Abstract
Background
Esophageal cancer is an aggressive malignant tumor, and its incidence rate is constantly increasing. Searching for a new biomarker is essential for the study of esophageal cancer.
Methods
The microRNA and messengerRNA expression levels were measured by employing RT-qPCR. The diagnostic value of miR-1303 in esophageal cancer was measured by the receiver operating characteristic curve. In TE-1 cells and KYSE520 cells, miR-1303 and CLDN18 were overexpressed or inhibited by transfection. The target of miR-1303 was forecasted by the TargetScan online database, which was identified by the dual-luciferase reporter system. The correlation between miR-1303 and CLDN18 in esophageal cancer was analyzed.
Results
MiR-1303 in esophageal cancer tissues increased and had a high diagnostic value in esophageal cancer. In TE-1 cells and KYSE520 cells, the overexpression of miR-1303 increased the proliferation and decreased the apoptosis, and the inhibition of miR-1303 decreased the proliferation and increased the apoptosis. The overexpression of miR-1303 significantly increased MMP2 and MMP9 and decreased E-cadherin in TE-1 cells and KYSE520 cells, and the inhibition of miR-1303 decreased MMP2 and MMP9 and elevated E-cadherin in TE-1 cells and KYSE520 cells. In esophageal cancer tissues, the expression of CLDN18 was significantly reduced, and CLDN18 was negatively correlated with miR-1303. In TE-1 cells and KYSE520 cells, the up-regulation of CLDN18 alleviated the progression of esophageal cancer caused by miR-1303.
Conclusion
MiR-1303 was increased in esophageal cancer tissue and had a high diagnostic value in esophageal cancer. MiR-1303 promoted the invasion and metastasis of esophageal cancer. In esophageal cancer, the inhibition of miR-1303 suppressed the development by targeting CLDN18.
Introduction
Esophageal cancer (EC) is a common malignant tumor and ranks sixth among cancer-related mortality rates worldwide. 1 EC is classified into esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC) in histopathology. 2 EC has had a serious impact on patients’ quality of life and increased the social burden. Therefore, it is essential to find a new biomarker for the early prevention and pathogenesis of EC.
In recent years, the focus on microRNA (miRNA) has intensified. MiRNA targets oncogenes or tumor suppressor genes to regulate processes such as proliferation and apoptosis. 3 The focus of our research was mainly on miR-1303. Some evidence has indicated that miR-1303 relates to proliferation and apoptosis in cancer. In prostate cancer (PCa), miR-1303 is highly expressed and promotes the proliferation, migration, and invasion of PCa. 4 In liver cancer (LC), miR-1303 is related to the progression of LC. 5 In addition, miR-1303 is related to the metastasis of osteosarcoma. 6 These studies indicate that inhibiting miR-1303 may suppress the proliferation and invasion of cancer cells. Therefore, it is conjectured that the down-regulation of miR-1303 may inhibit the development of EC.
Claudins are a family of transmembrane proteins and an important component for tight cellular connection, and among them, Claudin-18 (CLDN18) is located on the surface of the cell membrane and is a crucial factor in maintaining cell polarity and barrier function. 7 The subtype of CLDN18, CLDN18.2, is activated and overexpressed in many primary cancers, especially in digestive system malignancies. 8 Much evidence has shown that CLDN18.2 is closely connected with EC. 9 It can be reasonably speculated that CLDN18 also plays an important role in EC.
In this study, our focus is on the expression and role of miR-1303 in EC to provide a new biomarker for EC. Meanwhile, the targeting relationship between miR-1303 and CLDN18 in EC has also been studied.
Materials and methods
Patients and specimens
From September 2021 to June 2024, 70 patients from Nanjing Drum Tower Hospital Clinical College of Nanjing Medical University with EC underwent esophagectomy (36 males and 34 females). A total of 70 pairs of EC tissues and the adjacent tissues were obtained through the operation. The adjacent tissues were more than 5 cm away from the tumor edge. All the subjects had no previous history of malignant tumors, chemotherapy, or radiotherapy. This experiment was approved by the Ethics Committee of Nanjing Drum Tower Hospital Clinical College of Nanjing Medical University and adhere to the tenets of the Declaration of Helsinki. All the subjects signed the written consent form.
Cell culture
The 293T cells, human EC cell lines TE-1 and KYSE520, and normal esophageal epithelial cell line HET-1A were obtained from iCell Bioscience Inc (Shanghai, China). HET-1A cells are a type of epithelial cell derived from normal human esophageal tissue through the SV40 virus gene, which are widely used as a “normal” control cell model in EC research. TE-1 and KYSE520 are classic cell models derived from patients with EC and are particularly suitable for reflecting the disease characteristics of the Asian population. All cells were cultured in Dulbecco modified Eagle medium + 10% fetal bovine serum (Thermo Fisher Scientific, Waltham, MA, USA) and maintained at 37°C and 5% CO2.
Cell transfection
To up-regulate or down-regulate miR-1303, miR-1303 mimic or miR-1303 inhibitor was transfected into TE-1 cells and KYSE520 cells. To up-regulate miR-1303 and CLDN18, miR-1303 mimic and the pcDNA3.1-CLDN18 (OE-CLDN18) or the pcDNA3.1 vector (OE-NC) were transfected together into the TE-1 cell line and the KYSE520 cell line. All plasmids from GenePharma (Shanghai, China) were employed.
Dual-luciferase reporter assay
The binding sequence of the CLDN18 3′ UTR for miR-1303 was amplified and inserted downstream of the luciferase gene in the pmirGLO plasmid (Promega, Madison, WI, USA). This created a wild-type double luciferase reporter plasmid (CLDN18 Wt) and a mutant double luciferase reporter plasmid (CLDN18 Mut). The CLDN18 Wt and CLDN18 Mut plasmids were co-transfected into 293 T cells with either the miR-1303 mimic or the miR-1303 inhibitor. After 48 h, the cells were harvested, lysed, and centrifuged to collect the supernatant. Finally, luciferase activity was then measured.
RT-qPCR
All RNA was extracted. SYBR qPCR Mix and RT-qPCR system were employed to detect the expression of miR-1303, CLDN18, E-cadherin, and matrix metalloproteinase (MMP) 2 and 9. The expression of miR-1303 and other genes was normalized to U6 and GAPDH. All gene expression levels were calculated using the 2−ΔΔCT method.
Cell counting kit-8 assay
A suspension of TE-1 cells and KYSE520 cells was prepared, with a cell count of 2 × 103 cells/mL being achieved. A 100 μL cell suspension was cultured in a 96-well plate for 24 h. 10 μL of cell counting kit-8 (CCK-8) solution was added to the plate and incubated for 1 h. The absorbance of a 450 nm wavelength was measured using an enzyme-labelled instrument with a 96-well plate.
Flow cytometry
Apoptosis of the TE-1 cells and KYSE520 cells was detected using a flow cytometer and an Annexin V-FITC/PI apoptosis detection kit. The cells were suspended three times in cold phosphate-buffered saline, adjusted to a concentration of 1 × 106 cells/mL, and stained with the staining solutions. Flow Jo V10 software was then used to analyze the apoptosis.
Western blot
Cells were harvested and lysed, and total protein was extracted and calculated by the BCA method. The separation gel and the concentrated gel were formulated. After loading, the concentrated glue was run at a constant pressure of 80 V, and the separated glue was run at 120 V until the target strip reached the middle and lower parts of the separated glue. After activating and balancing the PVDF membrane, the membrane transfer was carried out. The membranes were blocked with 5% skim milk for 2 h at room temperature and washed three times with 0.05% TBST. They were then incubated overnight with the primary antibody at 4°C. The blot was then incubated with the HRP-labeled secondary antibody at room temperature for 2 h. Finally, the imprints were visualized using ECL reagents (BD Bioscience, Woburn, MA,USA). All antibodies were purchased from Abcam (Cambridge, UK).
Statistical analysis
Data were handled by SPSS 23.0. The t-test and one-way analysis of variance were utilized to analyze the differences between the groups. To analyze the diagnostic value, the receiver operating characteristic (ROC) curve was utilized. The TargetScan online database was used to prove the downstream target of miR-1303 and identify the binding sites of CLDN18. Pearson correlation analysis was utilized to confirm the correlation.
Results
Effects of miR-1303 in EC
MiR-1303 was significantly increased in EC tissues (Figure 1(a)). The ROC curve showed that miR-1303 had significant diagnostic value in EC, with an AUC of 0.839, and sensitivity and specificity were 74.29% and 78.57% (Figure 1(b)).
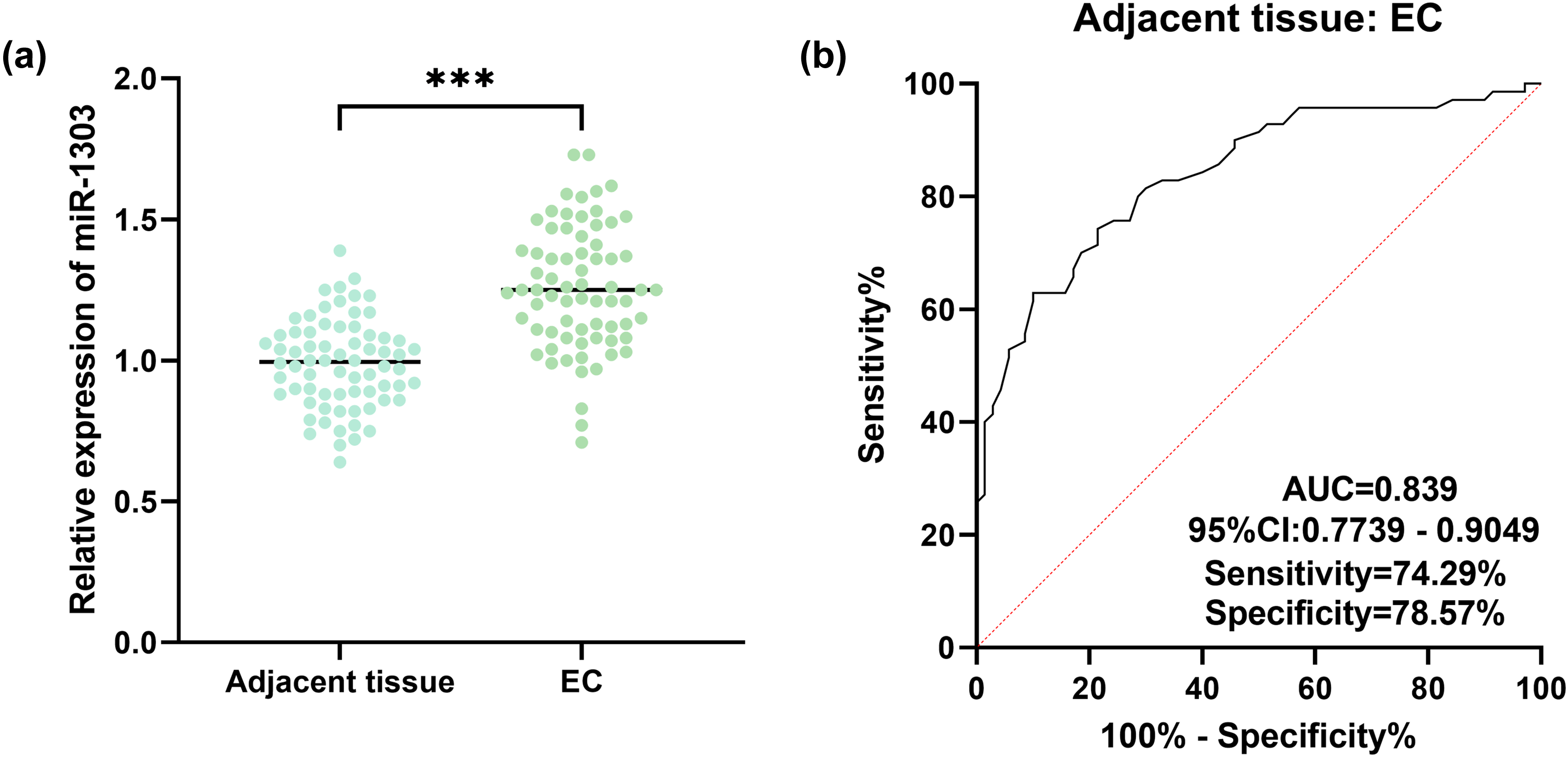
Effects of miR-1303 in EC. (a) Relative expression of miR-1303 in EC tissues. (b) The diagnostic value of miR-1303 in EC. ***
Effects of miR-1303 on proliferation and apoptosis of TE-1 cells and KYSE520 cells
MiR-1303 mimic or miR-1303 inhibitor was transfected into TE-1 cells and KYSE520 cells, which was confirmed by RT-qPCR (Figure 2(a) and (d)). As demonstrated in Figure 2(b), in TE-1 cells, the overexpression of miR-1303 obviously elevated proliferation, and the down-regulation of miR-1303 reduced proliferation. Conversely, in TE-1 cells, the up-regulation of miR-1303 obviously decreased apoptosis, while the inhibition of miR-1303 obviously elevated apoptosis (Figure 2(c)). Similarly, in KYSE520 cells, the overexpression of miR-1303 significantly increased proliferation and reduced apoptosis, while the inhibition of miR-1303 weakened proliferation and elevated apoptosis (Figure 2(e) and (f)).
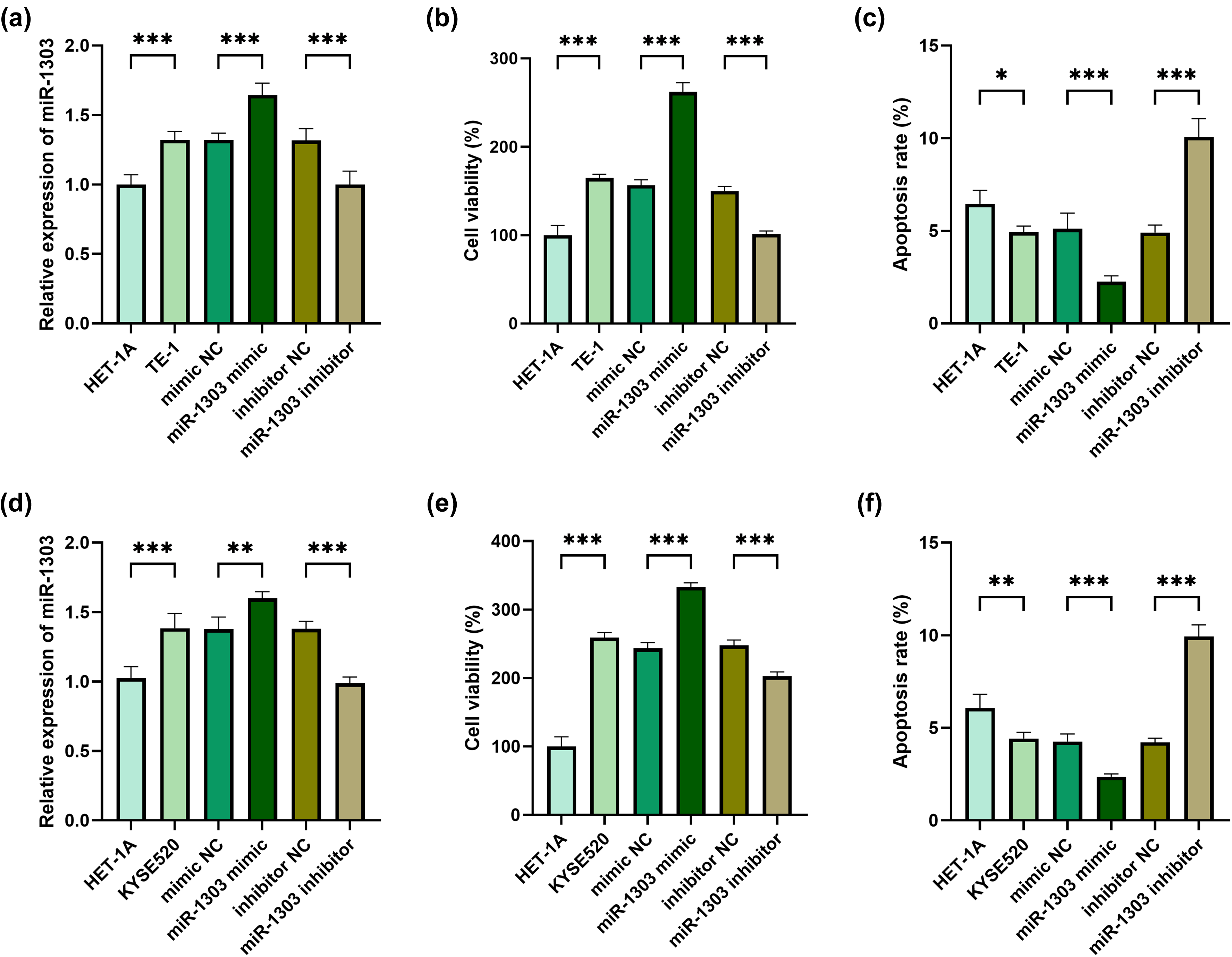
Effects of miR-1303 on proliferation and apoptosis of TE-1 cells and KYSE520 cells. (a) Relative expression of miR-1303 in HET-1A cells and TE-1 cells after overexpression or inhibition of miR-1303. (b) OD value in HET-1A cells and TE-1 cells after overexpression or inhibition of miR-1303. (c) Apoptosis in HET-1A cells and TE-1 cells after overexpression or inhibition of miR-1303. (d) Relative expression of miR-1303 in HET-1A cells and KYSE520 cells after overexpression or inhibition of miR-1303. (e) OD value in HET-1A cells and KYSE520 cells after overexpression or inhibition of miR-1303. (f) Apoptosis in HET-1A cells and KYSE520 cells after overexpression or inhibition of miR-1303. *
Effects of miR-1303 on invasion and metastasis of EC cells
MMP2 and MMP9 are important factors involved in cancer progression, which may affect the invasion of cancer.10,11 The overexpression of miR-1303 significantly elevated MMP2 and MMP9 in TE-1 cells, and the inhibition of miR-1303 reduced MMP2 and MMP9 in TE-1 cells (Figure 3(a) and (b)). E-cadherin regulates the intensity of cell adhesion within tissues and is related to cancer progression.12,13 In TE-1 cells, the overexpression of miR-1303 significantly reduced E-cadherin, while the inhibition of miR-1303 significantly increased E-cadherin (Figure 3(c)). In the same way, in KYSE520 cells, the overexpression of miR-1303 significantly elevated MMP2 and MMP9, while the inhibition of miR-1303 weakened MMP2 and MMP9 (Figure 3(d) and (e)). The overexpression of miR-1303 significantly reduced E-cadherin in KYSE520 cells, and the inhibition of miR-1303 obviously elevated it (Figure 3(f)). The protein expression trends of MMP9, MMP2, and E-cadherin are consistent with those of messengerRNA (mRNA; Figure 3(g)).
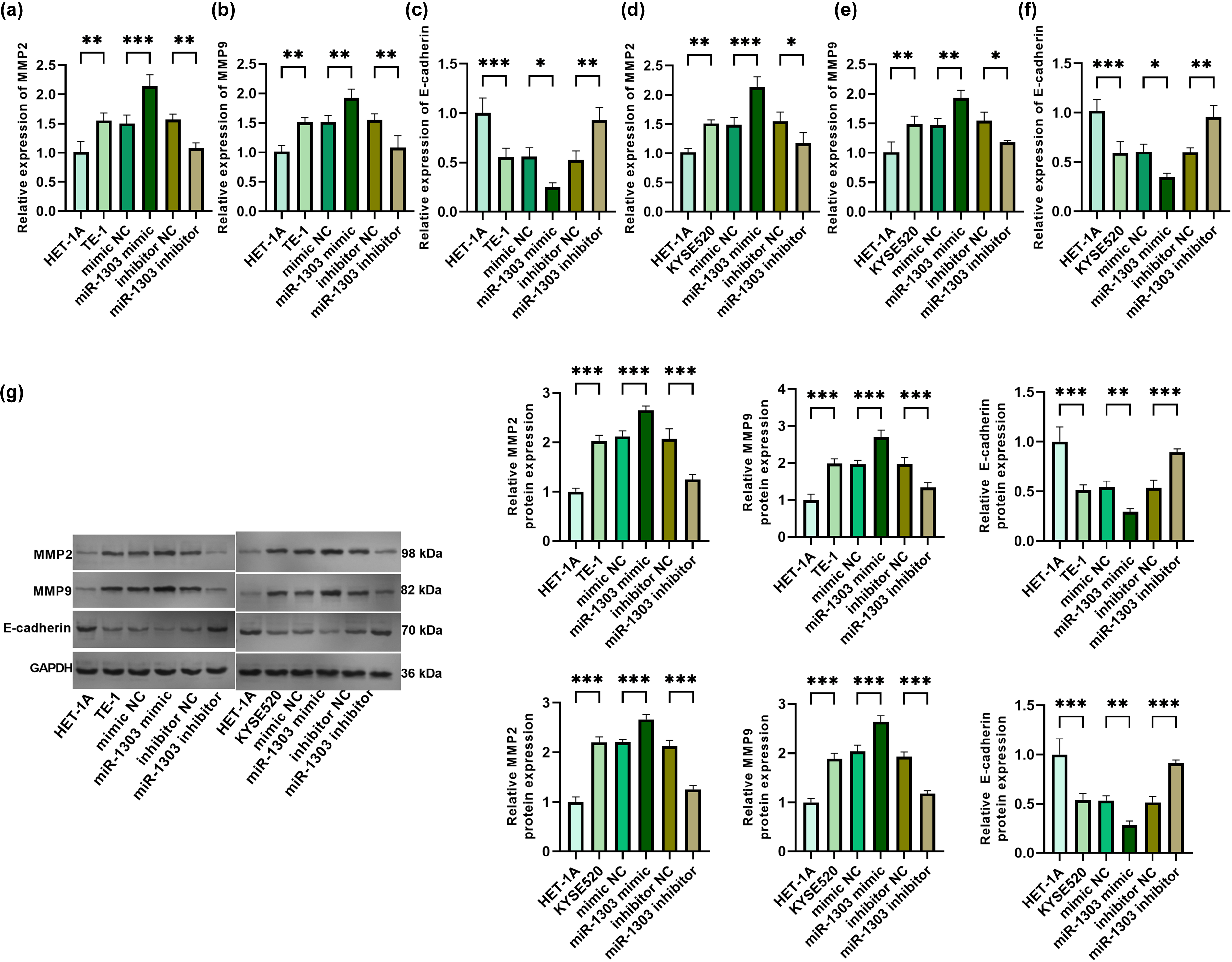
Effects of miR-1303 on invasion and metastasis of TE-1 cells and KYSE520 cells. (a) Relative expression of MMP2 in HET-1A cells and TE-1 cells after overexpression or inhibition of miR-1303. (b) Relative expression of MMP9 in HET-1A cells and TE-1 cells after overexpression or inhibition of miR-1303. (c) Relative expression of E-cadherin in HET-1A cells and TE-1 cells after overexpression or inhibition of miR-1303. (d) Relative expression of MMP2 in HET-1A cells and KYSE520 cells after overexpression or inhibition of miR-1303. (e) Relative expression of MMP9 in HET-1A cells and KYSE520 cells after overexpression or inhibition of miR-1303. (f) Relative expression of E-cadherin in HET-1A cells and KYSE520 cells after overexpression or inhibition of miR-1303. (g) Protein expression levels of E-cadherin, MMP9, and MMP2. *
The targeting relationship between miR-1303 and CLDN18
The targeting relationship between miR-1303 and CLDN18 was predicted, and the binding sites of them were obtained from TargetScan online database (Figure 4(a)). MiR-1303 obviously decreased the luciferase activity of the CLDN18 Wt reporter vector, whereas no such effect was observed with the CLDN18 Mut reporter vector (Figure 4(b)). CLDN18 was obviously reduced in EC tissues (Figure 4(c)). In EC tissues, CLDN18 was negatively correlated with miR-1303 (Figure 4(d)).
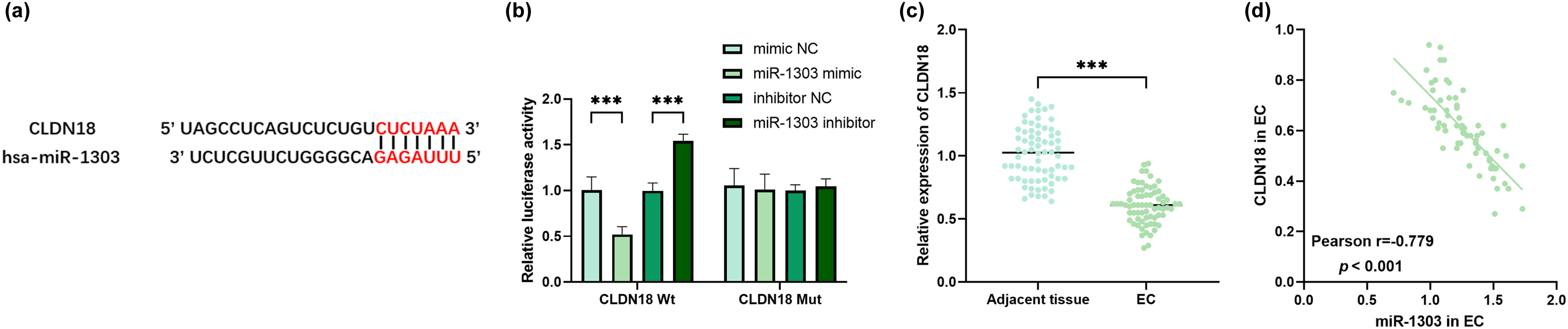
The targeting relationship between miR-1303 and CLDN18. (a) The binding sites between miR-1303 and CLDN18 from TargetScan online database. (b) The targeting relationship between miR-1303 and CLDN18. (c) Relative expression of CLDN18 in EC tissues. (d) The correlation of miR-1303 and CLDN18 in EC. ***
Co-effects of miR-1303 and CLDN18 in EC
MiR-1303 mimic and the pcDNA3.1-CLDN18 vector were co-transfected into TE-1 cells and KYSE520 cells, and successful transfection was confirmed (Figure 5(a) and (d)). The level of CLDN18 protein shows a trend consistent with that of mRNA (Figure 5(g)). In TE-1 cells, the up-regulation of CLDN18 obviously weakened the proliferation and increased the apoptosis (Figure 5(b) and (c)). And in KYSE520 cells, the up-regulation of CLDN18 also decreased the proliferation and increased the apoptosis (Figure 5(e) and (f)). In addition, in TE-1 cells, the overexpression of CLDN18 significantly reduced MMP2 and MMP9, and elevated E-cadherin (Supplementary Figure 1(a) to (c). Similarly, in KYSE520 cells, the overexpression of CLDN18 also reduced MMP2 and MMP9, and elevated E-cadherin (Supplementary Figure 1(d) to (f)).
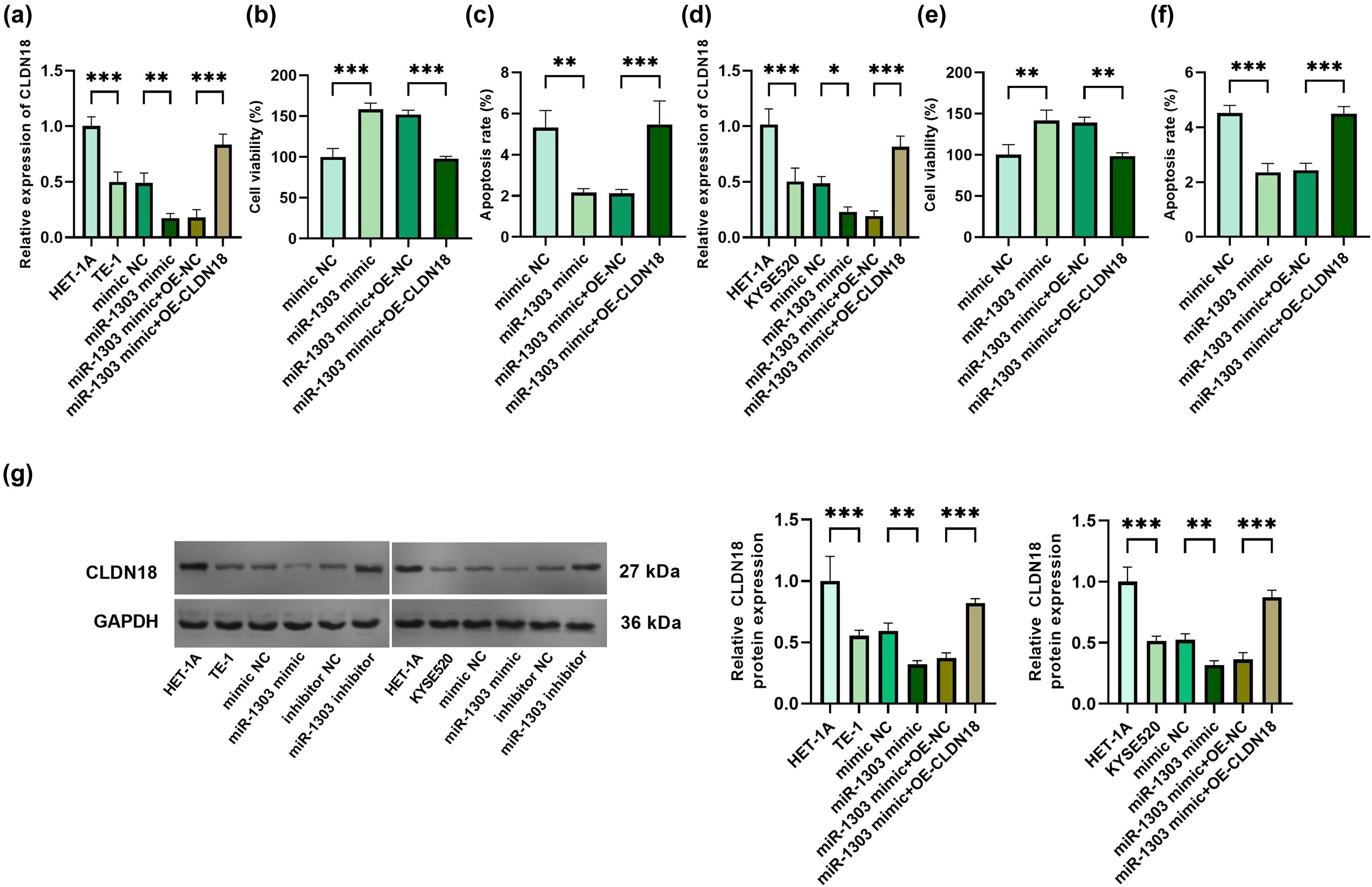
Verification of the targeting relationship between miR-1303 and CLDN18 in TE-1 cells and KYSE520 cells. (a) Relative expression of CLDN18 in HET-1A cells and TE-1 cells after overexpression of miR-1303 and CLDN18. (b) OD value in HET-1A cells and TE-1 cells after overexpression of miR-1303 and CLDN18. (c) Apoptosis in HET-1A cells and TE-1 cells after overexpression of miR-1303 and CLDN18. (d) Relative expression of CLDN18 in HET-1A cells and KYSE520 cells after overexpression of miR-1303 and CLDN18. (e) OD value in HET-1A cells and KYSE520 cells after overexpression of miR-1303 and CLDN18. (f) Apoptosis in HET-1A cells and KYSE520 cells after overexpression of miR-1303 and CLDN18. (g) Protein expression levels of CLDN18. *
Discussion
EC is an aggressive gastrointestinal cancer. 14 Most patients with EC have already shown advanced disease at the time of diagnosis, and the 5-year survival rate is merely 10–15%. 15 The main reasons for the low survival rate of EC are the late diagnosis and the early metastasis and spread of cancer cells. 16 EC imposes a serious social burden and has a negative impact on economic development. Pathological diagnosis is the “gold standard” for confirming tumors, and the comparison between tumor tissue and adjacent normal tissue is a key link in pathological diagnosis. With the development of precision medicine, molecular detection of paired tissues has become an important part of tumor diagnosis and treatment decisions. The core is to screen tumor-specific molecular targets and biomarkers. The diagnostic value of miR-1303 may provide new tumor-specific expression markers for clinical diagnosis at the molecular level.
At present, there are not many studies on miR-1303, and most of them are related to cancer. MiR-1303 significantly increased in adenomas and early-stage cancerous adenomas. 17 In ESCC, miR-1303 was obviously increased, and patients with a low expression level of miR-1303 had a lower recurrence rate. 18 Therefore, it was conjectured that the expression of miR-1303 increased in EC. Our results confirmed that miR-1303 increased in TE-1 cells, KYSE520 cells, and EC tissues, which was consistent with the previous research results.
Some evidence has confirmed that miR-1303 is closely connected with cell proliferation and apoptosis. In non-small cell lung cancer, the inhibition of miR-1303 expression inhibited the proliferation and invasion of cancer. 19 In neuroblastoma (NB), miR-1303 increases and can promote the proliferation of NB. 20 In acute myeloid leukemia, a clonal tumor disease, the increased expression of miR-1303 significantly reduces cell apoptosis. 21 The above evidence suggests that miR-1303 may promote proliferation and inhibit apoptosis in EC. In this study, in TE-1 cells and KYSE520 cells, the overexpression of miR-1303 obviously elevated proliferation and weakened apoptosis, while the inhibition of miR-1303 weakened proliferation and increased apoptosis, which verified the previous conjecture.
MMP, also known as matrix metalloproteinase, is the main regulatory factor of tissue remodeling and is closely related to tumor invasion. 22 Among them, MMP2 and MMP9 belong to the key members of the MMP family. 23 In basal cell carcinoma (BCC), the mRNA expression of MMP2 in invasive BCC is significantly higher than that in nodular BCC. 24 In thyroid cancer, MMP9 is usually high in metastatic tumor tissues and other samples with a high degree of malignancy. 25 These results indicate that MMP2 and MMP9 are connected with the invasiveness of cancer cells. Our results showed that in TE-1 and KYSE520 cells, the overexpression of miR-1303 significantly elevated the MMP2 and MMP9, while the inhibition of miR-1303 decreased MMP2 and MMP9, indicating that miR-1303 increased the invasiveness of EC.
Metastasis is the main driver of cancer death. 26 E-cadherin is an epithelial cell adhesion molecule and a key molecule for adhesion and connection between epithelial cells. 27 The loss of intercellular adhesion mediated by E-cadherin is a prerequisite for the formation of tumor cell invasion and metastasis. 28 Studies have shown that the absence of E-cadherin promotes the initial invasive behavior of certain epithelial-derived cancers. 29 In varieties of cancers, such as breast cancer, 30 PCa, 31 and hepatocellular carcinoma, 32 E-cadherin is associated with the metastasis of cancer cells. Our results confirmed that in TE-1 cells and KYSE520 cells, overexpressed miR-1303 significantly reduced the expression of E-cadherin, while inhibited miR-1303 significantly increased its expression. Therefore, it is speculated that miR-1303 will promote the metastasis of EC cells.
Studies have shown that different subtypes of CLDN are expressed in different human organs, and it has been determined that changes in their functions may cause tumor carcinogenesis in the corresponding tissues.
33
Among them, the subtype of CLDN18—CLDN18.2—is abnormally expressed in gastrointestinal cancers, including EAC.
34
In this study, in EC tissues, CLDN18 was significantly reduced and negatively correlated with miR-1303, and the targeting relationship between the two was also verified in the cell experiments. The ENCORI database shows that CLDN18 is significantly downregulated in EC. The data are derived from 162 EC patients and 11 normal controls, which is consistent with the trend of the results of this study (Supplementary Figure 2,
This study still has some limitations. The control tissue defined as “normal” is the adjacent tissue rather than the truly healthy esophageal tissue. The adjacent tissue may already have genetic or epigenetic alterations. Therefore, the area under the curve value of miR-1303 obtained in this study may be overestimated. However, this indicator still has significant statistical significance, showing that miR-1303 is a robust EC-related biomarker. In addition, the limited sample size and the absence of in vivo models also impose restrictions on the research results. In the future, its true potential for early screening needs to be further confirmed in larger-scale prospective studies that include healthy population cohorts.
All in all, miR-1303 was highly expressed in EC and had a high diagnostic value for EC. MiR-1303 promoted the proliferation, inhibited the apoptosis, and promoted the proliferation and invasion of EC cells. In EC, CLDN18 is downstream of miR-1303 and negatively regulated by miR-1303.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155261416110 - Supplemental material for Inhibition of miR-1303 suppresses the development of esophageal cancer by targeting CLDN18
Supplemental material, sj-docx-1-jbm-10.1177_03936155261416110 for Inhibition of miR-1303 suppresses the development of esophageal cancer by targeting CLDN18 by Xiaoming Shi, Chongliu Lin, Zirui Zhang, Ning Zhou, Zichen Jiao and Tao Wang in The International Journal of Biological Markers
Footnotes
Funding
This work was supported by Nanjing Drum Tower Hospital Clinical Research Special Fund Project 2024 (Grant numbers: 2024-LCYJ-PY-47).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplementary material
Supplemental material for this article is available online.
