Abstract
Background
Colon cancer (CC) is one of the most common malignancies worldwide. Characterization of new prognostic biomarkers for right-sided CC (RCC) and left-sided CC (LCC) may contribute to improving early detection. An association of human leukocyte antigens class II (HLA-II) with the predisposition to CC was suggested.
Aim of the study
We evaluated the association of DRB1 and DQB1 with the risk of LCC and RCC.
Patients and methods
Our study comprised 93 CC patients and 100 healthy controls. Genotyping of HLA class II alleles were performed by the Polymerase Chain Reaction Sequence-Specific Primers (PCR-SSP).
Results
DRB1*03 was positively associated with CC. In contrast, DRB1*11, DRB1*13, DQB1*03, and DQB1*05 were negatively linked to CC. Haplotype analysis revealed that DRB1*04-DQB1*04 and DRB1*09-DQB1*02 were positive, while DRB1*01-DQB1*05, DRB1*04-DQB1*03, DRB1*07-DQB1*02, DRB1*11-DQB1*03, DRB1*11-DQB1*05, and DRB1*13-DQB1*06 were negatively associated with CC. For sigmoid CC, DRB1*13, DRB1*11, and DQB1*05 were negative, while DRB1*04-DQB1*02, and DRB1*07-DQB1*03 were positively associated. DRB1*03 and DRB1*04-DQB1*04 were positive, while DRB1*11 and DQB1*03 were negatively linked to RCC. According to the LCC, DRB1*07, DRB1*11, DQB1*03, DQB1*05, and DRB1*07-DQB1*02 were negative. In contrast, DRB1*09-DQB1*02 was positively associated with LCC. Stratified analysis revealed that DRB1*11 is associated with higher risk of metastasis in CC and sigmoid CC, and tolerance to treatment in RCC. DQB1*03 was associated with lymph-node invasion in CC.
Conclusion
DRB1 and DQB1 polymorphisms could be used as future biomarkers for the early detection of subjects at a higher risk of developing RCC and LCC, metastasis in sigmoid CC, and tolerance to treatment in RCC.
Introduction
Colon cancer (CC) remains a major cause of morbidity and death worldwide with an estimated 576,858 deaths occurring in 2020 according to the GLOBOCAN database. 1 CC ranks fifth in terms of prevalence and mortality. The incidence of CC is 600,896 in men and 547,619 in women. 1 Most studies have revealed poorer survival in RCC than in LCC, 2 while recent findings have reported that RCC prognosis is better than that of LCC. 3 Clinical and molecular characteristics, survival outcomes, and therapy responses are different between LCC and RCC. 2 Adjuvant chemotherapies such as 5-fluorouracil (5-FU), and targeted therapies including anti-epidermal growth factor receptor (EGFR) are more efficient for LCC cases. 4
Characterization of new biomarkers for RCC and LCC may contribute to improving early detection. Few studies have focused on identifying novel prognostic variants associated with RCC and LCC susceptibility, which in turn may improve their clinical management. Certain immunological biomarkers such as Kisspeptins, which inhibit the metastasis of malignant cells, were reported to be associated with LCC. 5 A human glycoprotein—MUC16—was described to be highly expressed in RCC conferring poor prognosis. 5 Another recent study evaluated the role of microRNA to search for prognostic biomarkers associated with sigmoid CC. 6
Human leukocyte antigen (HLA) class II molecule has been reported as a prognostic biomarker for many cancers including gastric cancer, 7 and colorectal cancer (CRC).8,9 However, until now few studies have examined the polymorphism of HLA class II as a biomarker for LCC and RCC in different populations. Kanno et al., 10 reported that the expression of HLA class II genes is lower in RCC than in LCC. Another Italian study documented that HLA-DQA1*0201 was significantly less common in patients with CC than in controls. 7 A recent study highlighted an increased expression of HLA class II proteins in CC. 11 Due to the limited results, more studies are needed to examine the possible association of DRB1 and DQB1 gene polymorphisms with LCC and/or RCC. To the best of our knowledge, no study has been conducted to investigate the association of these genes and CC in the Tunisian population.
The aim of this study was to evaluate the association of HLA-DRB1 and DQB1 gene polymorphisms as potential prognostic biomarkers of LCC and RCC in Tunisia and patients’ response to treatment.
Patients and methods
Patients
A case-control study involved a total of 93 Tunisian patients with CC (mean age: 61 ± 12.89 years, 45 male and 48 female). Only 83 CC patients were analyzed according to tumor location, 33 LCC, 21 RCC, and 29 had sigmoid CC. They were recruited from the Oncology service of Saleh Azaeiz Institute (ISA) from February 2014 to October 2018, and the control group consisted of 100 healthy non-cancer individuals (mean age: 42.2 ± 10.5 years, 58 male and 42 female), recruited from the Immunology Service of the Military Hospital of Tunis, reported no personal or family history of CC and were matched to cases by gender and age.
Demographic features were collected for CC patients from the medical records using a coherent questionnaire. Clinical studies included gender; mean age; hypertension; diabetes; smoking and alcohol status; tumor location; resistance and tolerance to treatment; metastasis; and lymph node invasion. The study was approved by the Research & Ethic Committee of ISA (IRB: ISA/2016/03.1). All participants provided written informed consent before participating in the study.
The inclusion criteria were patients with CC. Diagnosis confirmation was established through histopathological examination. At the time of recruitment, none of the patients had undergone any form of therapy and were followed for at least 6 months after treatment using their medical records. Patients’ medical records were also reviewed at 6 months after sampling to monitor their response to first-line chemotherapy (CT) based on Folfox (oxaliplatin, leucovorin, and fluorouracil). Exclusion criteria were patients with rectal cancer, who had a history of previous cancer, had incomplete data, and those who were unable to provide consent to participate in the study.
DNA extraction and HLA genotyping
DNA was extracted from peripheral blood samples collected in EDTA anticoagulant using a QIAampR DNA Blood Mini kit. The genotyping of HLA class II DRB1-DQB1 was done by polymerase chain reaction single-specific-primer (PCR-SSP) according to Micro SSP™ DRB/DQB DNA Typing Trays. PCR-SSP was carried out on a Micro-SSPTM plate with 32 wells. Each well contained a pair of primers specific to one HLA class II allele or group of alleles and a pair of specific β-globin primers used as internal control. The left well, which was the negative control, contained only the pair of specific β-globin primers. After amplification, amplified DNA fragments were migrated on 2.5% ethidium bromide (EtBr) agarose gel electrophoresis and visualized under UV transillumination. One Lambda DNA/LMT Software version 3.0 was used to detect specific DRB1 and DQB1 alleles.
Statistical analysis
Statistical analysis was performed using SPSS software package (version 20.0). HLA allele and haplotype frequencies were estimated by the Arlequin software (version 3.5). Exact tests for Hardy–Weinberg equilibrium (HWE) for DRB1 and DQB1 loci were estimated under the known gametic phase. Pearson chi-square tests with Yate's correction were applied to compare HLA-DRB1 and DQB1 alleles frequencies among cases and controls and to assess the association between specific alleles and clinical features, odds ratio (ORs) and 95% confidence intervals (95% CIs) were determined using logistic regression analysis. The significance was considered at a P-value < 0.05.
Results
Demographic characteristics of CC patients
Overall, 93 CC patients were included in this study: 45 were male and 48 were female. The mean age was 61 ± 12.89 years. Among CC patients, 36 were smokers and 19 were alcoholics. Most CC cases consumed red meat (61.3%) and fats (90.3%). Among patients, 22 were hypertensive and 19 were diabetic. Data concerning tumor location were available for 83 patients: RCC accounted for 21 (25.3%); LCC and sigmoid CC accounted for 33 (39.8%); and 29 (34.9%), respectively. Twenty-three (24.73%) CC patients have developed metastasis, while 58 (62.37%) had lymph-node invasion. Moreover, 31 (33.33%) developed resistance and 90 (96.77%) developed tolerance to CT (Table 1).
Clinical data of CC patients and healthy controls.
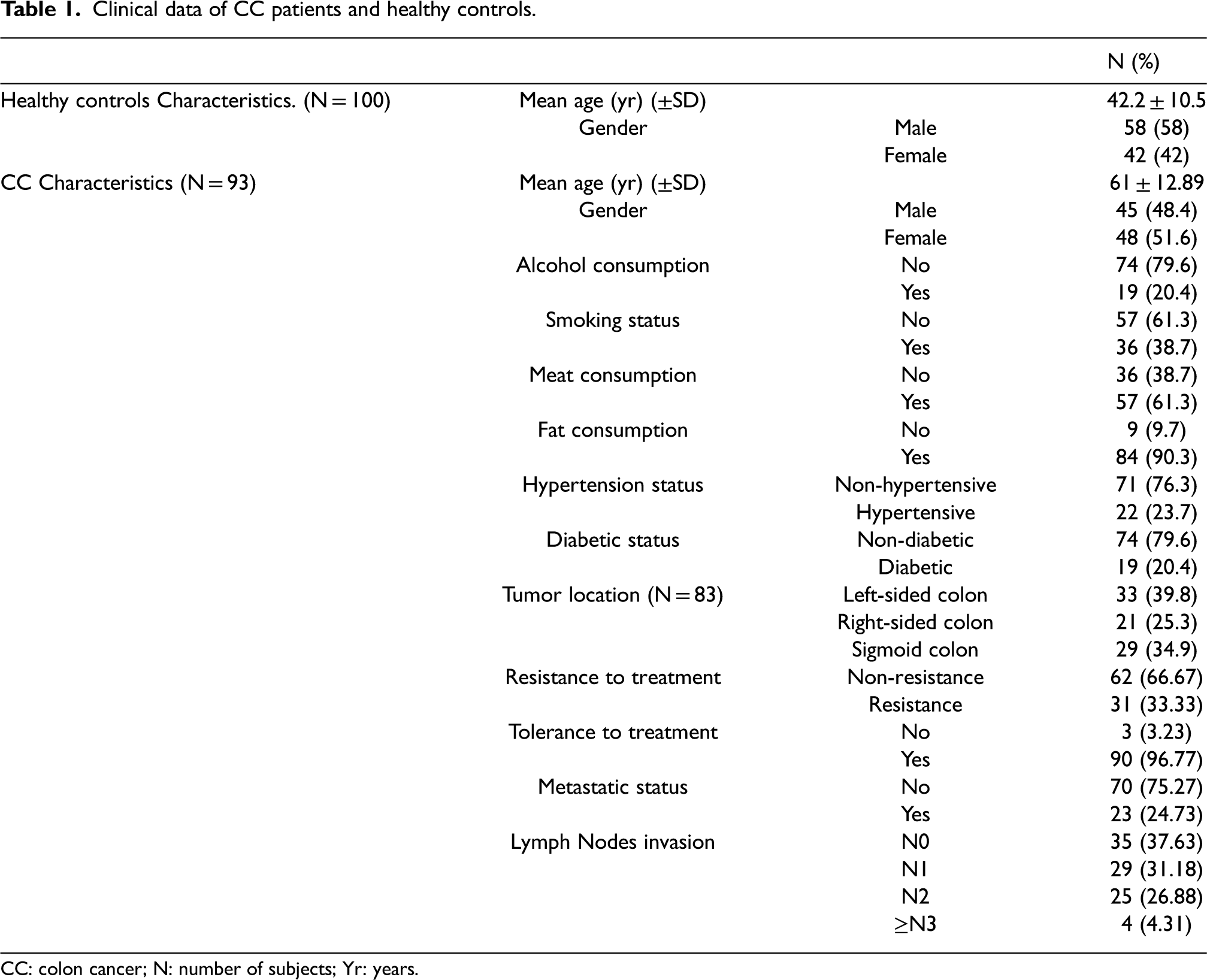
CC: colon cancer; N: number of subjects; Yr: years.
Distribution of DRB1/DQB1 alleles
HLA-DRB1 and -DQB1 typing were undertaken for 93 CC patients and 100 healthy controls. Genotype frequencies of DRB1 and DQB1 loci were consistent with HWE for cases and controls. For DRB1 loci, the results showed a total of 12 alleles in patients and controls. For DQB1 loci, 5 alleles were found. The most frequent alleles in patients were DRB1*03 (21.5%) and DQB1*02 (26.8%), while DRB1*11 (18%) and DQB1*03 (40.5%) were the most frequent alleles recorded for controls. DRB1*03 was positive (P = 0.04, OR (95% CI) = 1.825 (0.978, 3.404)), while DRB1*11 (P = 0.0001, OR (95% CI) = 0.234 (0.108, 0.508)), DRB1*13 (P = 0.012, OR (95% CI) = 0.456 (0.239, 0.873)), DQB1*03 (P = 0.0002, OR (95% CI) = 0.222 (0.120, 0.408)) and DQB1*05 (P = 0.003, OR (95% CI) = 0.346 (0.165, 0.726)) were negatively linked to CC (Table 2).
Distribution of HLA-DRB1* and DQB1* alleles and haplotypes frequencies in CC patients and healthy controls.
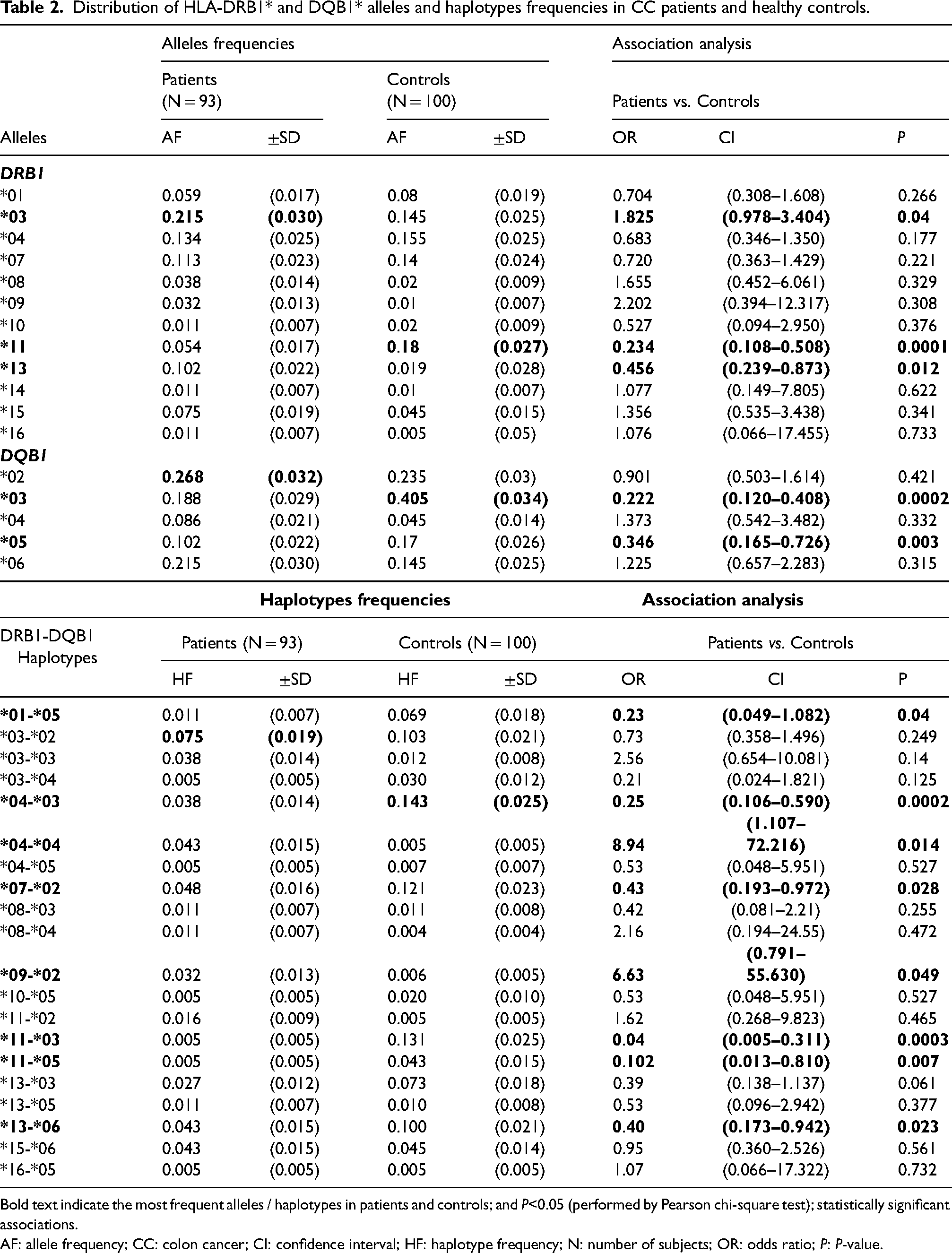
Bold text indicate the most frequent alleles / haplotypes in patients and controls; and P<0.05 (performed by Pearson chi-square test); statistically significant associations.
AF: allele frequency; CC: colon cancer; CI: confidence interval; HF: haplotype frequency; N: number of subjects; OR: odds ratio; P: P-value.
DRB1-DQB1 haplotype analysis
DRB1-DQB1 haplotype analysis identified 20 common haplotypes in patients and controls. Identification of common haplotypes was achieved through advanced genetic analysis facilitated by Arlequin software, 12 based on the concept of linkage disequilibrium (LD), which evaluates the non-random pairing of alleles at different loci. 13 LD plays a pivotal role in reconstructing haplotypes by measuring the degree of association among alleles within the HLA-DRB1 and DQB1 loci.
The most frequent haplotype in patients was DRB1*03-DQB1*02 (7.5%), while DRB1*04-DQB1*03 (14.3%) was the most frequently found in controls. A positive association of DRB1*04-DQB1*04 and DRB1*09-DQB1*02 with CC susceptibility was observed (P = 0.014, OR (95% CI) = 8.94 (1.107, 72.216); P = 0.049, OR (95% CI) = 6.63 (0.791, 55.630), respectively. In contrast, DRB1*01-DQB1*05 (P = 0.04, OR (95% CI) = 0.23 (0.049, 1.082)), DRB1*04-DQB1*03 (P = 0.0002, OR (95% CI) = 0.25 (0.106, 0.590)), DRB1*07-DQB1*02 (P = 0.028, OR (95% CI) = 0.43 (0.193, 0.972)), DRB1*11-DQB1*03 (P = 0.0003, OR (95% CI) = 0.04 (0.005, 0.311)), DRB1*11-DQB1*05 (P = 0.007, OR (95% CI) = 0.102 (0.013, 0.810)) and DRB1*13-DQB1*06 (P = 0.023, OR (95% CI) = 0.40 (0.173, 0.942)) were negatively linked to CC (Table 2).
Association of DRB1/DQB1 alleles with tumor location
We tested the possible association of DRB1 and DQB1 alleles with CC according to tumor location. Of the included CC patients, 33 were LCC, 21 were RCC, and 29 were sigmoid CC. For RCC, the results showed a total of 10 DRB1 and 5 DQB1 alleles in patients and controls. DRB1*03 (P = 0.03, OR (95% CI) = 2.879 (1.09, 7.606)) was positive, while DRB1*11 (P = 0.019, OR (95% CI) = 0.204 (0.045, 0.929)) and DQB1*03 (P = 0.0002, OR (95% CI) = 0.132 (0.041, 0.424)) were negatively linked to RCC (Table 3).
HLA-DRB1 and DQB1 alleles frequencies in patients with LCC, RCC and sigmoid CC in comparison with healthy controls.
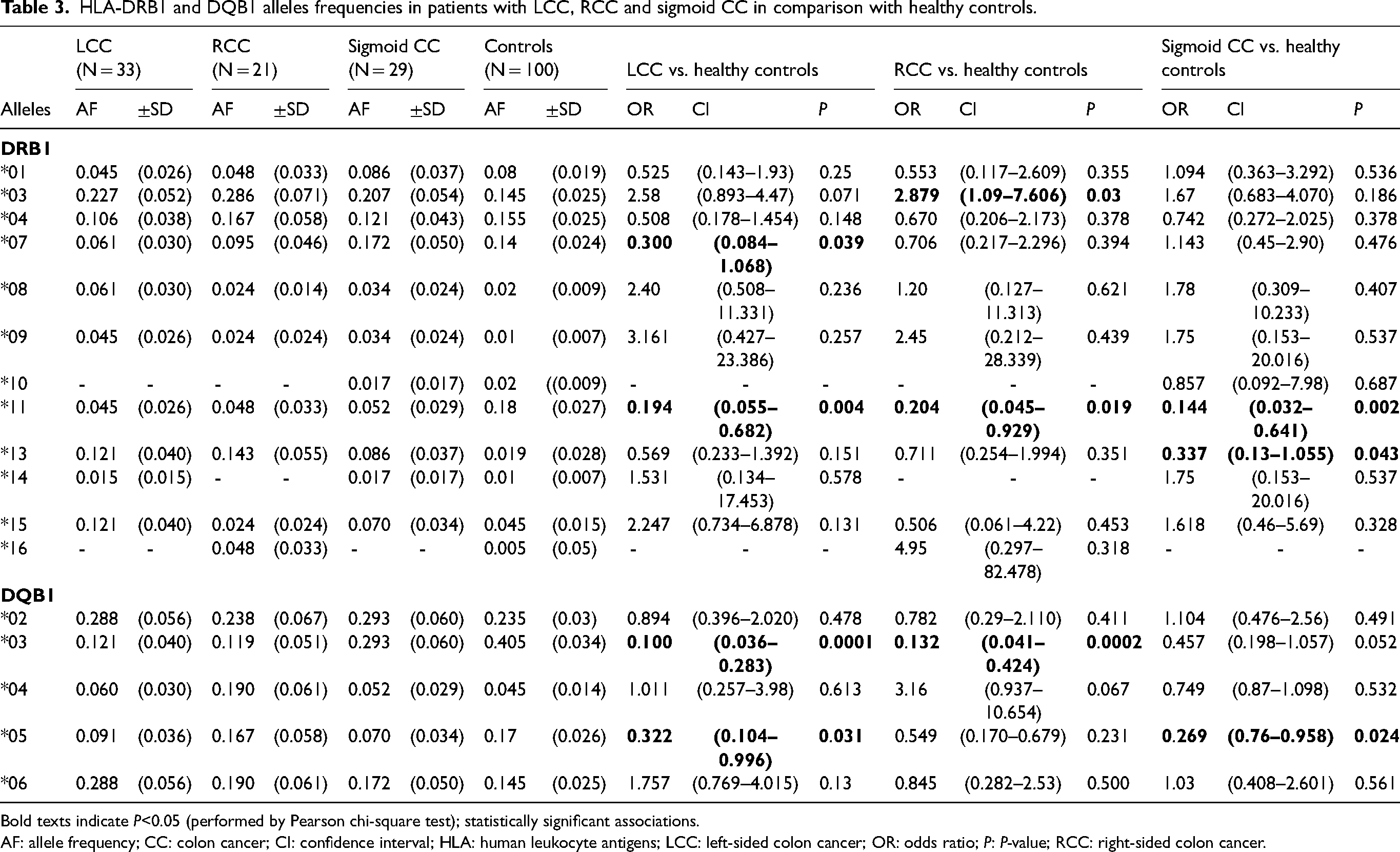
Bold texts indicate P<0.05 (performed by Pearson chi-square test); statistically significant associations.
AF: allele frequency; CC: colon cancer; CI: confidence interval; HLA: human leukocyte antigens; LCC: left-sided colon cancer; OR: odds ratio; P: P-value; RCC: right-sided colon cancer.
For LCC, 10 DRB1 and 5 DQB1 alleles in patients and controls were noted. For DRB1 loci, DRB1*07 (P = 0.039, OR (95% CI) = 0.300 (0.084, 1.068)) and DRB1*11 (P = 0.004, OR (95% CI) = 0.194 (0.055, 0.682)) were negatively associated with LCC. For DQB1 loci, DQB1*03 (P = 0.0001, OR (95% CI) = 0.100 (0.036, 0.283)) and DQB1*05 (P = 0.031, OR (95% CI) = 0.322 (0.104, 0.996)) were negatively linked to LCC (Table 3).
According to sigmoid CC, the results showed a total of 11 DRB1 and 5 DQB1 alleles in sigmoid CC and controls. DRB1*11 (P = 0.002, OR (95% CI) = 0.144 (0.032, 0.641)), DRB1*13 (P = 0.043, OR (95% CI) = 0.337 (0.13, 1.055)) and DQB1*05 (P = 0.024, OR (95% CI) = 0.269 (0.76, 0.958)) alleles were negatively associated with increased risk of sigmoid CC (Table 3).
Association of DRB1-DQB1 haplotype with tumor location
We investigated the possible associations of DRB1-DQB1 haplotypes with LCC, RCC, and sigmoid CC. Results from Table 4 demonstrated 17 common haplotypes in LCC patients and controls. DRB1*09-DQB1*02 (P = 0.048, OR (95% CI) = 9.47 (0.968, 92.527)) was positively associated with LCC. While, DRB1*07-DQB1*02 (P = 0.016, OR (95% CI) = 0.14 (0.018, 1.050)) was negative. Fourteen common haplotypes in RCC patients and controls were shown. Only the DRB1*04-DQB1*04 haplotype was positively associated with RCC (P = 0.0004, OR (95% CI) = 33.17 (3.876, 283.764)). According to sigmoid CC, results illustrated 14 common haplotypes in patients and controls. DRB1*04-DQB1*02 (P = 0.036, OR (95% CI) = 10.85 (1.107, 106.421)) and DRB1*07-DQB1*03 (P = 0.048, OR (95% CI) = 3.67 (1.027, 13.183)) were positively associated with sigmoid CC (Table 4).
HLA-DRB1 and DQB1 haplotype frequencies in patients with LCC, RCC, and sigmoid CC in comparison with healthy controls.
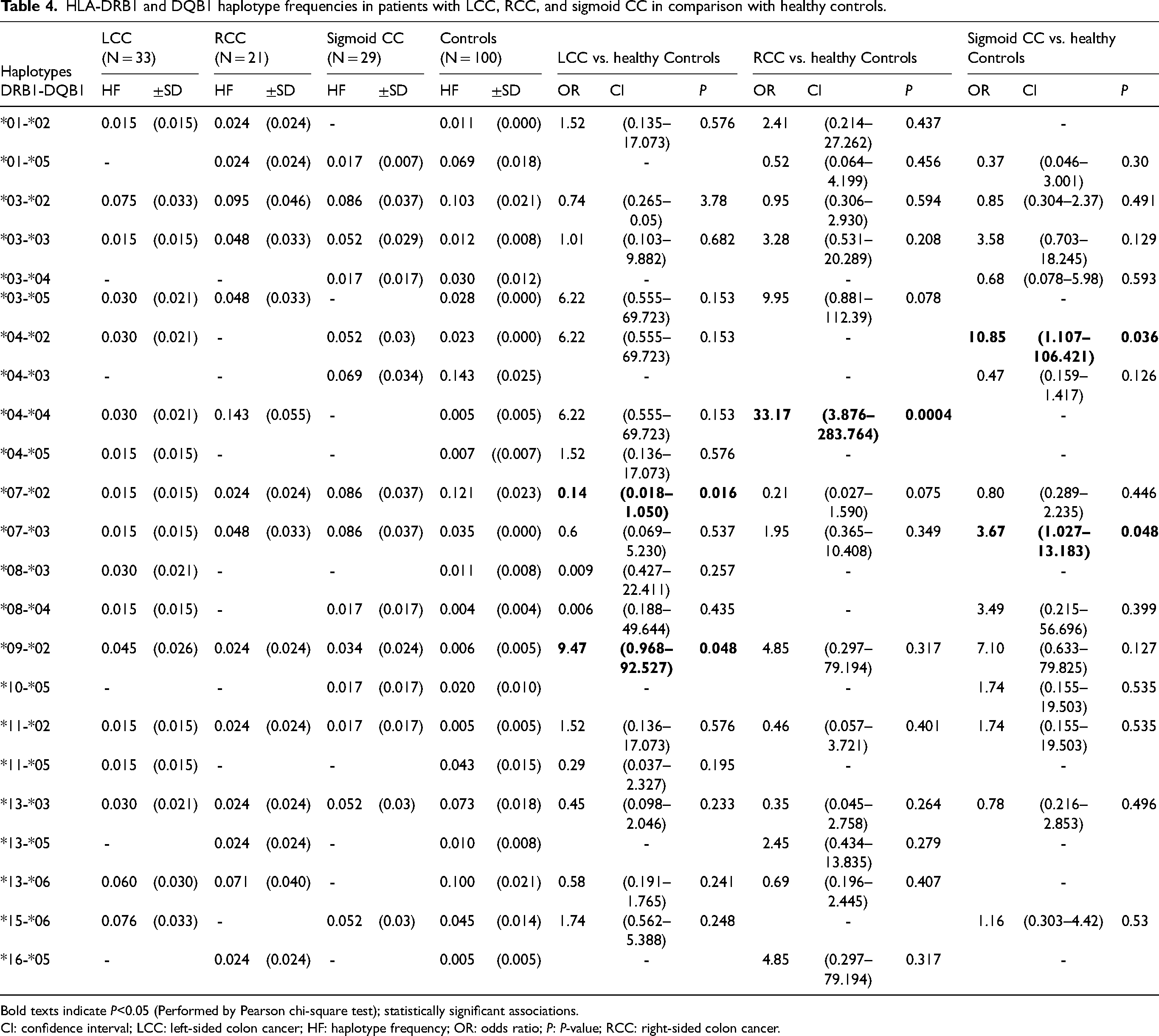
Bold texts indicate P<0.05 (Performed by Pearson chi-square test); statistically significant associations.
CI: confidence interval; LCC: left-sided colon cancer; HF: haplotype frequency; OR: odds ratio; P: P-value; RCC: right-sided colon cancer.
Association of DRB1/DQB1 specific alleles with metastatic status, lymph nodes invasion, and response to treatment
According to metastatic status, our results revealed that HLA-DRB1*11 was positively associated with metastasis in CC (P = 0.002, OR (95% CI) = 9.14 (1.363, 103.53)), and sigmoid CC (P = 0.004, OR (95% CI) = 47 (1.976, 111.7)). However, we did not find any significant associations between DRB1 and DQB1 specific alleles and the development of metastasis in LCC and RCC (Table 5).
Association of HLA-DRB1 and -DQB1 specific alleles with metastasis, lymph-node invasion, and response to treatment in CC, LCC, RCC, and Sigmoid CC.
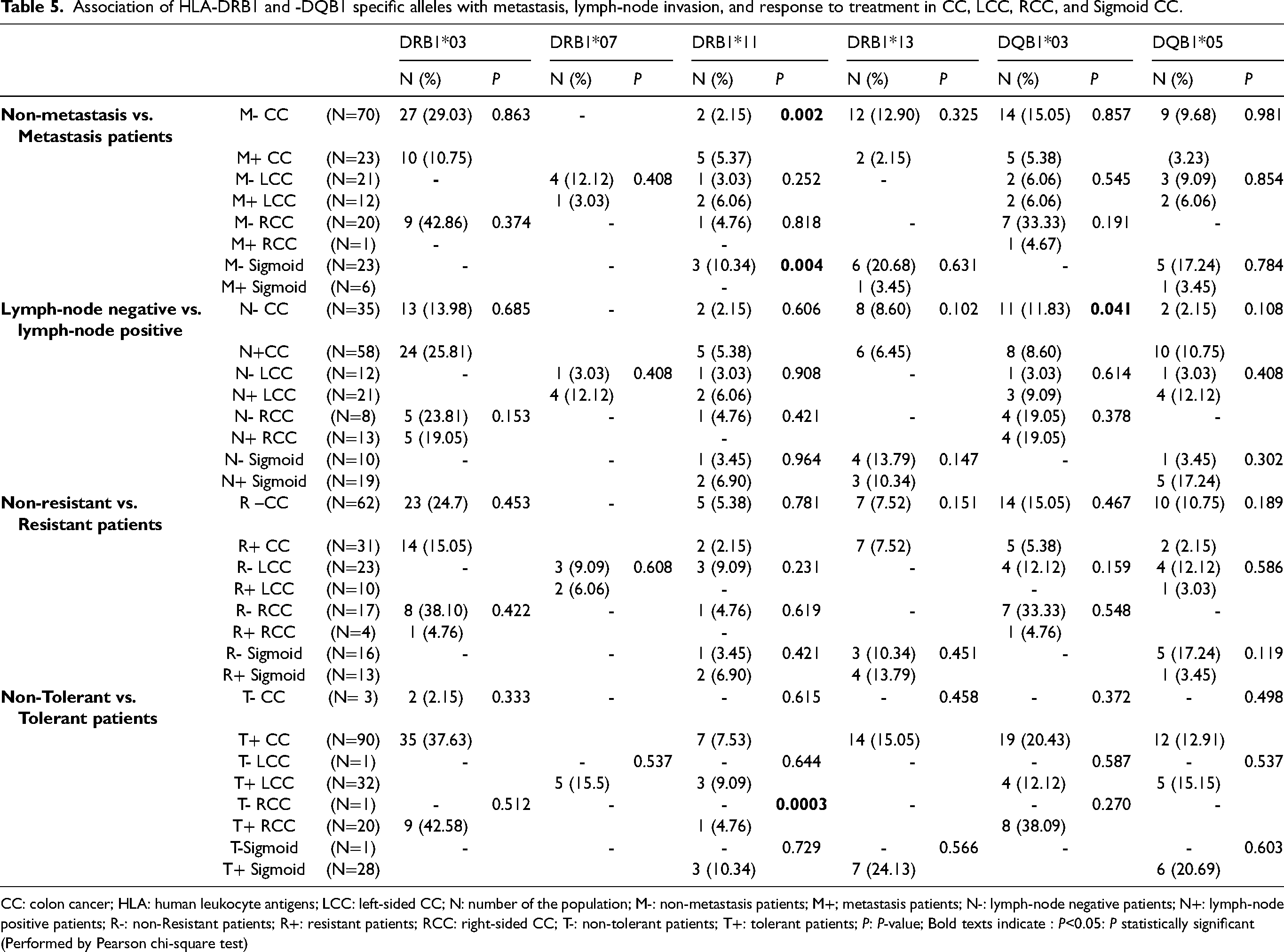
CC: colon cancer; HLA: human leukocyte antigens; LCC: left-sided CC; N: number of the population; M-: non-metastasis patients; M+; metastasis patients; N-: lymph-node negative patients; N+: lymph-node positive patients; R-: non-Resistant patients; R+: resistant patients; RCC: right-sided CC; T-: non-tolerant patients; T+: tolerant patients; P: P-value; Bold texts indicate : P<0.05: P statistically significant (Performed by Pearson chi-square test)
For lymph nodes invasion, a negative association between HLA-DQB1*03 and CC with lymph-node invasion was revealed (P = 0.041, OR (95% CI) = 0.35 (0.107, 1.107)) (Table 5).
Results from Table 5 demonstrated the association between specific alleles DRB1 and DQB1 and tolerance, as well as resistance to CT. Positive association of DRB1*11 with tolerance to CT in RCC (P = 0.0003, OR (95% CI) = 12.3 (5.172, 292.5)) was found (Table 5).
Discussion
CC is one of the most common malignancies worldwide. 1 Previous studies have highlighted the link between HLA class II gene polymorphisms and several diseases including type I diabetes 14 and aplastic anemia, 15 as well as several cancers including nasopharyngeal carcinoma, 16 glioblastoma, 17 CRC,8,18 breast cancer, 19 and cervical cancer. 20 To the best of our knowledge, the association of DRB1 and DQB1 alleles/haplotypes with CC is not yet investigated. Although some studies have suggested that HLA-DRB1 and -DQB1 gene polymorphisms play a role in the pathogenesis of CRC, they have not enrolled stratified analysis according tumoral location.
The HLA class II antigen-processing pathway plays a critical role in presenting antigenic peptides to CD4+ T-cells, primarily derived from exogenous proteins within the endocytic compartment. CD4+ T-cells function as both helper and effector cells, identifying tumor cells expressing HLA class II antigens. 21 Consequently, the expression of these antigens by tumor cells impacts the immune responses specific to tumor antigens. 21 Notably, HLA class II antigen expression in normal rectal and colon and CRC cells can be induced through stimulation with interferon gamma (IFNγ). However, the expression of HLA class II antigens in CRC cells is regulated by the methylation of class II transactivator-isoform-PIV. Furthermore, this expression is inhibited by somatic mutation in the Regulatory Factor X5 gene, a crucial transcriptional activator involved in regulating MHC class II genes. 21
Our study aimed to explore the association between HLA-DRB1 and DQB1 gene polymorphisms and the occurrence of CC, LCC, RCC, and sigmoid CC, and the development of metastasis, lymph nodes invasion, and response to treatment.
The results of our study documented the positive association of DRB1*03 with an increased risk of CC. Otherwise, negative associations were revealed for DRB1*11, DRB1*13, DQB1*03, and DQB1*05 alleles. An Italian study enrolled only on HLA-DQA polymorphism reported an association of HLA-DQA1*0201 alleles with the risk of CC. 7 Otherwise, regarding the expression of HLA class II antigens in CC compared to normal colonic mucosa, inconsistent findings were reported. In fact, a reduced expression of HLA class II antigens in tumoral tissue of CC from New York patients was found. 22 Conversely, a recent study highlighted an increased expression of HLA class II antigens in CC of patients from Boston, USA. 11
Haplotype association analysis showed that DRB1*04-DQB1*04 and DRB1*09-DQB1*02 were positive, while DRB1*01-DQB1*05, DRB1*04-DQB1*03, DRB1*07-DQB1*02, DRB1*11-DQB1*03, DRB1*11-DQB1*05, and DRB1*13-DQB1*06 were negatively linked to CC, suggesting a protective effect of these haplotypes. Several studies addressed the association of HLA class II gene polymorphisms with different types of cancer development such as CRC; HLA-DRB1*0301 was reported to be associated with an increased risk in patients with a clinical history of ulcerative colitis, while DRB1*0701 and DQB1*0501 were negatively linked to CRC in the USA. 23 Another Italian study suggested that DRB1*13:01 is positively associated with CRC, and DRB1*11:01 is negatively associated it. 24 The HLA-DQB1*02 allele has also been reported to be positively associated with CRC risk in the Chinese population. 25 To the best of our knowledge, this is the first study to report the association of DRB1 and DQB1 gene polymorphisms with CC susceptibility.
DRB1 and DQB1 gene polymorphisms were shown to be related to the tumor location of CC in this study. For sigmoid CC, DRB1*13, DRB1*11, and DQB1*05 alleles were negatively related to increased risk, while DRB1*04-DQB1*02 and DRB1*07-DQB1*03 haplotypes were positively linked to sigmoid CC. For RCC, DRB1*03 allele and DRB1*04-DQB1*04 haplotype were positive, while DRB1*11 and DQB1*03 were negatively linked to RCC. According to LCC, DRB1*07, DRB1*11, DQB1*03, and DQB1*05 alleles and DRB1*07-DQB1*02 haplotype were negative, while DRB1*09-DQB1*02 haplotype was positively associated with LCC development. The expression of HLA class II was reported to be lower in RCC than in LCC. 10
We analyzed the association of DRB1- and DQB1-specific alleles with metastasis, lymph-node invasion, and response to treatment in patients. Our findings indicated that DRB1*11 was positively associated with the development of metastasis in CC and sigmoid CC, implying their potential role in the disease progression. However, no significant associations were observed for LCC and RCC. These findings are novel and have never been reported in CC. These results contrasted with observations in other diseases (e.g., breast cancer); HLA-DRB1*11 showed no association, while the deletion of this allele correlated with obesity. 19
Lymph nodes are crucial for several biological processes, including the development of metastasis and immune modulation. 26 Lymph-node invasion is a prognostic factor for many cancers such as CRC 27 and pancreatic cancer. 28 In our study, DQB1*03 was negatively associated with lymph-node invasion in CC. Lee et al. 27 reported that an increased retrieval of lymph nodes was linked to better survival in CRC. HLA-DQB1 has been suggested to be positively associated with early-stage in lung adenocarcinoma 29 and immune response in cervical cancer. 20 HLA-DQB1*03 correlated with increased lymph node metastasis and advanced stage in patients with gastric cancer, suggesting its contribution to a poor prognosis. 30 DQB1*03:01 was associated with worse survival in chemo resistant patients with urothelial cancer. 31
Drug resistance presents a significant challenge in achieving effective cancer treatment; combination chemotherapy became a new paradigm for cancer therapy that led to the development of increasingly complex regimens. 32 In this study, we evaluated the association between specific alleles with resistance and tolerance to CT in CC, RCC, LCC, and sigmoid CC. Our analysis revealed a positive association between DRB1*11 and tolerance to treatment for RCC. However, we did not identify any associations between specific alleles and resistance to CT. To the best of our knowledge, this is the first study to report the association of DRB1*11 with response to treatment in CC. It has been suggested that this allele is associated with increased risk for disease recurrence in patients with melanoma 33 and interstitial aseptic pneumonitis in cancer patients receiving PD-1/PDL1 immune-checkpoint blockade. 34
These results suggested that RCC and LCC presented several differences. They are not only located on different sites of the colon but also present different molecular mechanisms, proposing the role of the HLA class II system in patient outcome. Our study shows the differences in associations between HLA class II gene polymorphisms and LCC and RCC: DRB1*03 and DRB1*04-DQB1*04 were positive, while DRB1*11 and DQB1*03 were negatively linked to RCC. Moreover, DRB1*11 was associated with tolerance to CT. According to LCC, DRB1*07, DRB1*11, DQB1*03, and DQB1*05 alleles, and DRB1*07-DQB1*02 haplotype were negative. In contrast, DRB1*09-DQB1*02 was positively associated with LCC. To the best of our knowledge, no other studies were reported to examine the association of DRB1 and DQB1 gene polymorphisms with RCC and LCC. DRB1 and DQB1 polymorphisms can be considered as prognostic biomarkers for early detection of LCC and RCC.
The present study had several strengths. In fact, CC cases and controls were matched according to ethnicity (only Tunisian Arabic-speaking cases and controls), thereby minimizing the possibility of ethnic differences inherent in genetic association studies. However, our study has a critical point since it had a modest sample size. Additional larger designed studies are needed to better evaluate the effect of these variants on CC susceptibility.
Conclusion
Our study shows a new association between HLA class II gene polymorphisms and CC and proves that it would be important to investigate the association of these biomarkers with RCC and LCC in different populations worldwide. HLA class II genes may constitute potential predictive biomarkers for CC susceptibility, metastasis, and response to treatment, and tumor location should be considered a prognostic factor for the early identification of individuals at high risk of developing CC.
Footnotes
Acknowledgments
We thank all blood donors and the patients with CC who participated in the present study. We are grateful to the medical team of Tunisian Saleh Azaiez Institute and Tunisian Military Hospital for their help in the collection of blood samples.
Author contributions
Besma YACOUBI-LOUESLATI and Imene Namouchi designed the study. Amani Attia analyzed, interpreted the data, drew the tables and drafted the manuscript. Imene Namouchi and Amani Attia performed experiments and the statistical analysis of the data. Amel Mezlini was involved in the collection of the data. Awatef Lagha and Ezzedine Ghazouani participated in the methodology. Besma YACOUBI-LOUESLATI helped with the final revision of the article. Besma Yacoubi-Loueslati and Imene Namouchi contributed equally to this work. All authors have read and approved the final manuscript.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval
The study was approved by the National Ethics Board and that all participants have provided informed consent, and the approval code is: ISA/2016/03.1, granted in April 2016.
Funding
The study was supported by the Tunisian Ministry of Higher Education and Scientific research.
