Abstract
The study was carried out to determine, by PCR-RFLP, the magnitude of drug resistance in Mycobacterium tuberculosis. The study was carried out on 221 random sputum samples collected from patients and 120 suspected cases of drug resistance. Genetic variation in drug-resistant strains was evaluated through PCR-RFLP for isoniazid, ethambutol, streptomycin, and ofloxacin. Out of 341 patients, 91.5% were confirmed as M. tuberculosis complex infected on the basis of PCR. The random samples revealed resistance in 8.2% cases, while 73.3% of those with suspected drug resistance were found resistant. Among drug-resistant isolates, 56.1% were resistant to a single drug, 33.3% to two drugs, and 10.6% to more than two drugs. Ofloxacin resistance was observed along with isoniazid, ethambutol, and streptomycin in 6.5% cases. Resistance to isoniazid was observed in 61% cases, to ethambutol in 50.4%, and to streptomycin in 43.1% cases. It was concluded that PCR-RFLP is a useful molecular technique for the rapid detection of mutations in drug-resistant tuberculosis patients and may be used to diagnose drug resistance at the earliest.
Introduction
Pakistan is placed among 22 high burden countries in terms of estimated tuberculosis (TB) cases with the frequency of 620/100,000 persons. 1 According to the World Health Organization (WHO), 55% of the TB patients worldwide are located in Asia, 30% in Africa, 7% in the eastern Mediterranean region, 4% in Europe, and 3% in the Americas. 1 In addition to the prevalence of TB the emergence of drug-resistant tuberculosis is the significant problem. Multiple drug-resistant tuberculosis (MDR-TB) results when Mycobacterium tuberculosis becomes resistant to most of the baseline anti-TB drugs such as isoniazid (INH) and rifampicin (RIF). 2 Extensively drug-resistant tuberculosis (XDR-TB) is resistant to RIF, INH, streptomycin (SM), and pyrazinamide, as well as to aminoglycosides (kanamycin, amikacin, capreomycin) and fluoroquinolones (ofloxacin, moxifloxacin). XDR-TB is the most challenging form of drug resistance because of limitations in treatment options. M. tuberculosis drug resistance has been characterized by a number of mutations in genes involved in drug metabolism. 3 Mutations in the rpoB and katG genes have been shown to be responsible for resistance to RIF and INH, respectively. 3 Most (95%) of the mutations in cases of RIF resistance were located in the 81-bp region of the rpoB gene that codes for the β-subunit of RNA polymerase. INH resistance is coupled with a variety of mutations affecting one or more than one gene such as those encoding katG gene (catalase-peroxidase). 4 Mutations in the katG gene at codon 315 is associated with resistance to INH in 60–70% of the cases 3 while mutations at codon 306 of the embB gene is specifically associated with ethambutol (EMB) resistance. 5 Fluoroquinolone resistance is caused by mutations in gyrA and gyrB genes. 6 SM is the drug responsible for inhibiting the synthesis of protein. The genes rpsL encoding S12 ribosomal protein are involved in SM resistance. 7 There is a lack of gene base data about the existence of drug resistance strains of M. tuberculosis in Pakistan; therefore, the present study was carried out to study the PCR-RFLP based drug resistance against INH, EMB, SM, and ofloxacin in M. tuberculosis complex isolates/strains from a local population.
Materials and methods
The study was carried out on sputum samples from 120 individuals with suspected drug-resistant TB cases and 221 random samples collected from local TB hospitals. They are considered as suspected drug-resistant TB cases due to persistent clinical symptoms and their response to first line anti-TB drugs. The suspected drug resistant cases were included in the study to observe the mutation in selected genes for drug resistance. The pattern of random TB cases was matched and compared with the suspected cases of drug resistance. These patients belonged to District TB Hospital Faisalabad, General and Civil Hospital Faisalabad, Gulab Devi Chest Hospital Lahore, and Mayo Hospital Lahore. The experimental work was carried out in the Molecular Pathology research lab in the Department of Pathology, University of Agriculture Faisalabad, Pakistan. The study was formally approved by the synopsis scrutiny/ethical committee of the University of Agriculture, Faisalabad. The patients were informed about the study before collecting samples and data. The sputum samples were collected from random cases, while the culture isolates were obtained from provincial TB reference laboratory, Lahore-Punjab.
Sputum samples were decontaminated by using sodium hydroxide by the modified Petroff method. 3 The phenol chloroform method was used for the extraction of bacterial DNA from sputum samples and boiling method was used for obtaining DNA from colonies of the organism on culture media. 8 Direct PCR was performed for the identification of bacteria from sputum samples of all study participants by amplification of specific sequences. Visualization of amplified DNA was done by running on 1% agarose gel and bands were seen under ultraviolet light transilluminator (Biorad, Gel Doc Ez. Imager). Oligonucleotide primer sets specific to 16S rRNAwas used for the identification of Mycobacterium genus and MPB70 for identification of M. tuberculosis complex. 9 The oligonucleotide primers set used for the amplification of the katG, embB, rpsL, and gyrA genes were selected from earlier studies.10–13 The PCR was carried out in a total volume of 25 μL with 5 uL of the template, 17 μL of PCR-EZ D-PCR master mix (Biobasic, Cat. No. BS294; 10 mM KCl, 10 mM (NH4)2 SO4, 20 mM Tris HCl, 0.1% Triton X-100, 0.1 mg/mL BSA, 2 mM MgCl2, 200 μM dNTPs), 1 μL Taq DNA polymerase (Biobasic, Cat. No. B0089; 5 u/μL) and 1 μL each of the primers (Forward + Reverse). The tubes were placed into the thermal cycler (Qantarus) for the amplification of specific targets. The M. tuberculosis control was run with these samples, which was already identified in the lab from previous study. 14 Amplified PCR products were electrophoresed using 1% agarose gel with100 bp plus DNA ladder (Vivantis) at 90 V for 1.5 h (Biorad) and visualized under Gel Doc Ez Imager (Biorad).
Restriction fragment length polymorphism (RFLP) was performed on the PCR products for evaluating the drug resistance. For restriction analysis, 10 µL of PCR product, 1.0 µL of specific restriction enzyme, and 2.0 µL of the buffer supplied with the enzyme were used. For the katG (MSP1; GeneMark), embB306 (NIaIII, Fat I; GeneMark), embB497 (A1wNI, GeneMark), rpsL (MobII; GeneMark), and gyrA (Taq1; GeneMark) genes respective restriction enzymes were used as described earlier.10–13 The restricted products along with 100 bp ladder and negative control were run on 2% agarose gel at 90 volts for 2 h and visualized.
The data obtained were analyzed by applying the Chi-square test and 95% confidence limits were worked out by using SAS software. 15
Results
In the present study, 91.5% (312/341) cases were found to be positive by PCR for M. tuberculosis complex. The collected sample from each individual in the study was analyzed by PCR. Initially 341 PCR were performed in order to screen all the study individuals suspected of TB infection. The results revealed that 39.4% of isolates were resistant to one or more drugs (Table 1). The prevalence of drug resistance observed in random samples was 18.2% and in suspected drug resistant cases was 73.3%. Among drug resistant isolates, 56.1% were resistant to a single drug, 33.3% to two drugs, and 10.6% to more than two drugs (Table 2). Among isolates those were resistant to a single drug, INH resistance was noted in 10.9%, to EMB in 6.41%, and to SM in 4.81%cases. Resistance to INH+EMB was noted in 5.13%, INH+SM in 3.84%, and EMB+SM in 4.47% cases. Resistance to INH+EMB+SM was noted in 1.60% cases and to all these plus ofloxacin in 2.56% cases. The results, regardless of resistance to one or more drugs, found INH resistance in 61% cases, EMB resistance in 50.4%, and SM resistance in 43.1% cases. Among INH and EMB susceptible cases, SM resistance was noted in 12.2% cases (Table 3). In INH-susceptible and EMB-resistant cases, SM resistance was noted in 10.6% cases. Among INH-resistant and EMB-susceptible cases, SM resistance was noted in 9.8% cases. In both INH- and EMB-resistant cases, SM resistance was noted in 10.57% cases.
Overall frequency of drug-resistant TB cases.

Frequency of drug-resistant TB cases against each drug.

Frequency of multiple drug susceptible/resistant cases in different combinations.

The patterns of PCR-RFLP of katG (INH), embB (EMB), rpsL (SM), and gyrA (ofloxacin) are presented in Figures 1–4 with description.

PCR-RFLP results of katG gene for INH resistance. Lanes 1 and 15 DNA Ladder 100 bp plus; Lane 2 negative control; Lanes 3, 5, 7, 9, 11, and 13 (resistant strains with single band of 620 bp after digestion with enzyme indicates mutation), Lanes 4, 6, and 8 (sensitive strains with three bands of about 150, 200, and 230 bp after digestion with enzyme); while in Lanes 10, 12, and 14 (sensitive strains with two bands of 160 and 240 bp after digestion with enzyme).

PCR-RFLP result of embB gene for EMB. Lane 1 and 15 DNA ladder 100 bp plus; Lane 2 negative control; Lanes7, 8, 11, and 12 (resistant strains with single band of about 167 bp after enzymic digestion); Lanes 13 and 14 (sensitive strains with three bands of about 50, 70, and 240 bp); Lanes 3–6, 9, and 10 (no clear banding pattern appears after enzymic digestion).
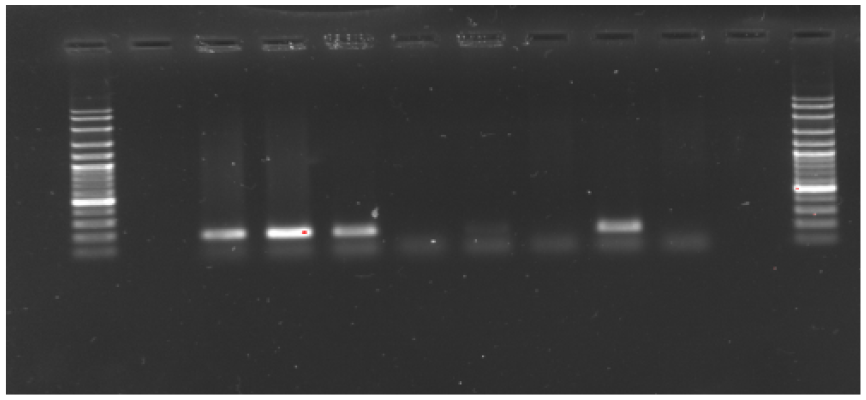
PCR-RFLP result of rpsL gene for SM. Lanes 1 and 12 DNA ladder 100 bp plus; Lanes 2 and 11 negative control; Lanes 3–5 and 9 (resistant strains with single band of about 267 bp); Lanes 6–8 and 10 (no clear band appears after digestion with restriction endonuclease).

PCR-RFLP result of gyrA gene for ofloxacin. Lane 15 DNA ladder; Lanes 1 and 14 negative control. Two types of patterns were observed. Pattern 1, in Lanes 4, 9, and 10 (resistant strains with single band of 260 bp); pattern 2 in Lanes 2, 3, 5, 6, 12, and 13 (sensitive strains with two bands of about 100 and 160 bp) while in Lanes 7, 8, and 11 (no clear band appears after digestion with restriction endonuclease).
Discussion
Sputum smear microscopy by Ziehl Neelsen’s (ZN) staining is a common and widely used technique for the diagnosis of tuberculosis in the developing countries. 16 The limitation of ZN staining is that it can only detect and identify the AFB when 1000 bacilli are present in 1 mL of a sputum sample. 17 The second most widely used technique for the diagnosis of TB is culture isolation. This is currently the gold standard for TB diagnosis because of its sensitivity and specificity, but has the drawback of a longer period (weeks to months) required for the growth of M. tuberculosis isolates. It has been reported that the application of DNA-PCR has improved the Mycobacteria species detection rate. 18 The use of PCR for the detection of MTB complex in clinical specimens has been found to be the most promising tool for the early detection of MTB complex. 19 As ZN staining is widely practiced in Pakistan for initial screening, the results are not confirmatory and should not be used solely as a diagnostic tool and patients should not to be put on expensive and toxic drugs unless diagnosis is confirmed by another means. Therefore, during the present study, all the acid fast bacilli (AFB) positive samples were analyzed through PCR. In the present study, PCR analysis showed that 91.5% cases were found positive for M. tuberculosis complex, while 8.5% cases were positive for Mycobacterium other than MTB complex.
It has been reported that poverty, overcrowding, poor nutrition, addiction, homelessness, and lack of healthcare facilities are the major reasons in the persistence and spread of TB infection and the emergence of MDR and XDR-TB strains. 20 Drug resistance in M. tuberculosis originates from the spontaneous and/or random mutations in chromosome resulting in decreased susceptibility to specific agents. 21 The molecular basis of drug resistance to TB was first reported by Zhang et al. in 1992. 22 Zhang et al. 22 found that mutations in the katG gene were linked to the resistance to INH in M. tuberculosis isolates. Aarts et al., in their study, used PCR-RFLP technique for mutation detections associated with antibiotic resistance on the basis of the fact that mutations can increase or decrease the recognition sites for restriction enzymes. 23 In the present study, we found that 39.4% isolates were resistant to one or more drugs. Among them, 56.1% were resistant to a single drug, 33.3% to two drugs, and 10.6% to more than two drugs. A study reported an overall first line drug resistance in 59.0% of the isolates and 13.2% of the cases were found resistant to two or more than two drugs. 24 They also reported that 12.5% of the isolates were XDR strains. INH is considered as the first-line anti-TB drug which is active against the growing tubercle bacilli. The 64% of isolates have been reported to be found resistant to INH in one study, 25 while 75% were found resistant in another. 26 In our case, we found 60.98% cases were found resistant to INH. This suggests that the resistance to INH is higher than other drugs. Leung et al. observed higher frequency of INH resistant strains. 27 Similarly, a study reported an increased incidence of katG mutations in INH-resistant strains. 28
EMB is another first line anti-TB drug which interferes with the cell wall biosynthesis. EMB resistance was observed in 50.41% cases during the present study. Other studies have reported 32% resistant cases to EMB, while another study reported 71.5% resistance to EMB.29,30 We studied the EMB gene codon 306 and 497 by digesting the amplified product with NIaIII (Fat I) and A1wNI, respectively, as a result multiple banding patterns were obtained.
SM is an antibiotic used for anti-TB drug therapy that kills the growing tubercle bacilli. It prevents the synthesis of protein causing misinterpretation of the mRNA message in the translation process. The resistance to SM during the present study was 43.09%. Another study reported 55% resistance to SM, while 25% resistance is also reported.29,30 Nachamkin et al. 31 reported that 76.2% of clinical isolates of M. tuberculosis were resistant to refampicin, 44.4% to INH, and 28.1% to SM.
Fluoroquinolones such as gatifloxacin and moxifloxacin, as well as aminoglycosides and cyclic peptides, are used for the treatment of MDR-TB. Mutation in the quinolone resistance-determining region (QRDR) of the gyrA gene is the principal mechanism of fluoroquinolone resistance in M. tuberculosis. The first report published of gyrA gene mutation for the detection of MDR-TB in Kuwait concluded that molecular methods were useful tools for the diagnosis of MDR/XDR-TB. 32 We also studied the gyrA gene mutation through PCR-RFLP in the present study and found different banding patterns associated with ofloxacin resistance. Zhao et al. 13 studied the gyrA gene mutation through PCR-RFLP and found it much more sensitive than the conventional phenotype method. In the current study 1.6% of strains were MDR and 2.6% XDR. Another study from Pakistan reported MDR strains to be 9.2%; 33 however, a much higher prevalence of 66.1% of MDR strains has also been reported. 34 These results indicate a variation in the incidence of MDR and XDR strains, but the important fact is that these strains are present which should be considered as a worrying factor.
We concluded that PCR-RFLP is a useful diagnostic tool for the rapid diagnosis of drug-resistant tuberculosis. It enables the early detection of mutations in the drug resistant genes. It can be concluded that molecular methods may be used in future for the rapid diagnosis of mycobacterial infection and drug resistance.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This article was funded by Higher Education Commission, Islamabad through project grant 20-1519/R and D/09.
