Abstract
While the rapid development of personal computers and high-throughput recording systems for circadian rhythms allow chronobiologists to produce huge amounts of data, the software to analyze them often lags behind. Here, we announce newly developed chronobiology software that is easy to use, compatible with many different systems, and freely available. Our system can perform the most frequently used analyses: actogram drawing, periodogram analysis, and waveform analysis. The software is distributed as a pure Java plug-in for ImageJ and so works on the 3 main operating systems: Linux, Macintosh, and Windows. We believe that this free software raises the speed of data analyses and makes studying chronobiology accessible to newcomers.
Nowadays, most chronobiological data recordings are based on computerized systems. The amazing development of computer technology has certainly promoted the speed of our research by enabling high-throughput recording of circadian rhythms. For instance, in our group, the main experimental organism is Drosophila melanogaster, and the locomotor or eclosion rhythms of hundreds of animals are simultaneously recorded either by our in-house recording system (Rieger et al., 2007) or by a commercially available one (Trikinetics, Waltham, MA). The recording systems have improved along with the computer technology, with the result that the quantities of data produced have been increasing year by year. Both of these systems produce numeric data that are saved in a text file in chronological order. There are several commercial and noncommercial software packages available to analyze such chronobiological data (e.g., chronobiological software by Refinetti [http://www.circadian.org/main.html], Brandeis Rhythm Package [Levine et al., 2002], ClockLab from Actimetrics [Wilmette, IL], El Temps by Diez-Noguera [http://www.el-temps.com], BRASS by Millar [http://millar.bio.ed.ac.uk], and free chronobiology software by Hut [http://hutlab.nl]). While these packages offer excellent analysis methods, we have often faced some inconveniences. 1) Many of them are not free, which imposes the burden of high costs particularly when they have to be installed on multiple computers. 2) Most of these systems run only on a particular operating system, often requiring the researcher to find additional alternative hardware to run them on. 3) Some of them can perform quite sophisticated analyses, with many parameters that must be set or tuned. While this is an advantage from one point of view, it typically makes the software more difficult to use, particularly for new users such as students. We believe that such problems slow down our daily data analyses and deter students from joining the field of chronobiology. 4) Furthermore, we have found that the available software solutions do not draw actograms from high-resolution time series data completely accurately. We discuss this in more detail below.
Here, we present our software that overcomes these shortcomings. Our package is implemented using the platform-independent Java programming language, so that it runs identically on Linux, Macintosh, and Windows, both in 32-bit and 64-bit modes. Additionally, we chose an open-source program, ImageJ (http://rsb.info.nih.gov/ij/), a freely available package for image processing in the life sciences, as an underlying platform. ImageJ and its descendant Fiji (Schindelin, 2008) (http://pacific.mpi-cbg.de) are actively developed by an international community and used by thousands of users, guaranteeing the long-term maintenance of our package. Development based on ImageJ enables easy distribution not only among chronobiologists but also in any other bioscience field. To address the third issue, we tried to keep the software self-explanatory. We have deliberately omitted rarely used functions, which in our opinion render available software packages complicated to use. We rather focused on the 3 most frequently used analyses: drawing actograms, periodogram analysis, and waveform analysis.
Actogram
Actogram plots are a common way of displaying time series data in circadian rhythm research. Because we particularly focused on this function, we called this plug-in “ActogramJ”. Drawing an actogram is often the first step to analyze data in circadian rhythm research. This visual representation of circadian data helps us to see overall characteristics of the rhythms. Importantly, the interpretation of the actogram influences following studies. Therefore, the actogram must be clearly and precisely drawn.
Current personal computers allow us to record the rhythms with short time intervals because of their high data-storage capacity. The high-resolution time series recording can reveal different characteristics of behavior such as sleep in Drosophila (Shaw et al., 2000; Hendricks et al., 2000). However, there is a crucial problem with drawing actograms of high-resolution time series data on a computer screen. Due to the limited resolution of typical computer monitors, it is difficult to display a whole actogram on the screen without losing data lines. This problem becomes more severe when the actogram has to be printed as a small figure in an article. To avoid imprecise drawing of high-resolution data, ActogramJ displays the actogram in a form that is adapted to the chosen zoom factor. For example, if the data are collected in 1-minute bins and its actogram is zoomed out by 5 times, the averaged values for 5-minute intervals are drawn on the actogram, which allows us to display all data with a lower resolution. This precise drawing of actograms for the high-resolution data is one special feature of ActogramJ.
Four calculation functions for actogram drawing are implemented in ActogramJ. 1) An average (pure arithmetic) actogram can be drawn from selected actograms. 2) Similarly, the sum of the selected actograms can also be drawn. 3) Data can be smoothed by a convolution with a uniform or Gaussian kernel. 4) By choosing a reference actogram, other actograms can be normalized. This function helps to evaluate the characteristics of the rhythm itself without considering the different activity levels of animals. For normalizing, the total activity for each actogram including the reference actogram is calculated. The individual activities of the actograms are then scaled to match the total activity of the reference actogram.
Figure 1 shows 3 example actograms drawn by ActogramJ, raw and smoothed data from one individual fly, and averaged data of a group of flies. The actograms drawn by ActogramJ can be exported as PDF files for further use.
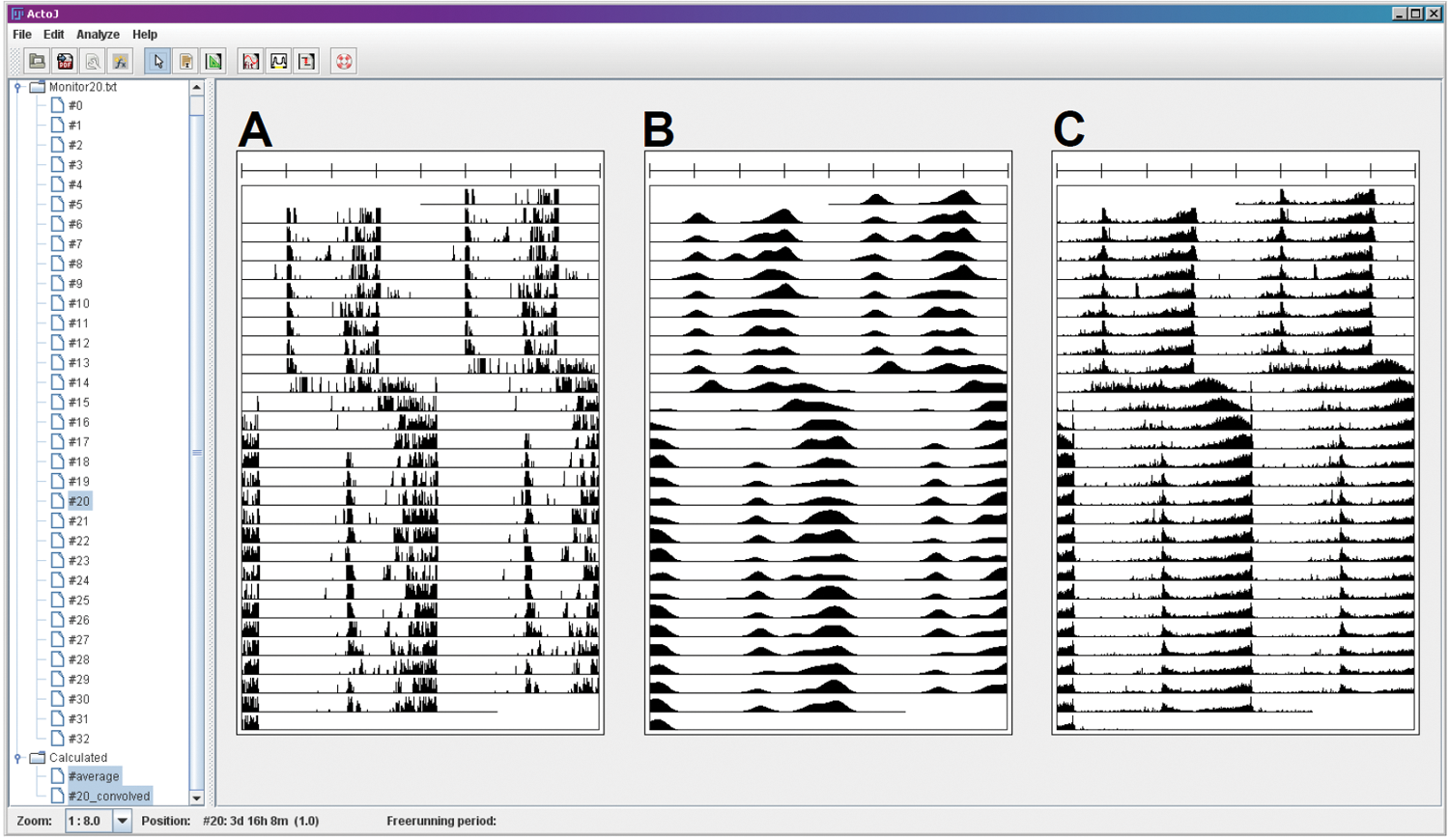
Actograms drawn by ActogramJ. The actograms are of an 8-hour phase shift experiment of light-dark cycles (16 h:8 h) in cry mutant flies (Dolezelova et al., 2007). The locomotor data were collected in 1-minute intervals. (A) The actogram is derived from one individual, (B) the actogram is smoothed by a convolution with a Gaussian kernel (sigma of 50), and (C) the average of 29 flies.
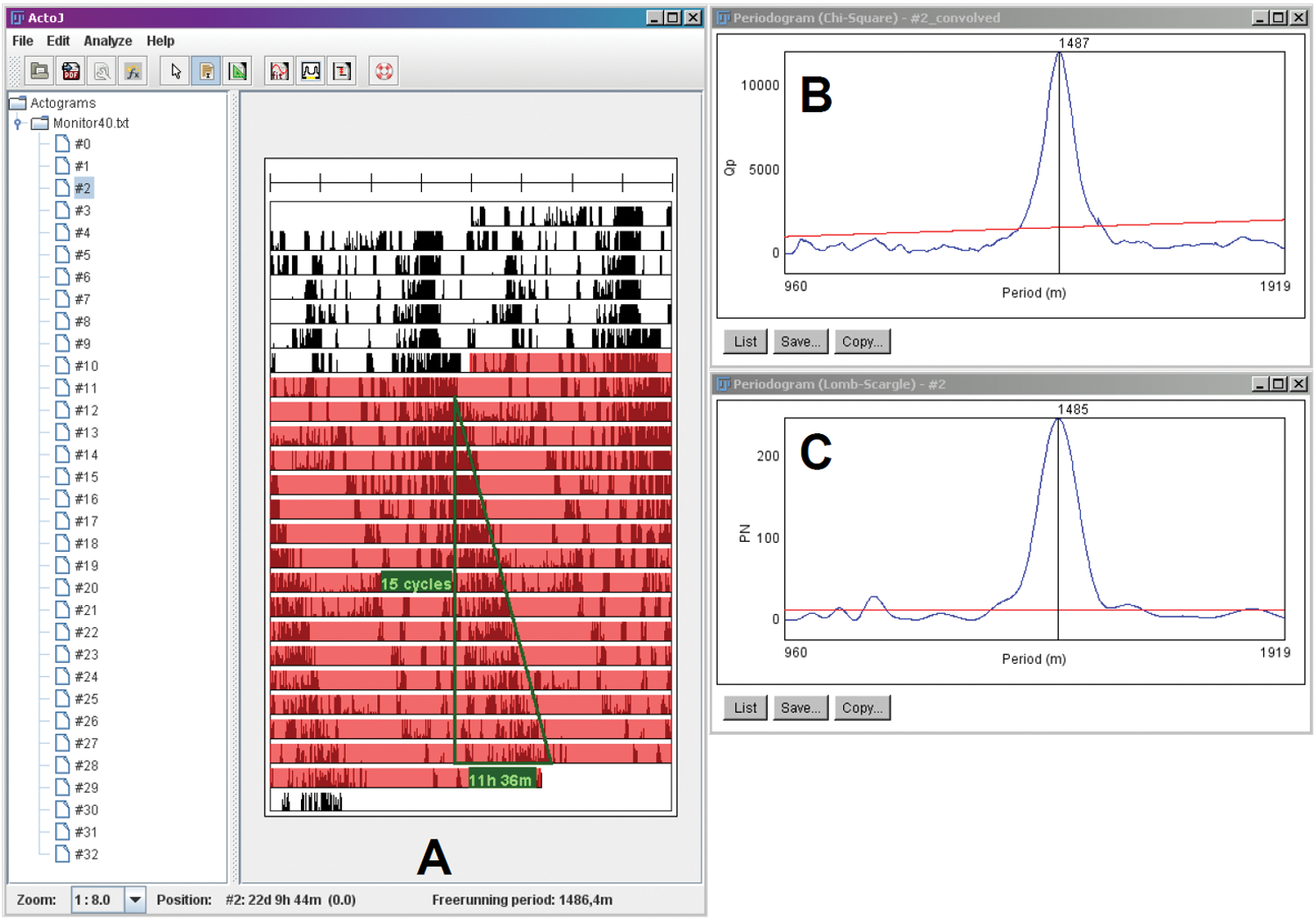
The periodogram analyses by ActogramJ. (A) The free-running period (1486.4 minutes) estimated by the green eye-fitted line, which is drawn on the actogram. (B) The result of the χ2 periodogram, calculated for the range that is highlighted in red on the actogram. Before applying periodogram analysis, the raw data were smoothed by a Gaussian kernel (sigma of 50). (C) The result of the Lomb-Scargle periodogram. All 3 methods show very similar results.
Periodogram Analysis
With the help of an excellent review of time series analysis in circadian rhythm research (Refinetti et al., 2007), we implemented several periodogram analyses in ActogramJ. First, a classic eye-fit line can be drawn on an actogram, and the estimated period is automatically calculated (Fig. 2A). As well as this manual method to determine periodicity, we implemented the 3 most frequently used methods to analyze time series data automatically: Fourier analysis (power spectrum) (Fourier, 1822), χ2 periodogram (Enright, 1965; Sokolove and Bushell, 1978) (Fig. 2B), and Lomb-Scargle periodogram (Ruf, 1999; Van Dongen et al., 1999) (Fig. 2C). To confirm the correctness of our implementations, we compared the results of the χ2 and Lomb-Scargle periodograms with those obtained by 2 other software packages (El-Temps by Diez-Noguera and the chronobiological software by Refinetti). The period lengths estimated by all 3 programs are very close (Suppl. Tables S1-S3). Additionally, we applied our χ2 and Lomb-Scargle periodograms to artificially generated random signals. In this case, no significant period was detected (Suppl. Table S4).
Waveform Analysis
Waveform analysis is useful for judging and comparing the characteristic shapes of rhythms as well as to determine the phase of the peak(s). Chronobiologists often calculate average waveforms of the rhythms for several cycles and display it on a graph. ActogramJ can easily calculate the waveforms from selected areas on an actogram.
Summary
Because ActogramJ uses ASCII text files to enter numeric data, it can be used with a wide range of data sources. Although ActogramJ does not cover all analysis methods for chronobiology, keeping the usage as simple as possible is certainly a benefit of this software and was one of our design goals. We implemented the 3 main analyses methods that meet the needs of the majority of researchers. We have provided detailed instructions for the usage of ActogramJ on our project home page (http://actogramj.neurofly.de). In summary, we believe that ActogramJ removes a barrier to doing chronobiological analyses and that the software will benefit not only many chronobiologists but also people planning to enter the field. We hope that they enjoy analyzing chronobiological data with ActogramJ.
Footnotes
Appendix
The calculations of the periodogram analyses were programmed exactly as described in Refinetti et al. (2007). More detailed information about the χ2 periodogram and Lomb-Scargle periodogram can be found in the original publications (Enright, 1965; Sokolove and Bushell, 1978; Ruf, 1999; Van Dongen et al., 1999).
The easiest way to set up ActogramJ is to use Fiji because itext-1.3.jar (a Java library) is already installed. Detailed instructions are given on the project home page (http://actogramj.neurofly.de), together with the relevant references. To support developers who wish to extend ActogramJ, we include the source code in the jar archive and provide API documentation on the website. Further support is available from the authors upon request.
Acknowledgements
The authors thank Wolfgang Engelmann, Christiane Hermann, and Irina Stahl for the comments on a previous version of the article, as well as Mark Longair for final proofreading. C.H.-F. acknowledges financial support from the European Community (the 6th Framework Project EUCLOCK no. 018741).
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
