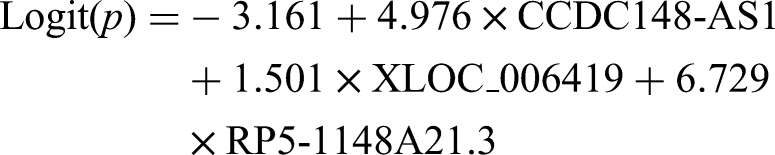Abstract
Purpose
We aimed to exploit a urine exosomal long non-coding RNAs (lncRNAs) fingerprint to facilitate the early diagnosis of bladder cancer.
Methods
Microarray differential expression profiling of lncRNAs was for the first time employed in urine exosomes from 10 non-muscle-invasive bladder cancer (NMIBC) patients and 10 healthy controls to screen out candidate exosomal lncRNA biomarkers, which were then verified by quantitative real-time polymerase chain reaction in three independent phases including bladder cancer cells, culture fluid and 200 NMIBC participants. Logistic regression was performed to construct a diagnostic model—the diagnostic potency of which was assessed.
Results
The profile of three exosome-derived lncRNAs (CCDC148-AS1, XLOC_006419, and RP5-1148A21.3) was screened and further verified to be notably over-expressed in NMIBC patients and bladder cancer cell lines, and exhibited area under the receiver-operating characteristic curve values of 0.873, 0.825, and 0.834, respectively, in training, validation, and double-blind validation phases. The profile was superior to urinary cytology in discriminating NMIBC from healthy controls (P < 0.0001). A significant correlation existed between a higher level of CCDC148-AS1 and a higher tumor grade (P < 0.001), and up-regulated CCDC148-AS1 as well as XLOC_006419 were statistically related with tumor node metastasis stage (P = 0.004 and P = 0.031, respectively). These three identified lncRNAs were confirmed to originate from bladder cancer cells and be packaged within exosomes, thus staying sufficiently stable in urine.
Conclusions
Tumor-originated urine exosomal lncRNAs, as fingerprint in NMIBC, exhibited satisfying clinical significance in early diagnosis of bladder cancer.
Keywords
Introduction
Globally, bladder cancer (BC) is responsible for an estimated 573,000 new cases and 213,000 deaths, ranking in 10th place for incidence, and in 2nd place in genitourinary tract malignancies regarding mortality. 1 Roughly three-quarters of patients have non-muscle invasive BC (NMIBC), about 50% of which are low grade; one-quarter have muscle-invasive BC or metastatic disease, most of which are high grade with poor prognosis. 2 Currently, cystoscopy and urine cytology as the major diagnostic and monitoring tools for BC have been proven to be far from clinically satisfactory. The cystoscopic approach is invasive, uncomfortable, costly, and hard to detect carcinoma in situ. 3 Urine cytology had low sensitivity (especially for low-grade tumors) and poor interobserver and intraobserver reproducibility. 2 Since there has been no improvement in survival rates for BC for three decades, 4 there is an imperative need to explore worthy non-invasive biomarkers with high accuracy for early diagnosis of BC.
Long non-coding RNAs (lncRNAs), transcripts of greater than 200 nucleotides without evident protein-coding function, could regulate gene expression at the levels of transcription, RNA processing, and translation.5,6 It is reported that many lncRNAs are dysregulated in human cancers in a tissue-specific fashion, which could therefore serve as biomarkers for tracing circulating cancer cells or cancer metastases to their origins. 7
Exosomes are small extracellular vesicles (<150 nm in diameter) that can shuttle bioactive molecules between cancer cells and the surrounding cells in the tumor microenvironment, inducing the exchange of genetic message as well as the reprogramming of the recipient cells.8,9 Increasing evidence suggests that exosomes derived from tumor cells may influence all stages of cancer progression including tumor initiation, growth, progression, metastasis, and therapy resistance. 10 lncRNAs may be selectively packaged into exosomes by a specific RNA sorting mechanism, and thus be well-protected and not degraded in body fluids. 11 Exosome-derived lncRNAs accomplish various remarkable biological functions such as regulating tumor apoptosis, proliferation, and migration, as well as inducing angiogenesis, and are expected to become diagnostic and prognostic biomarkers of different cancers.6,12 Nevertheless, research on circulating exosome-derived lncRNAs as biomarkers for diagnosing BC is not nearly enough.
For the sake of exploring novel circulating exosomal lncRNA biomarkers for BC, our study was devised as follows.
1. Candidate exosomal lncRNAs overexpressed in urine of NMIBC was first screened out by genome-wide lncRNA profiling analysis and then validated by quantitative real-time polymerase chain reaction (qRT-PCR).
2. Furthermore, a mathematic model based on the lncRNA profile was set up—its diagnostic potential for NMIBC was assessed and compared with urine cytology.
3. In addition, expression levels of candidate lncRNAs were also determined in cell lines, tumor tissues, and exosomes from paired pre-and post-surgery urine.
Materials and methods
Patients, clinical samples, and study design
The study was performed according to the principles of the Declaration of Helsinki and authorized by the Ethics Committee of Linyi People's Hospital. All participants offered written informed consent and were divided into three independent classifications as follows: NMIBC patients were histopathologically diagnosed, staged, and graded on the basis of the 2009 Union for International Cancer Control TNM system 13 and the 2004 WHO classification system. 14 Participants with benign bladder disease (BBD) consist of those calling for cystoscopy for symptoms implicating BC (e.g., hematuria), such as those with bladder calculi, bladder infection, or benign neoplasm of bladder, etc. Healthy controls were enrolled from people who underwent routine physical examination and had results in the normal range. There was no significant difference in age and gender ratios among the three classifications of participants. Urine samples of NMIBC patients were collected 2 days before tumor resection and 7 days after surgery. None of these patients had received radiotherapy or chemotherapy before the first and second urine sampling. Tissues samples were collected and processed as we reported previously. 15
A multi-stage, case-control study was framed to recognize a panel of urinary exosomal lncRNA biomarkers for NMIBC. In the pilot phase, Human LncRNA Array v4.0 (Arraystar, Agilent, Santa Clara, CA, USA) was exploited for 20 urine samples (10 preoperative urine from NMIBC patients and 10 from healthy controls) to identify the candidate exosomal lncRNAs upregulated in NMIBC cases. In the training phase, candidate lncRNAs were further detected in a small cohort of urine exosome samples (50 paired pre- and post-operative urine from NMIBC, 50 from BBD, 50 from healthy controls), tissue from NMIBC (n = 50), and culture medium of BC cells by qRT-PCR. The upregulated exosomal lncRNAs were finally ascertained; a diagnostic mathematical model based on the lncRNA panel was constructed and evaluated; and the origin of urinary exosomal lncRNAs was investigated. In the validation phase, diagnostic efficacy of the urinary exosomal lncRNA-based panel was further accessed in another larger cohort of 100 NMIBC patients, 100 BBD patients, and 100 healthy controls. Finally, in the double-blind validation phase, diagnostic performance of the exosomal lncRNA panel and urine cytology were verified in another set of 100 randomly selected urine samples with the double blind method.
Additional methodological information can be found online in the Supplemental materials and methods section.
Statistical analysis
SPSS 16.0 (IBM, Armonk, NY, USA), MedCalc 15.2 (MedCalc Software, Ostend, Belgium), GraphPad Prism 5.0 (GraphPad Software, San Diego, CA, USA), and Microsoft excel were employed to perform all statistical analyses and draw diagrams with P < 0.05 (two-tailed test) indicating significant difference. The Kolmogorov–Smirnov test was carried out for normality test of data. The non-parametric Mann–Whitney U test, the Kruskal–Wallis test, or the Student's t-test were applied to determine differential expression of exosomal lncRNAs between independent groups, as the case might be. Spearman's rank correlation analysis and the Wilcoxon test were exploited for paired tissue versus urine and pre- versus post-operative urine, respectively. Multivariate binary logistic regression was performed to set up a mathematical model of the relationship between identified exosomal lncRNAs and BC. The validity of the mathematical model and diagnostic potency of the identified exosomal lncRNAs individually or in combination were appraised by receiver operating characteristic (ROC) analysis as well as the area under the ROC curve (AUC), with the optimal cutoff threshold being reckoned by the Youden index. 16 Comparison of the AUC from different diagnostic markers was performed via the Z test. The diagnostic model established in the training stage was directly applied to the following validation stages.
Results
Distinguishing features of exosomes extracted from urine
We verified that exosomes could be successfully harvested from urine with relatively high reproducibility in view of the following evidence. Transmission electron microscopy (TEM) revealed that exosomes assumed intact flattened spheres with a lipid bilayer, ranging from 30 to 150 nm in diameter (Figure 1(a)). Western blotting proved the presence of exosomal marker proteins (CD63 and Hsp70) in exosomes rather than in exosome-depleted supernatant (EDS) (Figure 1(b)).
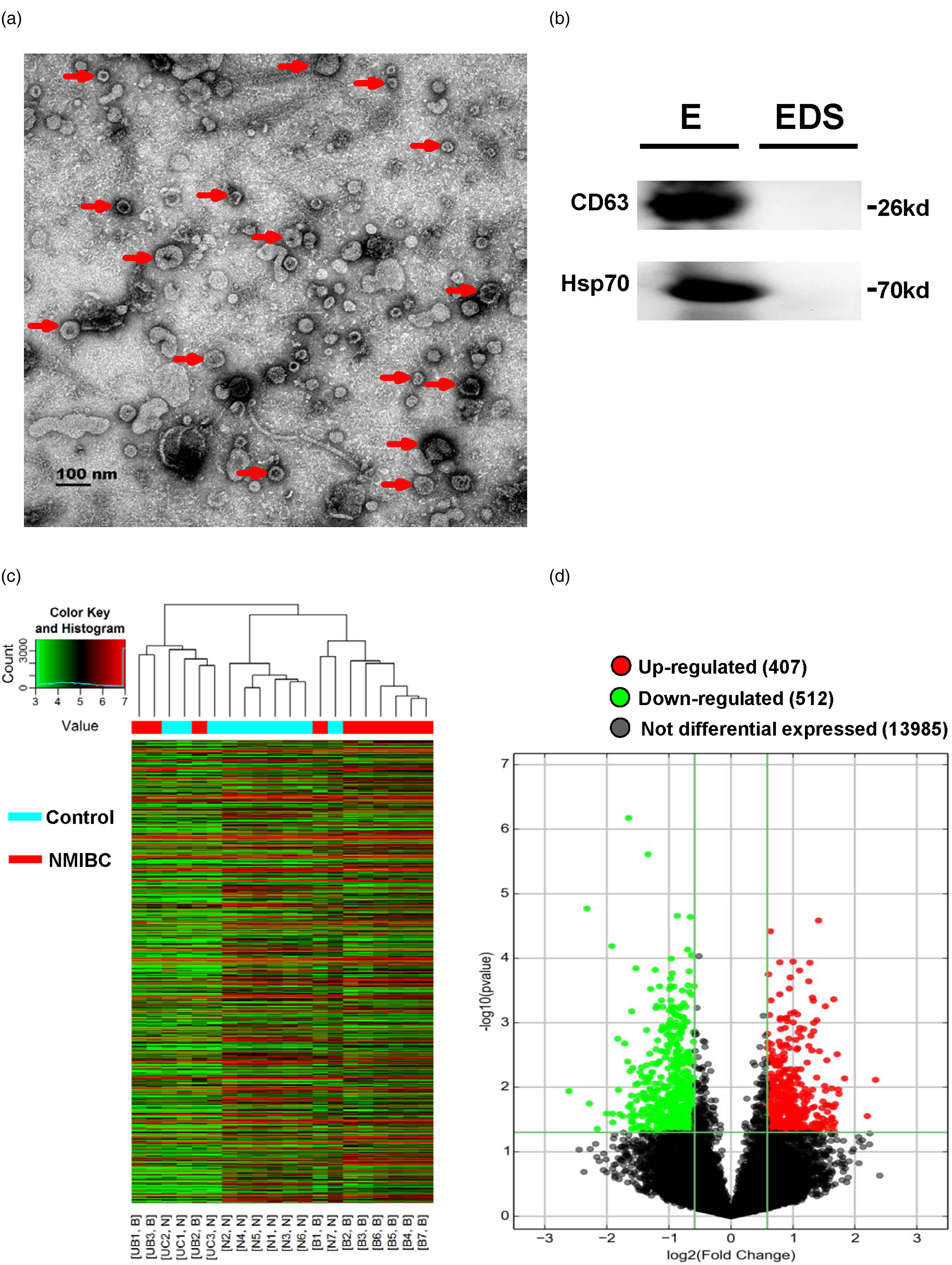
Aberrantly expressed, NMIBC-specific urine exosomal lncRNAs were discerned by microarray profiling. (a) Classic images of urine exosomes observed under TEM. Red arrows marked the representative exosome, and the scale bar was 100 nm. (b) Western blot detection of CD63 and Hsp70 in exosomes (E) and EDS. (c) Hierarchical cluster analysis of differentially expressed urine exosomal lncRNAs between 10 NMIBC patients and 10 healthy controls. (d) Volcano plot displayed the specifically dysregulated, NMIBC-specific urine exosomal lncRNAs.
Genome-wide microarray-based screening for NMIBC-associated urine exosome-derived lncRNAs
Exosomal RNA extracted from urine of preoperative NMIBC patients (n = 10) and healthy controls (n = 10) was analyzed via genome-wide microarray to identify NMIBC-specific urine exosome-derived lncRNAs. Specifically dysregulated exosomal lncRNAs between NMIBC patients and healthy controls were sorted by hierarchical cluster analysis as demonstrated in a heatmap (Figure 1(c)). Figure 1(d) displayed that 407 exosomal lncRNAs were found to be up-regulated in NMIBC patients compared with controls (P < 0.05 as well as fold change ≥1.5). Further screening of the 407 aberrantly expressed lncRNAs on the condition of fold change ≥3.0 eventually yielded 10 lncRNA candidates on which we would focus in the next part of this study.
Identification of three significantly up-regulated exosomal lncRNA, establishment of diagnostic model for NMIBC, and its performance assessment by training phase and validation phase
We further assayed these 10 dysregulated exosomal lncRNAs by qRT-PCR in a training specimen set (50 BC patients, 50 BBD patients, and 50 healthy controls). As a result, this phase engendered a panel of three exosomal lncRNAs (CCDC148-AS1, XLOC_006419, and RP5-1148A21.3), which were significantly over-expressed in the NMIBC group compared with the BBD group and with controls as demonstrated in Figure 2(a) to (c). Then, in a larger validation specimen set (100 NMIBC patients, 100 BBD patients, and 100 healthy controls), we verified that the urine expression levels of these three exosomal lncRNAs in NMIBC patients were significantly higher than those in BBD patients and healthy controls, which was concordant with the results in the training set (Figure 2(d) to (f)).
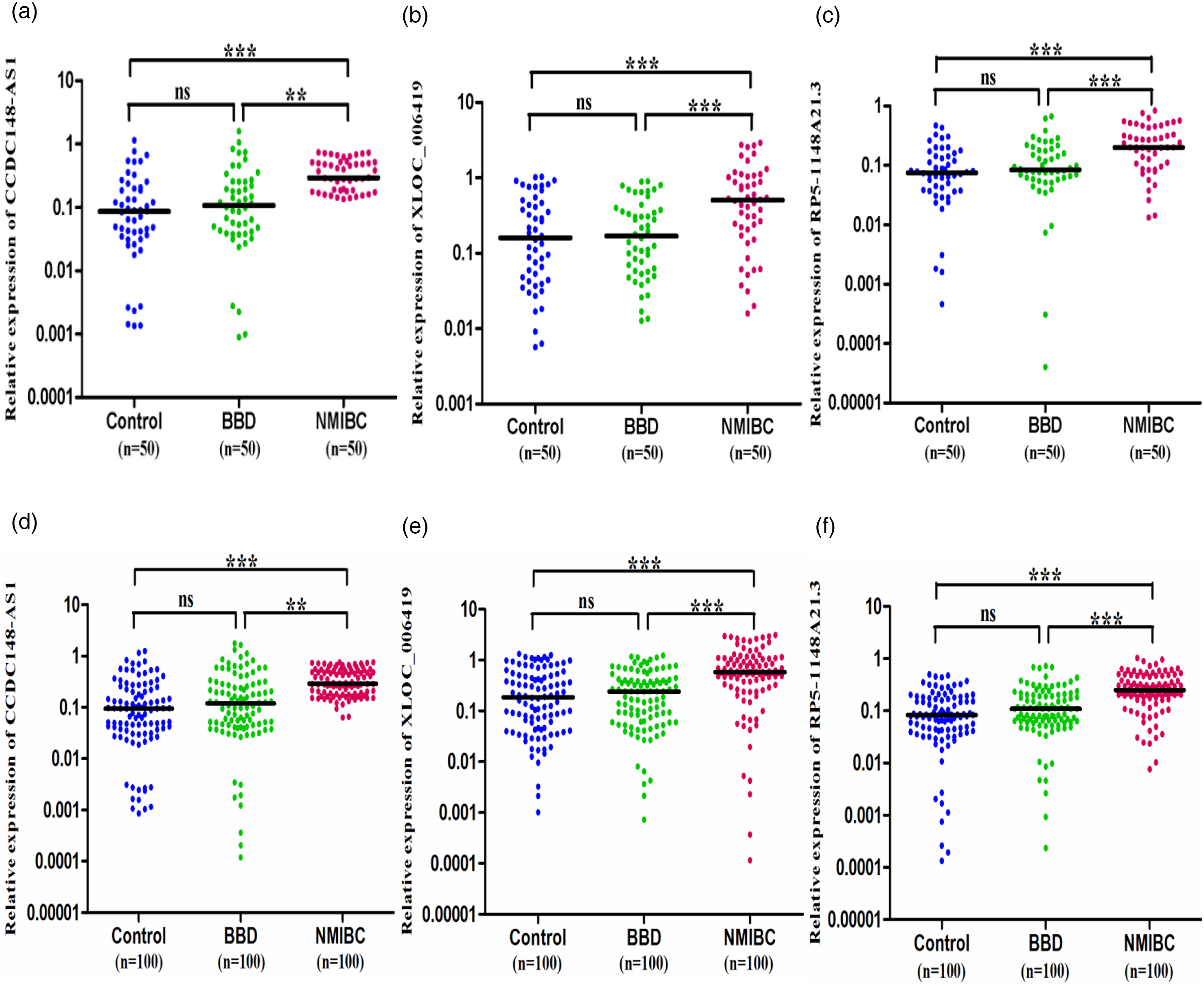
Quantitative testing of the three exosomal lncRNAs in NMIBC in training and validation phases. (a) to (c) Urine expression levels of CCDC148-AS1, XLOC_006419, and RP5-1148A21.3 in patients with NMIBC, patients with BBD, and healthy controls in the training phase. The horizontal lines indicate median values. **P < 0.01, ***P < 0.001. (d) to (f) Urine expression levels of CCDC148-AS1, XLOC_006419, and RP5-1148A21.3 in patients with NMIBC, patients with BBD, and healthy controls in validation phase. The horizontal lines indicate median values. **P < 0.01, ***P < 0.001.
In the training phase, we constructed the three-lncRNA panel-based mathematical model for identifying NMIBC by utilizing multivariate logistic regression to ascertain the weighting of each exosomal lncRNA (Supplemental Table 1). Probabilistic forecasting of developing BC for a patient could be implemented by the formula below:
Then, a ROC curve analysis was carried out to estimate the capability of recognized single lncRNA and the three-lncRNA panel for NMIBC discernment from healthy controls. As displayed in Figure 3(a), the diagnostic efficacy (denoted as AUC) of CCDC148-AS1, XLOC_006419 and RP5-1148A21.3, and the three-lncRNA signature was 0.830 (95% CI = 0.743–0.918), 0.706 (95% CI = 0.606–0.807), 0.772 (95% CI = 0.679–0.864), and 0.873 (95% CI = 0.804–0.942), respectively, in the training phase. The diagnostic efficacy was 0.808 (95% CI = 0.744–0.872), 0.723 (95% CI = 0.652–0.794), 0.801 (95% CI = 0.738–0.863), and 0.825 (95% CI = 0.766–0.883), respectively, in the validation phase (Figure 3(b)). At a cutoff value of 0.403, the sensitivity and specificity of the three-lncRNA signature were 92.00% and 74.00%, respectively, in the training phase; and were 93.00% and 70.00%, respectively, in the validation phase. Similar to the results above, in the validation set, this three-lncRNA panel exhibited a relatively high AUC of 0.800 (95% CI = 0.757–0.912) in distinguishing 100 NMIBC cases from 200 cancer-free controls with a sensitivity of 93.00% and a specificity of 67.00% (Figure 3(c)). Taken together, we proved that this three-lncRNA signature qualified as a powerful diagnostic biomarker for distinguishing NMIBC.
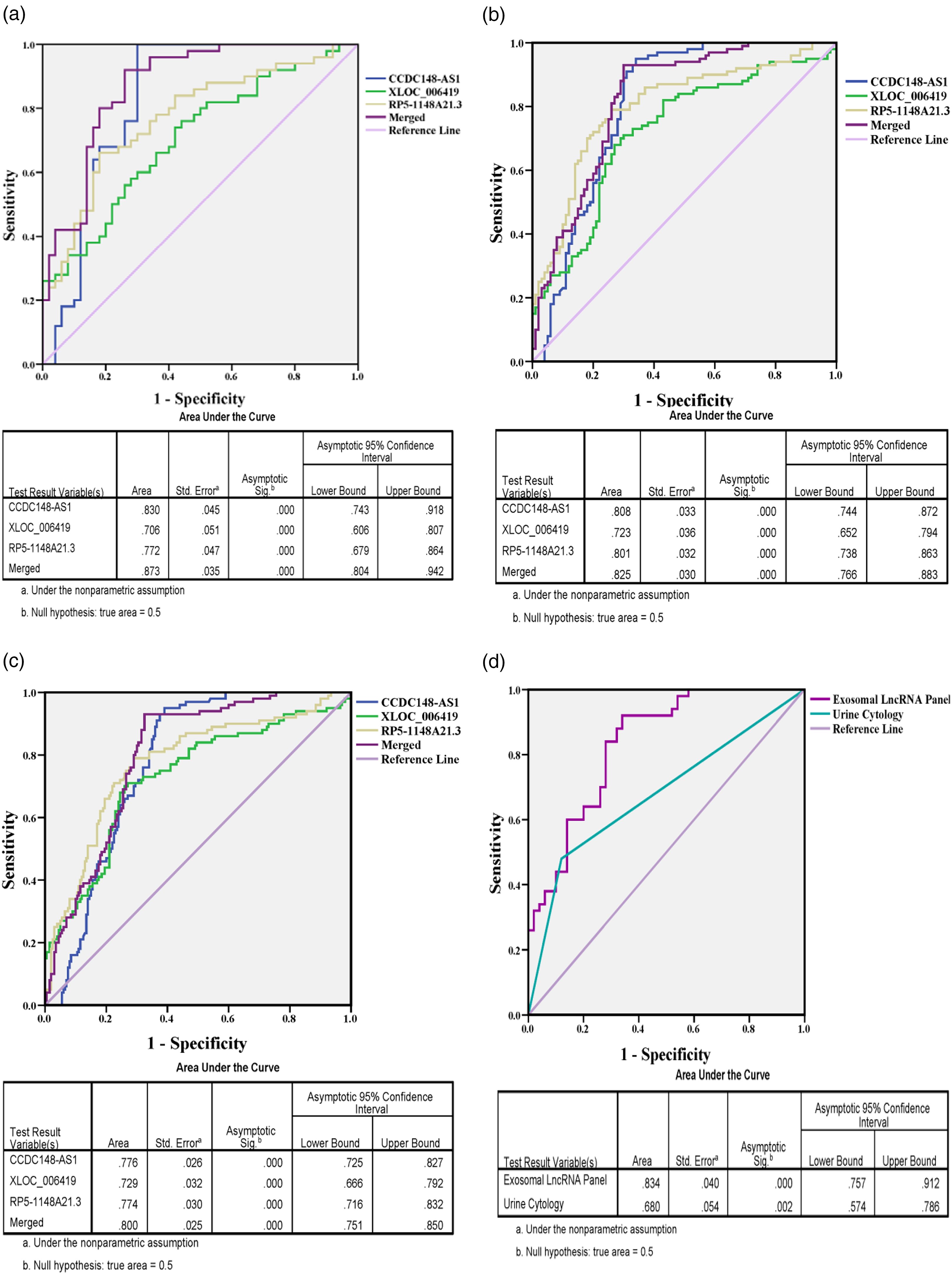
Diagnostic accuracy of the three exosomal lncRNAs in NMIBC in the training, validation, and double-blind phases. (a) ROC curves for CCDC148-AS1, XLOC_006419, RP5-1148A21.3, and the three-lncRNA signature to distinguish 50 NMIBC cases from 50 healthy controls in the validation phase with the AUC shown below. (b) ROC curves for CCDC148-AS1, XLOC_006419, RP5-1148A21.3, and the three-lncRNA signature to distinguish 100 NMIBC cases from 100 healthy controls in the validation phase with the AUC shown below. (c) ROC curves for CCDC148-AS1, XLOC_006419, RP5-1148A21.3, and the three-lncRNA signature to distinguish 100 NMIBC cases from 200 cancer-free controls (including 100 BBD patients and 100 healthy controls) in the validation phase with the AUC shown below. (d) ROC curves for the three-lncRNA signature and urine cytology to distinguish NMIBC cases from healthy controls in a double-blind phase with the AUC shown below.
Double-blind trial for verifying the diagnostic efficacy of the exosomal lncRNA panel
The expression levels of these three exosomal lncRNAs were measured in urine specimens from 100 participants with or without NMIBC in a double-blind mode and the previously built logistic regression formula based on the three-lncRNA panel was applied to classify these participants. Surprisingly, NMIBC was explicitly identified from participants and the three-lncRNA panel as a NMIBC biomarker exhibited a precision rate of 78.00%, a sensitivity of 90.00%, a specificity of 66.00%, and an AUC of 0.834 (95% CI = 0.757–0.912) (Figure 3(d)). Since urine cytology is one of the main diagnostic means for BC, we assessed its diagnostic accuracy and made a comparison between it and this panel in a double-blind set. Our results revealed that diagnostic accuracy of this three-lncRNA panel was superior to that of urine cytology (AUC = 0.620) (95% CI = 0.510–0.730) (Z = 6.668, P < 0.0001) (Figure 3(d)).
Clinicopathological relevance analysis in NMIBC patients
The statistics suggested that concentrations of CCDC148-AS1 were found to be significantly associated with tumor grade and tumor node metastasis (TNM) stage (P < 0.001 and P = 0.004, respectively). Moreover, our data showed that over-expressed XLOC_006419 was statistically related to higher TNM stage (P = 0.031) (Supplemental Table 2).
General characterization of the lncRNA panel in urine exosomes
We confirmed that these three discerned lncRNAs in urine primarily existed in exosomes, since their expression levels in exosomes were observed to be significantly higher than those in EDS (Figure 4(a)).
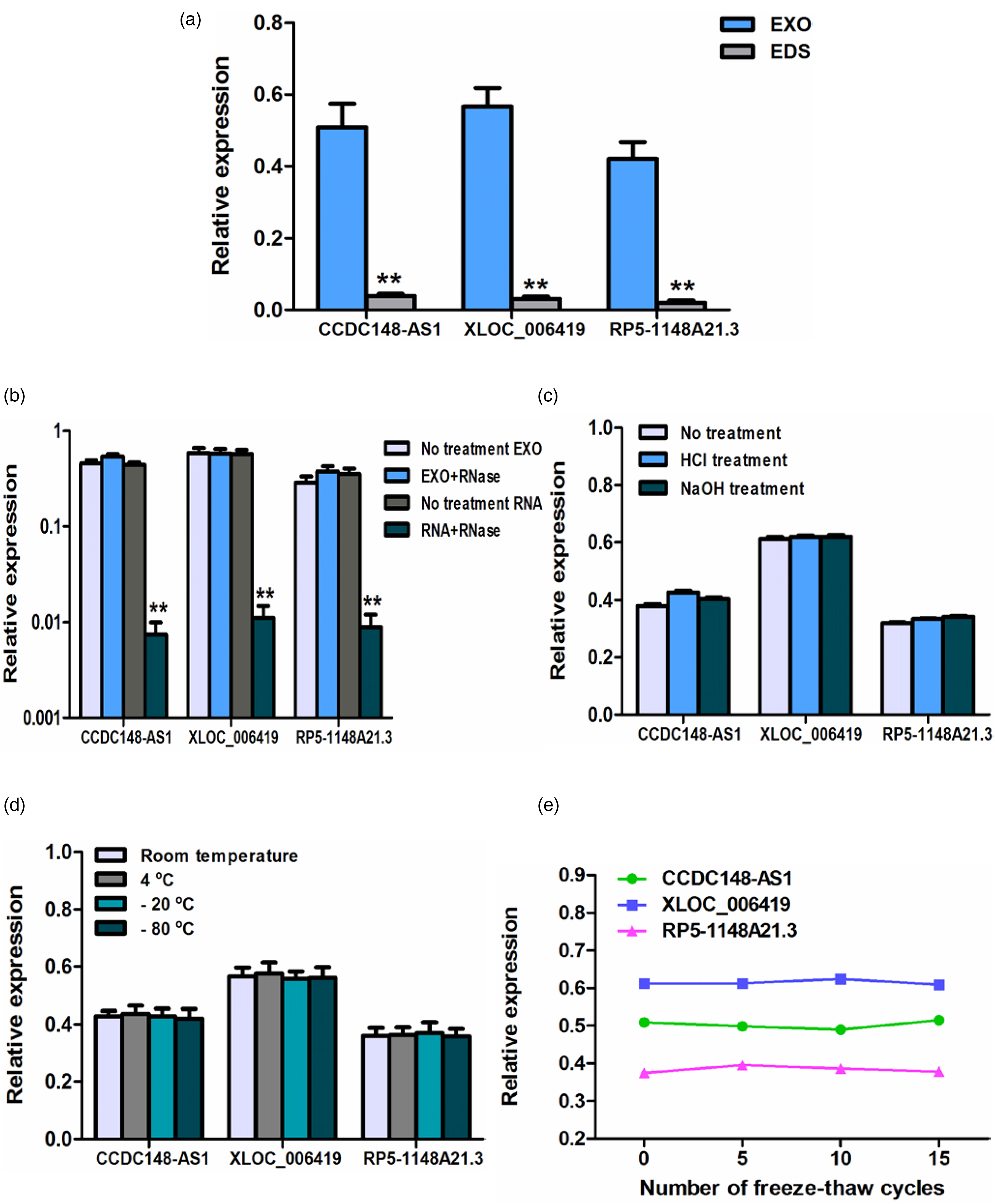
Basic characteristics of urine exosomal CCDC148-AS1, XLOC_006419, and RP5-1148A21.3. (a) Expression levels of CCDC148-AS1, XLOC_006419, and RP5-1148A21.3 detected in EDS and urine exosome (EXO). **P < 0.01. (b) Expression levels of CCDC148-AS1, XLOC_006419, and RP5-1148A21.3 in urine exosome and exosome-isolated RNA digested by RNase A . **P < 0.01. Level changes of exosomal CCDC148-AS1, XLOC_006419 and RP5-1148A21.3 in urine samples subjected to (c) acid solution (pH = 1)/base solution (pH = 13) treatment, (d) incubation at different temperature (−80 °C, −20 °C, 4 °C, and room temperature), and (e) repeated freeze–thaw cycles.
To attest whether the exosome membrane could effectively defend these exosomal lncRNAs in urine against erosion, urine samples were subjected to severe conditions including RNase A digestion for 3 h, acid/base treatment for 3 h, incubation at different temperatures (−80 °C, −20 °C, 4 °C, and room temperature) for 48 h, and multiple freeze–thaw cycles (5, 10, and 15 cycles). As illustrated in Figure 4(b), RNase A exerted no effect on levels of these lncRNAs in urine exosomes, while it almost completely digested these lncRNAs in exosome-isolated RNA. Additionally, other treatments imposed minimal influence on the concentrations of the three exosomal lncRNAs (Figure 4(c) to (e)). In brief, our data verified that these exosomal lncRNAs stayed sufficiently stable in urine, making them qualified biomarkers for NMIBC diagnosis.
Inquiry into the origin of the urine exosomal lncRNAs
We also unraveled that these three urine exosomal lncRNAs indeed originated from BC cells through the following three experiments.
1. In this first experiment, expression levels of these three exosomal lncRNAs in the culture solution of BC cells significantly rose in a cell count- and time-dependent mode, while their expression levels scarcely varied in a culture solution of SV-HUC-1, implying they were discharged by tumor cells into circulation (Figure 5(a) and (b)).
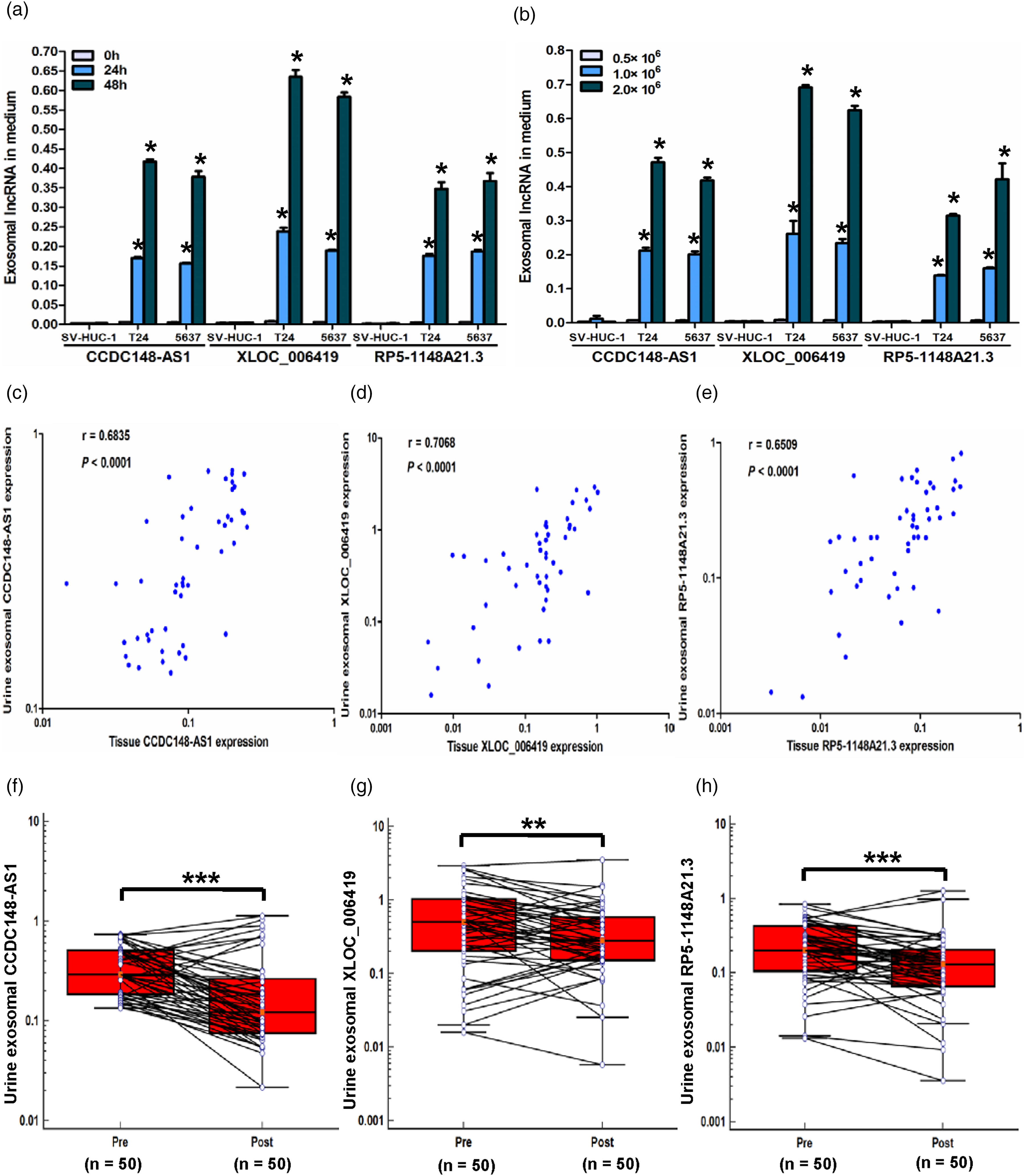
Urine exosomal lncRNAs originated from NMIBC tumor. (a) and (b) Expression levels of exosomal CCDC148-AS1, XLOC_006419 and RP5-1148A21.3 in the culture solution of T24 and 5637 cells elevated with longer incubation periods and increased cell count. *P < 0.05. (c) to (e) Spearman's correlation analysis between exosomal levels of CCDC148-AS1, XLOC_006419, and RP5-1148A21.3 in urine and paired tumor tissues of NMIBC. The x-axis and y-axis both use log10 scale. (f) to (h) Shifts in urine levels of exosomal CCDC148-AS1, XLOC_006419 and RP5-1148A21.3 before surgery (Pre) and 7 days after tumor resection (Post) in NMIBC patients. **P < 0.01, ***P < 0.001.
2. In the second experiment, a statistically positive correlation was evident between levels of these lncRNAs in urine exosomes and in paired tumor tissues (Figure 5(c) to (e)).
3. In the third experiment, expression levels of these three exosomal lncRNAs dramatically dropped in postoperative urine in comparison with matched preoperative urine (Figure 5(f) to (h)).
Discussion
In the current research, we discerned a NMIBC-specific over-expressed signature consisting of three urine exosomal lncRNAs, CCDC148-AS1, XLOC_006419, and RP5-1148A21.3, based on which the diagnostic model was constructed. Furthermore, we verified that this exosome-derived lncRNAs profile could serve as a worthy diagnostic biomarker that significantly outperformed urine cytology in diagnostic efficacy. Moreover, our results demonstrated that these exosomal lncRNAs could stably present in urine and their dysregulation seemed to be in a tissue-specific fashion. The novelties of our study lie in genome-wide lncRNA profiling performed for the first time in urine exosome samples followed by a multistage validation, a relatively large sample size, and the setting up of a urine exosomal lncRNA-based diagnostic model.
It has been gradually uncovered that exosomes are important tools for exchanging intercellular material such as lncRNAs. 11 Increasing evidence reveals that circulating exosomal lncRNAs could serve as minimally invasive and convenient diagnostic biomarkers and are emerging stars in tumor research. As for BC, there have been a few studies investigating the diagnostic value of circulating exosomal lncRNAs. A urinary exosome-derived lncRNA expression signature (MALAT1, PCAT-1, and SPRY4-IT1) was developed and could serve as a novel non-invasive biomarker for diagnosis of BC with an AUC of 0.854. 17 The following diagnostic panels based on urinary exosomal lncRNAs have also been set up: lnc-UCA1-201, UCA1-203, MALAT1, and LINC00355 (AUC = 0.960) 18 ; lnc-MIR205HG and GAS5 (AUC = 0.842) 19 ; and lnc-RMRP, UCA1, and MALAT1 (AUC = 0.875), 20 all manifesting satisfying diagnostic value for BC. Moreover, other urine exosome-derived lncRNAs such as ANRIL (AUC = 0.723), 21 PCAT-1 (AUC = 0.729), 21 ELNAT1 (AUC = 0.800), 22 TUG-1 (AUC = 0.780), 23 TERC (AUC = 0.836), 24 SNHG16 (AUC = 0.791), 25 and RMRP (AUC = 0.830) 26 have also been testified to exhibit their potential as non-invasive diagnostic biomarkers for BC. In terms of diagnostic efficacy (e.g., AUC values) alone, our urine exosomal lncRNA-based diagnostic model did not seem to perform better than other diagnostic biomarkers above. However, the lncRNAs explored in all of these reports were simply picked from previous documents, certainly resulting in omission of most of worthy lncRNAs. What set us apart from previous studies17–26 is that we mainly implemented the first NMIBC-specific genome-wide exosomal lncRNA profiling in urine samples that would identify biomarkers more comprehensively and accurately. Moreover, for the first time, we investigated the origin and tissue-specificity of urine exosome-derived lncRNAs in BC, which was a necessary condition for them to become qualified markers.
Our assay clarified the characteristics of these three exosomal lncRNAs, including their distribution, measurability, steadiness, and origin. First, exosomes extracted from urine in our study were verified to present fairly high pureness, abundance, and reproducibility by TEM and Western blot. Second, these urinary lncRNAs of BC patients were observed to mainly consist in exosomes instead of EDS and could be determined by qRT-PCR, revealing that they seem to be incorporated into exosomes by a particular RNA selecting mechanism. 11 Moreover, these exosomal lncRNAs were confirmed to be almost unaffected under extremely cruel circumstances (including RNase A digestion, strong acid, strong alkali, repeated freeze-thaw cycles, etc.), proving that exosomes could vigorously defend them from being spoiled. Furthermore, our data suggested that these exosomal lncRNAs were tumor-derived and their expression in urine could specifically represent loads of lncRNAs in tumor tissues for these reasons: (a) Levels of exosomal lncRNAs in culture liquid of BC cells instead of epithelial cells significantly rose along with incubation time and the initial cell counts; and (b) expression levels of the lncRNAs signature in urine exosomes from NMIBC patients before an operation were statistically positively relevant to that in paired tumor tissues and were found to decline significantly after tumor resection. Urine supernatant is commonly acknowledged to be an especially satisfying source of biomarkers for BC because of its convenience to obtain, minimal protein interference, and high exposure to exosomes from tumor tissue. 15 In addition, the expression levels of GAPDH and UBC in urine exosomes were validated to be fairly constant regardless of any bladder disease status; thus, they were exploited as internal references for normalization. All our findings, above, highlighted the applicability and vivid specificity of urine exosomal lncRNAs as forceful diagnostic biomarkers for NMIBC.
Our data also demonstrated that elevated exosomal CCDC148-AS1 and XLOC_006419 in urine were statistically related to clinicopathological indexes predicting an adverse outcome (e.g., higher tumor grade as well as stage), hinting that they may play a critical part in tumorgenicity and progression. Based on this up-regulated exosomal lncRNA panel, a logistic regression model was set up in the training phase and yielded forceful diagnostic efficacy with an AUC of 0.873. The diagnostic model was further evaluated in expanding the validation stage and exhibited relatively satisfying distinguishing power (AUC = 0.825). Similar results were also observed in the subsequent double-blind validation stage, and the diagnostic efficacy of the model (AUC = 0.834) overmatched that of urine cytology (AUC = 0.620) (P < 0.0001). Although our diagnostic model might cost slightly more money and time than urine cytology, the latter has poorer interobserver and intraobserver reproducibility 2 and requires professional pathologists with rich morphological experience. Further, in view of cost effectiveness and ease of use, our diagnostic model seems to be superior to cystoscopy, which is costly, invasive, and uncomfortable. 3 The above data verified the high diagnostic accuracy, cost effectiveness, and ease of use of the exosomal lncRNA panel as a reliable non-invasive biomarker for NMIBC detection, which would be helpful for clinics to adopt early intervention, thereby improving the survival of patients with BC.
Growing evidence has proven that exosomal lncRNAs are intercellular signal mediators. The biological function of the exosome-derived lncRNA panel presented in this paper has rarely been investigated so far. Thus, systematic and embedded research is of critical importance to clarify the roles in cell-to-cell information transfer as well as possible underlying carcinogenic mechanisms of these differentially expressed exosomal lncRNAs. This would shed new light on developing candidate therapeutic targets for BC.
Although the present study identified a meaningful novel exosome-derived lncRNA signature for the early diagnosis of BC, there were some deficiencies. The clinical data included in the study were merely from local, Han residents. Ethnicity might be a feasible source of heterogeneity for exosomal noncoding RNAs-assays in BC, which could be ascribed to different living environments and genetic elements between different races. 12 Therefore, large-scale verification should be carried out in various ethnic populations before clinical practice for this exosome-derived lncRNA profile as diagnostic biomarker for NMIBC. Also, since there are no functional data on how these lncRNAs might contribute to tumorigenesis or progression so far, our ongoing work should be focused on clarifying the biological function of the exosomal lncRNAs in pathogenesis of BC by in vitro and in vivo experiments.
Conclusions
Through the first genome-wide urine exosomal lncRNA profiling followed by multi-phase verification, we discerned a three exosome-derived lncRNA panel (CCDC148-AS1, XLOC_006419, and RP5-1148A21.3) that may serve as novel non-invasive biomarker for an early diagnosis of BC. Our preliminary innovative findings may expand the pipeline of ideas for comprehensive clinical treatment of BC, while it is also imperative to carry out more extensive validation and functional investigation of these exosomal lncRNAs.
Supplemental Material
sj-doc-1-jbm-10.1177_03936155251317551 - Supplemental material for Urine exosomal lncRNAs as novel biomarkers for early diagnosis of bladder cancer based on microarray differential expression profiling
Supplemental material, sj-doc-1-jbm-10.1177_03936155251317551 for Urine exosomal lncRNAs as novel biomarkers for early diagnosis of bladder cancer based on microarray differential expression profiling by Jun Li, Liming Zhao, Luning Li, Xiaohua Wang, Yisheng Gao, Yongli Gao and Jinfeng Wang in The International Journal of Biological Markers
Footnotes
Acknowledgments
The authors are grateful to Aksomics Biotechnologies LLC (Shanghai, China) for performing the microarray experiments.
Author contributions
Jinfeng Wang proposed the research design. Jun Li, Jinfeng Wang, Luning Li, and Xiaohua Wang prepared material and conducted experiments. Jun Li, Liming Zhao, Yisheng Gao, and Yongli Gao collected clinical samples. Jun Li, Liming Zhao, Jinfeng Wang, Yisheng Gao, and Yongli Gao carried out the data analysis. Jinfeng Wang wrote the first draft of the manuscript and conducted manuscript revisions. All authors contributed to the article, reviewed the manuscript, and approved the submitted version.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The data that support the findings of this study are available on request from the corresponding author upon reasonable request.
Ethics approval and consent to participate
The studies involving human participants were reviewed and approved by the Ethics Committee of Linyi People's Hospital (No. KY2017020) in accordance with the Declaration of Helsinki and its later amendments or comparable ethical standards. Written informed consent was obtained from all patients/participants included in the study.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by China Postdoctoral Science Foundation (2018M642671); Shandong Provincial Key Research and Development Project (2019GSF108275); Natural Science Foundation of Shandong Province (ZR2020MH315).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.