Abstract
We aimed to evaluate the expression profiles of five circulating lncRNAs (HOTAIR, MALAT-1, XIST, SNHG15, and H19) in DLBCL patients and explore potential associations between their expression and different clinicopathological features. Diffuse large B-cell lymphoma (DLBCL), the most common non-Hodgkin lymphoma (NHL), exhibits marked genetic and clinical heterogeneity, emphasizing the need for improved tools for risk stratification. Long non-coding RNAs (lncRNAs) emerged as regulators in different cellular processes and have been linked to cancer pathogenesis. Real-time quantitative PCR (qRT-PCR) was used to evaluate lncRNA expression in the plasma of 65 newly diagnosed adult DLBCL patients and 30 age-matched controls. HOTAIR expression was significantly elevated in DLBCL patients, while SNHG15 was significantly downregulated. Interestingly, both HOTAIR and SNHG15 demonstrated robust discriminatory power between DLBCL and healthy individuals, achieving area under the curve (AUC) values of 69% and 71%, respectively. H19 expression displayed a significant association with early-stage (stage I) DLBCL. While upregulated HOTAIR was a significant independent predictor of poor prognosis, high SNHG15 expression appeared to have a protective effect on mortality rates. Our findings suggest that circulating lncRNA expression patterns are promising tools as non-invasive biomarkers for diagnosis of DLBCL. Specific lncRNAs, such as HOTAIR, SNHG15, and H19, could offer potential for disease staging and patient prognosis. Long-term follow-up studies are recommended to further elucidate the interplay between these lncRNAs and survival rates, as well as their interactions with other genetic and pathological features of DLBCL.
Introduction
Diffuse large B-cell lymphoma (DLBCL) constitutes approximately one-third of all non-Hodgkin lymphomas (NHL), with a prevalence ranging from 20% to 50% across different countries. Globally, the incidence of DLBCL varies from 2.3 to 13.8 cases per 100,000 person per year.1,2 DLBCL is characterized by significant heterogeneity, diverse genetic profiles, clinical presentations, and therapeutic responses, ultimately leading to variable patient outcomes.2–4 Due to the marked heterogeneity observed in DLBCL, the development of prognostic tools has become increasingly important. Clinical scoring systems have been devised to enhance patient risk stratification and facilitate the selection of appropriate treatment strategies. The International Prognostic Index (IPI), introduced over two decades ago, remains a cornerstone for risk assessment in clinical trials.5–7 This, along with the revised IPI (R-IPI) and the National Comprehensive Cancer Network IPI (NCCN-IPI), utilize readily available clinical parameters like age, lactate dehydrogenase levels, disease stage, and performance status. 8 While the IPI effectively stratifies patients into risk groups with varying survival outcomes, some patients with favorable IPI scores experience poor clinical courses. This highlights the ongoing need for additional prognostic markers to enhance DLBCL patient classification and guide treatment response prediction.
To address this limitation, molecular prognostic tools have been developed. These tools can effectively distinguish between two major DLBCL subtypes with distinct clinical outcomes, independent of IPI scores: the germinal center B-cell-like (GCB) and the activated B-cell-like (ABC) DLBCL. Notably, patients diagnosed with GCB DLBCL exhibit a significantly better prognosis, with a reported 5-years survival rate of 60% compared to 35% for patients with ABC DLBCL.9,10
Long non-coding RNAs (lncRNAs) constitute a heterogeneous group of non-protein-coding transcripts exceeding 200 nucleotides in length. These molecules perform regulatory functions affecting gene expression at various levels, including transcription, subcellular localization, mRNA stability, translation, and post-transcriptional processes. 11 Notably, lncRNAs have also been implicated in the pathogenesis of various human diseases. Underscoring these diverse functions, numerous studies have highlighted the significant contributions of lncRNAs to cancer development and progression.12–15
The growing interest in lncRNA research in recent years has prompted investigators to identify several differentially expressed lncRNAs in DLBCL, both in cell lines and in clinical samples. 11 Their distinct expression patterns in DLBCL make them promising candidates for diagnostic biomarkers or therapeutic targets. 16
To select candidate lncRNA biomarkers for this study, we adopted a two-step approach. The primary focus was on lncRNAs with documented roles in either DLBCL or other cancers, prioritizing those with robust supporting evidence. The Lnc2Cancer 3.0 database, a comprehensive resource for experimentally validated lncRNAs associated with human cancers (https://bio-bigdata.hrbmu.edu.cn/lnc2cancer/), served as the foundation for identifying these lncRNAs. Additionally, relevant published literature was reviewed to select these biomarkers.
Within this framework, we prioritized lncRNAs with well-defined functions in DLBCL, such as HOTAIR, MALAT-1, and XIST, based on evidence retrieved from either Lnc2Cancer 3.0 or the relevant literature. To broaden the scope, minimize redundancy with the previously published studies, and potentially identify novel DLBCL biomarkers, we additionally included two other candidates: H19, which has a controversial role in cancer and was studied in some hematological malignancies, and SNHG15, which was primarily studied in solid tumors. Notably, research has linked these lncRNAs to migration, invasion, and metastasis in various cancers,17,18 but their impact on DLBCL remains uninvestigated. This selection strategy ensured a data-driven approach, prioritizing lncRNAs with validated functions in DLBCL while incorporating potentially informative candidates with less established roles in this specific context.
We aimed at evaluating the expression profiles of these five lncRNAs in the plasma, and to explore the potential correlations between their expression levels and various clinicopathological parameters, including morphological classifications, diagnostic markers, treatment response, and patient prognosis.
Patients and methods
Patients
This study enrolled newly diagnosed adult patients with DLBCL. All patients were recruited from the Medical Oncology Department of the National Cancer Institute (NCI), Cairo University, between March 2020 and December 2022. Written informed consent was obtained from all participants prior to study. The study protocol was approved by the ethical committee of the NCI, Cairo University (approval no MO2305-505-051).
Inclusion and exclusion criteria
Patients were eligible for inclusion if they had a newly diagnosed, histologically confirmed DLBCL, were 18 years or older, and had an Eastern Cooperative Oncology Group (ECOG) Performance Status of 0-2. Patients were excluded if they had double primary cancer or secondary malignancy, poor performance status or organ dysfunction that would prevent active standard treatment, or primary central nervous system lymphoma.
Sample size
The sample size was calculated using Minitab 17.1.0.0 for Windows (Minitab Inc., 2013, Pennsylvania, USA). Globally, DLBCL comprises 20%–50% of all non-Hodgkin lymphomas. 1 In Egypt, a previous study from the National Cancer Institute, Cairo University, reported that DLBCL account for approximately 49% of NHL cases presenting to the institution. 19 Based on the data from this previous study, with a 10% margin of error and a 90% confidence level, a minimum sample size of 65 cases was determined.
Methods
At diagnosis, peripheral blood samples were collected in EDTA tubes under sterile conditions. Total RNA, including miRNA, was extracted using the miRNeasy Mini Kit (Cat no. 217,004) from QIAGEN, following the manufacturer’s instructions. Reverse transcription was carried out using the miScript II RT kit, according to the manufacturer’s instructions. To determine the relative expression levels of five specific long non-coding RNAs, quantitative real-time polymerase chain reaction (qRT-PCR) was performed using the miScript SYBR Green kit from Qiagen.
For qRT-PCR, 1 µL of diluted cDNA was used as a template in a 10 µL PCR reaction containing 1X SYBR Green master mix, 200 nM of the forward primer specific to each lncRNA, and 200 nM of the reverse primer. The qRT-PCR amplification was carried out using the following conditions: an initial denaturation step at 95°C for 10 min, followed by 40 cycles of denaturation at 95°C for 15 s and annealing/extension at 60°C for 30 s. All qRT-PCR reactions were performed on a ViiA seven real-time PCR system from Applied Biosystems.
To normalize the expression data, the expression of glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was measured. For each qRT-PCR experiment, we performed three technical replicates. The primers for qRT-PCR were designed using the Primer-BLAST tool available on the NCBI website (https://www.ncbi.nlm.nih.gov/tools/primer-blast/). The design criteria included selecting primers with optimal melting temperatures (Tm) between 58 and 60°C, GC content between 40%–60%, and amplicon sizes ranging from 100 to 250 base pairs to ensure efficient amplification and specificity. The designed primers were also checked to avoid secondary structures such as hairpins and dimers, and to ensure specificity to the target sequences by avoiding cross-reactivity with non-target genes. The primers were synthesized and obtained from Eurofins Genomics in Hamburg, Germany, which provided high-purity oligonucleotides suitable for quantitative real-time PCR applications. Sequences of primers used for gene amplification ae listed in supplemental table 1. Data analysis was performed using relative quantification, and results were expressed using the ΔΔCT method. 20
The efficiency of each primer used in the qRT-PCR was determined by generating a standard curve using a serial dilution of cDNA. The efficiency (E) was calculated using the formula:
Treatment protocol
A standardized protocol incorporating clinical assessment, laboratory investigations, and radiological evaluation was employed for all patients. While R-CHOP (rituximab, cyclophosphamide, doxorubicin, vincristine, and prednisone) chemoimmunotherapy served as the first-line induction regimen for most cases, individualization was implemented for some patients: two with high HCV PCR viremia received CHOP therapy excluding rituximab, and two with cardiac comorbidities received an anthracycline-free induction regimen. Stratification of subsequent treatment plans (cycle number and potential radiotherapy inclusion) was guided by disease stage and additional clinical risk factors identified as adverse prognostic features, such as bulky disease, elevated LDH levels, and poor Eastern Cooperative Oncology Group performance status.
Statistical analysis
Statistical analysis for this study was performed using Minitab 17.1.0.0 for Windows (Minitab Inc., 2013, Pennsylvania, USA). Data normality was assessed using the Shapiro-Wilk test. Continuous data were presented as mean and standard deviation (SD), or median and interquartile range (IQR), as appropriate. Categorical data are presented as frequencies and percentages. Kruskal Wallis test was used to compare between two groups with non-normality distribution, and chi square test was used to compare two or more groups of categorical characters.21,22 The receiver operating characteristic (ROC) curve was used to evaluate the utility of each lncRNA expression in discriminating DLBCL from healthy subjects. An area under the curve (AUC) above 0.6 was considered acceptable. The Pearson correlation coefficient was used to examine the linear relationship between numerical data. The sign before “r” denoted the direction of the relationship. A general linear model (GLM) with and without stepwise selection was used to evaluate the factors affecting the fold changes of lncRNAs. The sign of the coefficient indicates the direction of the correlation. Kaplan Meier test with subsequent Cox regression analysis was used to estimate the survival probability of patients, and the independent risk factors associated with mortality. 23 All tests were two-sided, and a p-value of less than 0.05 was considered statistically significant.
Results
General characteristics of the study groups
This study enrolled a total of 65 newly diagnosed adult patients with DLBCL and 30 age-matched healthy controls. Supplemental table 2 summarizes the clinical and demographic characteristics of the patients. DLBCL patients had a mean age of 55.56 years (SD ± 13.88), with a balanced gender distribution (male: female ratio = 1:1). Hepatic comorbidities were observed frequently, with 26% of patients diagnosed with hepatitis C virus (HCV) infection and 20% presenting with hepatic cell failure.
Comparison between the expression of lncRNAs in patients and controls
No significant age or gender differences were observed between DLBCL patients and healthy controls. Age comparison yielded a mean of 55.56 ± 13.88 years for patients and 54.61 ± 12.91 years for controls (p = 0.51), and both groups maintained near-equal male-to-female ratios (p = 0.99). However, distinct lncRNAs expression patterns emerged. As shown in Figure 1, HOTAIR lncRNA expression was significantly upregulated in DLBCL patients compared to controls (p = 0.004), while SNHG15 lncRNA displayed a marked downregulation (p < 0.001). These lncRNAs demonstrated substantial discriminatory power, with AUC values reaching 69% for HOTAIR and 71% for SNHG15 (p = 0.004 and <0.001, respectively) as presented in Figure 2. Comparison between the plasma expression of HOTAIR, MALAT-1, XIST, SNHG15 and H19 long noncoding RNA in patients and controls. ROC curve of fold changes for HOTAIR and SNHG15 long noncoding RNA.

Utility of HOTAIR and SNHG15 long noncoding RNA in detecting patients with DLBCL.
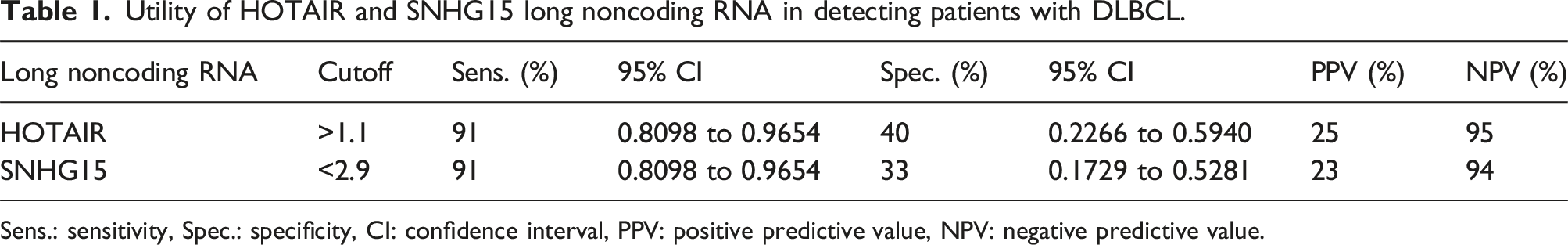
Sens.: sensitivity, Spec.: specificity, CI: confidence interval, PPV: positive predictive value, NPV: negative predictive value.
Factors affecting the expression of lncRNAs
Supplemental Figure 1 shows a significant positive correlation between the fold change of HOTAIR and other long non-coding RNAs (MALAT-1, XIST, and SNHG15), as well as between MALAT-1 and SNHG15 and H19, and between XIST and SNHG15.
Univariate analysis revealed differential associations between lncRNA expression and disease features, as detailed in Supplemental Table 3. Notably, SNHG15 expression emerged as potentially influenced by hepatitis C virus (HCV) status, chronic liver disease (CLD), hypertension (HTN), and diabetes mellitus (DM). Positive HCV status demonstrated a weak association with upregulated SNHG15 (coefficient = −6.37, p = 0.05), while CLD displayed a negative correlation (coefficient = 7.13, p = 0.03). HTN positively impacted SNHG15 expression, while DM showed an inverse relationship (p = 0.001). H19 expression only linked positively with hemoglobin (Hb) levels (p = 0.05). We observed no significant association between the IPI score and the expression levels of the investigated lncRNAs.
The GLM with stepwise selection technique identified the independent factors significantly influencing lncRNA expression, as summarized in Supplemental Table 4. For HOTAIR, BM abnormal lymphoid percentage emerged as a positive regulator (coefficient = 0.24, p = 0.01). Patient age and Hb levels positively impacted MALAT-1 expression (coefficients = 0.38 and 2.41, respectively, p = 0.02). Age had an inverse effect on XIST levels (coefficient = −0.16, p = 0.04). SNHG15 expression was affected by a complex interplay of factors: positive HCV status without CLD, and DM without HTN. Additionally, Hb level and DLBCL stage I independently influenced H19 expression (coefficients = 9.25 and 39.3, respectively; p = 0.01).
Factors affecting mortality
Independent predictors of mortality in DLBCL patients.

The test of fitness: Hosmer-Lemeshow, X2 = 3.4, p = 0.44, the test of significant: Multiple logistic regressions with backward elimination methods, p < 0.05 was considered significant.
Treatment response and survival analysis
The study observed a high overall response rate to the treatment regimen, with 69.23% of patients achieving complete response (CR) and 18.46% achieving partial response (PR). Progressive disease (PD) and stable disease (SD) were less frequent, occurring in 9.23% and 3.08% of cases, respectively. Supplemental Table 5 highlights a significant correlation between bone marrow infiltration and treatment response. Patients with a bone marrow biopsy (BMB) positive for infiltration exhibited a lower CR rate (55%) compared to those with a negative BMB (75.56%, p = 0.05). Conversely, a higher proportion of patients with positive BMB achieved PR (35%) compared to the negative BMB group (11.11%, p = 0.02). Furthermore, the study revealed a concerning association between positive BMB and mortality. Notably, 20% of patients with positive BMB died during treatment, underscoring the severity of bone marrow involvement in this patient population.
Figure 3 shows long-term outcomes for the study population, with Kaplan-Meier curves illustrating high survival and disease-free probabilities at 36 months post-treatment. The mean overall survival (OS) reached 35 months with a standard error of approximately 1 month, translating to an 87% survival probability at 36 months. Similarly, the mean disease-free survival (DFS) time was 34 months with a standard error of roughly 1 month, corresponding to a 92% DFS probability at 36 months. Kaplan-Meier survival analysis: Assessing overall survival (OS) and disease-free survival (DFS) over time.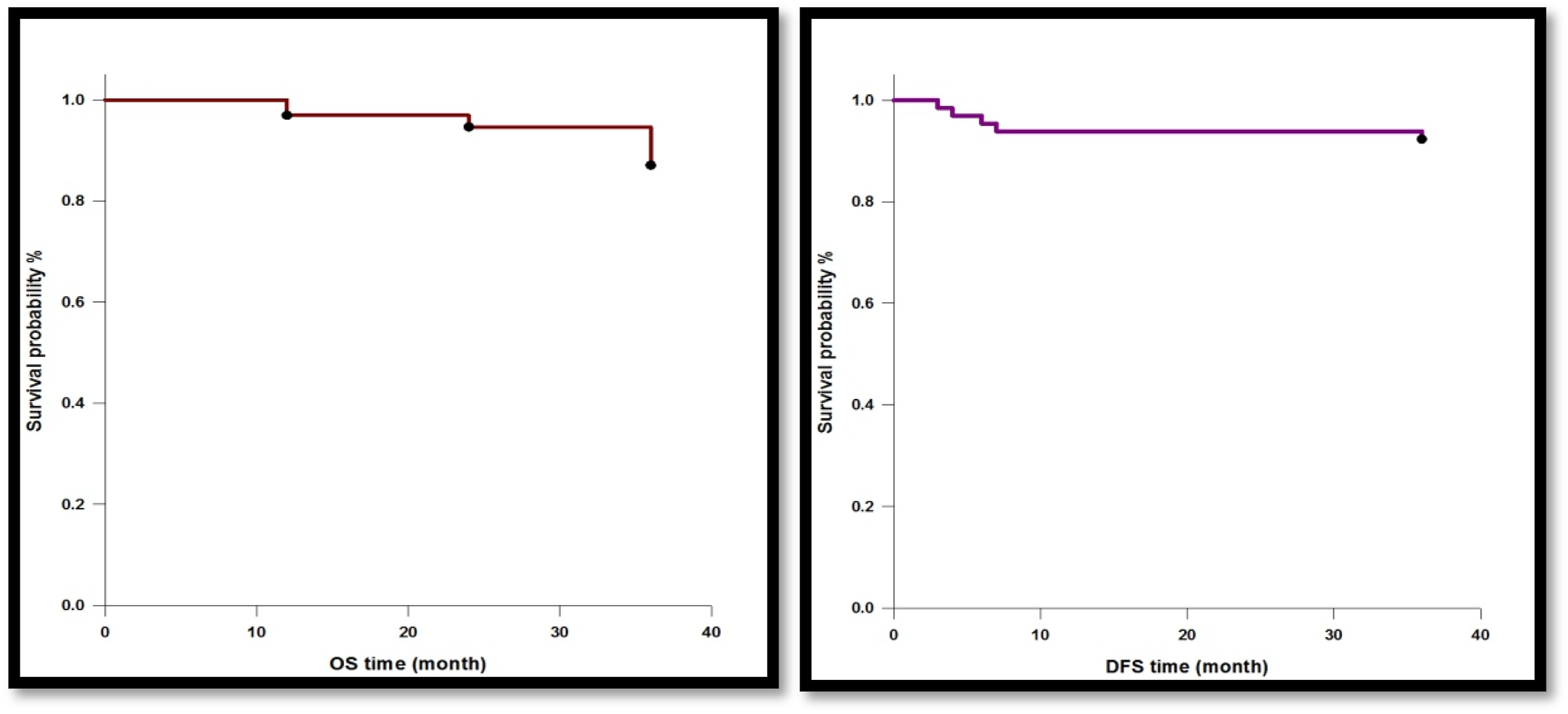
Cox regression analysis of lncRNA expression and overall survival in DLBCL.

HR: hazard ratio, CE: Coefficient, SE: standard error, CI-L: confidence interval low, CI-U: confidence interval high, The test of significant: Cox regression analysis model, p < 0.05 considered significant.
Supplemental Table 7 summarizes the analysis of correlations between HOTAIR, MALAT-1, XIST, SNHG15, and H19 lncRNAs expression levels and treatment response. No statistically significant associations were identified.
Discussion
Despite the promising initial response of 69.23% complete remission (CR) achieved in our study using standard R-CHOP, the heterogeneous treatment response in DLBCL remains a significant challenge. 24 While this CR rate aligns with previous reports, 25 the poor prognosis associated with relapse after R-CHOP failure necessitates further investigation into risk stratification for this aggressive malignancy.24,26 This study presents circulating lncRNAs as potential non-invasive biomarkers for DLBCL diagnosis. These markers may hold additional value in disease staging and assessment of bone marrow infiltration in these patients.
HOTAIR has been implicated in several carcinogenic processes across various solid tumors and hematological malignancies, including enhanced proliferation, epithelial-to-mesenchymal transition (EMT), invasion, aggressive tumor behavior, and metastasis.27-32 Our study observed a significant upregulation of HOTAIR expression in the plasma of DLBCL patients compared to healthy controls. This finding suggests HOTAIR’s potential as a non-invasive biomarker for DLBCL screening. Furthermore, a positive correlation was observed between HOTAIR levels and abnormal lymphoid cell counts in the bone marrow, implying its potential utility in monitoring disease infiltration. These observations are consistent with previous reports demonstrating elevated HOTAIR levels in DLBCL plasma 33 and lymph nodes. 34 Notably, both these studies associated HOTAIR upregulation with a significant impact on mortality rate and identified it as an independent predictor of poor prognosis or treatment response,33,34 further strengthening the case for its potential clinical application.
In contrast to HOTAIR, SNHG15 expression showed a significant downregulation in DLBCL patient plasma compared to healthy controls. Interestingly, this downregulation emerged as a protective factor against mortality. Although previously linked to unfavorable prognoses in solid tumors,35,36 SNHG15, a member of the SNHG family implicated in cellular stress responses, 37 seems to exhibit a distinct protective role in DLBCL. Notably, and to the best of our knowledge, this study represents the first investigation into SNHG15 expression in DLBCL patient plasma, suggesting a potentially unique function within this hematological malignancy. These findings underscore the necessity for further research to unravel the multifaceted and context-dependent roles of lncRNAs like SNHG15 across different cancer types.
H19 exemplifies the context-dependent nature of lncRNAs. It can function as either a tumor suppressor or an oncogenic factor, depending on the specific tumor microenvironment and its intricate interplay with cellular factors.38,39 While H19 upregulation has been associated with processes that promote tumorigenesis, including motility, growth, migration, invasion, and metastasis, 40 it paradoxically exhibits anti-oncogenic properties in specific contexts, such as pituitary adenomas. Additionally, H19 demonstrates stage-specific roles in certain cancers, like thyroid carcinoma. 40 These observations illuminate the remarkable plasticity and context-dependency inherent to lncRNAs like H19, emphasizing the need for further exploration of their multifaceted functions across diverse disease landscapes.
Previous research has documented elevated H19 expression in hematological malignancies, particularly in both B-cell and T-cell acute lymphoblastic leukemia (ALL). 41 This upregulation extends to BCR-ABL-transformed cell lines, where H19 acts as a crucial factor for BCR-ABL-driven tumor growth. The BCR-ABL protein is the oncogenic driver of chronic myeloid leukemia (CML) and Philadelphia chromosome-positive ALL. 39 Interestingly, our study observed a unique association between H19 upregulation and early-stage DLBCL.
Our study has some limitations. First: Although we included all eligible patients diagnosed during the study period, the relatively small sample size may limit the applicability of our findings to a broader population. Second: While our study analyzed several key features, some important parameters, such as the molecular subtypes of DLBCL, were not included. This limits the comprehensiveness of our analysis and the potential for deeper insights into specific patient subgroups. Finally: the investigation solely examined circulating lncRNA levels in plasma. Integrating this data with tissue expression levels of the same lncRNAs could have provided valuable validation and potentially revealed more expression patterns across different tissue compartments.
Conclusions
Our findings highlight the potential of using circulating lncRNAs for diagnosing DLBCL and provide preliminary insights into their association with disease characteristics such as staging and prognosis. Certain lncRNAs, including HOTAIR, SNHG15, and H19, may hold promise for determining disease stage and predicting patient outcomes.
Recommendations
To establish robust correlations between the expression of different lncRNAs, patient survival, and other genetic and pathological parameters, further long-term follow-up studies are warranted. Such a comprehensive investigation would pave the way for the development of personalized medicine approaches in DLBCL, ultimately leading to improved patient care and management strategies.
Supplemental Material
Supplemental Material - Plasma long non-coding RNAs as biomarkers for bone marrow infiltration and stage in diffuse large B-cell lymphoma
Supplemental Material for Plasma long non-coding RNAs as biomarkers for bone marrow infiltration and stage in diffuse large B-cell lymphoma by Ahmed S Abdelhafiz, Reem Nabil, Mohammed Ghareeb, Dalia Ibraheem, Asmaa Ali, Samar S. Elshazly, Asmaa Mohamed Soliman and Yasser M Bakr in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgments
In this study, we build upon findings initially presented in a poster at British Society for Hematology Annual Scientific Metting on 28-30 April 2024, where preliminary results were discussed.
Authors’ contributions
ASA, RN, and YMB Conceived and designed the experiments. RN, MG, DI, SSE, and AMS collected samples and data. YMB Performed the experiments: AA analyzed the data. ASA, MG, and AA participated in writing the manuscript.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be considered as a potential conflict of interest.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical statement
Data availability statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

