Abstract
Objective:
This meta-analysis aims to systematically evaluate the associations of four specific Single Nucleotide Polymorphisms (SNPs)—rs712829, rs712830, rs11568315, and rs884225—located in the promoter, intronic, and 3’ untranslated regions (3’UTR) of the EGFR gene, with lung cancer risk.
Introduction:
The associations between EGFR gene polymorphisms and lung cancer risk is a topic of ongoing debate, which is still deemed controversial. Despite numerous studies, results are inconsistent.
Methods:
We conducted a comprehensive literature search across the PubMed, Science Direct, and Web of Science databases to identify relevant case-control studies examining the association between EGFR gene polymorphisms and lung cancer risk.
Results:
From an initial pool of 26,959 articles, 10 case-control studies were included, involving 2471 lung cancer patients and 4489 controls. A significant association between rs712829 and increased lung cancer risk was found across multiple genetic models. Under the allelic contrast model (G vs T), the OR was 1.31 (95% CI = [1.02; 1.68],
Conclusion:
The findings from the current meta-analysis confirm that rs712829 within the EGFR gene is significantly associated with lung cancer risk according to the allele, dominant, homozygote and heterozygote models.
Introduction
Lung cancer remains one of the most prevalent forms of cancer globally, accounting for the highest number of cancer-related deaths among men and ranking as the third most common cancer in women.1,2 As of 2020, lung cancer was responsible for approximately 1.8 million deaths worldwide, highlighting its significant public health burden and the urgent need for research to address its high mortality rate (WHO, GLOBOCAN, 2020). 3
Non-Small Cell Lung Cancer (NSCLC) accounts for about 85% of all lung cancer cases, presenting a more favorable prognosis compared to Small Cell Lung Cancer (SCLC). However, it still poses major challenges, including low long-term survival rates and reduced quality of life for patients.4,5
The etiology of lung cancer is multifactorial, influenced by both environmental and genetic factors. 6 In addition to the well-known role of tobacco smoking, other risk factors include occupational exposures, air pollution, and genetic susceptibility.7–10 Recent research shows that approximately 10%–15% of lung cancer cases occur in individuals who have never smoked, suggesting that genetic factors, such as polymorphisms, play a critical role in determining disease risk.1,4,11,12 Studies have confirmed that genetic variations significantly contribute to lung cancer susceptibility, and an increasing focus has been placed on understanding the impact of specific gene polymorphisms on the development of the disease.4,5,11
The epidermal growth factor receptor (EGFR) gene has emerged as a key player in lung cancer research. 13 EGFR is a transmembrane tyrosine kinase receptor involved in several critical cellular processes, including cell proliferation, differentiation, and survival.14,15 It is encoded by a gene located on chromosome 7, and its activation triggers signaling pathways that promote cellular growth and division. 16 Aberrations in the EGFR gene, such as mutations or overexpression, have been identified in various cancers including non-small-cell lung cancer, metastatic colorectal cancer, glioblastoma, head and neck cancer, pancreatic cancer, and breast cancer, making it a prominent target for cancer therapeutics.11,14,15
Among the most significant sources of human genetic diversity are single nucleotide polymorphisms (SNPs), which can influence an individual’s susceptibility to diseases such as cancer. 17 Numerous studies have demonstrated associations between specific SNPs in the EGFR gene and lung cancer risk.4,12,18
One notable example is the highly polymorphic dinucleotide repeat in intron 1, known as CA simple sequence repeat 1 (CA-SSR1, rs11568315), which has been shown to correlate with increased transcription and expression of the EGFR protein.19–23 Two single nucleotide polymorphism (SNPs) in the promoter region, 216 (G/T or T/T) and 191 (C/A or A/A), have been linked to promoter activity and EGFR expression.12,21,24–28 Additionally, a polymorphism (G2607A) within the EGFR tyrosine kinase domain has been identified as associated with the risk of lung cancer. 29 These genetic variations highlight the multifaceted implications of EGFR alterations in the development of lung cancer, which are modulated by factors such as smoking history, ethnic background, and gender. 18
Despite the growing body of research, the relationship between EGFR gene polymorphisms and lung cancer risk remains controversial.15,17,18,30–33 This premise provides a critical foundation for the subsequent comprehensive meta-analysis, which synthesizes data from previous studies investigating the relationship between EGFR gene polymorphisms and lung cancer risk. Specifically, we focused on evaluating the associations of four SNPs (rs11568315, rs712829, rs712830, and rs884225) distributed across various regions of the EGFR gene.
Our meta-analysis offers a comprehensive and updated evaluation of the impact of EGFR polymorphisms on lung cancer susceptibility. Unlike previous meta-analyses that have focused on a limited number of polymorphisms or treatment outcomes, our study examines EGFR variants associated with lung cancer risk, including the rs884225 polymorphism, which has not been addressed in previous meta-analyses. While other studies, particularly those focusing on the relationship between EGFR polymorphisms and TKI treatment outcomes,5,34,35 have provided valuable insights, our work specifically targets genetic susceptibility and offers a more complete understanding of the link between EGFR polymorphisms and lung cancer risk. By incorporating recent studies and individual patient data, we ensure our findings reflect the latest research, providing more precise estimates of the associations between EGFR variants and lung cancer susceptibility. Our focus on key polymorphisms such as rs712829, rs11568315, rs712830, and the previously unexamined rs884225, sets our study apart from earlier meta-analysis that either included fewer variants or concentrated primarily on treatment-related outcomes.
Materials and methods
Search strategy
A comprehensive literature search was carried out in the ScienceDirect, PubMed, and Web of Science databases for papers published using the combination of the following keywords: (1) Susceptibility
Selection criteria
Inclusion criteria
Relevant literature was identified based on the following inclusion criteria: (1) Peer-review articles written in English or French were exclusively considered; (2) Case-control studies examining the association between mutations or polymorphisms in the
Exclusion criteria
Review articles, systematic reviews, meta-analyses, case reports, abstracts, conference papers, GWAS studies, and other non-case-control study types were excluded. Additionally, studies in languages other than English or French and those lacking sufficient data were also excluded. Furthermore, studies focusing on diseases other than lung cancer were omitted, particularly those that did not specifically investigate the impact of mutations or polymorphisms in the
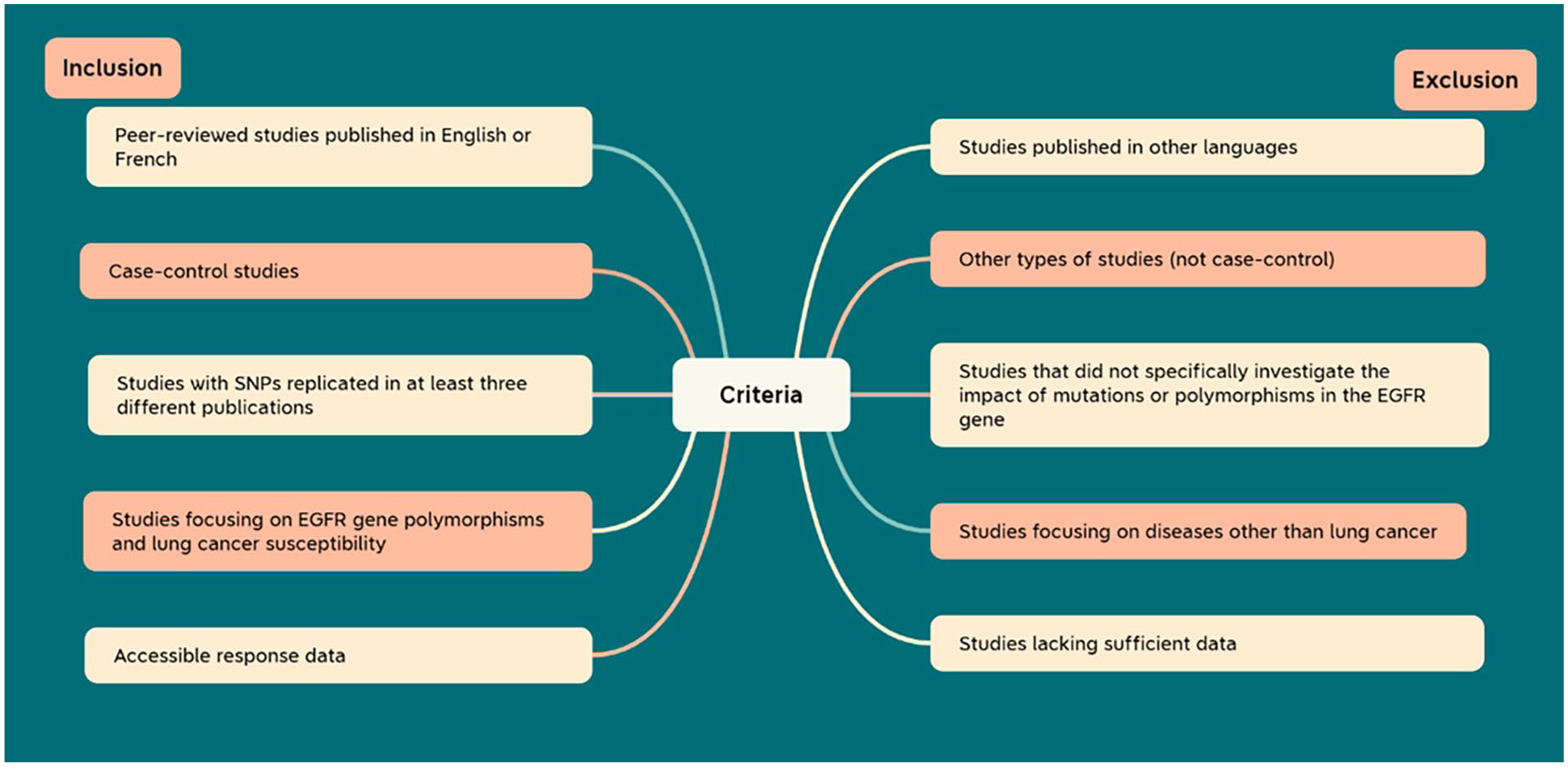
Diagram illustrating the inclusion and exclusion criteria.
Data extraction
The data extraction process involved capturing specific details from the selected articles, encompassing the following key elements: the first author’s name, publication year, countries involved in the studied population, study design, sample sizes for both case and control groups, participant age, genotyping methodology, smoking status, histopathology, and the genotypic and allelic frequencies of EGFR polymorphisms in both cases and controls. Data from all eligible studies were extracted independently by two investigators. Any disagreement was discussed with the third investigator to reach a consensus.
Statistical analysis
Genetic association studies
The genetic association studies investigating EGFR polymorphisms and lung cancer were analyzed using the MetaGenyo web tool (https://metagenyo.genyo.es/). This tool facilitated a comprehensive and standardized approach to synthesizing the data from various studies, ensuring the reliability and accuracy of our findings.
Hardy-Weinberg equilibrium (HWE) evaluation
The Hardy-Weinberg equilibrium (HWE) was assessed in both patients and controls using the chi-square (χ2) test. A
Relationship evaluation: Odds ratio and confidence interval computation
To evaluate the relationship between EGFR gene polymorphism and lung cancer susceptibility, the odds ratio (OR) and its corresponding 95% confidence interval (CI) were computed. This statistical approach quantified the strength and direction of the association, providing insights into the potential risk or protective effects of specific EGFR polymorphisms.
Heterogeneity assessment
Heterogeneity across studies was evaluated using the chi-squared-based Q statistic. Significance in heterogeneity was acknowledged when the
Model application
The fixed-effects model was employed to calculate the pooled odds ratio (OR) in instances where there was no observed heterogeneity among studies. In the presence of heterogeneity, the random-effects model was utilized. Specifically, for EGFR polymorphisms rs712829 and rs884225, the fixed-effects model was applied. For CA-SSR1 (rs11568315) and rs712830, the random-effects model was used, accommodating the variability in study results.
Genetic models examined
Various genetic models were examined to calculate pooled ORs for the studied polymorphisms. These models included allele comparison, recessive model, dominant model, homozygote comparison, heterozygote comparison, and over-dominant model. A genotype with an OR < 1 indicated a protective role, while an OR > 1 suggested a risk factor. Statistical significance was denoted by a
Sensitivity analysis
A sensitivity analysis was conducted to assess the stability of the overall effect in our meta-analysis. This required evaluating the ORs and their corresponding 95% CIs before and after systematically excluding each eligible study or adjusting the combination model, ensuring the robustness and reliability of our findings.
Publication bias evaluation: Egger regression asymmetry test
The presence of publication bias was evaluated using the Egger regression asymmetry test, with statistical significance determined at a threshold of
Results
Eligible studies
A total of 26,959 records were initially identified through the search strategy. After excluding 18,936 studies that were not research articles, 8023 research articles in English and French were assessed for screening. A total of 405 duplicate articles in English and 16 in French were removed, as well as 3811 records excluded based on title and abstract screening. The remaining records underwent further exclusion: 955 were review or systematic review articles, 53 were meta-analyses, 19 were GWAS studies, 84 were case reports, 737 lacked healthy controls, and 463 were other study types. Additionally, 1280 studies focused on diseases other than lung cancer, 170 did not involve the EGFR gene, and 29 lacked sufficient data. This study focused on polymorphisms that were investigated in a minimum of three separate publications.
Ultimately, after thorough screening, 10 case-control studies met the eligibility criteria and were selected for analysis. The study flow diagram, following the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines, is presented in Figure 2.
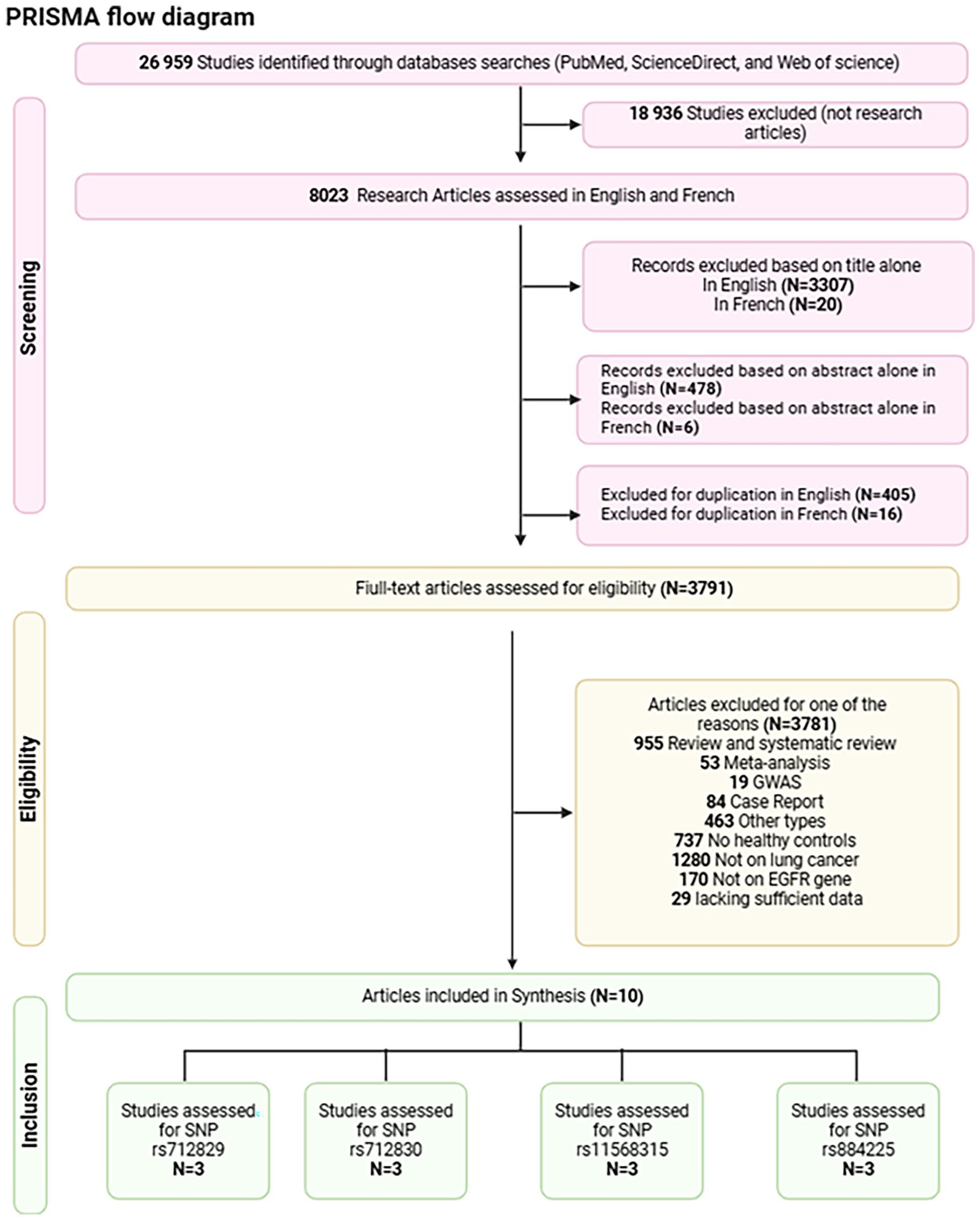
Search strategy flow diagram created with BioRender.com.
Study characteristics
This study emphasized polymorphisms that were investigated in at least three publications, specifically rs712829, rs712830, rs11568315, and rs884225 (Figure 3). In addition, 13 EGFR gene polymorphisms that were documented in fewer than three publications within the databases were excluded from the analysis. These polymorphisms include rs763317, rs6965469, rs2233947, rs6721961, G2607A, G1380A, 8227G>A, rs2293347, R776G, rs2227983, rs2227984, rs1050171, and Exon 19 deletion.

Illustration of EGFR polymorphisms created with BioRender.com.
This meta-analysis comprised 10 case-control articles, encompassing a total of 2471 individuals diagnosed with lung cancer. Among the patients, 1752 were males, and 819 were females, with mean ages ranging from 54.37 + 10.77 to 65.5 ± 8.5 years. The control group included 4489 subjects, with 2422 males and 2067 females, and mean ages varying from 54.25 ± 10.82 to 64.1 ± 10.3 years. A detailed screening process is outlined in Table S1. Notably, two studies were performed within the South Korean, and Jordanian populations,12,17,22,36 and one study each was carried out in Japanese, Serbian, Indian, Iranian, Caucasian, and Chinese populations23,26,27,28,37,38 (Figure 4).

Geographic distribution of selected case-control studies investigating the association between EGFR polymorphisms and lung cancer risk.
Ultimately, 10 out of the 24 case-control studies were included in this meta-analysis, with a specific focus on exploring the association between EGFR polymorphisms and lung cancer. The selection of these articles was conducted meticulously to ensure the reliability and coherence of the data analyzed in this study.
The details of each case-control study incorporated in this meta-analysis, including allelic and genotypic distributions, along with their HWE status, country, ethnicity, Sample size, and Genotyping method are presented in Table S1.
In most studies, the genotype distribution aligns with HWE, except for a singular study group (Serbian study for the EGFR gene rs712830 polymorphism). To assess the robustness of the meta-analysis, a sensitivity analysis was conducted, wherein the aforementioned study was systematically excluded. The analysis was iteratively repeated, each time omitting one study, to discern the impact of each individual study on the overall statistical outcomes (see Table S1).
Quantitative analysis
Heterogeneity test
Tables 1–4 showcase the results derived from the comprehensive investigation into the correlation between EGFR gene polymorphisms (specifically, rs712829, rs712830, rs11568315, and rs884225) and the susceptibility to lung cancer. Examination of the data revealed substantial heterogeneity among studies (heterogeneity < 0.05) for the two polymorphisms (rs712830, rs11568315), with the exception of these two polymorphisms (rs884225 and rs712829) across all genetic models. There was no significant heterogeneity observed in Allele contrast (C vs T,
Association analysis of allele/genotype of the EGFR rs11568315 (CA repeat) polymorphism and lung cancer risk using a random effects model.

Association analysis of allele/genotype of the EGFR rs712829 polymorphism and lung cancer risk using a fixed model effects.

Association analysis of allele/genotype of the EGFR rs712830 polymorphism and lung cancer risk using a random model effects.

Association analysis of allele/genotype of the EGFR rs884225 polymorphism and lung cancer risk using a fixed model effects.

Association test
Association between EGFR gene polymorphism rs11568315 (CA repeat) and susceptibility to Lung cancer
Table 1 provides a detailed overview of the findings from the comprehensive analysis investigating the association between the rs11568315 (CA repeat) polymorphism in the EGFR gene and lung cancer risk. Three studies, involving a total of 882 lung cancer patients (510 males, 372 females) with a mean age of 62.03 ± 8.7, and 2903 healthy subjects (1200 males, 1703 females) with a mean age of 60.33 ± 9.8, were included in this analysis. The outcomes indicated no significant association between the rs11568315 polymorphism and lung cancer risk across all tested models: Allele contrast (S vs L): OR = 1.2177, 95% CI = [0.8371; 1.7713],
Association between EGFR gene polymorphism rs712829 and susceptibility to Lung cancer
We conducted an analysis involving three studies with a total of 285 patients diagnosed with lung cancer (217 males, 68 females), with an average age of 57.45 ± 0.9, alongside 282 healthy subjects (200 males, 82 females), averaging 53.5 ± 0.8 years. The investigation focused on the association between rs712829 and lung cancer susceptibility. The results, detailed in Table 2, indicated a significant association for several models: Allele contrast (G vs T): OR = 1.3072, 95% CI = [1.0151; 1.6833],
Association between EGFR gene polymorphism rs712830 and susceptibility to Lung cancer
In this meta-analysis of the EGFR polymorphism rs712830, data were pooled from three studies, encompassing a total of 285 lung cancer patients (214 males, 171 females) with a mean age of 59.05 ± 8.35 years, and 282 healthy subjects (203 males, 79 females) with a mean age of 52.41 ± 5.81 years. As outlined in Table 3, the analysis revealed no significant association between rs712830 and lung cancer across all tested models: Allele contrast (A vs C): OR = 2.1911, 95% CI = [0.6163; 7.7903],
Association between EGFR gene polymorphism rs884225 and susceptibility to Lung cancer
Three studies were included in the investigation of the association between the EGFR polymorphism rs884225 and the risk of lung cancer. The analysis involved a total of 1019 lung cancer patients (811 males, 208 females) with a mean age of 62.7 ± 9.65, and 1022 healthy subjects (819 males, 203 females) with a mean age of 62.26 ± 9.85. The findings, as outlined in Table 4, revealed no significant association between rs884225 and lung cancer across all tested models: Allele contrast (C vs T): OR = 1.1117, 95% CI = [0.9768; 1.2653],
The analysis of the data showed considerable variability among the studies in their overall comparisons. Consequently, random effects were selected for EGFR polymorphisms rs11568315 and rs712830, while fixed effects were applied to rs712829, and rs884225. Forest plots illustrating these findings are depicted in Figure 5 (OR = 1.22,

Forest plot illustrating the association between the EGFR rs11568315 polymorphism and lung cancer risk.
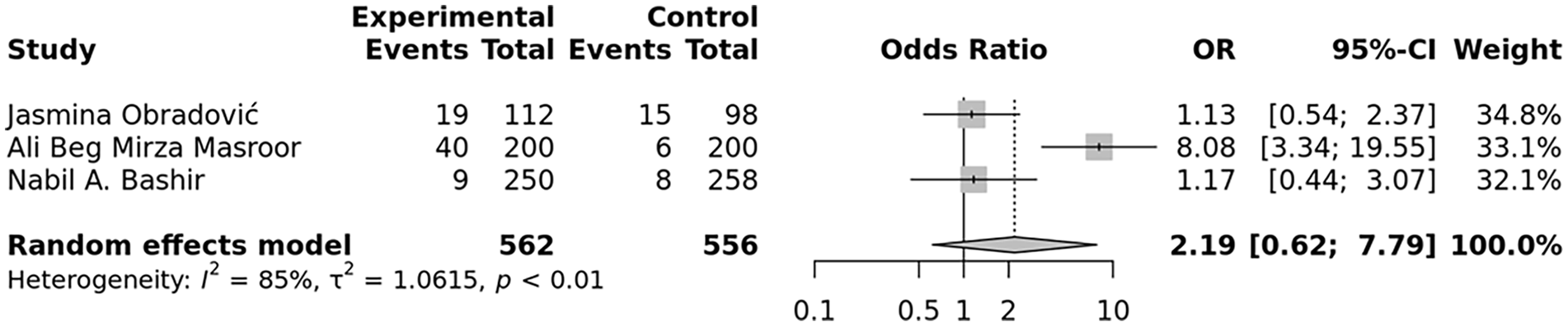
Forest plot illustrating the association between the EGFR rs712830 polymorphism and lung cancer risk.
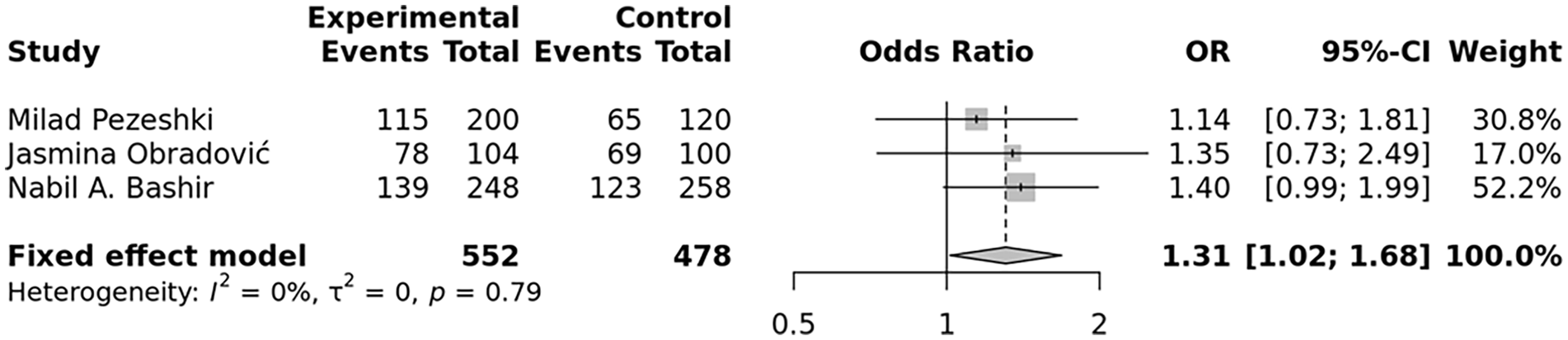
Forest plot illustrating the association between the EGFR rs712829 polymorphism and lung cancer risk.

Forest plot illustrating the association between the EGFR rs884225 polymorphism and lung cancer risk.
Sensitivity analysis
To evaluate the consistency and robustness of our results, we conducted a sensitivity analysis for each polymorphism. This involved removing each study individually and re-estimating the pooled Odds Ratios (ORs). The pooled ORs for the four polymorphisms (rs884225, rs712829, rs712830, and rs11568315) remained largely unchanged, as illustrated in Figures 9–12 respectively. This observation indicates the statistical robustness of our findings.

Sensitivity analysis in the overall populations for rs884225 (allele model: C vs T).

Sensitivity analysis in the overall populations for rs712829 (allele model: T vs G).

Sensitivity analysis in the overall populations for rs712830 (recessive model: AA vs AC + CC).

Sensitivity analysis in the overall populations for rs11568315 (allele model: S vs L).
Publication bias
To assess potential publication bias across all genetic models, Egger’s regression test was employed. Our examination of the link between EGFR gene polymorphisms and lung cancer risk revealed no significant asymmetry. Specifically, the Egger’s regression test results for rs712830, rs712829, rs884225, and rs11568315 suggested that there was no significant publication bias detected under all genetic models (

Funnel plot of Egger’s test assessing publication bias in the overall population for (a) rs11568315 (allele model: S vs L), (b) rs712829 (allele model: T vs G), (c) rs712830 (allele model: A vs C), (d) rs884225 (allele model: C vs T).
Discussion
Lung cancer poses a major global health challenge as it remains the top cause of cancer-related mortality across the world. While smoking is the predominant culprit, accounting for around 85% of cases, genetics indeed contribute to lung cancer development.39–41 Nonetheless, recent research has indicated that approximately 10%–15% of lung cancer cases occur in individuals who have never smoked, suggesting that genetic polymorphisms and other factors may contribute to disease risk. 12
Among the myriads of genetic variants implicated in lung cancer, EGFR polymorphisms emerge as potential risk factors for lung cancer susceptibility. EGFR, a gene frequently dysregulated in many cancers, including NSCLC, 42 not only serves as a prognostic marker but also exhibits potential as a therapeutic target.12,43 Various studies have highlighted the impact of EGFR polymorphisms on lung adenocarcinoma. However, substantial interethnic disparities exist, and the findings lack consistency.4,44
Further research is imperative to elucidate the multifaceted role of genetic variables in the development of lung cancer, with a particular focus on understanding the implications of EGFR polymorphisms for early detection and prevention strategies.
Numerous studies highlighted the complex role of EGFR mutations in lung cancer development and therapy, with some studies emphasizing the heterogeneity in treatment responses depending on specific EGFR mutations (e.g., G719X, L861Q, S768I) and co-occurring mutations such as TP53, which can impact the efficacy of EGFR inhibitors like afatinib. 45 Additionally, polymorphisms in the EGFR gene, have been shown to contribute to lung cancer susceptibility, further supporting the need for personalized therapeutic strategies. 11 The interaction between EGFR mutations and environmental carcinogens, like tobacco smoke, further complicates lung cancer pathogenesis, necessitating the consideration of genetic factors in risk assessments.17,46
Statistical analysis of the present meta-analysis conducted on a total of four polymorphisms from 10 studies, indicates that according to the allele, dominant, homozygote and heterozygote models the rs712829 polymorphism in the EGFR gene is closely linked to an increased risk of lung cancer. Research has shown that this specific SNP affects the outcome of tyrosine kinase inhibitor (TKI) based treatment in NSCLC patients, influencing overall survival (OS) and progression-free survival (PFS), compared to those with the wild genotype, indicating that this polymorphism was a significant risk factors for unfavorable outcomes.5,47 This is discordant with a meta-analysis, that showed that the presence of the -216T variant allele of rs712829 is linked to improved overall survival outcomes in NSCLC patients treated with TKIs, as indicated by a lower risk of death (hazard ratio of 0.80) and a statistically significant association (
Analysis of the EGFR promoter region and exon 1 in 18 pulmonary carcinoma patients unveiled an association between the -216 G/T variant and 58.82% of NSCLC cases, particularly those with squamous carcinoma. Notably, homozygote (T/T) variants were predominantly observed. 49 Additionally, research on Serbian NSCLC patients revealed a strong association between the -216GG genotype and lung cancer risk, especially among smokers, 26 highlighting its potential as an informative diagnostic marker for early detection of the disease. Furthermore, the -216G/T polymorphism, along with other EGFR gene variants, has been associated with increased risks of pleural metastasis in lung adenocarcinoma patients, potentially due to enhanced EGFR protein expression induced by these polymorphisms. 50
Two well-studied EGFR single nucleotide polymorphisms (SNPs), -216G/T (rs712829) and -191C/A (rs712830), are positioned within the promoter region of the EGFR gene, suggesting their potential involvement in the regulation of EGFR expression.5,51 Specifically, the -216G/T variant situated within the EGFR promoter’s transcription factor Sp1 binding site, constituting one of the four Sp1 recognition sites in the promoter. 44 Consequently, it is conceivable that these polymorphisms could exert a modulatory effect on promoter activity. Linkage disequilibrium (LD) analysis revealed that the -216G/T polymorphism is not associated with other key EGFR SNPs in this genomic region, including the (CA)n polymorphism in intron 1, suggesting its independent role in gene regulation. 51 Expression studies demonstrated that the substitution of G by T at this position notably enhances both promoter activity and mRNA expression levels.21,52 Specifically, studies have shown that this substitution leads to a significant increase of 30% and 40% in promoter activity and gene expression, respectively. 52
In contrast, no significant correlations were observed for rs11568315 (CA repeat), rs712830, and rs884225 across all models. Furthermore, the EGF + 61 polymorphism (AG and AG + GG) was found to be independently associated with stable disease in lung cancer patients, although it was not linked to the overall response rate to first-generation TKIs or patient outcome. 53 Contrary to our findings, studies have indicated that the rs712830 CC genotype is associated with unfavorable outcomes in lung cancer patients, correlating with poorer overall survival. 47 Additionally, the rs712830 A allele has been linked to reduced progression-free survival and increased cancer risk in a lung adenocarcinoma cohort from India. 28 The CA repeat polymorphism (rs11568315), among others, emerges as a significant determinant in the treatment response of NSCLC patients undergoing TKI therapy. Meta-analysis confirms that the CA repeat polymorphism significantly impacts OS and PFS in NSCLC patients treated with gefitinib or erlotinib, These findings suggest that the length of the CA repeat in EGFR intron 1 could serve as an additional predictive marker for the efficacy of TKI-based therapy in NSCLC patients. 5 Others have reported that shorter CA repeats are associated with EGFR mutations, particularly in exon 19, and may predict better clinical outcomes for NSCLC patients treated with EGFR-TKIs. 54 Additionally, shorter CA repeats have been linked to increased EGFR protein expression levels in lung carcinoma, 55 and a systematic review found an association between the EGFR Short genotype of cytosine adenine repeat number polymorphism and increased lung cancer risk. 13 Contradictory findings have emerged from these investigations that concur with our results. Some studies have found no significant association between the CA repeat polymorphism and lung cancer risk or histological subtypes. 22
The rs884225 polymorphism in the EGFR gene, a 3′-untranslated region variant c.*774T>C affects EGFR expression levels by altering the binding site of miR-103a-3p, leading to increased EGFR expression. This SNP has been associated with TKI toxicities in NSCLC Chinese patients.34,56 Due to lack or not enough data was available from other studies for comparison regarding this polymorphism. There is a lack of association between rs884225 and lung cancer development.
The findings of this study shed light on the intricate relationship between genetic polymorphisms in the EGFR gene and their implications for lung cancer susceptibility and treatment outcomes. While several polymorphisms, such as rs712829 and rs712830, have been extensively studied and shown to impact patient prognosis, others remain relatively understudied.
Among these less-explored polymorphisms is rs884225, situated in the 3′-untranslated region of the EGFR gene. Existing research indicates that this variant, known as c.*774T>C, modulates EGFR expression levels by altering the binding site of miR-103a-3p, thereby potentially influencing cancer progression. 57 Notably, studies have linked rs884225 to TKI toxicities in Chinese patients with NSCLC.34,56 However, the lack of sufficient data from other populations impedes a comprehensive understanding of its role in lung cancer development.
To address this knowledge gap, future research efforts should prioritize the investigation of rs884225’s association with lung cancer susceptibility. Large-scale studies involving diverse populations are warranted to gather robust data and ascertain the significance of this polymorphism. By expanding our understanding of less-studied EGFR polymorphisms such as rs884225, we can enhance personalized treatment approaches and improve patient outcomes in lung cancer management.
In this meta-analysis, we identified a significant overall association between the polymorphism rs712829 and lung cancer, with an overall
The significant overall
However, subgroup analysis was not applicable due to the small number of studies, which limited statistical power. The lack of significant associations within individual regions might be due to smaller sample sizes in these subgroups, which reduces the statistical power to detect associations. It’s possible that the effect size of the polymorphism is modest, requiring larger sample sizes to achieve statistical significance in regional analyses.58–60 Therefore, the overall significant
Additionally, the potential for gene-environment interactions should be considered. The polymorphism’s effect on disease risk might be modulated by region-specific factors.61–63 Such interactions could mask the association in individual regions while still contributing to the overall significance.
Differences in allele frequencies, linkage disequilibrium patterns, and phenotype specificity across populations can lead to variations in associations, contributing to heterogeneity in studies.64–66.
Other limitations include a small number of studies per SNP, language bias from excluding non-English/French studies, and restriction to case-control studies. Future research should broaden sample sizes, incorporate diverse study designs, and explore gene-environment interactions to enhance understanding of EGFR polymorphisms in lung cancer risk.
Conclusions
The findings from the current meta-analysis confirm that rs712829 within the EGFR gene is significantly associated with lung cancer risk. Future well-designed research, including larger sample sizes and representation from diverse populations and ethnic groups, such as Genome Wide Association Studies, is necessary to validate this finding. Further functional investigations into these single nucleotide polymorphisms may advance our understanding of the mechanisms underlying the development and progression of lung cancer.
Supplemental Material
sj-docx-1-iji-10.1177_03946320251316731 – Supplemental material for Meta-analysis of EGFR gene polymorphisms and lung cancer risk
Supplemental material, sj-docx-1-iji-10.1177_03946320251316731 for Meta-analysis of EGFR gene polymorphisms and lung cancer risk by Meryem Fakhkhari, Ikram Salih, Khaoula Errafii, Salsabil Hamdi and Khalid Sadki in International Journal of Immunopathology and Pharmacology
Supplemental Material
sj-docx-2-iji-10.1177_03946320251316731 – Supplemental material for Meta-analysis of EGFR gene polymorphisms and lung cancer risk
Supplemental material, sj-docx-2-iji-10.1177_03946320251316731 for Meta-analysis of EGFR gene polymorphisms and lung cancer risk by Meryem Fakhkhari, Ikram Salih, Khaoula Errafii, Salsabil Hamdi and Khalid Sadki in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgements
None.
Author contributions
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Consent to participate
Not applicable.
Data availability statement
All data are contained within the article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
