Abstract
The impact of interleukin-10 (IL-10) gene promoter polymorphisms (SNPs) on treatment response in HCV patients was dissimilarly estimated. Hence, the aim of this meta-analysis was to robustly assess the effect of IL-10 SNPs on treatment response in HCV patients. An electronic literature search was carried out through PubMed, EMBASE, Web of science, and Scopus databases. Studies assessing the association between IL-10 polymorphisms and treatment response in HCV patients were included. Studies were excluded if genotype frequencies are not consistent with the Hardy–Weinberg Equilibrium (HWE) or in case of including patients with hepatitis B virus coinfection. Risk of bias in included studies was assessed using the Newcastle–Ottawa Scale. Meta-analyses were performed for the influence of IL-10 gene promoter SNPs (rs1800896 (-1082 A/G), rs1800871 (-819 C/T), and rs1800872 (-592 C/T)) and haplotypes on treatment response in HCV patients. Subgroup analyses, meta-regressions, publication bias assessment, and sensitivity analyses were also conducted. Overall, 32 studies with a total of 5943 HCV cases and 2697 controls were included in the present study. The -1082*G allele was significantly associated with increased risk of non-response (NR) to treatment, OR [95% CI] = 1.29 [1.1–1.51], p = .002. Besides, the rs1800872 -592*C allele was significantly associated with increased NR risk, OR [95% CI] = 1.22 [1.02–1.46], p = .03. Subgroup analysis showed that this association remained significant only in patients treated with PEG-IFN alone, p = .01. The -1082*G/-819*C/-592*C (GCC) haplotype was significantly associated with increased NR risk, OR [95% CI] = 1.62 [1.13–2.23], p = .009. Our results suggest that the IL-10 rs1800896 was associated with NR risk especially in North-African and Asian populations. Moreover, the IL-10 gene promoter -1082*G/-819*C/-592*C (GCC) haplotype which has been associated with higher production of IL-10, was significantly associated with increased NR risk.
Introduction
Hepatitis C virus (HCV) infection remains a serious worldwide health problem despite the use of oral direct-acting antivirals (DAA). In fact, the World Health Organization (WHO) estimated its prevalence as high as 58 million persons in 2019 with 290 000 deaths, mostly from cirrhosis and hepatocellular carcinoma (HCC). 1 Acute HCV infections are usually asymptomatic and most do not lead to a life-threatening disease. Around 30% (15–45%) of infected persons spontaneously clear the virus within 6 months of infection without any treatment. 1 The remaining 70% (55–85%) of persons will develop chronic HCV infection. Of those with chronic HCV infection, the risk of cirrhosis ranges from 15% to 30% within 20 years. 1 The remarkable medical breakthrough in hepatitis C treatment using direct acting agents (DAAs) has simplified the care cascades through its high efficacy in achieving sustained virological response (SVR) and tolerable side effects. 2 Indeed, DAA therapy did benefits beyond SVR, in preventing further worsening of liver status and the occurrence of cirrhosis and HCC. 2 Although DAA can cure 95% of patients, the problem lies in poor access to diagnosis and treatment in several low- and middle-income countries (LMIC). 1 In fact, universal DAA prescription may pose socioeconomic burdens for underprivileged LMIC. 2
Several host factors, including the cytokine network (TNF, IFN-γ, IL-1, IL-6, IL-28, IL-22, IL-27, IL-10, IL-12, IL-35, and IL-17) and the NLRP3 inflammasome, can influence the outcome of HCV infection and treatment response. 3 In addition, genetic polymorphisms of some cytokines such as IL-10 have been associated with response to conventional treatment with pegylated-IFN (PEG-IFN),4,5 or the PEG-IFN/Ribavirin (RIB) combination,6–8 or even DAAs. 9
IL-10 is a pleiotropic cytokine that potently inhibits the production of proinflammatory cytokines such as TNF, IL-1, and IL-6 in diverse immune-cell populations and prevents dendritic cell maturation. 10 Furthermore, IL-10 inhibits the expression of MHC and costimulatory molecules which are critical for cell-mediated immunity against pathogens and tumor cells. 10 The IL-10 gene is highly polymorphic, with most notably 3 promoter polymorphisms (SNPs), the rs1800896 (-1082 A/G), the rs1800871 (-819 C/T), and the rs1800872 (-592 C/T).11,12 The GCC (-1082/-819/-592) haplotype has been associated with high IL-10 production whereas the ATA haplotype has been associated with significantly lower IL-10 level.11,12
The aim of this meta-analysis was to summarize existing data on the influence of IL-10 promoter SNPs on treatment response in HCV patients by comparing NR risk across IL-10 genotypes and haplotypes.
Material and methods
Search strategy
This study was performed according to the PRISMA guidelines for systematic reviews and meta-analyses (supplemental file 3). 13 An electronic literature search for eligible studies among all papers published prior to November 30, 2023, was conducted through PubMed, EMBASE, Web of science, and Scopus databases. The following search string was used: (((Interleukin-10) OR (IL-10) OR (IL10) OR (Cytokine)) AND ((Polymorphism) OR (SNP) OR (Variant) OR (Mutation) OR (Allele) OR (Allelic) OR (Genotype) OR (Genotypic) OR (rs1800896) OR (-1082 A/G) OR (rs1800871) OR (-819 C/T) OR (rs1800872) OR (-592 C/T)) AND ((Hepatitis C) OR (HCV))). The literature search was carried out without any language restriction. A detailed search strategy is available within the supplemental file 1.
Selection criteria
All studies were independently assessed and evaluated by 2 reviewers (Tarak Dhaouadi, T.D. and Imen Sfar, I.S.) for the inclusion and the exclusion. The following selection criteria were adopted:
Inclusion criteria
• Studies of case-control (retrospective) or cohort (prospective) design. • Studies assessing the association between IL-10 polymorphisms and response to treatment in HCV patients. • Full manuscript with genotypes or alleles or haplotypes frequencies. • Precise definition of the treatment and the treatment response.
Exclusion criteria
• Genotype frequencies are not consistent with the Hardy–Weinberg Equilibrium (HWE) as recommended by Trikalinos et al.
14
• Studies including patients with hepatitis B virus coinfection. • Case series of subjects, narrative or systematic review, comments, or meta-analysis. • If many studies have been carried out using duplicate cases, only the study with complete data and the largest sample size was included.
Definition of the treatment response
SVR was defined as an undetectable level of HCV-RNA 24 weeks after the treatment completion. Relapse (REL) was defined as a detectable HCV-RNA during follow-up in patients with undetectable HCV-RNA at the end of treatment. Non-response (NR) was defined as detectable HCV-RNA at the cessation of treatment.
Data extraction
Data were extracted using a predeveloped form and entered in an Excel datasheet. Two investigators (T.D. and I.S.) independently extracted the following information: first author, year of publication, study design, study duration (months), country, continent, mean or median age, and gender ratio, The number of patients, treatment (PEG-IFN or PEG-IFN/RIB or DAA), the numbers of SVR, REL and NR, the number of controls (if applicable), HCV genotypes (if specified), IL-10 SNPs genotyping method, the frequencies of genotypes, alleles, and haplotypes for each of the following SNPs: rs1800896 (-1082 A/G), rs1800871 (-819 C/T), and rs1800872 (-592 C/T). A third investigator (Awatef Riahi) compared the results of the extracted data for potential discrepancies.
Quality assessment
The quality of eligible studies was assessed independently by 2 reviewers (T.D. and I.S.) using the Newcastle–Ottawa Scale (NOS) 15 which is based on the following 3 general categories: selection (4 points), Comparability of the study groups (2 points) and ascertainment of outcome (3 points). Studies with a score ≥7 were classified as high-quality reports. Additionally, risk of bias was assessed for each included study through a generic form (Excel spreadsheet) and visualized via the Cochrane ROBVIS online tool (https://mcguinlu.shinyapps.io/robvis/). Two additional independent reviewers (Taieb Ben Abdallah and Yousr Gorgi) examined the quality-assessment results.
Study endpoints
The primary endpoint of this meta-analysis was the association of the IL-10 promoter SNPs with NR outcome. The secondary endpoint was to evaluate potential confounding factors that might influence the impact of IL-10 SNPs on response to treatment.
Statistical analysis
rs1800896 genotype frequencies and HWE assessment.

†HWE individual p-value in HCV patients; ‡HWE adjusted p-value in HCV patients.
aNA: not applicable.
rs1800871 genotype frequencies and HWE assessment.

†HWE individual p-value in HCV patients; ‡HWE adjusted p-value in HCV patients.
aNA: not applicable.
rs1800872 genotype frequencies and HWE assessment.
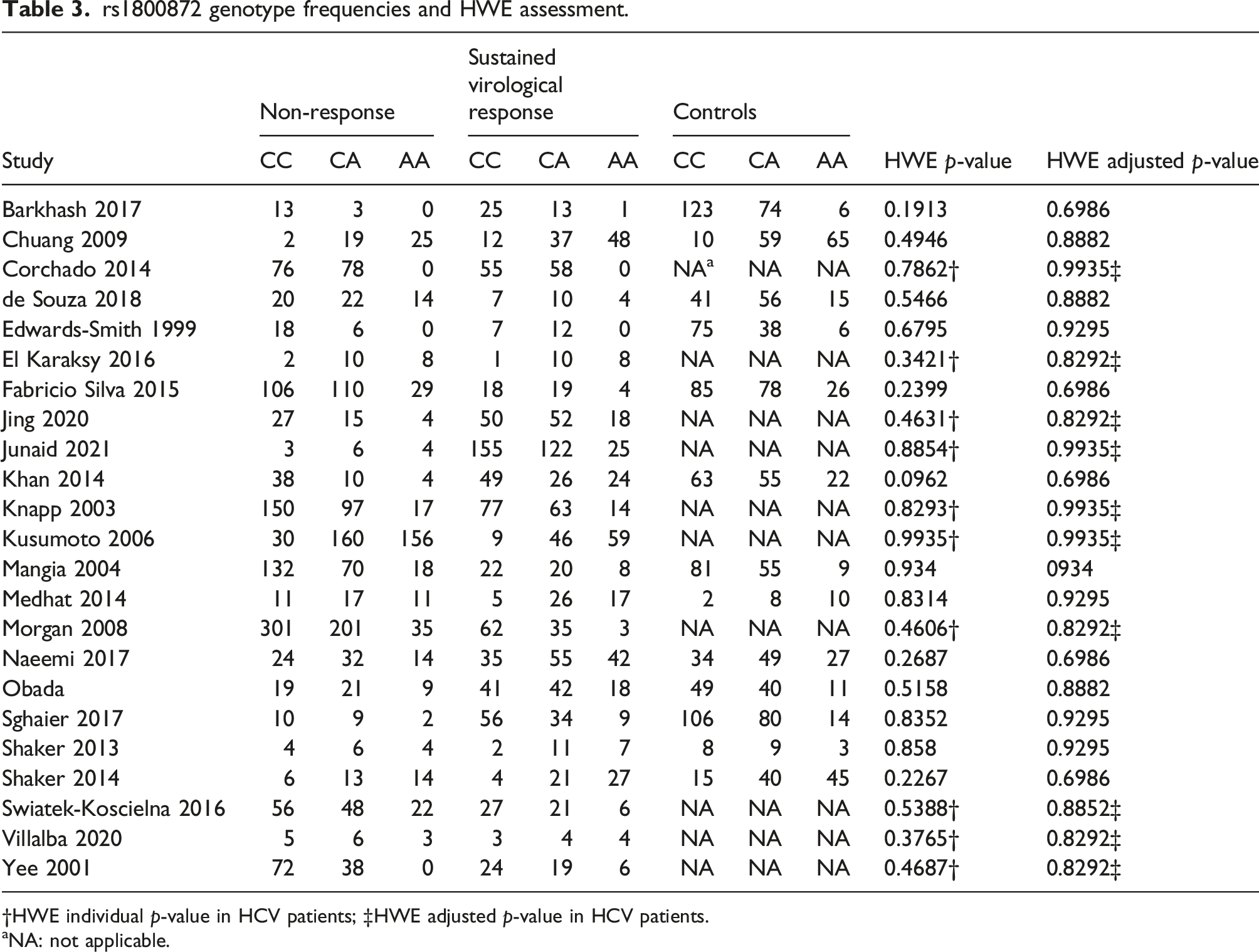
†HWE individual p-value in HCV patients; ‡HWE adjusted p-value in HCV patients.
aNA: not applicable.
A supplementary file with additional tables and figures is available with the full-manuscript.
Systematic review registration
This review has been registered on PROSPERO: CRD42024495977, Available from: https://www.crd.york.ac.uk/prospero/display_record.php?ID=CRD42024495977.
Results
Search results and study characteristics
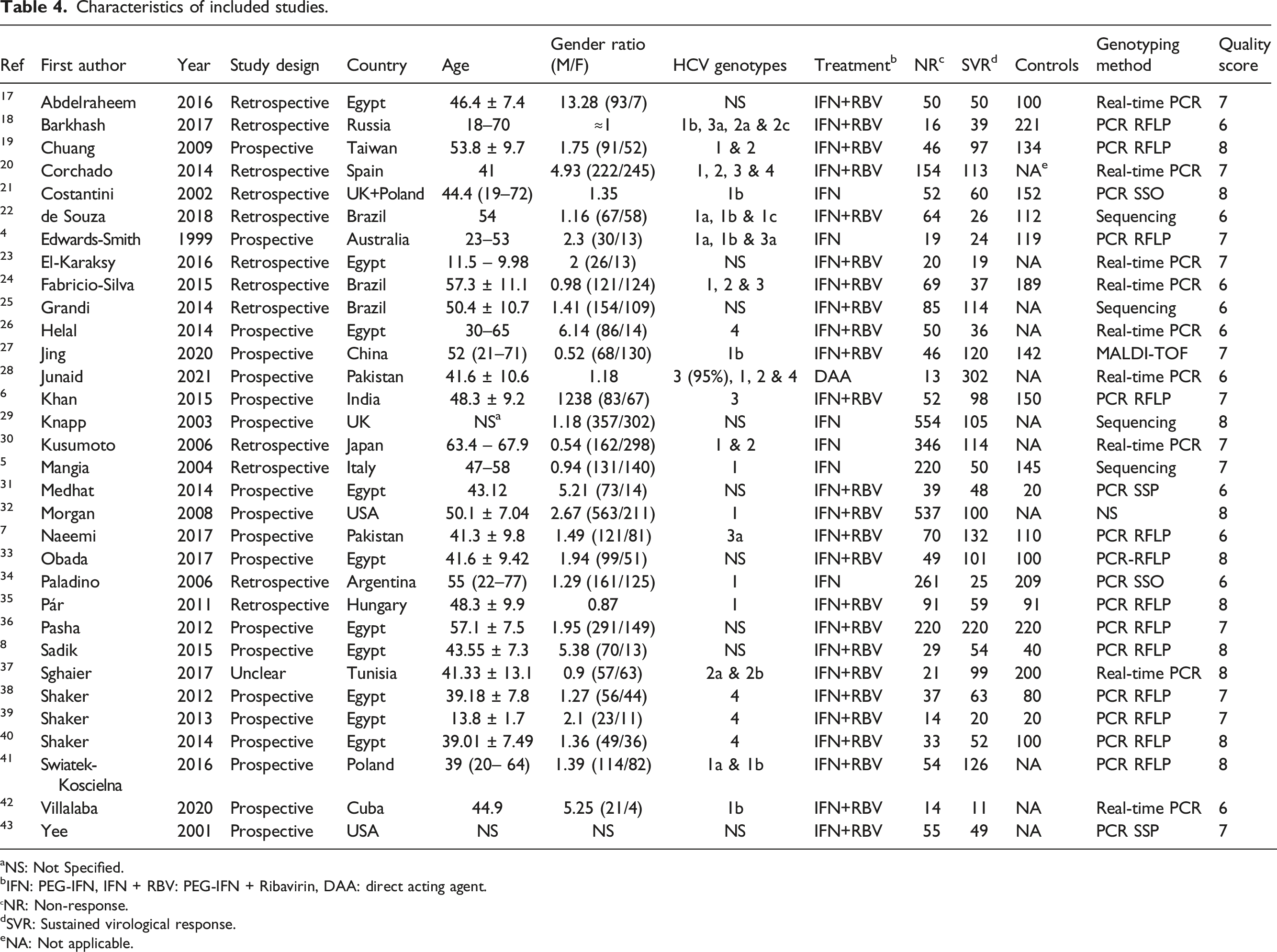
A PRISMA flow diagram was generated to depict the study selection process (Figure 1). Overall, 32 studies with a total of 5943 HCV cases (NR: 3380; SVR: 2563) and 2697 controls were included in the present study.4–8,17–43 Included studies characteristics are summarized in Table 4. Twenty-three studies were included for the rs1800896 (-1082 A/G) SNP,4–8,17,19,22–26,29–37,39,41 17 for the rs1800871 (-819 C/T) SNP,4–7,19,22,27,30,31,33,37–43 23 for the rs1800872 (C/T -592) SNP,4–7,18–20,22–24,27–33,37,39–43 and 9 for the IL-10 haplotype analysis .4,5,7,19,21,29,30,34,41 The NOS quality score results for each included study are shown in Table 4. Risk of bias is summarized in Figure 2. PRISMA flow diagram for study selection. Characteristics of included studies. aNS: Not Specified. bIFN: PEG-IFN, IFN + RBV: PEG-IFN + Ribavirin, DAA: direct acting agent. NR: Non-response. dSVR: Sustained virological response. eNA: Not applicable. Summary of study risk of bias.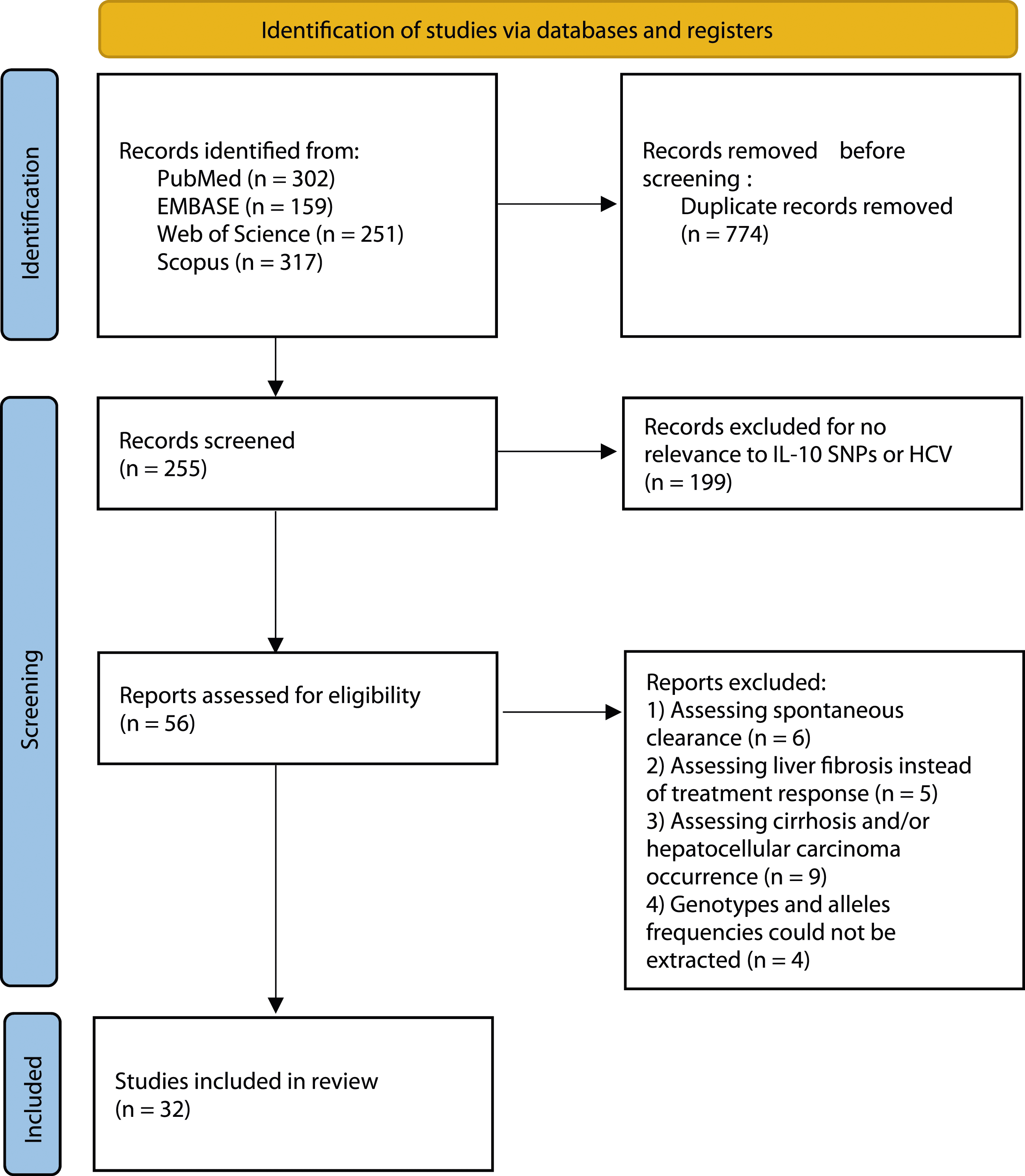

IL-10 rs1800896 meta-analysis
The -1082*G mutated allele was significantly associated with increased risk of NR in HCV patients, OR [95% CI] = 1.29 [1.1–1.51], p = .002 (Figure 3). This association with the codominant model was also significant when recessive and dominant models were applied (Table 5). However, there was a significant amount of between-studies heterogeneity, I2 = 47%, Tau2 = 0.0646, 95% PI = [0.74–2.24] and p = .007. Subgroup analysis by continent revealed that the increase of NR risk conferred by the G allele was significant only in North-African and Asian populations; OR [95% CI] = 1.44 [1.1–1.9], p = .008 and OR [95% CI] = 1.64 [1.11–2.44], p = .013, respectively (Figure 4) (Table S1). Besides, the association between the G allele and the risk of NR remained significant only in patients treated with PEG-IFN/RBV combination (OR [95% CI] = 1.35 [1.11–1.63]) while no significant overall effect was observed in groups treated with PEG-IFN alone (Figure 5) (Table S2). Meta-regression revealed no significant association of the overall effect size with age, the gender ratio (M/F), the SVR/NR ratio, or the frequency of G allele in HCV patients, Omnibus p-value = .308 (Table S3). Forest plot for the association between the IL-10 rs1800896 (-1082 A/G) SNP and NR risk. Main results of the 3 IL-10 SNPs in NR risk. Subgroup analysis by continent for the IL-10 rs1800896 (-1082 A/G) SNP. Subgroup analysis by treatment for the IL-10 rs1800896 (-1082 A/G) SNP.



IL-10 rs1800871 meta-analysis
The integrated analysis did not show any association between the IL-10 -819 C/T SNP and the risk of NR, OR [95% CI] = 1.23 [0.96–1.57], p = .09 (Figure 6). All the genetic models did not show any association with the NR risk (Table 5). Nevertheless, the heterogeneity between included studies was substantial, I2 = 67%, Tau2 = 0.1635, 95% PI = [0.50–3.03] and p < .0001. Subgroup analysis by continent revealed a significant higher NR risk conferred by the C allele in populations from America (OR [95% CI] = 1.89 [1.06–3.36], p = .03) but the 95% PI was [0.01–509.4] attributable to a significant amount of heterogeneity (I2 = 41%) and the small number (n = 3) of studies from America (Figure S2) (Table S4). Besides, stratified analysis by type of treatment showed a significant association between C allele and NR risk in patients treated with PEG-IFN alone (OR [95% CI] = 1.56 [1.02–2.37], p = .039) with, however, a large 95% PI of [0.02–103.71] as a consequence of both heterogeneity (I2 = 47%) and small number (n = 3) of studies (Figure S2) (Table S5). Subsequent meta-regression (Table S6) revealed a positive correlation of the effect size with the gender ratio (Male/Female), p = .039 (Figure 7(A)). This result indicates that the C allele was associated with a greater risk of non-response mainly in men. Inversely, a negative correlation was observed between the effect size and the SVR/NR ratio, p = .013 (Figure 7(B)). Thus, the association between the C allele and the NR risk could be missed in some studies when the SVR rate is high. Forest plot for the association between the IL-10 rs1800871 (-819 C/T) SNP and NR risk. A: Meta-regression for the rs1800871 with gender ratio (M/F). B: Meta-regression for the rs1800871 with the SVR/NR ratio.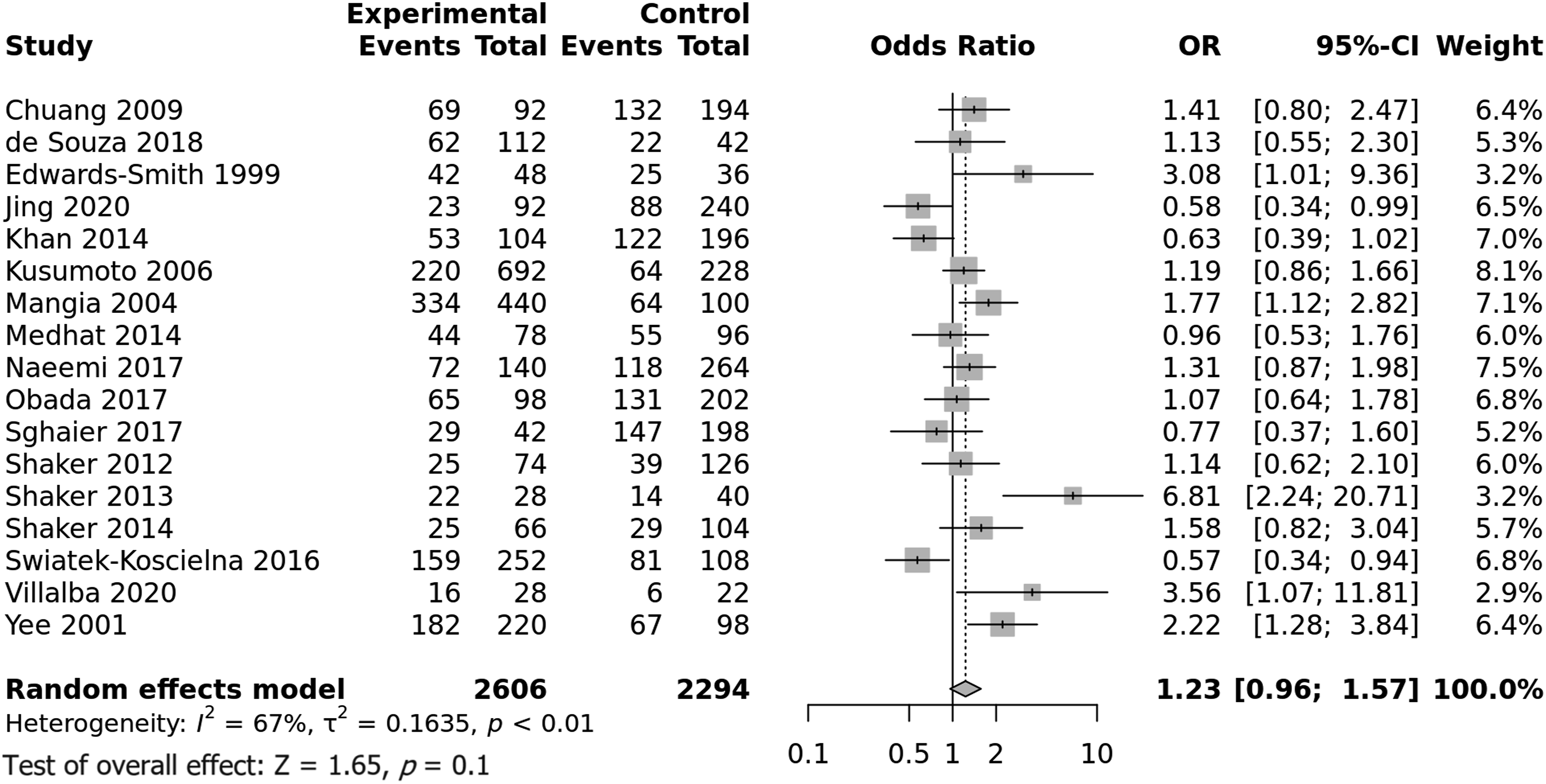

IL-10 rs1800872 meta-analysis
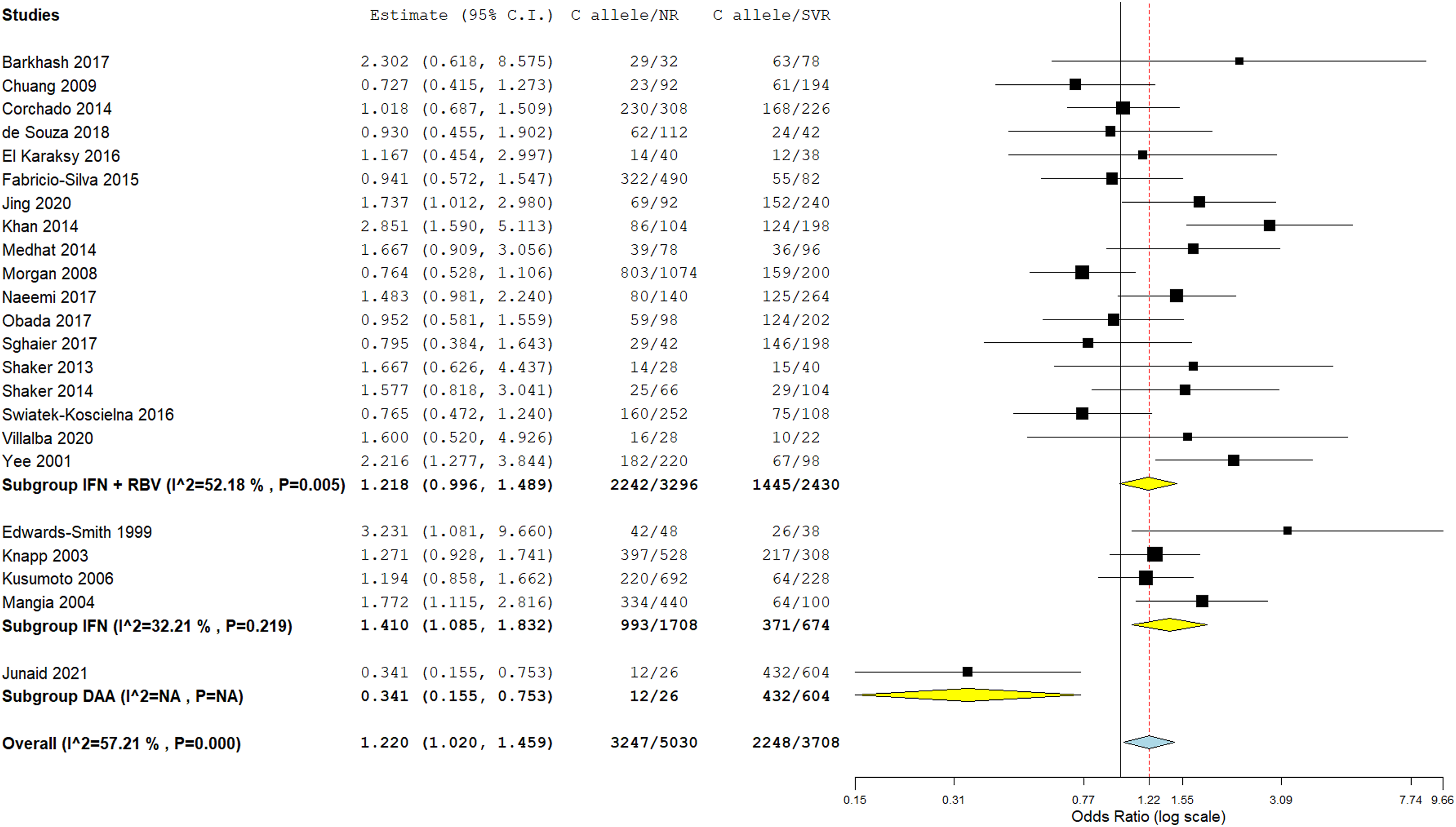
The -592*C wild allele was significantly associated with increased risk of NR in HCV patients, OR [95% CI] = 1.22 [1.02–1.46], p = .03 (Figure 8). This association was also noted when the recessive model was applied while no association was observed in the dominant and overdominant models (Table 5). Nonetheless, there was a significant amount of between-studies heterogeneity, I2 = 57%, Tau2 = 0.0979, 95% PI = [0.62–2.4] and p = .0004. Subgroup analysis by continent revealed no significant associations (Figure S4) (Table S7). In contrast, subgroup analysis by treatment showed that the C allele conferred an increased risk of NR in patients treated with PEG-IFN alone (OR [95% CI] = 1.41 [1.09–1.83], p = .01, 95% PI = [0.61–3.25]), whereas the only study in which patients were treated with DAA
28
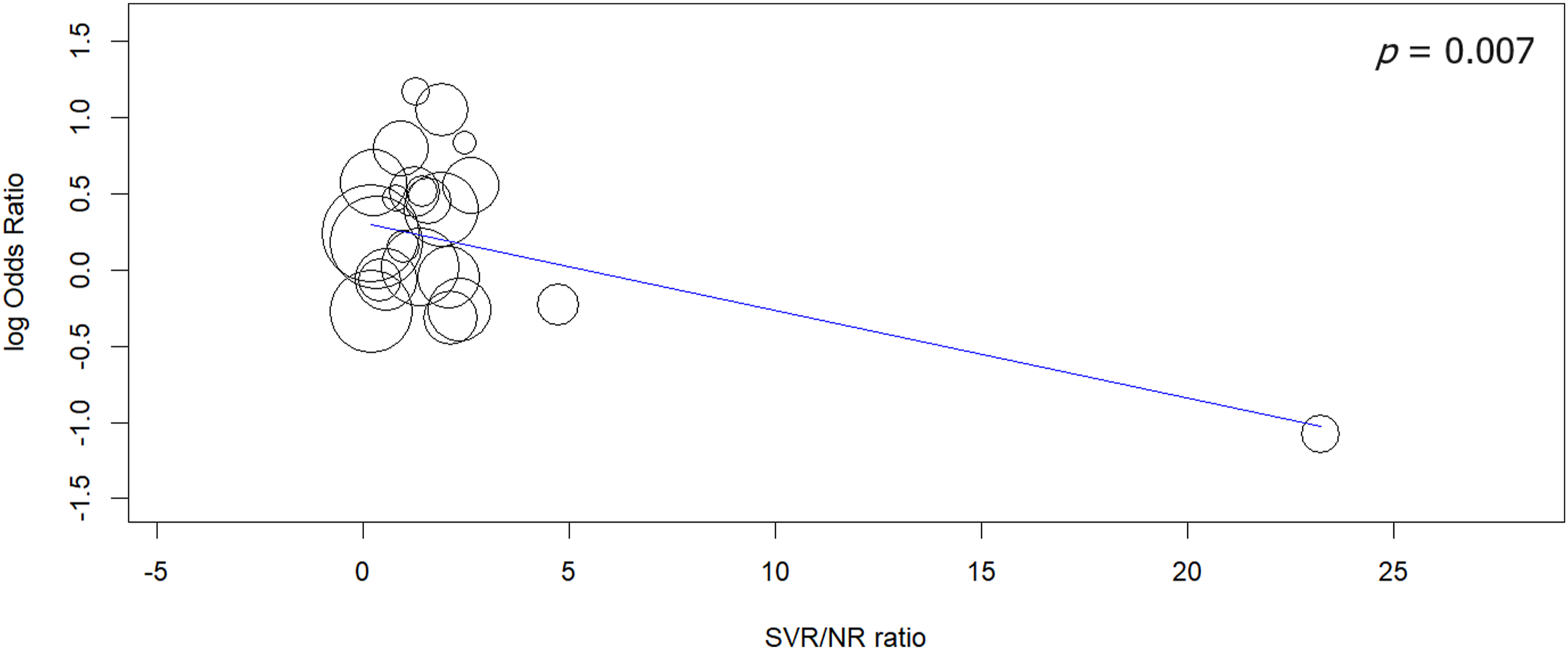
showed that the C allele favored SVR (Figure 9) (Table S8). Besides, meta-regression (Table S9) revealed that the SVR/NR ratio was negatively correlated with the effect size (p = .007) which indicates that a high SVR rate could hide the effect of the C allele on response to treatment (Figure 10). Forest plot for the association between the IL-10 rs1800872 (-592 C/T) SNP and NR risk. Subgroup analysis by treatment for the IL-10 rs1800872 (-592 C/T) SNP. Meta-regression for the rs1800872 with the SVR/NR ratio.
IL-10 haplotype meta-analysis
As the IL-10 gene promoter -1082*G/-819*C/-592*C (GCC) haplotype has been associated with higher production of IL-10,11,12 a meta-analysis was performed to investigate its association with NR risk. The pooling estimate showed a significant association with NR, OR [95% CI] = 1.62 [1.13–2.33], p = .009 (Figure 11). However, there was a significant amount of between-studies heterogeneity, I2 = 68%, Tau2 = 0.1841, 95% PI = [0.54–4.89] and p = .001. Subgroup analysis by continent showed no significant association for studies carried out in Asia and Europe, while a single study carried out in America and another in Australia showed that the association between the GCC haplotype and NR was significant, p = .01 and p = .0017, respectively (Figure S5) (Table S10). Conversely, the subgroup analysis by treatment revealed a significant association of the GCC haplotype with NR risk in patients treated with PEG-IFN alone, p = .035 (Figure 12) (Table S11). Subsequent meta-regression (Table S12) revealed a significant positive correlation between the gender ratio (M/F) and the overall effect size, p = .014 (Figure 13). Hence, the effect of the GCC haplotype on NR outcome seems to be greater in men than in women. Forest plot for the association between the IL-10 promoter GCC haplotype and NR risk. Subgroup analysis by treatment for the IL-10 GCC haplotype. Meta-regression for the IL-10 GCC haplotype with gender ratio (M/F).


Sensitivity analysis
The sensitivity analysis revealed that association results were stable for all performed meta-analyses (Figures S6, S7, S8, and S9), signifying a high level of integrity with reliable results.
Publication bias
Generated funnel plots (Figure 14) were found to be overall symmetrical and Egger’s tests confirmed these findings with non-significant p-values for the 3 IL-10 SNPs (0.1557, 0.1366, and 0.4579) and the GCC haplotype (0.3429), which indicated that results were not weakened by publication biases. Funnel plots assessing publication bias.
Discussion
In untreated 44 HCV-infected patients, the high frequency of chronicity outcome is striking as HCV is an RNA virus with no DNA intermediate in its cycle and no ability to integrate into the host genome. Hence, it is obvious that HCV has developed extraordinary mechanisms to insure its persistence despite innate and adaptive immune anti-HCV responses. 45 In treated patients, there are many factors including viral, host, and treatment-related factors that determine treatment response. 46 Viral factors include HCV genotypes, subtypes, and baseline viral load. 46 Indeed, SVR rates are higher in case of genotypes 2, 3, 5, and 6 than in 1 and 4. 46 Furthermore, in patients under triple therapy including DAAs, 1a subtype has been associated with higher frequencies of viral breakthrough and drug resistance comparatively to 1b subtype. 46 Besides, several host factors including age, obesity, HIV coinfection, insulin resistance, vitamin D levels, the extent of liver fibrosis, and the genetic compound impact negatively SVR rate. 46 Regarding the host genetic compound, IL-28B SNPs were found to be the strongest pre-treatment SVR predictor. 46 However, IL-28B variants account for roughly 50% of the host-genetic influence on treatment response. Indeed, numerous other genetic factors including IL-10 SNPs were found to influence treatment response. 47 It is of note that HCV has been shown to induce IL-10 production, and increased IL-10 levels were associated with persistent HCV infection, higher inflammation grade, and an increased risk of hepatocellular carcinoma occurrence. 47
Over the past two decades, several studies have focused on the relationship between 3 IL-10 SNPs, namely rs1800896, rs1800871, and rs1800872, and treatment response. However, published data were mostly inconclusive. Although a previous meta-analysis 48 was published in 2016, the literature search cut-off time in our study was fresher. In addition, while the study of Guo et al. 48 involved only 14 studies, our meta-analysis included 32 reports which could provide a more comprehensive and accurate estimation of IL-10 SNPs influence on NR risk.
The present study revealed that the IL-10 -1082*G allele was associated with approximately 29% increase in risk of NR. However, heterogeneity between studies was substantial and the true effect in 95% comparable populations could vary from 26% decreased risk to 124% increased risk. Subgroup analysis by continent revealed that NR risk conferred by the -1082*G allele was significant only in North-African and Asians populations, but again there was a large amount of heterogeneity between studies. Regarding the treatment, the association between the -1082*G allele and NR risk remained significant only in patients treated with PEG-IFN/RBV combination whereas no significant influence was noted in case of PEG-IFN alone use. A subgroup analysis by HCV genotype could not be performed, as many studies reported a varied mixture of genotypes, and many did not even provide information on the involved HCV genotypes. Besides, univariate and multivariate models of meta-regression did not show any influence of age, gender ratio, SVR/NR ratio, and the G allele frequency on the relationship between IL-10 rs1800896 SNP and NR risk.
In this study, the rs1800871 (-819 C/T) was not found to be associated with NR risk. As there was a large amount of between-studies heterogeneity subgroup analysis by continent and treatment together with a meta-regression were performed. Subgroup analysis by continent revealed a significant increased NR risk conferred by the -819*C allele in patients from the American continent, though the 95% PI was too large. Similarly, the significant increased NR risk in patients treated with PEG-IFN alone was hampered by the considerable amount of heterogeneity between studies and the wide 95% PI. As for the rs1800896 SNP, a subgroup analysis by HCV genotype for the rs1800871 SNP was unfeasible. Interestingly, meta-regression revealed a significant positive correlation of NR risk with the gender ratio (M/F) indicating that the -819*C allele conferred a higher risk of NR in male patients. This peculiar finding corroborates those of previous reports in which female patients had significantly higher SVR rates49,50 and a more frequent spontaneous HCV clearance [51, 52]. Besides, -819*C allele conferred NR risk was negatively correlated to the SVR/NR ratio. In this regard, the SVR/NR ratio varied considerably between studies from 0.095 to 23.23. Hence, the relationship between the -819*C allele and NR risk could have been missed in studies with high SVR rate.
IL-10 -592*C allele was found to be associated with 22% increase in NR risk in the present study. Nevertheless, heterogeneity of between-studies was important and the true effect in 95% comparable populations could diverge from 38% decreased risk to 140% increased risk. Subsequent subgroup analysis by continent showed that the association between the rs1800872 SNP and NR risk remained significant only in studies from Asia and America, though with high levels of heterogeneity and wide 95% PI. Regarding treatment, the -592*C allele was significantly associated with increased NR risk only in patients treated with PEG-IFN alone, but with decreased NR risk when DAA were used. It is of note that only one study investigated the role of IL-10 rs1800872 SNP in patients treated with DAAs. 28 Hence, this result needs to be replicated in independent large cohorts. Besides, the 592*C allele effect on NR risk was negatively correlated with the SVR/NR ratio. Thus, a low NR rate in some studies could have hidden the association with rs1800872 SNP.
In this study, only 9 eligible reports were included in the IL-10 haplotypes meta-analysis. The GCC haplotype was associated with 62% increase of NR risk. Nonetheless, due to a high level of heterogeneity, the true effect of the GCC haplotype in 95% comparable populations could fluctuate from 46% decreased risk to 389% increased risk. The GCC haplotype association with NR risk was only significant in patients treated with PEG-IFN alone. Interestingly, meta-regression showed a significant negative correlation of the GCC effect on NR risk with the gender ratio (M/F). This finding is in line with literature data which indicated a better HCV outcome in female patients.49–51.
In summary, the present meta-analysis revealed that IL-10 promoter SNPs could play a significant role in treatment response in HCV infected patients. However, it is noteworthy to admit the substantial between studies heterogeneity. To our knowledge, this study was the first to perform several meta-regressions in order to explore the aforementioned heterogeneity. Meta-regression provided the study with some interesting findings such as a positive correlation of the effect size with the gender ratio (M/F) and a negative correlation with the SVR/NR ratio. However, there are some limitations that need to be acknowledged. Firstly, the treatment response depends on several confounding factors such as the HCV genotype, the extent of liver fibrosis, the body mass index, etc., and as the present study analyses derived from pooling IL-10 SNPs aggregate findings without any access to raw data, there was a lack of further adjustment for baseline characteristics. Secondly, there was a diverse mixture of HCV genotypes in many studies without any raw data on the interaction between IL-10 SNPs and HCV genotypes, whereas several other studies did not even specify the HCV genotype. This issue prevented us from performing a robust subgroup analysis by HCV genotype. Thirdly, there was no published study from sub-Saharan Africa. Hence, the results of the present meta-analysis could not be generalized to sub-Saharan African populations. Fourthly, there was only one included study that investigated IL-10 SNPs influence on response to DAAs. Further studies are therefore required to accurately estimate the impact of IL-10 SNPs on treatment response in the DAA era.
In spite of the above-mentioned limitations, our results suggest that the IL-10 rs1800896 SNP was associated with NR risk especially in North-African and Asian populations. Moreover, the IL-10 gene promoter -1082*G/-819*C/-592*C (GCC) haplotype which has been associated with higher production of IL-10 was significantly associated with increased NR risk. Of note, some associations with NR risk were influenced by study-gender-ratios, SVR/NR ratios, and the type of treatment given to HCV patients.
Conclusions
This study demonstrated that IL-10 promoter SNPs could play a significant role in HCV treated infection outcome even though there was a substantial heterogeneity between studies. Further studies investigating their influence on response to DAA are needed.
Supplemental Material
Supplemental Material - Impact of IL-10 gene promoter polymorphisms on treatment response in HCV patients: A systematic review, a meta-analysis, and a meta-regression
Supplemental Material for Impact of IL-10 gene promoter polymorphisms on treatment response in HCV patients: A systematic review, a meta-analysis, and a meta-regression by Tarak Dhaouadi, Awatef Riahi, Taïeb Ben Abdallah, Yousr Gorgi and Imen Sfar in International Journal of Immunopathology and Pharmacology.
Footnotes
Author contributions
Conceptualization: Tarak Dhaouadi, Awatef Riahi, Taieb Ben Abdallah, Yousr Gorgi, Imen Sfar. Data curation: Tarak Dhaouadi, Imen Sfa.Formal analysis: Tarak Dhaouadi, Awatef Riahi, Imen Sfar. Investigation: Tarak Dhaouadi, Imen Sfar. Methodology: Tarak Dhaouadi, Awatef Riahi, Taieb Ben Abdallah, Yousr Gorgi, Imen Sfar. Supervision: Awatef Riahi, Taieb Ben Abdallah, Yousr Gorgi, Imen Sfar. Writing - original draft: Tarak Dhaouadi. Writing - review & editing: Tarak Dhaouadi.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was granted and supported by the Research Laboratory in Immunology of Renal Transplantation and Immunopathology (LR03SP01), Charles Nicolle Hospital, Tunis El Manar University, Tunisia.
Ethical statement
Data Availability Statement
The data that support the findings of this study are available within the article [and/or] its supplementary materials.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
