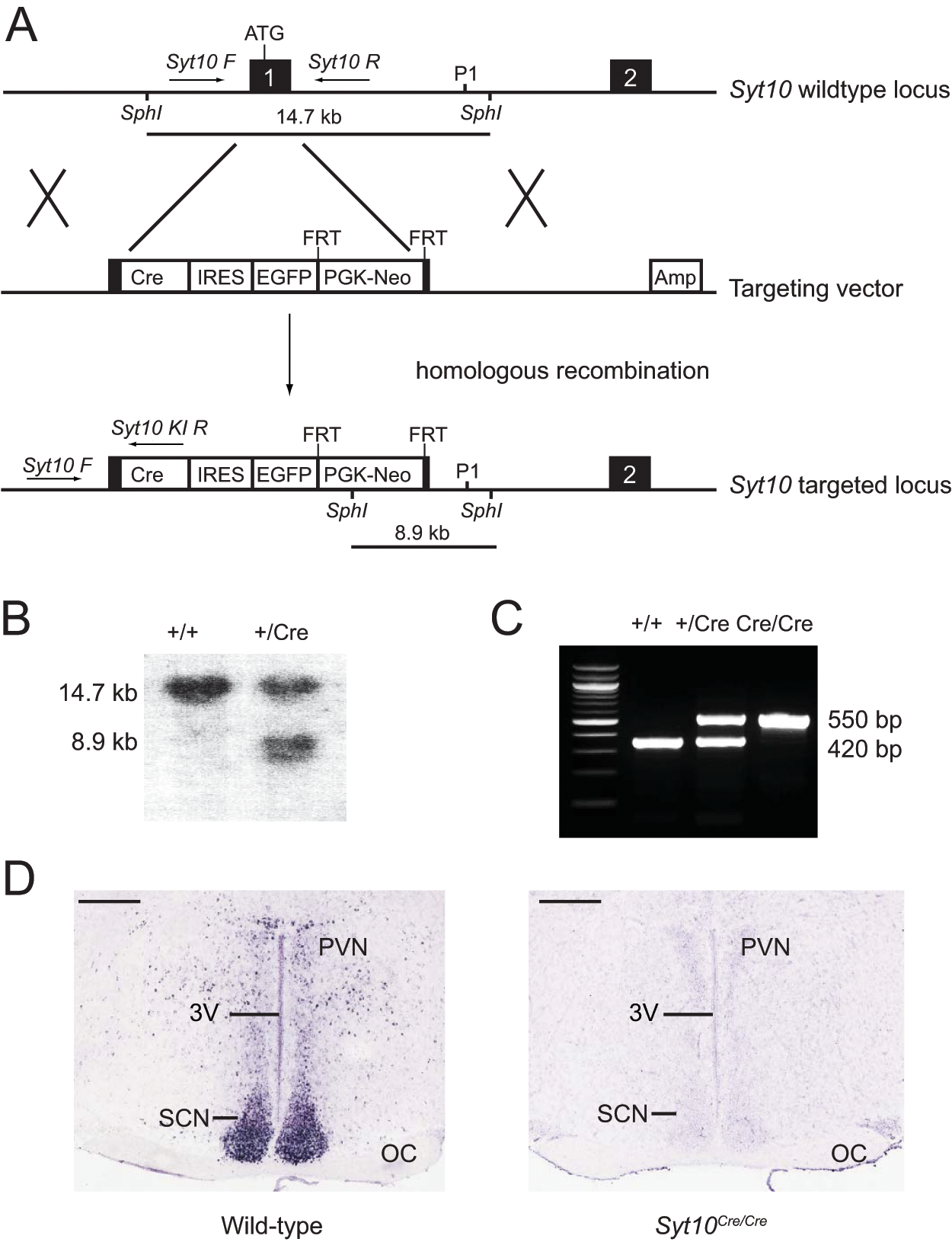
Generation of Syt10Cre knock-in mice. (A) Schematic of the Syt10 wild-type locus, the targeting vector, and the resulting targeted locus. Exons are indicated in black. Position of genotyping primers (Syt10 F, Syt10 R, and Syt10 KI R) are depicted by arrows. Southern fragments are shown as lines. The probe used for Southern genotyping is marked as P1. (B) Southern genotyping of F1 littermates obtained from chimera-C57BL/6 crosses. A 474-bp probe (P1) detected a 14.7-kb wild-type and an 8.9-kb mutant SphI fragment. (C) PCR genotyping of F1 littermates obtained from Syt10Cre/+ intercrosses using the genotyping primers Syt10 F, Syt10 R, and Syt10 KI R. A 420-bp wild-type and a 550-bp mutant band are separated on a 1.5% agarose gel. (D) Syt10 expression in the SCN as determined by in situ hybridization in wild-type and Syt10Cre/Cre mice. Scale bar: 0.5 mm. Abbreviations: 3V, third ventricle; Amp, ampicillin resistance gene; Cre, Cre recombinase; FRT, Flippase recognition target; EGFP, enhanced green fluorescent protein; IRES, internal ribosomal entry site; Neo, neomycin resistance; OC, optic chiasm; PGK, phosphoglycerine kinase A promoter; PVN, paraventricular nucleus of the hypothalamus; SCN, suprachiasmatic nucleus.
β
-
Galactosidase Staining
Syt10Cre/+ R26RLacZ/+ mice were sacrificed by cervical dislocation; brains were quickly removed and frozen in O.C.T (Tissue-Tek). Then, 25-µm cryosections were stained for β-galactosidase as described (Sakurai et al., 2005) and counterstained with Nuclear Fast Red or DAPI. Quantification of recombination efficiency was performed as follows: 10-µm cryosections were first stained with DAPI followed by colorimetric detection of β-galactosidase activity. The SCN was outlined and the number of DAPI-stained nuclei (a measure of cell number) and of β-galactosidase dots were counted, and the ratio of β-galactosidase dots/nuclei was computed. Two sections per SCN region (rostral, central, caudal) from 4 different animals were included. An identical analysis was carried out on SCNs of animals in which β-galactosidase activity is seen throughout the SCN (see Suppl. Table S1 for further information). Finally, the ratio of these two counts was calculated to obtain the percentage of SCN cells expressing Cre recombinase in Syt10Cre/+ R26RLacZ/+ mice.
In Situ Hybridization
Automated in situ hybridization on 25-µm frozen sections was performed as described (Visel et al., 2007; Yaylaoglu et al., 2005). A 1019-bp Syt10 template was used for riboprobe generation (NM_018803, nts 279 to 1298). Riboprobe concentration was 200 ng/µL.
Gene Expression Analysis by Reverse Transcription PCR
Syt10Cre/+ males were sacrificed and tissues were harvested in RNA Later solution (Ambion). RNA was Trizol-extracted (Invitrogen) and DNAse treated (TURBO DNA-free Kit, Ambion). CDNA synthesis was performed (Superscript II, Invitrogen) with the Oligo-dT primer. A 490-bp PCR fragment encompassing the Cre sequence was amplified using the following primers: Cre RT forward: 5′-GTGGATGCTGGGGAGAGAGCCAAGC-3′ and Cre RT reverse: 5′-CAGACCAGGCCAGGTATCTCTGCCC-3′. From the same samples, Eef1a1 was amplified as an internal standard.
Gene Expression Analysis by Quantitative Real-Time PCR
Animals were sacrificed at zeitgeber time (ZT) 18, brains were harvested, and brain punches of the anterior ventral hypothalamus comprising the SCN were taken from 1-mm-thick brain sections. RNA and cDNA were prepared as described above. QPCR was performed using iQ SYBR Green Supermix on an iCycler thermocycler (Bio-Rad) according to the manufacturer’s protocol. Eef1a1 was used as a standard, and quantification was performed as described (Kiessling et al., 2010). Primer sequences were as follows: Bmal1 forward: 5′-TGACCCTCATGGAAGGTTAGAA-3′, Bmal1 reverse: 5′-CAGCCATCCTTAGCACGGT-3′, Eef1a1 forward: 5′-AATTCACCAACACCAGCAGCAA-3′, and Eef1a1 reverse: 5′-TGCCCCAGGACACAGAGACTTCA-3′. Sample sizes were 9 animals for wild-type and 3 animals for each of the other genotypes.
Immunohistochemistry
First, 10-µm frozen sections were fixed in ice-cold 4% PFA for 15 min, washed 3 times in TNT buffer ( 10 mM Tris-HCl, 150 mM NaCl, and 0.05% Tween), blocked in 10% normal goat serum for 1 h, and incubated with anti-BMAL1 antibody (1:1000 rabbit anti-MOP3; Novus Biologicals) overnight at 4 °C. The next day, sections were washed 3 times with TNT buffer, incubated with the secondary antibody (1:400 anti-rabbit AF488; Invitrogen) for 2 h at room temperature, washed again, and mounted with DAPI containing mounting medium. BMAL1-positive cells in a 240 × 187 µm rectangle in 3 medial SCN sections were counted. For each genotype, 3 animals were analyzed.
Behavioral Experiments
All animal experiments were carried out in compliance with the German Law on Animal Welfare. Breeding strategies can be found in Supplementary Table S1. Mouse housing and behavioral monitoring were performed as described (Jud et al., 2005). Males on a mixed 129Sv/C57BL/6 background of 2 to 5 months of age were used. Controls: where feasible, littermates were used. Mice were kept on a 12:12 light dark (LD, 350-lux) cycle before transfer to constant darkness (DD) or constant light (LL, 100 lux). A 15-min 350-lux light pulse was given manually at circadian time (CT) 14. Behavioral data were analyzed using ClockLab acquisition and analysis software package (Actimetrics). Period and amplitude were calculated using χ2 periodogram analysis. Phase shifts in an Aschoff type I protocol were calculated as described (Jud et al., 2005). Onset error was calculated as mean deviation of real onset from a least squares–fit regression line over a period of 10 consecutive days in DD. Ultradian (period of 5-10 h) amplitudes were calculated using χ2 periodogram analysis of 5 consecutive days of the second week in DD.
Data Analysis
Statistical comparisons were made in GraphPad Prism, and p values below 0.05 were considered significant. Normality tests revealed that not all behavioral data followed a Gaussian distribution, and hence nonparametric analyses were performed throughout: Mann-Whitney U test for comparison of two groups and Kruskal-Wallis test for comparison of more than two groups. QPCR and immunohistochemistry data were analyzed using t tests.
Results
Generation of Syt10Cre Mice
The construct used for targeting the Syt10 locus replaced the endogenous Syt10 ATG located in exon 1 by the Cre cDNA sequence (Fig. 1A). This targeting should result in a knock-out of endogenous Syt10. We generated Syt10Cre mice by 129Sv ES cell targeting and subsequent blastocyst injections. We verified the genotype of F1 animals by Southern blotting and PCR (Fig. 1B,C). Subsequently, the FRT-flanked neomycin resistance cassette was deleted. The deletion of Syt10 was confirmed by in situ hybridization to coronal SCN sections. Syt10 transcripts were not detectable in homozygous Syt10Cre/Cre mutants, neither in the SCN (Fig. 1D) nor in any of the other Syt10 expression sites in the brain (not shown).
Syt10Cre heterozygous and homozygous mice are viable and fertile with no obvious morphological abnormalities. Breeding experiments revealed that Cre is active in the male germline. Hence, offspring of a father who is Syt10Cre positive and also carries a conditional allele of the gene to be deleted will be completely deficient in the paternally derived allele of the gene of interest. In the case of the reporter R26RLacZ, the resulting embryos will stain for β-galactosidase ubiquitously (for further information, see Suppl. Table S1).
Activity of Cre Is Brain Specific and SCN Enriched
We crossed Syt10Cre mice to the R26RLacZ reporter line that expresses β-galactosidase after Cre-mediated excision of a stop cassette (Soriano, 1999). Consistent with the in situ hybridization data, β-galactosidase staining was very strong throughout the SCN (Fig. 2A). To estimate the percentage of SCN cells expressing Cre recombinase, the number of β-galactosidase dots in the SCN of Syt10Cre/+ R26RLacZ/+ mice was determined relative to the SCN of mice that ubiquitously express β-galactosidase (see Materials and Method). We found that ~90% of the SCN cells in Syt10Cre/+ R26RLacZ/+ mice were β-galactosidase positive (Fig. 2B). The variation between rostral, central, and caudal levels (93%, 92%, and 84% respectively) is not significant. The β-galactosidase negative cells could be glia cells that do not express Syt10 (Zhang et al., 2004).
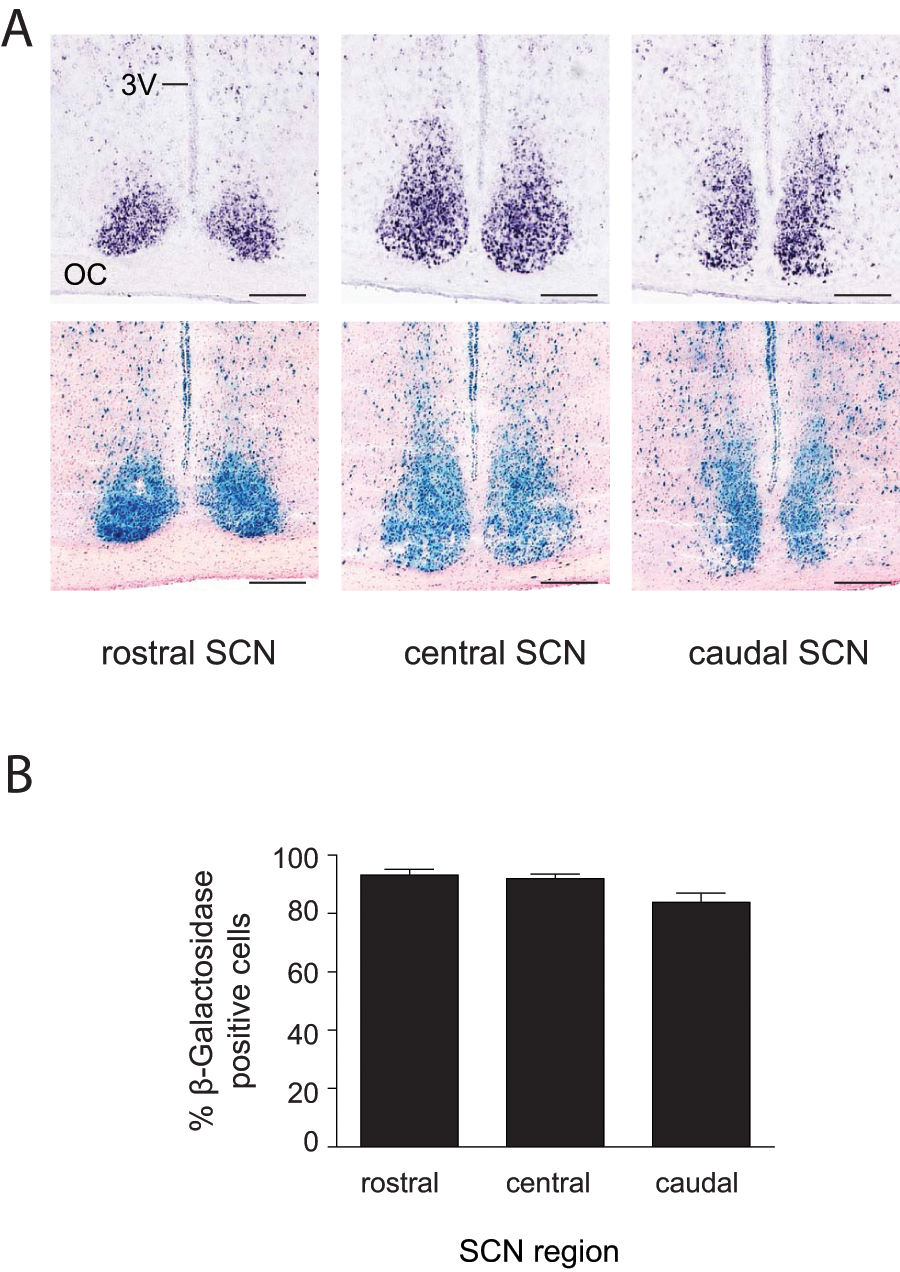
Strong Cre activity in the SCN of Syt10Cre/+ R26RLacZ/+ mice. (A) Syt10 expression and Cre activity at different levels along the rostrocaudal axis of the SCN as determined by in situ hybridization (upper panels) or β-galactosidase staining (lower panels). Sections shown for the two techniques are 25 µm apart. Scale bar: 0.2 mm. (B) Percentage of β-galactosidase positive cells in the SCN. No significant differences were detected between the 3 SCN axial levels (n = 4). Abbreviations: 3V, third ventricle; OC, optic chiasm.
Cre activity was detected in a variety of other brain structures. Coronal sections at the level of the SCN showing Syt10 expression detected by in situ hybridization (Suppl. Fig. S1A) and Cre activity detected by β-galactosidase reporter analysis (Suppl. Fig. S1B) give a good indication of the extent to which Cre is active in non-SCN areas. Supplementary Figure S2 shows β-galactosidase staining in a variety of brain tissues. We observed that Cre-expressing cells are embedded in numerous non-Cre-expressing cells. By contrast, in the SCN, the majority of cells are β-galactosidase positive (Fig. 2A). The β-galactosidase reporter expression pattern was also seen using the human alkaline phosphatase reporter line (Lobe et al., 1999). Phosphatase staining pattern in these mice was very similar to that of β-galactosidase (data not shown).
To examine whether Cre-induced recombination occurred outside the CNS, we performed β-galactosidase staining on sections of various tissues isolated from Syt10Cre/+ R26RLacZLacZ/+ mice (Fig. 3A). We did not detect any Cre activity in peripheral tissues except for the seminiferous tubules of the testis, an expression site presaged by the breeding experiments (Suppl. Table S1). To further confirm the lack of Cre expression in peripheral tissues, we performed Cre-specific reverse transcription PCR reactions on RNA isolated from 19 tissues of Syt10Cre/+ mice. A Cre PCR product was detected only in neuronal tissues, including the eye and the spinal cord, and in testis (Fig. 3B). In summary, the Syt10Cre driver is highly active in the SCN and, to a somewhat lesser extent, in other brain regions. Importantly, we did not detect any Cre activity in nonneuronal tissues with the exception of testis.
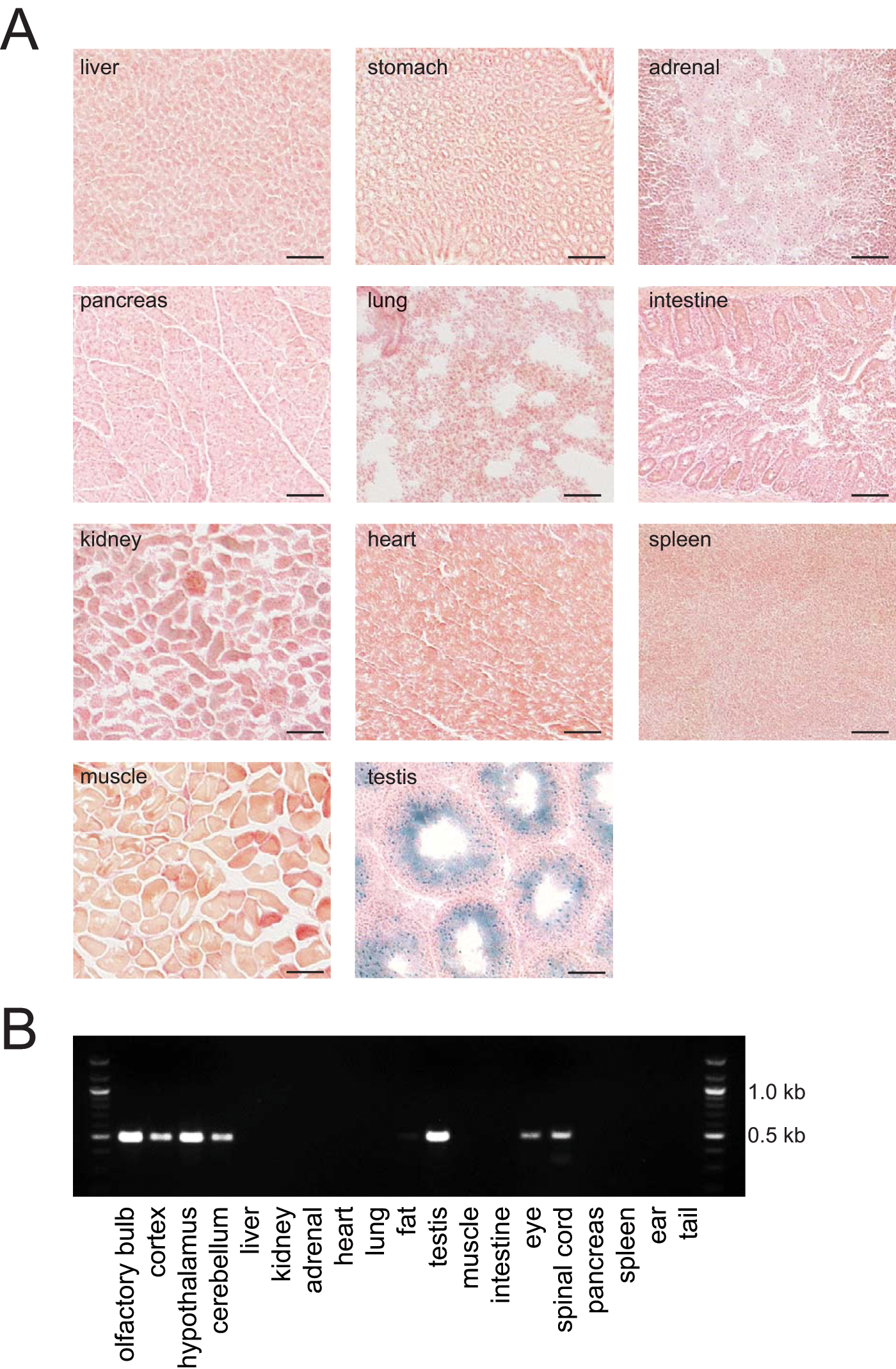
Cre activity is detected only in neuronal tissues and in testis. (A) β-galactosidase staining of various organs isolated from Syt10Cre/+ R26RLacZ+ /– mice. No Cre-mediated recombination resulting in β-galactosidase expression is detectable in peripheral organs with the exception of the seminiferous tubules of the testis. Scale bar: 0.1 mm. (B) Cre-specific reverse transcription PCRs on RNA isolated from various tissues of Syt10Cre/+ mice.
A Minor Light-Resetting Phenotype in Syt10Cre/Cre Homozygous Mice
We analyzed the circadian behavior of heterozygous and homozygous Syt10Cre mice. Male wild-type, Syt10Cre/+ andSyt10Cre/Cre littermates were tested for wheel-running behavior in a standard experimental setup. In a 12:12 LD cycle, Syt10Cre/+ as well as Syt10Cre/Cre mice entrained normally (Fig. 4A). Total activity levels as well as onset variability were not different between genotypes. When released into DD, all genotypes showed a similar endogenous period (Fig. 4B; 23.5 h, 23.4 h, and 23.7 h in wild-type, Syt10Cre/+, and Syt10Cre/Cre, respectively). Locomotor activity onset error as a measure of rhythm instability was also not different between genotypes (Fig. 4C).
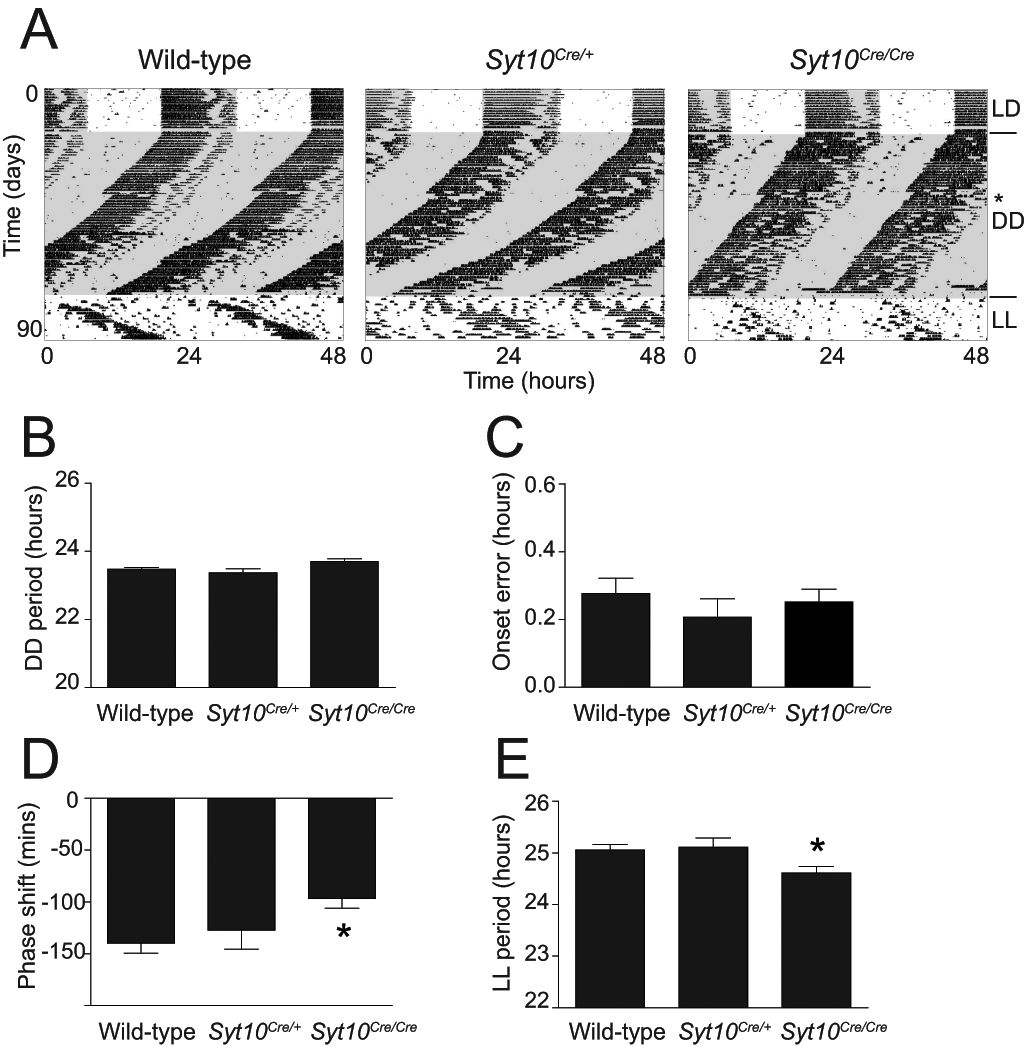
No major circadian locomotor impairments in Syt10Cre mice. (A) Representative double-plotted actograms of wild-type, Syt10Cre/+, and Syt10Cre/Cre mice kept in a 12-h light/12-h dark cycle (LD) and sequentially released in constant darkness (DD) followed by constant light (LL). The day of a 15-min light pulse is indicated with a star. (B) Magnitude of period in constant darkness by χ2 periodogram analysis. (C) Onset error in constant darkness. (D) Magnitude of phase shift after a 15-min light pulse at CT14. (E) Magnitude of period in constant light by χ2 periodogram analysis. All data are shown as mean and SEM and analyzed using a Mann-Whitney test, * indicates p < 0.05 tested against wild-type, n = 8 per genotype.
Next, we investigated the light response of the circadian system in all 3 genotypes by giving a 15-min light pulse at CT14. Wild-type as well as heterozygous mice showed the expected phase delay of approximately 140 min (Benloucif and Dubocovich, 1996). Syt10Cre/Cre mice, however, displayed a reduced phase delay of 90 min (Fig. 4D; wild-type vs. Syt10Cre/Cre; p = 0.006). Given this slightly reduced light response of the circadian system in homozygous mice, we determined their free-running period in LL. This parameter is influenced by both the endogenous circadian period and the light responsiveness of the circadian system (Daan and Pittendrigh, 1976). We did not find differences in the period between wild-type and heterozygous littermates. Homozygous mutants, however, showed a slightly shortened period in LL (Fig. 4E; 25.1 h and 24.6 h in wild-type and Syt10Cre/Cre, respectively; p = 0.03).
Therefore, the Syt10Cre driver line even in the homozygous state seems suitable for circadian experiments as it shows only a minor light-resetting phenotype. Both the circadian period and the stability of locomotor activity rhythms under LD and DD conditions are normal. Heterozygous mutants do not show any impairment in the tested circadian behavioral paradigms.
Cre Dosage-Dependent Circadian Phenotype in Conditional Bmal1-Deficient Mice
The only single-gene knock-out identified so far that produces a complete arrhythmic locomotor phenotype is Bmal1 (Bunger et al., 2000). Hence, we crossed the Syt10Cre line with a mouse line carrying a conditional allele of Bmal1 that allows Cre-mediated deletion of the exon encoding the BMAL1 basic helix loop–helix (bHLH) domain (Bmal1fl/fl) (Storch et al., 2007). We analyzed wheel-running behavior in 12:12 LD and DD conditions.
Syt10Cre/+ Bmal1fl/fl mice showed a shortened period in DD (Fig. 5A,B; 23.5 h and 22.9 h in wild-type and Syt10Cre/+ Bmal1fl/fl, respectively; p = 0.002). These mice were still rhythmic under both LD and DD conditions (Fig. 5A and Suppl. Fig. S3). However, rhythmicity in DD was less stable, and quantification of the onset error revealed increased onset variability in Syt10Cre/+ Bmal1fl/fl mice compared to wild-type controls (Fig. 5C; 0.38 h and 0.78 h in wild-type and Syt10Cre/+ Bmal1fl/fl, respectively; p = 0.038). It appears that in Syt10Cre/+ Bmal1fl/fl mice, the amount of Cre activity was insufficient to fully delete Bmal1 in all SCN cells.
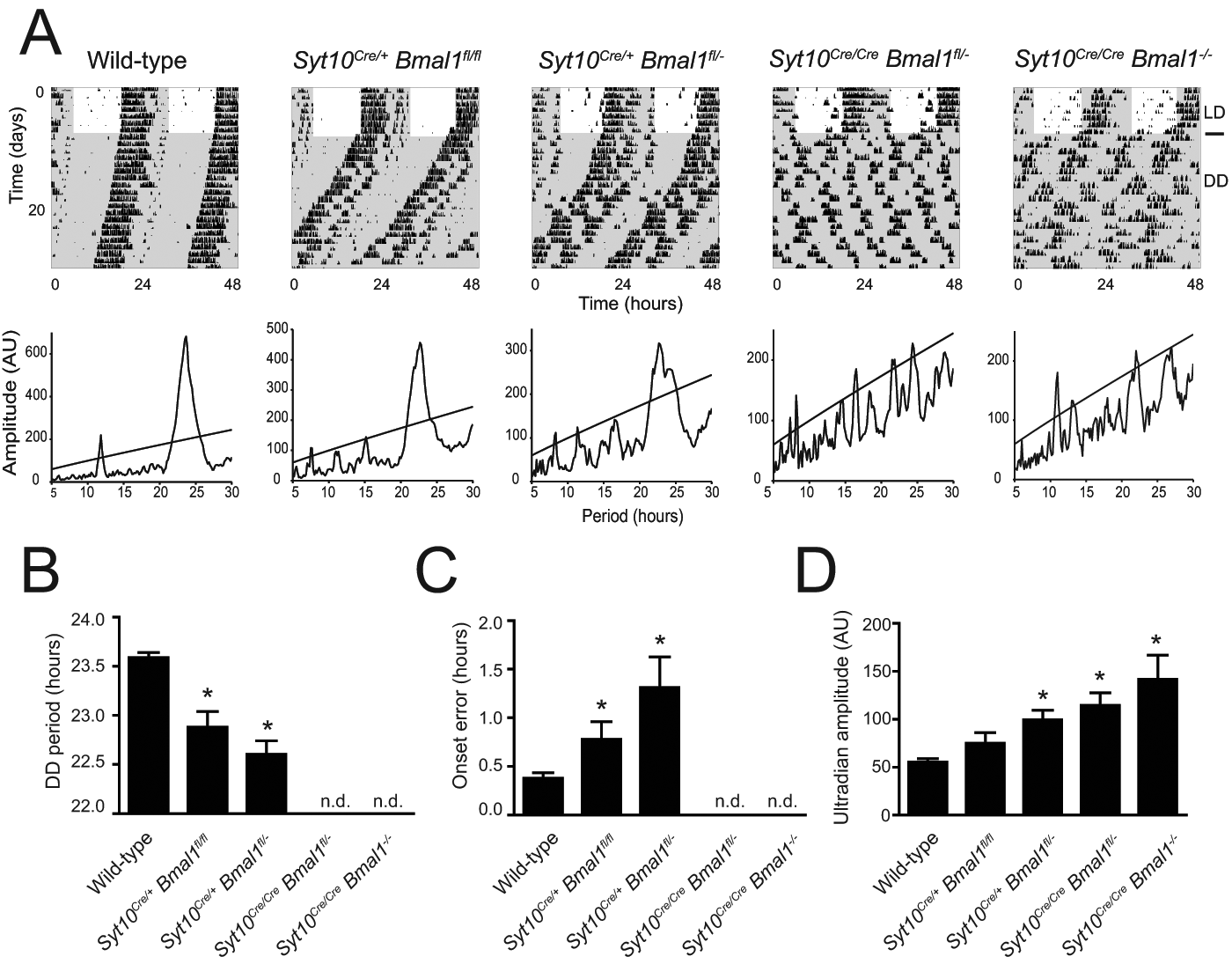
Circadian phenotypes in Syt10Cre-driven Bmal1 knock-outs. (A) Representative double-plotted actograms and periodograms of wild-type, Syt10Cre/+ Bmal1fl/fl, Syt10Cre/+ Bmal1fl/–, Syt10Cre/Cre Bmal1fl/–, and Syt10Cre/Cre Bmal1–/– mice. (B) Magnitude of period in DD determined by χ2 periodogram analysis. In Syt10Cre/Cre Bmal1fl/– and Syt10Cre/Cre Bmal1–/– mice, no circadian rhythmicity was detected (n.d.). (C) Onset error was determined for consecutive 10 days in DD. (D) Amplitudes in the ultradian range (5-10 h) were calculated from 5 days in DD. All data are shown as mean and SEM and analyzed using a Mann-Whitney test; * indicates p < 0.05 tested against wild-type.
Next we used a Bmal1fl/– background that, based on previous work, should in itself not produce a circadian phenotype (Bunger et al., 2000; Storch et al., 2007) but would clearly require less Cre activity since one Bmal1 allele is already mutated. Such Syt10Cre/+ Bmal1fl/– animals have a shorter period in DD compared to wild-type (Fig. 5A,B, Suppl. Fig. S3; 23.5 h and 22.6 h in wild-type and Syt10Cre/+ Bmal1fl/–, respectively; p = 0.0004). Onset variability analysis revealed that Syt10Cre/+ Bmal1fl/– animals have higher onset errors than wild-type controls (Fig. 5C; 0.38 h and 1.31 h in wild-type and Syt10Cre/+ Bmal1fl/–, respectively; p = 0.0024). Overall, Syt10Cre/+ Bmal1fl/– animals display impaired activity rhythms but are not completely arrhythmic. This led us to conclude that a single Cre allele is insufficient to produce enough Bmal1-deficient cells in the SCN to result in total circadian arrhythmicity.
We next examined Syt10Cre/Cre Bmal1fl/– mice and did get a fully arrhythmic phenotype (Fig. 5A and Suppl. Fig. S3). Ten of 11 animals were totally arrhythmic in DD; a χ2 periodogram analysis did not reveal any significant circadian rhythmicity. In LD conditions, these mice were still rhythmic; however, in the light phase, they were more active than wild-type controls (5.3% and 18.2% light activity in wild-type and Syt10Cre/Cre Bmal1fl/–, respectively; p = 0.046). One best appreciates the extent of the arrhythmic phenotype of Syt10Cre/Cre Bmal1fl/– mice by a comparison with pan-Bmal1 knock-out animals (Syt10Cre/Cre Bmal1–/–). The phenotype of Syt10Cre/Cre Bmal1fl/– is indistinguishable from the Syt10Cre/Cre Bmal1–/– phenotype in LD and in DD (Fig. 5 and Suppl. Fig. S3). Periodograms of both genotypes show no clear circadian peak (Fig. 5A). Thus, the Syt10Cre/Cre Bmal1fl/– model is a very efficient SCN knock-out and completely mimics the circadian behavioral phenotype of pan-Bmal1 knock-out mice. Overall, the onset variability, a measure for rhythm instability, was progressively increasing with increasing likelihood that both Bmal1 alleles were deleted (Fig. 5C). It has been shown before that in arrhythmic clock mutants, ultradian rhythms can become more prominent (Abraham et al., 2006). We therefore analyzed the periodogram amplitude in the ultradian range (5-10 h) in the different genotypes. Decreasing circadian amplitudes (Fig. 5A) clearly correlated with increasing ultradian amplitudes (Fig. 5D). In summary, the Syt10Cre evokes a variety of circadian phenotypes ranging from normal rhythmicity to totally arrhythmic phenotypes in a Cre dosage-dependent manner. Increasing the probability of a recombined Bmal1 allele by either using 2 Cre alleles or by working with a fl/– background increases the severity of the phenotype, eventually leading to a complete loss of circadian locomotor rhythmicity in Syt10Cre/Cre Bmal1fl/– mice.
To examine whether the Cre dose-dependent circadian defects correlate with the amount of Bmal1 expression in the SCN, we quantified Bmal1 mRNA and BMAL1 protein levels in wild-type, Syt10Cre/+ Bmal1fl/–, and Syt10Cre/Cre Bmal1fl/– mice at ZT18, which is the time point of maximal Bmal1 expression in the SCN (Oishi et al., 2000). QPCR analysis of SCN punches revealed a significant reduction of Bmal1 mRNA levels to less than 50% of wild-type levels in both conditional genotypes (Fig. 6A; 47% and 40% in Syt10Cre/+ Bmal1fl/– and Syt10Cre/Cre Bmal1fl/–; t test; p = 0.0081 and p = 0.004, respectively). Differences between Syt10Cre/+ Bmal1fl/– and Syt10Cre/Cre Bmal1fl/– genotypes were not significant. This analysis may underestimate the degree of knock-out in the SCN since SCN punches contain non-SCN tissue in which Bmal1 expression is normal. We thus performed immunohistochemistry with an anti-BMAL1 antiserum (for validation of the anti-BMAL1 immunoreactivity, see Suppl. Fig. S4). A progressive reduction of BMAL1 immunoreactivity with increasing Cre dosage was seen (Fig. 6B). Relative to wild-type, the SCN of Syt10Cre/+ Bmal1fl/– mice had a reduction of BMAL1-positive cells to 35%, and in the SCN of Syt10Cre/Cre Bmal1fl/– animals, merely 17% of the cells expressed BMAL1 (Fig. 6C; t test; p = 0.0012 and p = 0.0005, respectively, tested against wild-type). The number of BMAL1-positive cells in Syt10Cre/Cre Bmal1fl/– animals was significantly reduced compared to Syt10Cre/+ Bmal1fl/– animals (Fig. 6C; t test; p = 0.0077). The expression of BMAL1 thus correlated with the behavioral phenotypes of these conditional mutants (see Fig. 5).
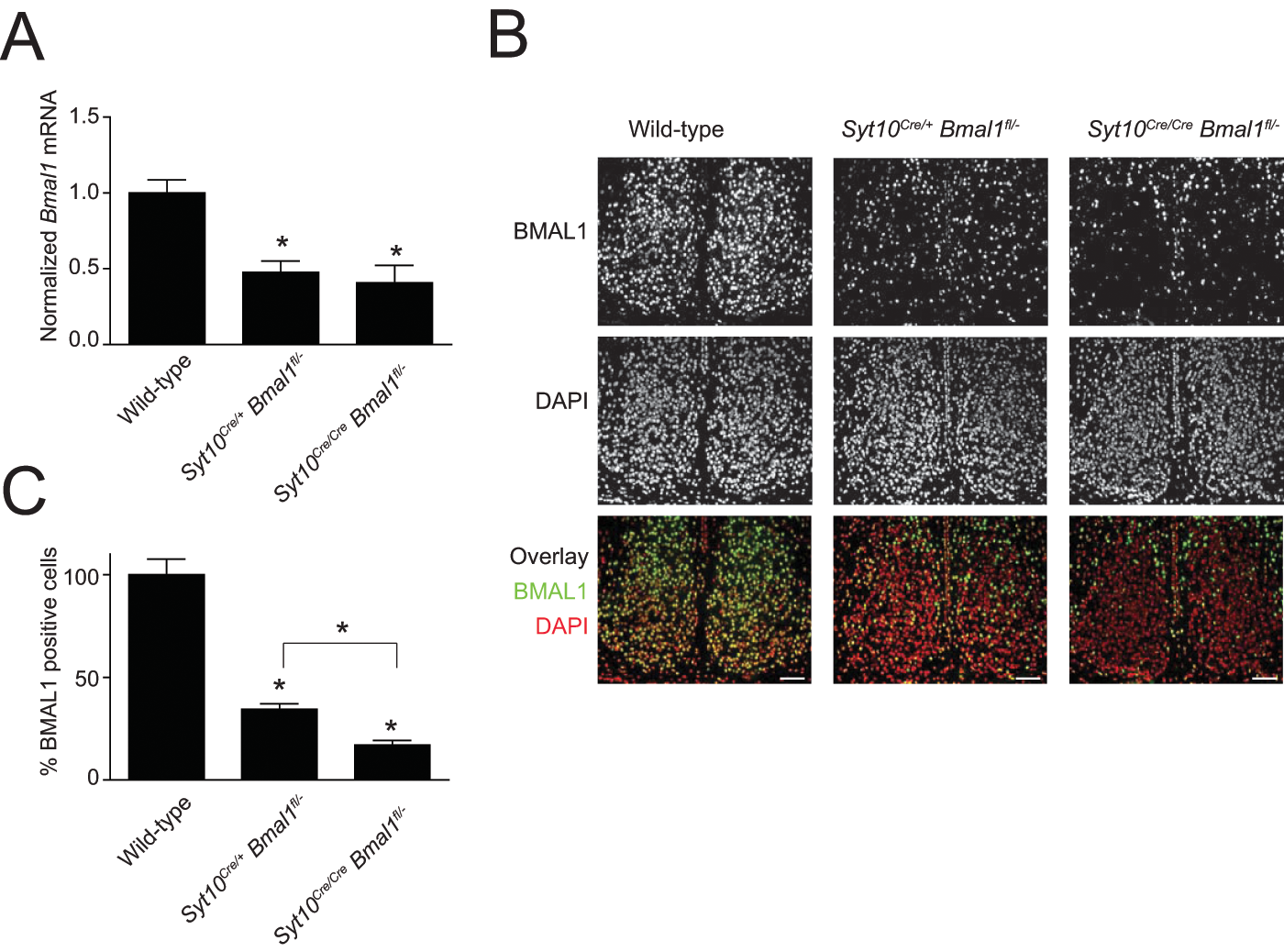
Quantification of Bmal1/BMAL1 levels in the SCN. (A) Relative expression levels of Bmal1 in SCN punches. (B) Representative images of SCN sections immunostained with anti-BMAL1 antibody (top row) and counterstained with DAPI (center). The lower row shows the overlay of BMAL1 immunoreactivity (green) and DAPI (red). Scale bar: 75 µm. (C) Percentage of BMAL1-positive cells in mutant SCN relative to wild-type. All data are shown as mean and SEM and analyzed using a t test; * indicates p < 0.05.
Discussion
The Syt10Cre driver mouse line will be useful to delete conditional alleles of clock or other genes that are expressed in the SCN. Cre activity is found only in neuronal tissues and seminiferous tubules of the testis. Cre activity is strong in the vast majority of SCN cells. We demonstrate that knocking out a conditional allele of the essential clock gene Bmal1 using the Syt10Cre driver renders animals arrhythmic as expected from an efficient SCN clock knock-out. Additionally, we show that the number of BMAL1-positive cells in the SCN correlates with the severity of the behavioral phenotype.
A significant benefit of a knock-in strategy is that it avoids typical problems seen with transgenics such as positional or copy number effects that might complicate an analysis of Cre-mediated effects. By expression of Cre from the Syt10 locus, we achieve Cre activity patterns that are highly similar to the endogenous Syt10 expression pattern. The driver is capable of targeting the majority of SCN cells. The cells that show no Cre-mediated β-galactosidase signal could be glial cells that are present in the SCN (Van den Pol, 1980) and do not to express Syt10 (Zhang et al., 2004).
Knocking out Syt10 has no major effect on circadian behavior, as the period in constant darkness and the entrainment to light-dark cycles remain unchanged. There is, however, a one-third reduction in the magnitude of light-induced phase delays and a slight decrease in the period in constant light. The fact that circadian impairments in Syt10Cre mice are minor may be due to the expression of several other Synaptotagmins in the SCN. The lack of a pronounced circadian phenotype in Syt10Cre mice is a prerequisite for making the Syt10Cre driver line suitable for circadian research.
We observed drastic effects on circadian rhythmicity when the Syt10Cre driver line is used to delete Bmal1. This result is in line with transplantation experiments and inducible expression of dominant negative CLOCK protein, both of which show that the genotype of the SCN determines locomotor period (Hong et al., 2007; Ko et al., 2010; Ralph et al., 1990; Sujino et al., 2003). Germline deletion of Bmal1 and restoring Bmal1 expression under the Secretogranin2 promoter in the brain causes not only circadian defects but also a reduction in life span, body weight, and overall activity levels (Bunger et al., 2005; Bunger et al., 2000; McDearmon et al., 2006). Such deficiencies are not seen in Syt10Cre/Cre Bmal1fl/– animals, an advantage that will facilitate further analysis of the circadian phenotype of these mice. Lesion and transplantation experiments led to one of the major advances in chronobiological research, the discovery of the SCN as the master pacemaker of the mammalian brain (Moore and Eichler, 1972; Ralph et al., 1990; Stephan and Zucker, 1972). The chief difference between deleting Bmal1 in the SCN and SCN lesion experiments is that in the former case, the SCN afferent and efferent neuronal connections are not impaired. The fact that SCN afferents and efferents are destroyed in such lesion experiments may complicate experiments investigating the role of neuronal connections between the SCN and the periphery.
We found gene dosage effects in that the number of BMAL1-positive cells in the SCN correlated with behavioral rhythmicity. This is reminiscent of what has been shown for the CLOCK mutation. It was found that the ratio of CLOCK mutant and wild-type cells in the SCN correlated with behavioral rhythmicity (Low-Zeddies and Takahashi, 2001). The ability to evoke gene deletion in a graded manner should be informative when examining how clock gene dosage in the SCN affects circadian rhythmicity or the expression of other (clock) genes. The benefit of such titrating experiments has also been demonstrated in a recent elegant in vitro study (Baggs et al., 2009). These authors downregulated clock genes in a dose-dependent manner by adding different amounts of small interfering RNAs to human osteosarcoma cells. This led them to uncover novel network features of the circadian timing system. In the context of a recent study showing stochastic rhythmicity in complete Bmal1 knock-out mice (Ko et al., 2010), it will be interesting to investigate whether and how a stepwise reduction of Bmal1 in the SCN affects oscillations of SCN neuronal networks.
The currently known canonical clock genes are not required for viability. However, it would not come as a surprise if yet unknown clock genes existed that are also required for a range of noncircadian functions, including viability. Potential examples are Creb or Glycogen synthase kinase-3β (Iitaka et al., 2005; Obrietan et al., 1999). In such cases, the Syt10Cre driver mouse can be useful to overcome lethality and produce mice suitable for circadian analysis.
When using the Syt10Cre driver line, one should consider the following points: First, the efficiency of Cre-mediated homologous recombination might vary depending on the targeted allele used. It has been shown that the exact location and distance between two loxP sites can affect recombination efficacy (Zheng et al., 2000). Thus, deletion of other conditional gene alleles might require less Cre to elicit a full penetrance of phenotype than in the case of Bmal1fl/fl. Hence, Syt10Cre/+ and Syt10Cre/Cre should be compared with respect to their recombination efficiency. Second, only females (and not males) that are carriers for Syt10Cre and the conditional allele of interest must be used for breeding. Otherwise, germline deletion of the conditional allele will occur, and all cells in the body will be deficient for the gene of interest. A guide for breeding strategies using the Syt10Cre is found in Supplementary Table S1. A caveat with using Syt10Cre in combination with a mouse in which one of the targeted alleles has already been removed (e.g., Bmal1fl/–) is that the heterozygous targeted animal may already have a phenotype. In summary, we believe that our Syt10Cre line will be a helpful tool to investigate the complexity of the mammalian circadian network.
Footnotes
Acknowledgements
We thank Axel Visel and Christina Thaller for their early work on expression patterns of Synaptotagmins. We thank Johanna Barclay for helpful comments on the manuscript. H.O. is an Emmy Noether Fellow of the German Research Foundation (DFG).
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
